Abstract
Background
N6-methyladenosine (m6A) methylation is known as the research hotspot for tumor epimodification, and its associated methyltransferase-like3 (METTL3) is significantly differentially expressed in gastric carcinoma, but its clinical value has not been summarized. This meta-analysis aimed to evaluate the prognostic significance of METTL3 in gastric carcinoma.
Material and methods
Databases, including PubMed, EMBASE (Ovid platform), Science Direct, Scopus, MEDLINE, Google Scholar, Web of Science, and Cochrane Library, were used to identify relevant eligible studies. The endpoints included overall survival, progression-free survival, recurrence-free survival, post-progression survival, and disease-free survival. Hazard ratios (HR) with 95% confidence intervals (CI) were used to correlate METTL3 expression with prognosis. Subgroup and sensitivity analyses were performed.
Results
Seven eligible studies involving 3034 gastric carcinoma patients were recruited for this meta-analysis. The analysis showed that high METTL3 expression was associated with significantly poorer overall survival (HR = 2.37, 95% CI 1.66–3.39, P < 0.01) and unfavorable disease-free survival (HR = 2.58, 95% CI 1.97–3.38, P < 0.01), as did unfavorable progression-free survival (HR = 1.48, 95% CI 1.19–1.84, P < 0.01)/recurrence-free survival (HR = 2.62, 95% CI 1.93–5.62, P < 0.01)/post-progression survival (HR = 1.53, 95% CI 1.22–1.91, P < 0.01). Subgroup analysis found that high METTL3 expression was associated with worse overall survival in patients with Chinese (HR = 2.21, 95% CI 1.48–3.29, P < 0.01), in studies with sample source from formalin-fixed, paraffin-embedded tissues (HR = 2.66, 95% CI 1.79–3.94, P < 0.01), and the reported directly from articles group (HR = 2.42, 95% CI 1.66–3.53, P < 0.01). The subgroup analysis that was performed based on sample size, detected method, and follow-up showed the same results.
Conclusions
High expression of METTL3 predicts poor prognosis in gastric carcinoma, indicating promise for METTL3 as a prognostic biomarker.
Introduction
Gastric carcinoma (GC) is the fifth most prevalent cancer and the third most deadly cancer in the world. GLOBOCAN statistics show that in 2020 there were 1.08 million new cases and 768,000 deaths due to GC worldwide. 1 In China, the average number of newly diagnosed GC patients is more than 400,000 cases per year, which is the third highest number of tumor-related diseases in terms of incidence and mortality; the 5-year survival rate of patients with advanced GC is only about 10%. 2 Current treatment of GC calls for the combination of surgery-based local treatment with targeted immune combination therapy, which has enabled an increasing number of patients with GC to control their disease. However, the prognosis of patients with cancer remains poor. Therefore, the identification of novel, precise biomarkers is important for the development of prognostic tools for cancer patients.
Methyltransferase-like 3 (METTL3) is a gene that is located on human chromosome 14. It encodes for a methyltransferase enzyme that plays a crucial role in regulating RNA methylation, particularly N6-methyladenosine (m6A) modification. 3 The m6A modification has been linked to several biological processes, including mRNA stability, 4 splicing, 5 and translation. 6
Recent studies have shown that METTL3 plays a crucial role in the development and progression of cancer. 7 Abnormal expression of METTL3 has been reported in various types of cancer, such as lung, 8 breast, 9 gastric, 10 and acute myeloid leukemia (AML). 11 In some cancers, high levels of METTL3 expression are associated with poor prognosis, 12 while in others, low levels of METTL3 expression are associated with worse outcomes. 13 However, whether its expression is associated with overall survival (OS) and disease-free survival (DFS) in GC patients remains uncertain. To improve overall understanding of the role of METTL3 in GC, to provide a reference for its future research as a biomarker and scientific basis for clinical practice, a meta-analysis was conducted to evaluate the relationship between METTL3 expression and the prognostic of patients with GC.
Materials and methods
This meta-analysis was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement. 14
Search strategy and literature inclusion criteria
To identify all literature related to the relationship between METTL3 and the prognosis of patients with GC we used PubMed, EMBASE (Ovid platform), Science Direct, Scopus, MEDLINE, Google Scholar, Web of Science, and the Cochrane Library, from the inception of the study to March 13, 2023. The search terms were the following: “Stomach Neoplasm” or “Gastric Neoplasm” or “Cancer of Stomach” or “Stomach Cancers” or “gastric cancer” and “METTL3” or “Methyltransferase-like 3” or “m6A Methyltransferase 3”. The references and relevant systematic reviews of the included studies were tracked.
