Abstract
Background
Globally, prostate cancer is the second most common malignancy in males. Serum microRNAs (miRNAs) may function as non-invasive and innovative biomarkers for various cancers. Our study aimed to determine potential miRNAs for prostate cancer screening.
Methods
A three-stage study was accomplished to ascertain crucial miRNAs as markers. In the screening stage, we searched PubMed for aberrantly expressed miRNAs relevant to prostate cancer and selected them as candidate miRNAs. In training and validation stages, with serum specimens from 112 prostate cancer patients and 112 healthy controls, expressions of candidate miRNAs were identified through quantitative reverse transcription-polymerase chain reaction. The diagnostic capabilities of miRNAs were determined by receiver operating characteristic curves. Bioinformatic analysis was utilized to explore the function of the critical miRNAs.
Results
Expression of six serum miRNAs (miR-34b-3p, miR-556-5p, miR-200c-3p, miR-361-5p, miR-369-3p, miR-485-3p) were significantly altered in prostate cancer patients contrasted with healthy controls. The optimal combination of critical miRNAs is a three-miRNA panel (miR-34b-3p, miR-200c-3p, and miR-361-5p) with good diagnostic capability. FLRT2, KIAA1755, LDB3, and NTRK3 were identified as the potential genes targeted by the three-miRNA panel.
Conclusions
The three-miRNA panel may perform as an innovative and promising serum marker for prostate cancer screening.
Introduction
Globally, prostate cancer (PC) is the second most frequently diagnosed cancer among men. There are significant ethnic and regional differences in the incidence of PC. In the USA, PC is the most common form of cancers in males, with an estimated 268,490 new cases and an estimated 34,500 deaths in 2022. 1 It is believed that the incidence of PC in Asian countries is much lower than that in Western countries. However, the incidence of PC has markedly increased in China. 2 It is estimated that, in 2020, PC ranked sixth in incidence and the seventh main cause of cancer-related death in China. 3 It is also obvious that the burden of PC is rising gradually. Therefore, it is crucial to implement PC screening and early diagnosis to enhance the rate of survival and reduce the burden of cancer.
Prostate-specific antigen (PSA), secreted by prostatic epithelial cells, has many clinical applications in PC such as early detection, staging, and surveillance of cancer progression and therapy effectiveness. 4 Detecting the blood concentration of total PSA is the most common method for PC screening. However, there is no recommended unanimous PSA threshold for all males. 5 The American Cancer Society found that the threshold of 4.0 ng/mL PSA can be used for PC screening with 21% of sensitivity and 91% of specificity, while reducing the PSA threshold to 3.0 ng/mL would lead to enhancing sensitivity to 32% and worsening specificity to 85%. 6 In addition, conditions like bacterial prostatitis, benign prostatic hyperplasia, and physical manipulation can explain falsely elevated PSA levels and is noteworthy. 5 Another way to screen PC is digital rectal examinations (DRE). It is a sign to perform PC biopsy with abnormal DRE findings. The obstacle to implementing DRE for PC screening is discomfort in men and low detection rate. 6 Biopsy and histological validation are the methods of definitive diagnosis. However, ultrasound-guided biopsy is generally the standard method for PC biopsies, which is invasive and can lead to bleeding and infection. 6 Therefore, a non-invasive, innovative, and credible screening method is imminently needed.
MicroRNAs (miRNAs) are non-coding and short (22 nucleotides in length) RNAs that play pivotal regulatory roles in gene expression through messenger RNA degradation and protein translation suppression. 7 Many studies lately have uncovered that miRNAs could be released into serum and may function as potential biomarkers for various tumors; they are also involved in the regulation of physiological and pathological processes.8–10 A review noted that miRNAs are differentially expressed in PC and participate in regulating its progression. 11 Therefore, serum miRNAs may serve as a promising marker for PC screening and early diagnosis.
The purpose of our study is to ascertain relevant serum miRNAs as markers for PC screening and diagnosis. The expression profile of serum miRNAs was verified through quantitative reverse transcription-polymerase chain reaction (RT-qPCR). Three-stage research was conducted to ascertain the diagnostic capability of key miRNAs. Moreover, we used bioinformatics analyses to explore potential targets of the critical miRNAs and their functional annotation and pathway analyses.
Materials and methods
Study population and statement of ethics
We recruited 112 PC patients and 112 healthy controls (HCs) from January 2018 to September 2020. All 224 subjects were from Peking University Shenzhen Hospital. The PC patients were diagnosed clinically based on prostate biopsy and received no treatment before serum specimen collection. The HCs were healthy male volunteers without cancers or other diseases. All the participants were fully informed and voluntarily signed informed consent forms. The consent of the Ethics Committee of Peking University Shenzhen Hospital was obtained for this research. Procedures for collecting the serum specimens abided by the relevant regulatory framework established by the committee.
Study procedure/research design
The three-stage study was undertaken to ascertain crucial miRNAs as markers for PC screening and diagnosis. First, in the screening stage, we searched PubMed for aberrantly expressed miRNAs relevant to PC and selected them as candidate miRNAs. The retrieval strategy was: “Prostate cancer” AND “MicroRNAs”[Mesh]. Then the 12 most potential miRNAs for which there was no literature on the expression in serum were selected for further investigation (Supplementary Table 1). Next, in the training stage, we utilized the RT-qPCR method to ascertain the expression profile of candidate miRNAs with serum specimens randomly chosen from 28 PC patients and 28 HCs. Finally, in the validation stage, serum specimens from 84 PC patients and 84 HCs were utilized for analyzing the expression profile and evaluating the diagnostic capabilities of crucial miRNAs by RT-qPCR. The optimal combination of miRNAs with optimal diagnostic capabilities were selected using backward stepwise logistic regression analysis. In addition, bioinformatic analysis was also included in our study.