The eligible studies included in this meta-analysis met the following criteria: patients were confirmed with gastric cancer through pathological or histological examination; the expression of METTL3 was explored in gastric cancer subjects; the relationship between METTL3 and OS and DFS was evaluated; there were sufficient data to estimate the hazard ratio (HR) and its 95% confidence interval (CI); and finally, the complete publication was written in English. Exclusion criteria included review articles, case reports, abstracts, editorials, letters, and meta-analyses. We also excluded articles that evaluated the molecular structure and function of METTL3, articles with insufficient data for analysis (even after contacting the study authors), and duplicate publications.
Data extraction
Two reviewers (ZCL and WQ) independently extracted relevant data from eligible studies using data tables designed for each study. Disagreements were resolved through discussion. Data retrieved from each publication included basic characteristics of each study, such as first author, publication year, country, sample size, sample source, detection method, and endpoint measures (OS, progression-free survival (PFS), recurrence-free survival (RFS), post-progression survival (PPS), DFS, and cutoff values). For OS, unless HRs were available through reading the publication, we recalculated HRs from published data using the number of patients at risk or the P value of the HRs. If this was not feasible, we used the HR digitizing software Engauge Digitizer 11.1 to extract and digitize survival data from Kaplan–Meier curves. 15 This involved importing clear and interpretable images from PDFs, plotting points, extracting data using Engauge Digitizer 11.1 software, and ultimately calculating HR estimates based on the obtained data.
Quality assessment
Quality assessment for each eligible study was performed by the same reviewers (ZCL and WQ) who extracted the data. They read and scored each publication independently according to the Quality in Prognosis Studies (QUIPS) tool. 16 The QUIPS tool is a reliable method for assessing the risk of bias (RoB) in studies of prognostic factors through six separate domains: (a) study participation; (b) study attrition; (c) prognostic factor measurement; (d) outcome measurement; (e) study confounding; and (f) statistical analysis and reporting. Studies were classified as low, moderate, or high RoB based on the QUIPS tool guidance for RoB judgments. Any disagreements were resolved through discussions.
Statistical analysis
HRs with 95% CIs were calculated for the association between METTL3 expression and the OS and DFS of GC patients. Heterogeneity in the HRs, OS, DFS, and DFS data was tested by using the Q statistic and I2 test. If the P value was more than 0.1 and (not or) I2 index less than 20%, all included studies lacked heterogeneity; thus, the Mantel–Haenszel method (fixed effect model) was used to merge the studies. Otherwise, the random effects model was adopted. Subgroup analyses were performed for different ethnicities, diagnoses, regions, cutoff values, control sources, and so on. Potential publication bias was also diagnosed and measured using funnel plots, and if significant asymmetry was revealed, the Begg's test and Egger's test were used for further validation. Statistical analyses were performed using STATA version 12.0 (StataCorp, College Station). Statistical significance for two-sided tests was set at P < 0.05.
Results
Literature search and study baseline characteristics
Through the search strategy described above, we conducted a literature search and initially retrieved 208 articles. A total of 52 articles remained after removing 156 duplicates. By further screening the titles and abstracts, we excluded 44 articles. After a full-text review, 1 article was excluded. Ultimately, we included 7 studies17–23 in this meta-analysis, which recruited a total of 3034 patients. The process of selecting studies is shown in Figure 1.
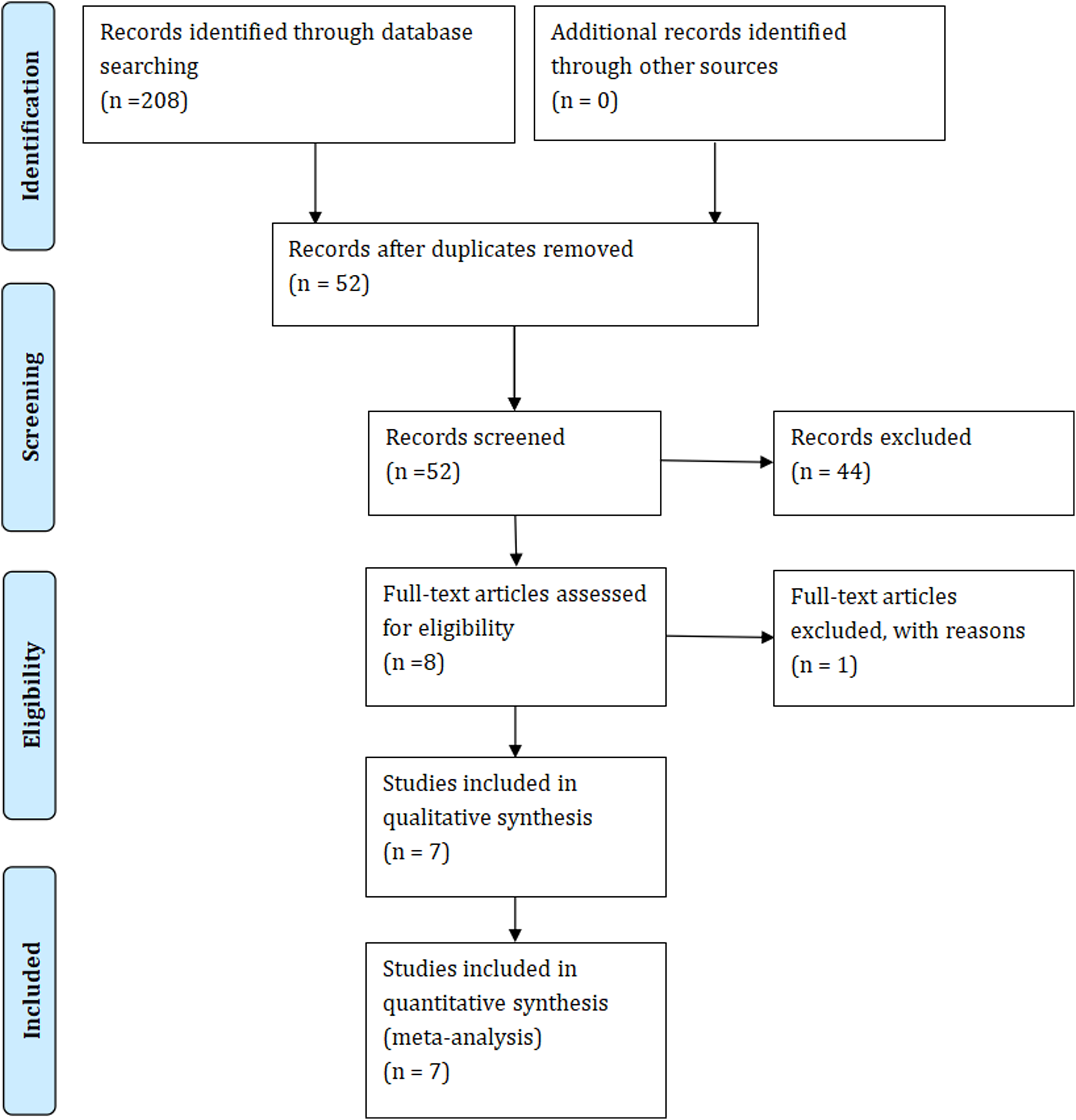
PRISMA flow diagram of the meta-analysis.
All 7 of these studies were prospective. The included studies were published from 2019 to 2022 and had a sample size of 57–875 cases. All 7 studies were from Asian countries, 4 of which were detected by reverse transcription-polymerase chain reaction (RT-PCR), and the other 4 were detected by immunohistochemistry. The patient characteristics and baseline data are presented in Table 1. The QUIPS risk of bias was low in 6 studies, moderate in 1, indicating high methodological quality of the included studies.
The characteristics of included studies.
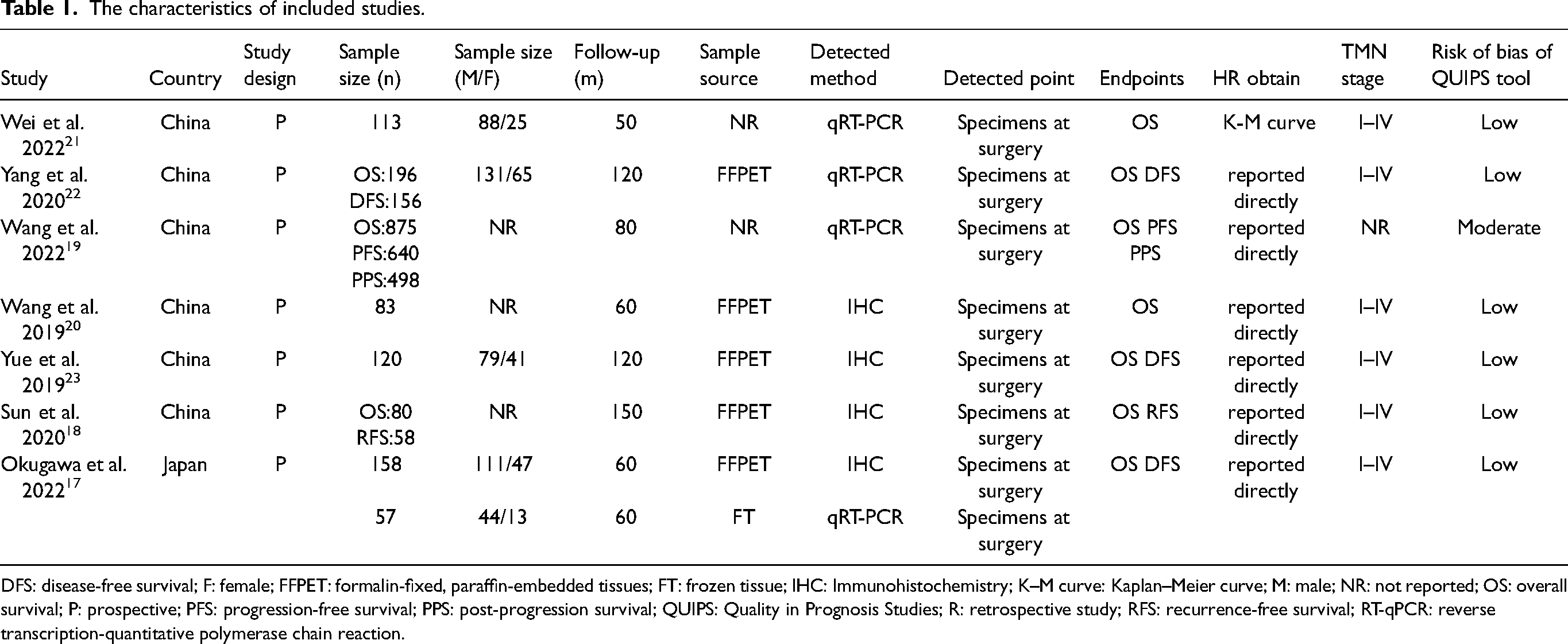