Serum sample collection and RNA isolation
None of the participants received any treatment before serum specimen collection. Once 10 mL of venous blood was taken, it was centrifuged within 2 h at 1000×g for 10 min and subsequently at 15,000×g for 5 min. For the purpose of normalizing the variability during the extraction procedure, we added 2uL miR-54 (cel-miR-54-5p) (10 nm/L, RiboBio, Guangzhou City, China) to each serum specimen in advance as an internal reference. Next, we used the TRIzol LS isolation kit (Thermo Fisher Scientific, Waltham, MA, USA) to complete total RNA extraction from serum in compliance with the manufacturer's manual. Following extraction, serum specimens were refrigerated at −80°C for purpose of further investigation. NanoDrop 2000c spectrophotometer (Thermo Scientific) was utilized to gauge the RNA concentration and purity.
RT-qPCR approach
The detection of miRNAs expression level was achieved through RT-qPCR. The amplification of miRNAs was implemented with the reverse transcription-specific primers from the Bulge-Loop miRNA RT-qPCR primer set (RiboBio). With Taqman probe, qPCR was completed on LightCycler480 Real-Time PCR System (Roche Diagnostics, Mannheim, Germany) under conditions at 95°C for 20 s, followed by 40 cycles of 95°C for 10 s, 60°C for 20 s, and 70°C for 10 s The analysis of relative miRNA expression was performed by the 2-ΔΔCq method. 12
Bioinformatic analysis
To explore potential function of critical miRNAs relevant to PC, miRWalk3.0 (http://mirwalk.umm.uni-heidelberg.de/) was used for predicting and experimentally validating information on miRNA-target interaction. 13 We considered those predicted genes targeted by two or three candidate miRNAs as potential target genes. Enrichment analysis and functional annotation for predicted target genes were performed through the Enrichr database (http://amp.pharm.mssm.edu/Enrichr/). 14 This study used Gene Ontology (GO) functional annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis.
Statistical analysis
The serum specimens of participants consisted of training and validation stage. Information of demographic and clinical features between groups was exhibited as percentages and counts. Continuous variables were demonstrated as mean values ± SD. The analysis regarding multiple comparisons between training and validation stages was performed by the Kruskal–Wallis test. The analysis regarding the differential expression of miRNAs in serum specimens between PC patients and HCs was performed by the Student’s t test or the Mann–Whitney test. Specificity, sensitivity, and diagnostic capability were evaluated by receiver operating characteristic (ROC) curves and the area under the ROC curve (AUC). The optimal diagnostic panel of miRNAs was determined by Youden index (computed as J = Sensitivity + Specificity – 1). All statistical processing were achieved through SPSS 20.0. When the P-value was < 0.05, the results were statistically significant.
Results
Characteristics of study subjects
After selection of candidate miRNAs in screening stage, a total of 224 men were recruited for further study including 112 PC patients and 112 HCs. The 112 PC patients who had primary PC without previous history of other tumors, received prostate biopsy and were pathologically diagnosed with PC according to the standards established by the World Health Organization. All 112 HCs consisted of those who had regular physical check-ups, people without previous history of cancer or disease, and patients with negative prostate biopsy. No significant difference was detected in age between PC patients and HCs (P-values > 0.05). Table 1 presented the tumor node metastasis staging, lymph node metastasis, International Society of Urological Pathology grade groups, and distant metastasis of PC.
Clinical and demographic information of 224 participants.
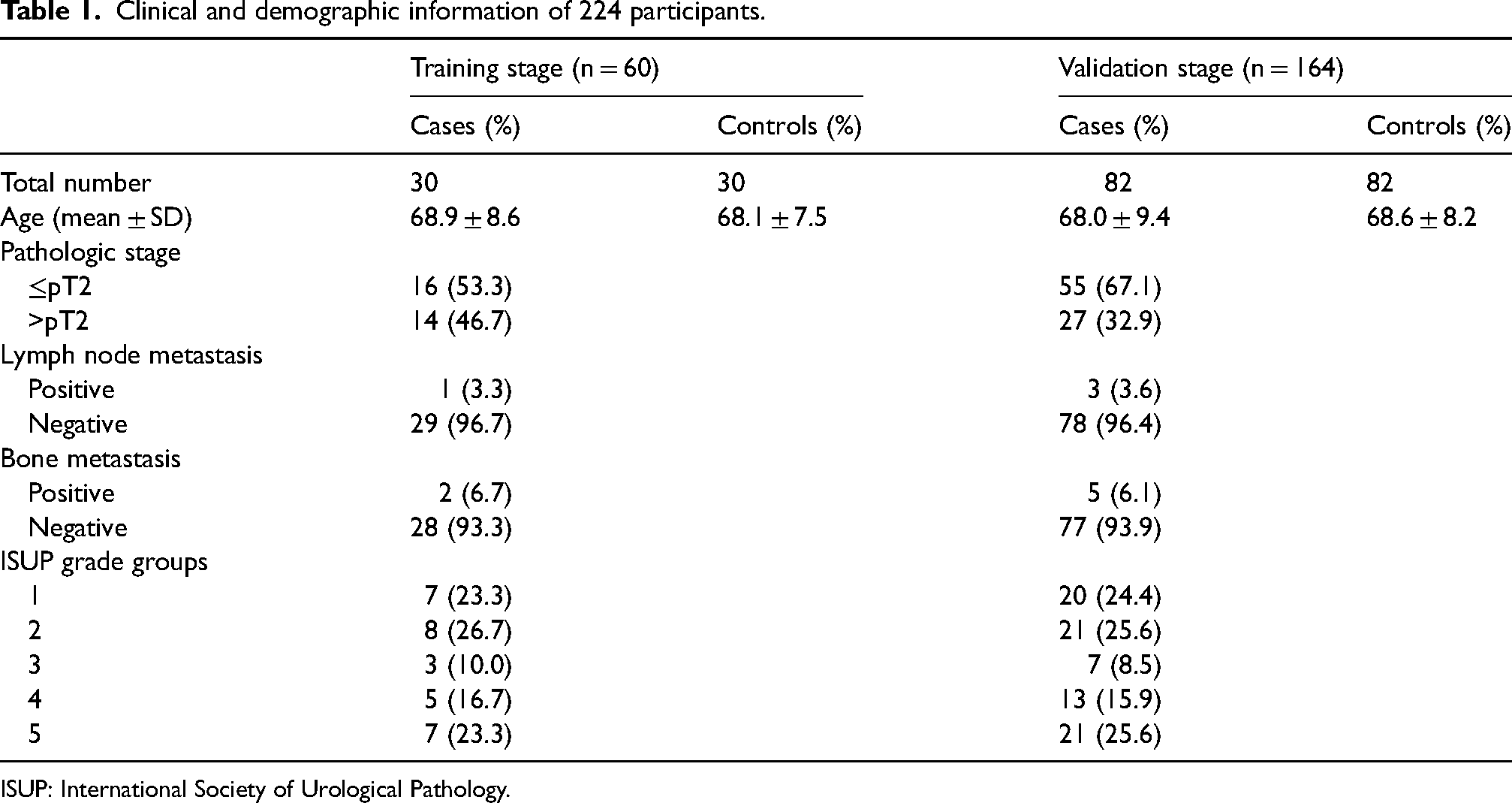
ISUP: International Society of Urological Pathology.
Selection of miRNAs in screening stage
We searched published articles on PubMed for miRNAs relevant to PC and eventually selected 12 miRNAs that were aberrantly expressed in PC as candidate biomarkers in the screening stage. They were miR-34b-3p, miR-27a-5p, miR-30a-5p, miR-556-5p, miR-200c-3p, miR-361-5p, miR-124-5p, miR-369-3p, miR-105-5p, miR-485-3p, miR-452-5p, and miR-642a-5p (Figure 1). Studies suggest that some miRNAs play tumor-suppressive roles in PC, such as miR-27a-5p, 15 miR-200c-3p, 16 miR-361-5p, 17 miR-485-3p, 18 and miR-642a-5p. 19 However, one study revealed that miR-556-5p function as onco-miRNA. 20 There was no literature on the expression of these specific miRNAs in serum. All these miRNAs were selected for subsequent research stages.
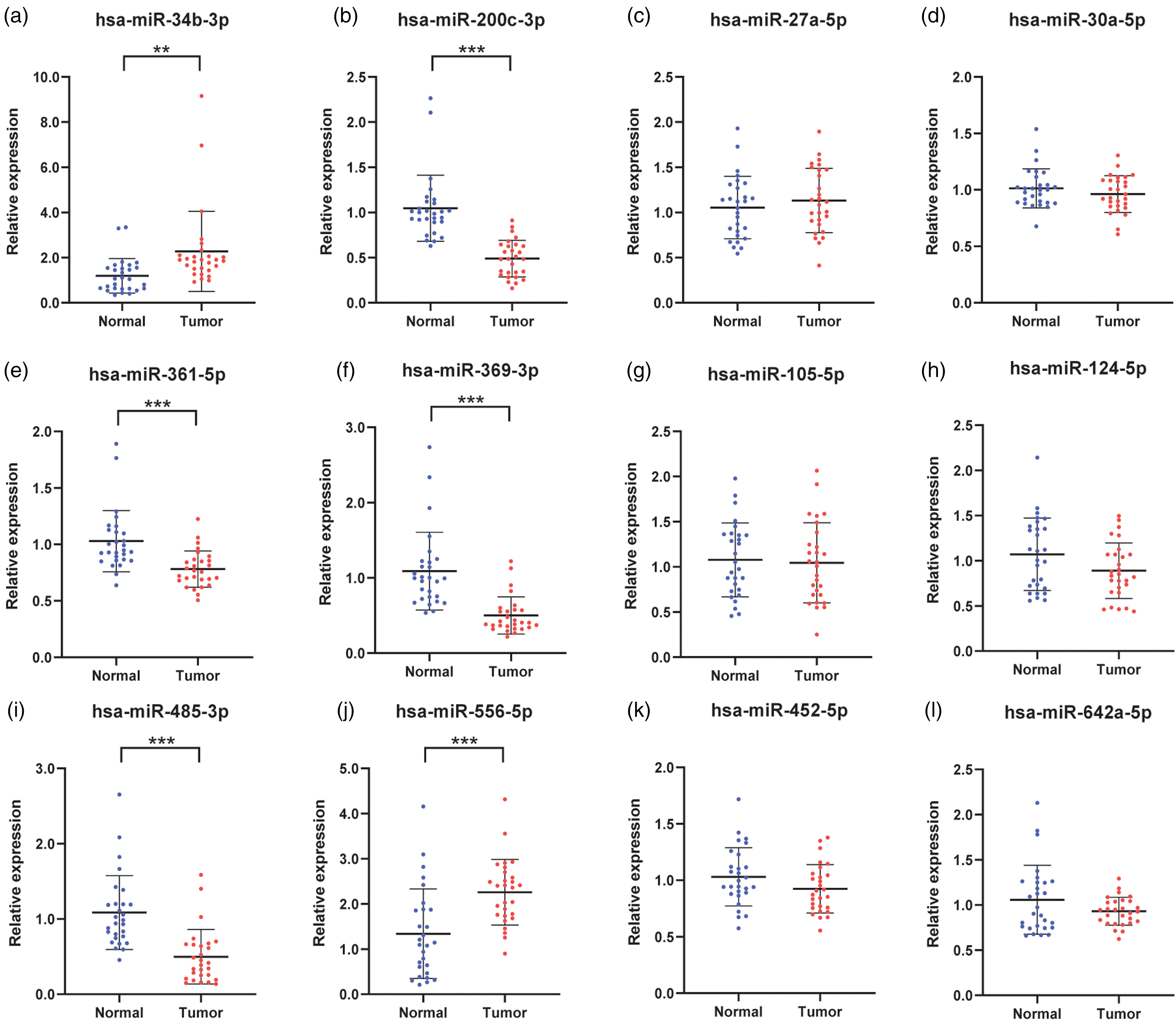
In the training stage, relative expression levels of (a) to (l) 12 candidate miRNAs in serum with 28 PC patients and 28 age- and stage- matched HCs. Six miRNAs that were aberrantly expressed were determined for the validation stage. They were (a) miR-34b-3p, (b) miR-200c-3p, (e) miR-361-5p, (f) miR-369-3p, (i) miR-485-3p, and (j) miR-556-5p. *P < 0.05, **P < 0.01, ***P < 0.001.