DFS: disease-free survival; F: female; FFPET: formalin-fixed, paraffin-embedded tissues; FT: frozen tissue; IHC: Immunohistochemistry; K–M curve: Kaplan–Meier curve; M: male; NR: not reported; OS: overall survival; P: prospective; PFS: progression-free survival; PPS: post-progression survival; QUIPS: Quality in Prognosis Studies; R: retrospective study; RFS: recurrence-free survival; RT-qPCR: reverse transcription-quantitative polymerase chain reaction.
High METTL3 expression and OS
Seven articles that involved a total of 1682 patients were used to explore the relationship between METTL3 expression and prognosis using OS. Since the I2 = 74.6% and P < 0.001, we used a random-effects model. For high METTL3 expression, the combined HR for OS was (HR = 2.37, 95% CI 1.66–3.39, P < 0.01), suggesting that high METTL3 expression is associated with significantly poorer prognosis in patients with GC. The forest plot is shown in Figure 2.
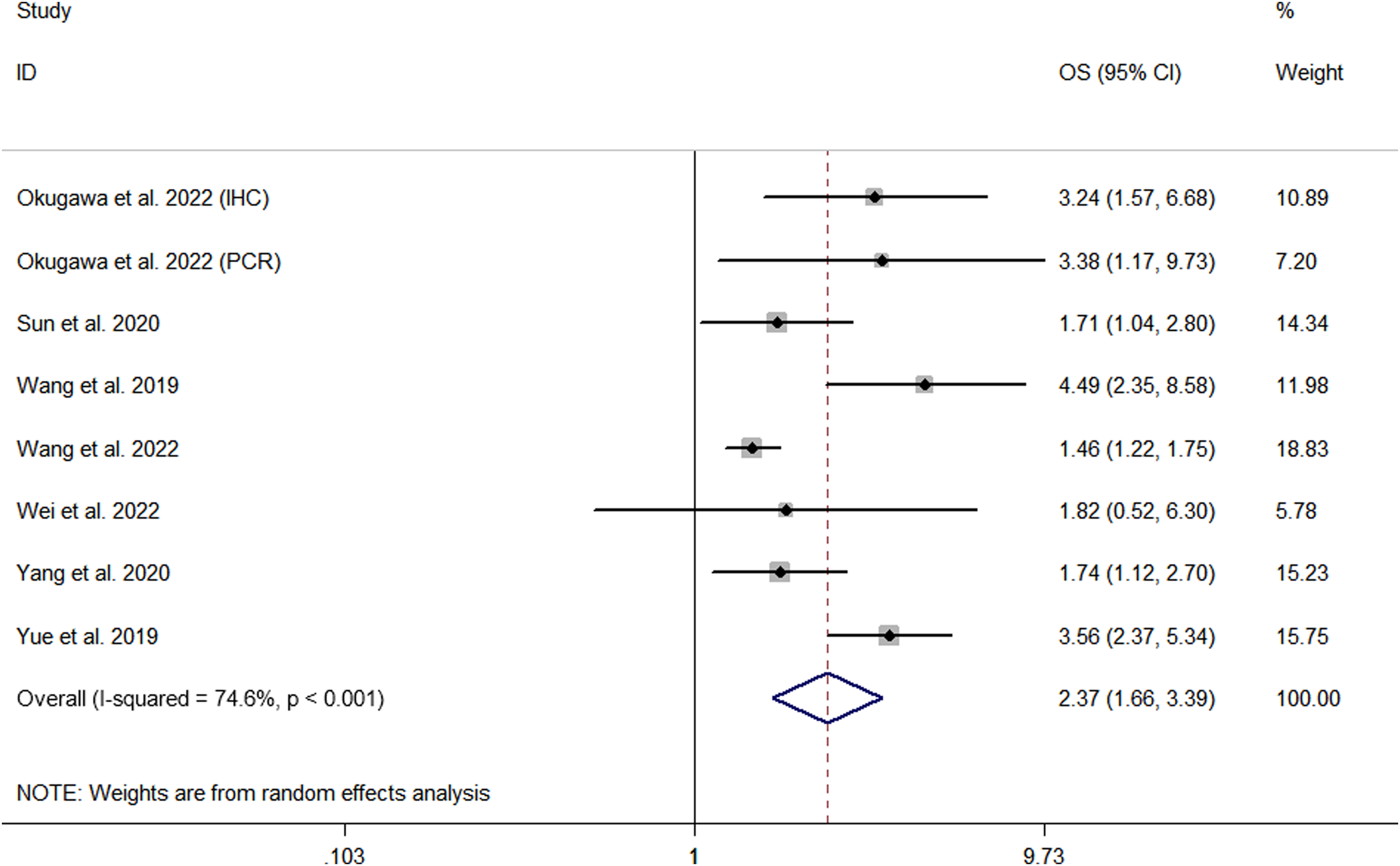
Forest plot of the relationship between high METTL3 expression and overall survival (OS).
High METTL3 expression and DFS
Four studies involving 491patients documented the relationship between METTL3 expression and prognosis using DFS. The I2 = 0.0% and P = 0.639, suggesting no significant heterogeneity among these studies. Therefore, a fixed-effects model was adopted. Meta-analysis showed that high METTL3 levels predicted unfavorable DFS (HR = 2.58, 95% CI 1.97–3.38, P < 0.01) in patients with GC (Figure 3).
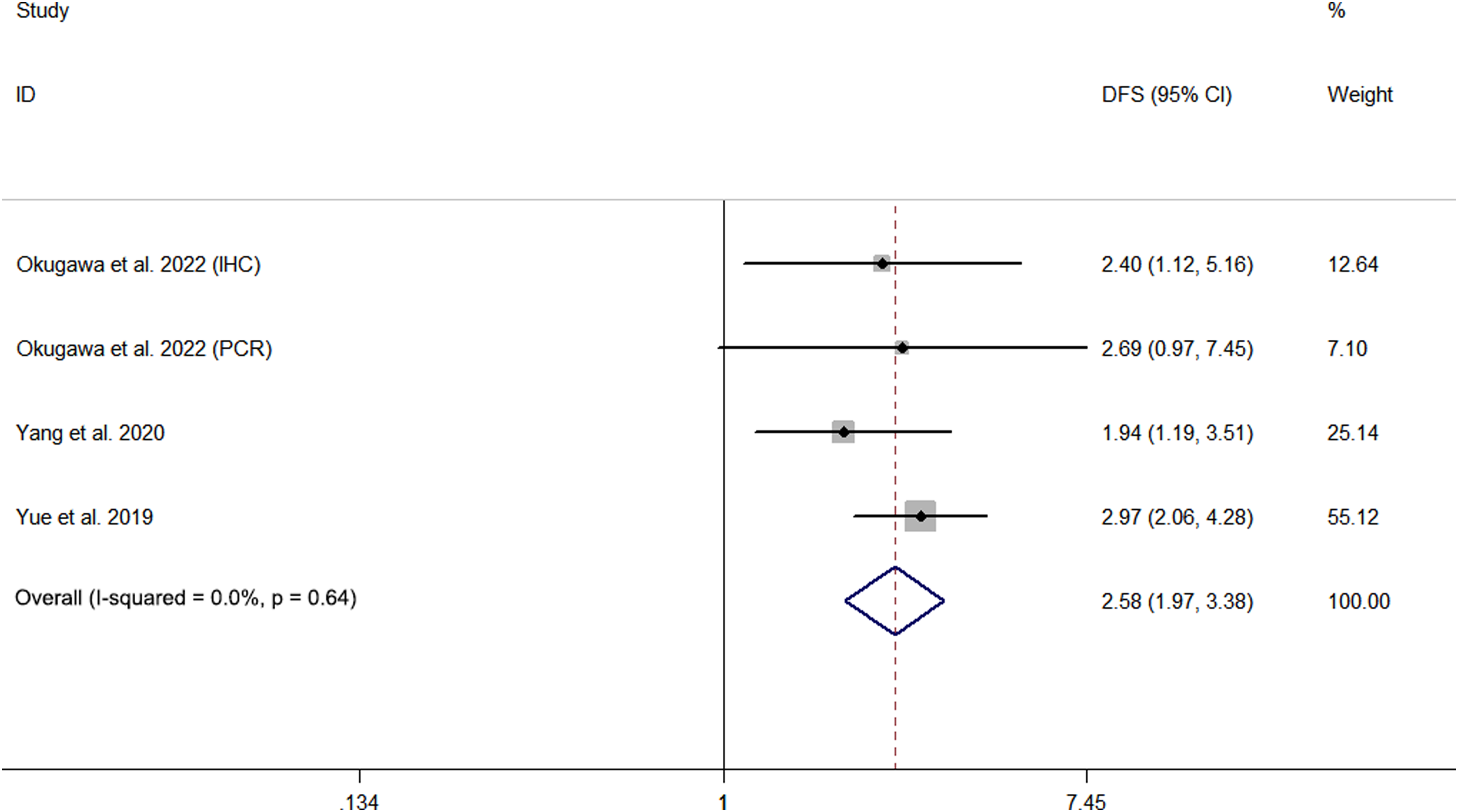
Forest plot of the relationship between high METTL3 expression and disease-free survival (DFS).