Expression of selected miRNAs in training stage
With serum specimens from 28 PC patients and 28 age- and stage- matched HCs, 12 candidate miRNAs that were selected in screening phase were evaluated by the RT-qPCR approach. In total, six miRNAs were still aberrantly expressed compared to HCs with a P-value of <0.05 (Figure 1). There were two upregulated miRNAs (miR-34b-3p, miR-556-5p) and four downregulated miRNAs (miR-200c-3p, miR-361-5p, miR-369-3p, miR-485-3p). Therefore, the above six miRNAs were determined for further investigation.
Expression profile and diagnostic capabilities of critical miRNAs in the validation stage
To further verify whether these six candidate miRNAs could be used as potential biomarkers with good diagnostic capability, serum specimens from 84 PC patients and 84 HCs were utilized in the validation stage. As shown in Figure 2, these six candidate miRNAs maintained significant alterations in PC patients contrasted with HCs (P<0.05). The relative expression levels of miR-34b-3p and miR-556-5p were significantly increased in PC, whereas the remaining miRNAs, namely, miR-200c-3p, miR-361-5p, miR-369-3p and miR-485-3p represented opposite results. These results were consistent with that in training stage.
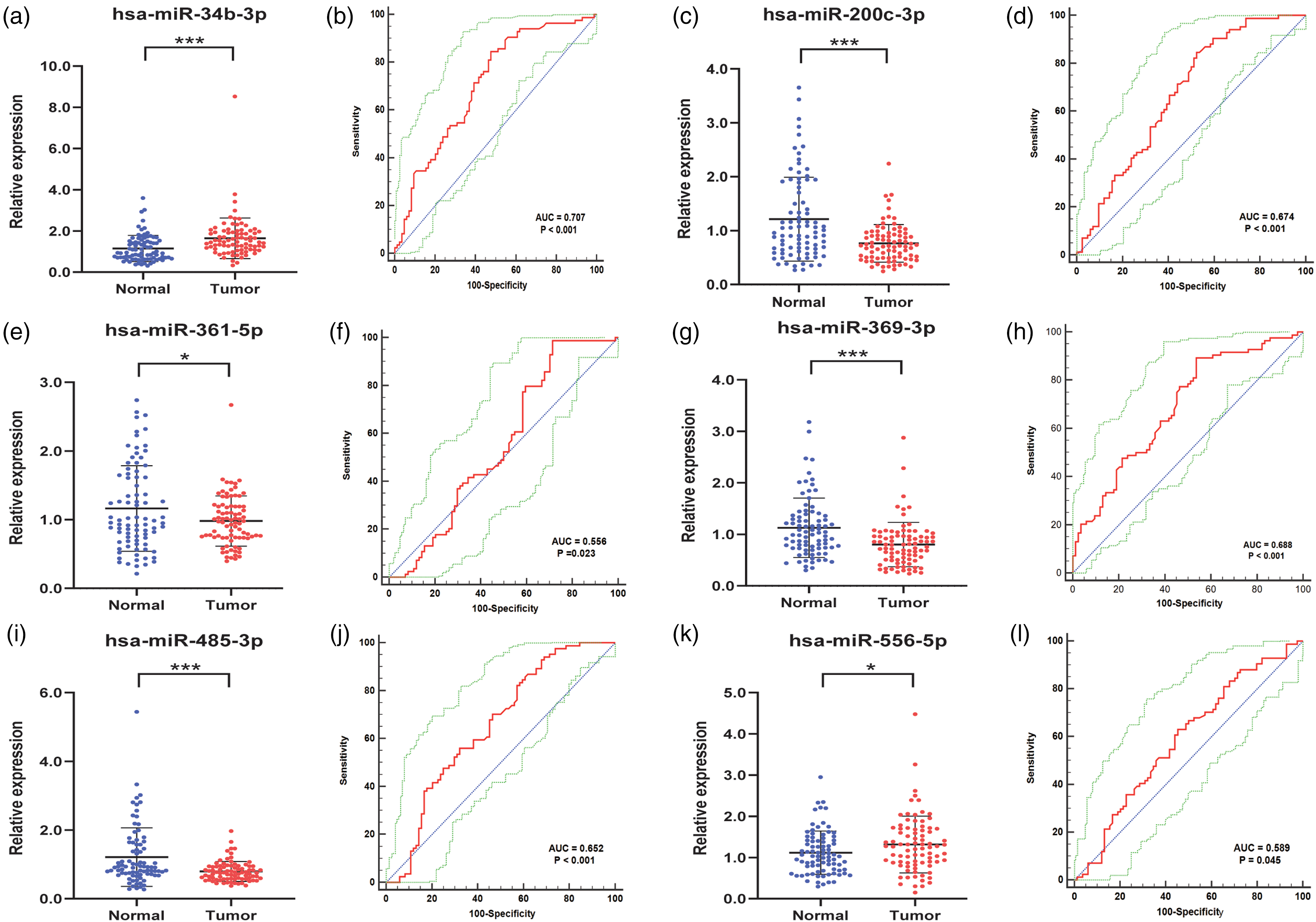
Serum expression levels and ROC curve analyses of six candidate miRNAs in validation stage with 84 PC patients and 84 HCs. The expressions of (a) miR-34b-3p and (k) miR-556-5p were significantly upregulated, while four other miRNAs, namely, (c) miR-200c-3p, (e) miR-361-5p, (g) miR-369-3p, and (i) miR-485-3p were significantly downregulated. ROC curve analyses of (b) miR-34b-3p, (d) miR-200c-3p, (f) miR-361-5p, (h) miR-369-3p, (j) miR-485-3p, and (l) miR-556-5p are shown in red curves. 95% ROC confidence intervals are shown in green curves. Diagonals are shown in blue lines. *P < 0.05, **P < 0.01, ***P < 0.001.
ROC curves analysis was conducted to assess the diagnostic capabilities of each candidate miRNA. Respectively, the AUCs were 0.707 (Figure 2(b)) for miR-34b-3p, 0.674 (Figure 2(d)) for miR-200c-3p, 0.556 (Figure 2(f)) for miR-361-5p, 0.688 (Figure 2(h)) for miR-369-3p, 0.652 (Figure 2(j)) for miR-485-3p, 0.589 (Figure 2(l)) for miR-556-5p. The 95% confidence intervals (CI) of each AUC are presented in Supplementary Table 2. In addition, calculation of the best cutoff values was performed by Youden index, and the optimum sensitivity and specificity of six critical miRNAs in diagnosing PC are shown in Supplementary Table 2.
Construction of the optimal miRNAs panel for PC
We combined several miRNAs to enhance diagnostic capability. With the stepwise logistic regression model, diagnostic panels were constructed by combining the expression data of six critical miRNAs. Ultimately, the final best combination comprised three miRNAs; namely, miR-34b-3p, miR-200c-3p, and miR-361-5p. The optimal logistic regression model was computed using the following formula, namely, logit (P) = −0.114 + 4.459 × miR-34b-3p - 4.678 × miR-200c-3p – 1.817 × miR-361-5p. The AUC of three-miRNA panel was 0.916 (95% CI 0.864, 0.953; sensitivity = 83.33%, specificity = 88.10%; Supplementary Table 2).
Bioinformatic analysis of the critical miRNAs
We utilized the MiRWalk 3.0 to predict target genes regulated by three candidate miRNAs (miR-34b-3p, miR-200c-3p, and miR-361-5p). A total of 700 genes targeted by two or three miRNAs were predicted (Figure 3(a)). The expression analysis of 74 target genes predicted by all miRNAs was performed through GEPIA with data from the GTEx and The Cancer Genome Atlas projects. 21 Among the 74 target genes, four were significantly aberrantly expressed in PC patients contrasted with HCs. With |log2Fold Change| cutoff >1.5, P < 0.01, FLRT2, KIAA1755, LDB3, and NTRK3 were recognized as potential genes targeted by the three-miRNA panel (Figure 3(b) to (e)).
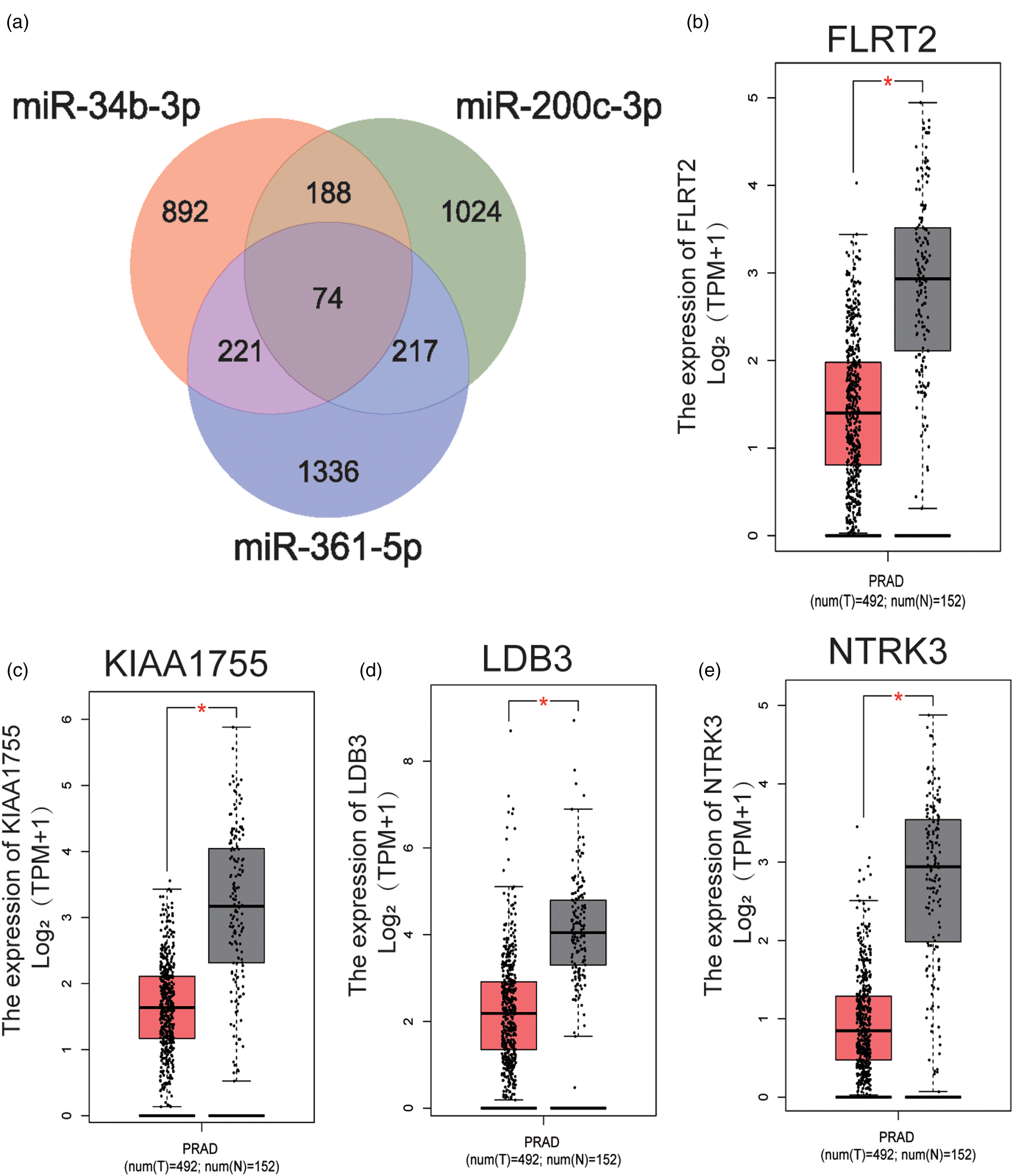
Target genes regulated by three critical miRNAs were predicted by miRWalk 3.0. (a) 700 genes targeted by two or three miRNAs were predicted. The expression analysis of 74 target genes predicted by all miRNAs in 492 PC patients and 152 HCs was performed through GEPIA with data from the GTEx and TCGA projects. (b) to (e) FLRT2, KIAA1755, LDB3 and NTRK3 were significantly aberrantly expressed with |log2Fold Change| > 1.5, P < 0.01, and were recognized as potential genes targeted by the three-miRNA panel. * P < 0.01;