High METTL3 expression and PFS/RFS/PPS
Three studies involving 1196 patients were used to investigate the relationship between METTL3 expression and prognosis using progression-free survival/post-progression survival/recurrence-free survival (PFS/PPS/RFS) as measures. The HR values of PFS/PPS/RFS were (HR = 1.48, 95% CI 1.19–1.84, P < 0.01), (HR = 1.53, 95% CI 1.22–1.91, P < 0.01), and (HR = 2.62, 95% CI 1.93–5.62, P < 0.01), respectively. It is indicated that the high expression of METTL3 has a predictive effect on unfavorable PFS/PPS/RFS in GC patients.
Subgroup analysis
The results of the subgroup analysis (based on the ethnicity, sample source, sample size, detected method, follow-up, and HR obtain method) are shown in Table 2. The subgroup analysis, classified by HR, validated the association of high METTL3 levels with unfavorable OS. Subgroup analysis found that high METTL3 expression was associated with worse OS in patients with Chinese (HR = 2.21, 95% CI 1.48–3.29, P < 0.01). This was the case for the group whose HR values were directly reported by the article (HR = 2.42, 95% CI 1.66–3.53, P < 0.01). As for the subgroup analysis of the sample source, high expression of METTL3 in FFPET (HR = 2.66, 95% CI 1.79–3.94, P < 0.001) indicated poor OS. Additional classification by sample size confirmed that high METTL3 expression was an adverse prognostic factor in subgroups of studies with a sample size <100 (HR = 2.83, 95% CI 1.43–5.59, P = 0.01), and ≥100 (HR = 2.18, 95% CI 1.40–3.39, P = 0.008). Subgroup analysis performed based on detected method, follow-up showed the same results.
Subgroup analysis of association between METTL3 and OS for gastric carcinoma.
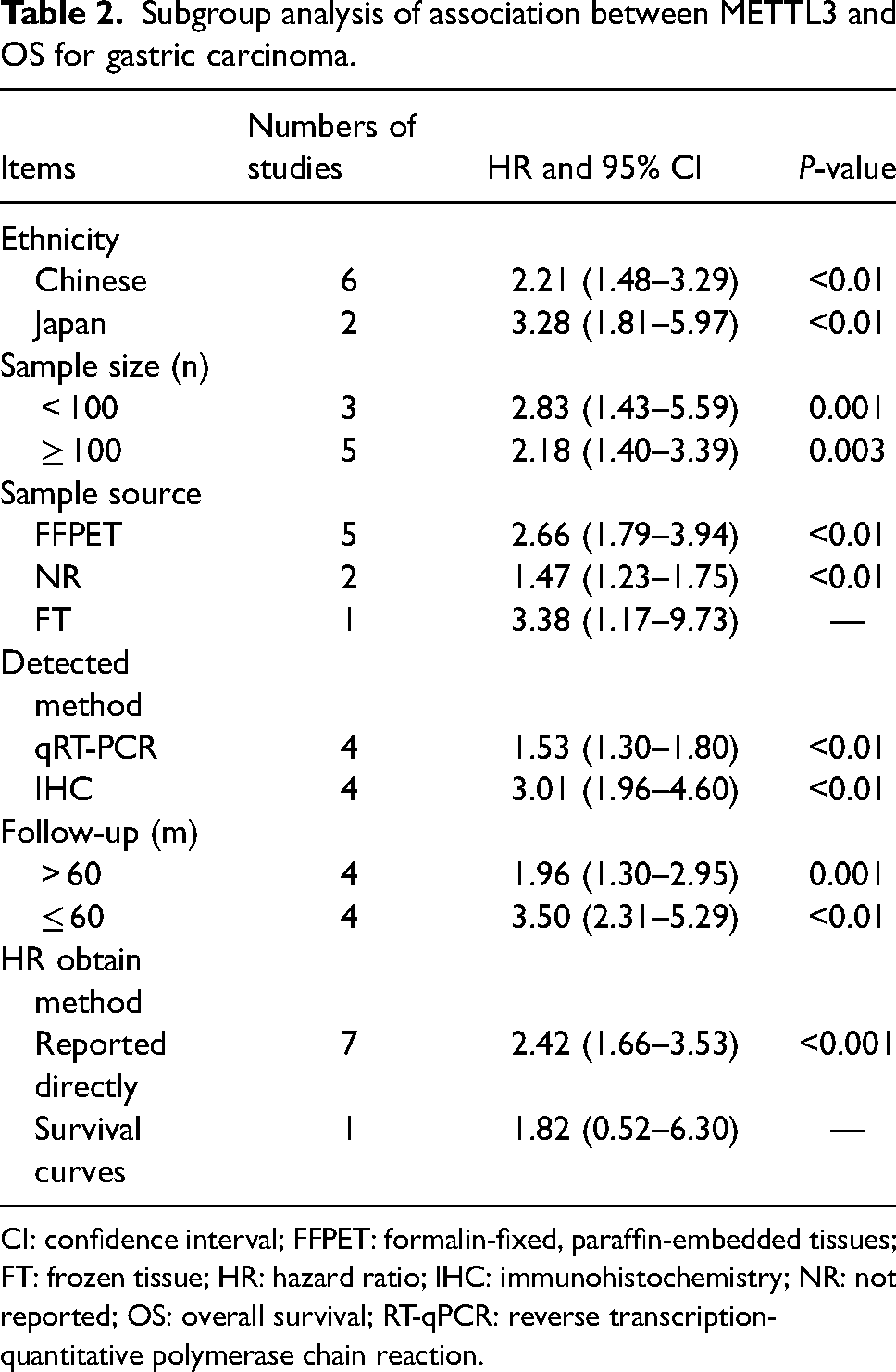
CI: confidence interval; FFPET: formalin-fixed, paraffin-embedded tissues; FT: frozen tissue; HR: hazard ratio; IHC: immunohistochemistry; NR: not reported; OS: overall survival; RT-qPCR: reverse transcription-quantitative polymerase chain reaction.
Sensitivity analysis and publication bias
To assess the reliability and stability of our results, we performed a sensitivity analysis by sequentially omitting any individual cohort analyses. The combined HR of OS was not affected, implying an increase in the confidence level (Supplementary Figure S1). Publication bias in the included studies was detected by drawing a funnel plot or by further performing Begg's and Egger's tests. For OS, the shape of the funnel plot showed asymmetry (Supplementary Figure S2), but the P-values of Begg's test and Egger's test were 0.902 and 0.073, respectively, indicating that there is no significant publication bias in our study.