To further investigate, GO functional annotation and KEGG pathway enrichment analysis of the total 700 target genes were performed by using Enrichr database. GO functional annotation comprises a biological process, a cellular component, and molecular function. The top five enriched GO items and KEGG items are shown in Figure 4, including regulation of transcription from RNA polymerase Ⅱpromoter (GO:0006357), peptidyl-serine phosphorylation (GO:0018105), and multicellular organism development (GO:0007275) in the biological process category; nucleus (GO:0005634), cytoplasm (GO:0005737), and cytosol (GO:0005829) in the cellular component category; protein binding (GO:0005515), protein serine/threonine kinase activity (GO:0004674), and protein kinase activity (GO:0004672) in the molecular function category; Wnt signaling pathway, Hippo signaling pathway, and adrenergic signaling in cardiomyocytes in KEGG.
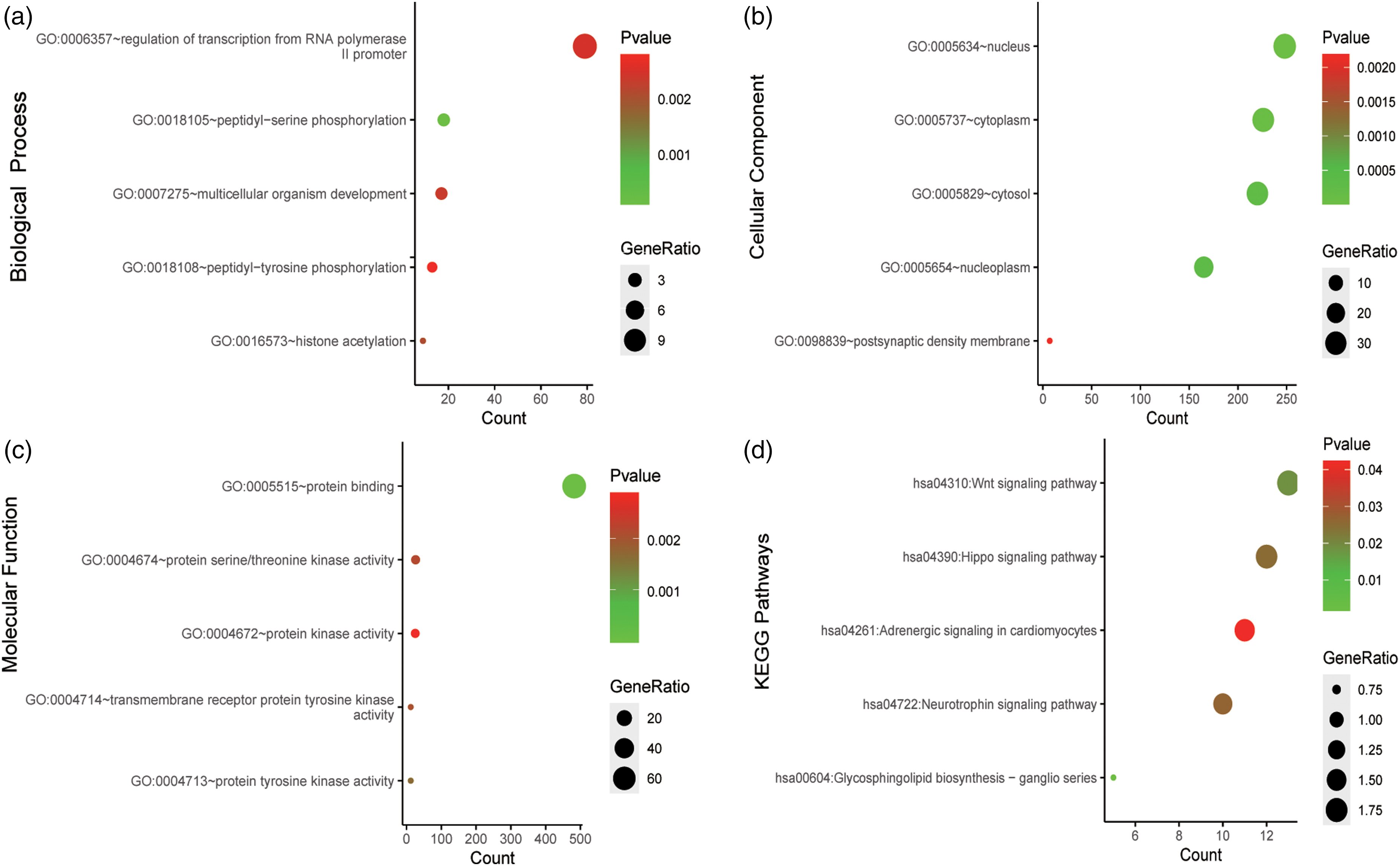
Go functional annotation and KEGG pathway enrichment analysis of the target genes of miR-34b-3p, miR-200c-3p, and miR-361-5p. The (a) biological process analysis, (b) cellular component analysis, (c) molecular function analysis, (d) KEGG pathway enrichment analysis are shown.
Discussion
PC is the second most common malignancy in males worldwide. Hence, it is critical to implement PC screening and early diagnosis aiming to enhance the rate of survival and reduce the burden of cancer. Detecting the blood concentration of total PSA is the most common method, while there is no recommended unanimous PSA threshold for the male population. 5 Another way is DRE, which is incompetent due to discomfort in men and its low detection rate. Biopsy and histological validation are the means of definitive diagnosis, which is invasive and contributes to bleeding and infection. 6 Numerous studies have revealed that serum miRNAs may function as non-invasive, innovative, and potential biomarkers for various cancers.8–10 In this study, three-stage research was conducted to ascertain the diagnostic capability of key miRNAs. Eventually, a three-miRNA panel composed of miR-34b-3p, miR-200c-3p, and miR-361-5p were constructed. This may perform as serum biomarker for PC screening with optimal diagnostic capabilities (AUC = 0.916, 95% CI 0.864, 0.953).
It has been established that miR-34b-3p is located on chromosome 11q23 and is a direct target gene of p53. 22 A study suggests that miR-34b could repress migration and invasion of PC cells through modulating the TGF-β signaling pathway. 23 Another study reports that miR-34b serves as a tumor suppressor in PC and has anti-proliferative and anti-invasive functions by impairing the AKT pathway and epithelial-to-mesenchymal transition (EMT) markers. 24 In this study, miR-34b-3p showed conspicuous up-regulation in PC patients compared to HCs in serum and could be a promising marker for PC screening (AUC = 0.707; 95% CI 0.632, 0.774; Figure 2(b)). However, miR-34b-3p is downregulated in PC cells and tissue samples.23,24 The reason why the expression levels of miR-34b-3p between tissue and serum was opposite requires more experimental research to explain.
miR-200c-3p, which belongs to miRNA-200 family with tumor suppressive functions, is recognized to inversely modulate EMT and metastasis in various cancers, including non-small cell lung cancer, 25 renal cell cancer, 16 and PC.26,27 Consistent with previous research, our results suggested that the expression of miR-200c-3p was downregulated in PC patients in contrast to HCs (AUC = 0.674; 95% CI 0.597, 0.744; Figure 2(d)).26,28 Xia et al. 28 reported that miR-200c-3p directly target type II-beta regulatory subunit of PKA (PRKAR2B), and eventually inhibits tumor growth and facilitates cell apoptosis. Patel et al. 29 unraveled that TBX2 represses expression of miR-200c-3p and drives neuroendocrine PC transdifferentiation through the miR-200c-3p/SOX2/N-MYC pathway. Zhang et al. 26 unraveled that miR-200c-3p was able to impair EMT via targeting ZEB2 while ZEB2 might suppress expression of miR-200c-3p simultaneously. Based on their study, this could be a promising treatment strategy of invasive and metastatic PC through targeting the miR-200c-3p/ZEB2 regulatory loop in the future.
Studies show that miR-361-5p is a considerable tumor suppressor in PC.17,30,31 One study suggested that miR-361-5p repressed the Sp1/PKM2 axis, hence suppressing PC cell proliferation, metabolism, and autophagy. 31 Liu et al. 17 found that miR-361-5p could impair the expression of the STAT6/Bcl-xL pathway, thereby triggering apoptosis and inhibiting cell proliferation. They also demonstrated that the expression level of miR-361-5p in castration-resistant prostate cancer (CRPC) was downregulated in contrast to androgen-dependent prostate cancer (ADPC), which means that miR-361-5p might restrain the development of ADPC to CRPC. Jiang et al. 30 found that exosomal HOXD-AS1, as a competing endogenous RNA (ceRNA) sponging miR-361-5p, facilitated FOXM1 expression, therefore promoting PC metastasis. Our research reveals that miR-361-5p(AUC = 0.556; 95% CI 0.477, 0.632; Figure 2(f)) was significantly downregulated, suggesting that miR-361-5p may play a critical role, such as suppressing proliferation or invasiveness in PC. To sum up, miR-361-5p may have crucial clinical value in preventing the development of PC.