Discussion
GC is a highly invasive and often fatal disease, and identifying prognostic markers can help identify which patients are at high risk of disease progression or recurrence, and which patients may benefit from more aggressive or personalized treatment approaches. Several potential prognostic markers for gastric cancer have been identified, including clinical factors such as tumor size, staging, and lymph node involvement, 24 as well as molecular markers such as HER2 overexpression, 25 KRAS mutations, 26 and microsatellite instability. 27 The main objective of this study was to explore the relationship between the expression levels of METTL3 and OS in GC patients and its potential as a prognostic marker.
METTL3 is an RNA methyltransferase that plays a key role in regulating gene expression through the methylation of RNA molecules. Recently, there has been growing evidence suggesting that METTL3 is an oncogenic factor in various types of cancer. 28 One proposed mechanism for the oncogenic role of METTL3 is through its ability to promote the translation of oncogenes. For example, Yang et al. 22 found that METTL3 promotes the translation of the oncogene MYC by enhancing the m6A modification of its mRNA, which leads to increased protein expression of MYC and subsequent cell proliferation. In addition, METTL3 has also been shown to regulate the splicing of pre-mRNA, which can impact the expression of oncogenes. Tang et al. 29 demonstrated that METTL3 promotes the splicing of the oncogene FLI1, resulting in increased FLI1 expression and enhanced tumorigenesis. Furthermore, METTL3 has been shown to regulate the stability of mRNA molecules, which can impact the expression of various genes, including oncogenes. Yang et al. 30 found that METTL3 promotes the stability of the oncogene HIF1α mRNA by enhancing its m6A modification, leading to increased protein expression of HIF1α and subsequent tumor growth. Overall, the oncogenic role of METTL3 in cancer is likely multifaceted, involving its ability to promote the translation of oncogenes, regulate splicing of pre-mRNA, and modulate mRNA stability.