FLRT2, KIAA1755, LDB3, and NTRK3, which were markedly aberrantly expressed between PC patients and HCs through GEPIA databases, were recognized as the potential genes targeted by the three-miRNA panel. FLRT2 belongs to FLRT family, and functions in cell adhesion and/or receptor signaling. Differential methylation of FLRT2 transcription start sites may affect FLRT2 and FGFR2 expression and interaction, subsequently regulating PC progression. 32 NTRK3 encodes TrkC that belongs to the neurotrophins receptor family. The inducible expression of TrkC in epithelial cells is involved in the abnormal growth of prostatic epithelium.33,34 Moreover, through the matching receptors, neurotrophins may contribute to the escape from the cell death mechanism occurring after androgen dissipation. 35 However, to the best of our knowledge, there are no studies unraveling the roles of KIAA1755 and LDB3 in PC. The functional involvement of KIAA1755 and LDB3 in PC needs to be further studied.
Currently, PSA is the most conventional biomarker for PC screening in clinical practice; however, its perfection is compromised by conditions like bacterial prostatitis, benign prostatic hyperplasia, and physical manipulation.5,36 Moreover, there is no recommended unanimous PSA cutoff point with synchronous favorable sensitivity and specificity. The American Cancer Society found that the sensitivity and specificity for PC screening were 21% and 91%, respectively, with a threshold of 4.0 ng/mL PSA; descending it to 3.0 ng/mL would lead to enhancing sensitivity to 32% and worsening specificity to 85%. 6 Similarly, the Prostate Cancer Prevention Trial elucidated that a PSA threshold of 1.1 and 4.1 ng/mL output sensitivities of 83.4% and 20.5%, and specificities of 38.9% and 93.8%, and AUC to distinguish PC patients versus health individuals were 0.678 (95% CI 0.666, 0.689). 37 In addition to PSA, other biomarkers are also gradually being developed. Many researchers suggested that 4Kscore Test (total, free and intact PSA, and human kallikrein 2) is of great value in detecting PC, especially high-grade PC.36,38 A meta-analysis suggested that the 4Kscore Test has reliable diagnostic capability with the AUC of 0.72 (sensitivity = 74%, specificity = 60%). 39 Moreover, prostate health index, calculated with the formula: [–2]proPSA/free PSA) × √PSA, was found to detect PC with the AUC of 0.76 (sensitivity = 89%, specificity = 34%). 39 Although the present study implied that the three-miRNA panel seems to perform better with the AUC of 0.916 (95% CI 0.864, 0.953; sensitivity = 83.33%, specificity = 88.10%), experiment with head-to-head comparisons is needed to verify this.
Although our results are remarkable, some limitations exist. First, the study specimen size is relatively small and all the participants are from Peking University Shenzhen Hospital. Although we aimed to determine potential miRNAs to discriminate PC patients and HCs; the value of the three-miRNA panel in different subgroups of patients with PC is unclear. Second, all the participants received no treatment before the collection of serum samples. Thus, whether the expression of each miRNA can be affected by treatment is obscure. Third, we did not analyze the relationship between the expression of serum miRNAs and the blood concentration of total PSA. Moreover, only 12 initial miRNAs were selected, but there were many aberrantly expressed miRNAs related to PC. These require further investigation.
Conclusion
We discovered a three-miRNA panel composed of miR-34b-3p, miR-200c-3p, and miR-361-5p. It may perform as a non-invasive, novel, and promising serum marker for PC screening and early diagnosis. More research is warranted to verify our results.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155231213660 - Supplemental material for The value of a three-microRNA panel in serum for prostate cancer screening
Supplemental material, sj-docx-1-jbm-10.1177_03936155231213660 for The value of a three-microRNA panel in serum for prostate cancer screening by Shengjie Lin, Chen Sun, Rongkang Li, Chong Lu, Xinji Li, Zhenyu Wen, Zhenjian Ge, Wenkang Chen, Yingqi Li, Hang Li and Yongqing Lai in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155231213660 - Supplemental material for The value of a three-microRNA panel in serum for prostate cancer screening
Supplemental material, sj-docx-2-jbm-10.1177_03936155231213660 for The value of a three-microRNA panel in serum for prostate cancer screening by Shengjie Lin, Chen Sun, Rongkang Li, Chong Lu, Xinji Li, Zhenyu Wen, Zhenjian Ge, Wenkang Chen, Yingqi Li, Hang Li and Yongqing Lai in The International Journal of Biological Markers
Footnotes
Author contributions
All authors devoted to the generalization and design of the study. Shengjie Lin and Chen Sun were responsible for investigating and storing the data and wrote the original manuscript. Rongkang Li and Chong Lu performed a formal analysis. Xinji Li, Zhenyu Wen, and Zhenjian Ge collected and visualized the serum specimens. Wenkang Chen and Yingqi Li assisted with the writing of the manuscript. Hang Li and Yongqing Lai were dedicated to the review, editing, and oversight work. All authors read and approved the final manuscript. Shengjie Lin and Chen Sun contributed equally to this work.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclose receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Development Fund Project of Shenzhen, Shenzhen High-level Hospital Construction Fund, Clinical Research Project of Peking University Shenzhen Hospital, (grant number JCYJ20180507183102747, LCYJ2017001, LCYJ2020002, LCYJ2020015, LCYJ2020020).
Ethical approval and consent to participate
The protocol of this study was approved by the ethics committee of Ethics Committee of Peking University Shenzhen Hospital (2019-043). All participants recruited in our study provided written informed consent.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