Increased expression of METTL3 is associated with the growth, invasion, and metastasis of GC. The exact mechanism by which METTL3 promotes the development of GC is not fully understood. However, several possible pathways have been proposed by research: 1. mRNA stability and translation regulation: METTL3-mediated m6A modification of specific mRNA may affect its stability and translation, resulting in changes in the expression of genes related to the pathogenesis of gastric cancer. 31 2. Epithelial-mesenchymal transition (EMT): EMT is a process by which epithelial cells acquire mesenchymal characteristics, making them more migratory and invasive. METTL3 has been shown to regulate the EMT process by altering the expression of EMT-related genes. 23 3. Wnt/PI3K-Akt pathway: The Wnt/PI3K-Akt pathway is a key signaling pathway involved in regulating cell proliferation and differentiation. METTL3 has been shown to activate this pathway by promoting the expression of Wnt ligands and downstream target genes. 32 4. Immune evasion: METTL3-mediated RNA modification may also play a role in immune evasion by regulating exons. 33
The present meta-analysis involved seven studies and revealed that a high expression level of METTL3 is associated with a poor OS in GC patients. This study has demonstrated that high METTL3 expression was associated with significantly poorer OS (HR = 2.37, 95% CI 1.66–3.39, P < 0.01) and unfavorable DFS (HR = 2.58, 95% CI 1.97–3.38, P < 0.01), as did unfavorable PFS (HR = 1.48, 95% CI 1.19–1.84, P < 0.01)/RFS (HR = 2.62, 95% CI 1.93–5.62, P < 0.01)/PPS (HR = 1.53, 95% CI 1.22–1.91, P < 0.01).
In addition, our subgroup analysis showed that race, sample source, sample size, detection method, follow-up status, and HR acquisition method did not significantly alter the pooled HR results for OS. Certainly, the quality of studies included may affect the results of a meta-analysis. In our study, we used the QUIPS tool to evaluate their quality. The result was that both the meta-analysis and subgroup analysis did not show that the quality score of included studies affected the heterogeneity between high and low expression levels of METTL3 in GC patients. We assessed publication bias using funnel plots and found that the shape of the funnel plots showed asymmetry, but the subsequent Begg's and Egger's tests did not demonstrate statistical significance; this suggested that there was no significant publication bias between METTL3 expression and endpoint indicators. Although we derived the prognostic significance of METTL3 in GC patients, further research is needed to elucidate its role in GC. Additionally, larger case-control studies are needed to confirm METTL3 as a clinical prognostic marker.
Limitations exist in this meta-analysis. First, one study did not provide survival data directly, and the HR and 95% CI calculated from the survival curve may lead to some errors. Second, since all included studies were from Asia, the data from this meta-analysis is not applicable to a global population. Third, due to the relatively small number and sample size of the included studies, we were unable to summarize the results by tumor classification. Therefore, there is an urgent need for larger, multicenter, high-quality studies to confirm our findings.
Conclusions
Based on existing evidence, our study indicates that upregulation of METTL3 is significantly associated with poor OS, DFS, and PFS/RFS/PPS in GC. Through a comprehensive review of previous studies, we found that METTL3 can interact with multiple tumor-signaling pathways and activate various targets. In summary, METTL3 can serve as a predictive and prognostic biomarker for GC. Larger scale prospective and standardized investigations should be conducted in the future to validate our results
Supplemental Material
sj-doc-1-jbm-10.1177_03936155231184908 - Supplemental material for Prognostic significance of N6-methyladenosine-modified related chemotransferase METTL3 in gastric carcinoma: Evidence from meta-analysis
Supplemental material, sj-doc-1-jbm-10.1177_03936155231184908 for Prognostic significance of N6-methyladenosine-modified related chemotransferase METTL3 in gastric carcinoma: Evidence from meta-analysis by Chenglou Zhu, Qiong Wu, Yan Xu, Jichun Ma, Yongli Hu, Junhong Wang, Zhenhua Gao and Mingxu Da in The International Journal of Biological Markers
Footnotes
Author’s contributions
ZCL, GZH, and DMX made substantial contributions to the conception and design for this work. ZCL and WQ collected all the data. ZCL, HYL, WJH, and MJC were the major contributors in writing the manuscript. XY and WQ performed the critical revision for this manuscript. ZCL, QW, and YX contributed equally to this work. All authors contributed to the article and approved the submitted version.
Availability of data and materials
Some or all data, models, or code generated or used during the study are available from the corresponding author by request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Not applicable. This article did not require an ethical board approval because the study data were downloaded from the open database. This is a secondary analysis of data from previous studies.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Gansu Province, National Natural Science Foundation of China, (grant number 21CX6FD163, 22JR11RA252, 82160588)
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
