Abstract
Background
A whole-exome or targeted cancer genes panel by next-generation sequencing has been used widely in assisting individualized treatment decisions. Currently, multiple algorithms are developed to estimate DNA copy numbers based on sequencing data, which makes a comprehensive global glance at chromosomal integrity possible. We aim to classify gastric cancers based on chromosomal integrity to guide personalized therapy.
Methods
We investigated copy number variations (CNV) across the entire genome of 124 gastric carcinomas via exome or targeted sequencing. Chromosomal integrity was classified as chromosomal stability (CS), chromosomal instability (CIN) and intermediate state (CIN/CS) based on CNV results. Chromosomal integrity was correlated to molecular features and clinical characteristics.
Results
According the states of chromosomal integrity, gastric carcinomas can be stratified into two cohorts: CS and CIN. Our results showed a significant relationship between CIN status and TP53 mutation, but not RB1, phosphatase and tensin homolog (PTEN), or other reported DNA damage repair genes. The mutation frequency of the TP53 gene had great relevance. Our study initially revealed clinical significance of chromosomal integrity that CIN patients were prone to HER2-positive and mucinous adenocarcinoma, while CS patients were a diffuse subtype and poorly differentiated but had longer overall survival.
Conclusions
We classified gastric carcinomas into two states of chromosomal integrity with clinical implications. The dichotomy is applicable to clinical transformation. We proposed that classifying gastric cancers based on chromosomal integrity would enable us to achieve personalized therapy for patients and may be beneficial to patient stratification in future clinical trials.
Introduction
Cancers of the stomach pose an overwhelming threat to human health. Gastric cancer is the fifth most common tumor and the third leading cause of cancer death worldwide. 1 Chinese cancer statistics show 679,000 new cancer diagnoses and 498,000 cancer deaths in 2015. Both newly diagnosed cancers and deaths were the second most common type of cancer among men and the total population. 2 Although many chemotherapies, targeted therapies, and immunotherapies have been approved for gastric cancer, further studies on stratifying gastric cancer and corresponding effective intervention strategies are urgently needed.
High-throughput molecular platforms ensure a genetic perspective to investigate cancer and have been widely used to guide precision medicine of clinical practice in lung cancer and other tumors.3,4 The gastric adenocarcinomas program of The Cancer Genome Atlas (TCGA) project developed a robust subgrouping model that classified cancers into four subtypes: EB+, microsatellite instability-high (MSI-H), genomic stability, and genomic instability tumors. 5 Another study also applied next-generation sequencing combined with other platforms to define a novel classification: the mesenchymal-like type, microsatellite-unstable tumors, TP53-active, and TP53-inactive types. 6 As each subtype described a comprehensive molecular landscape, how those molecular characteristics impacted on clinical implications—especially on tumor progression and response to therapy—had little consensus, which to some extent hampers clinical decisions guided by molecular subtypes.
In this study, we focused on investigating chromosomal integrity in the cancer genome, since widespread chromosomal instability (CIN) represents a staggerer for tumor cells, but its clinical relevance still needs to be better clarified. Whole-genome duplication and copy number variation (CNV) at the chromosomal, sub-chromosomal, and gene levels are very common in various types of cancer.7–10 Whole-genome sequencing of tumor cells identified special genetic material, circular extrachromosomal DNA (ceDNA), which represents the DNA-repaired product of chromosomal chromothripsis.11,12 The ceDNA taking along high oncogene copy number further enhances chromosomal complexity. In clinical medicine, CIN was usually associated with poor prognosis by means of triggering cytotoxic treatment resistance. 13 CIN can predict the transformation and aggressiveness of esophageal cancer earlier than current diagnostic guidelines, 14 and also contributes to ascertain a higher degree of malignant phenotype in multiple cancer types. 15 However, it has also been reported that CIN increased the sensitivity to genotoxicity of chemotherapy or radiotherapy. 16 A pan-cancer analysis revealed that a higher level of chromosome CNV was correlated with worse response of immunotherapy due to lower neoantigen amounts and restrained neoantigen presentation. 17
Gene mutations and copy number changes represent two different oncogenic processes, where chromosomal integrity is an important marker for distinguishing one from the other. 18 Our study aims to delineate the chromosomal integrity profiles of primary gastric cancers and their clinical implications. We will show the overview of CNV across the entire genome at single-base resolution using the Sequenza algorithm 19 and investigate how CIN shapes tumor pathological manifestations and responses to therapies.
Methods
Patients and tissue specimens
Fresh specimens including gastric carcinoma and adjacent normal tissue from newly diagnosed cases were subjected to whole-exome sequencing in the first phase of the study. Subsequently, we collected samples from 112 patients with newly diagnosed untreated gastric cancer. (There was overlap with samples previously published. 20 ) Tumor samples and corresponding peripheral blood were used for targeted sequencing with a panel of 612 cancer-associated genes (the detailed gene lists are provided in Table S2). All samples were collected from the Department of Gastrointestinal Surgery in Changhai Hospital (Shanghai, China) and confirmed by two pathologists. Informed consent was obtained from all patients for the study. The primary observational index included overall survival (OS), which was defined as the time from diagnosis until death from any cause or the most recent follow-up (final follow-up, May 30, 2021). Median of OS along with their 95% CI and Kaplan–Meier survival statistics were calculated using log-rank tests. For comparisons between chromosomal stability (CS) and CIN, the hazard ratio (HR) and 95% confidence interval (CI) were estimated using the Cox proportional hazards regression model. Detailed clinicopathological characteristics of 112 patients are listed in Table 1 and Table S1.
Clinical characteristics of the 112 gastric cancer patients.
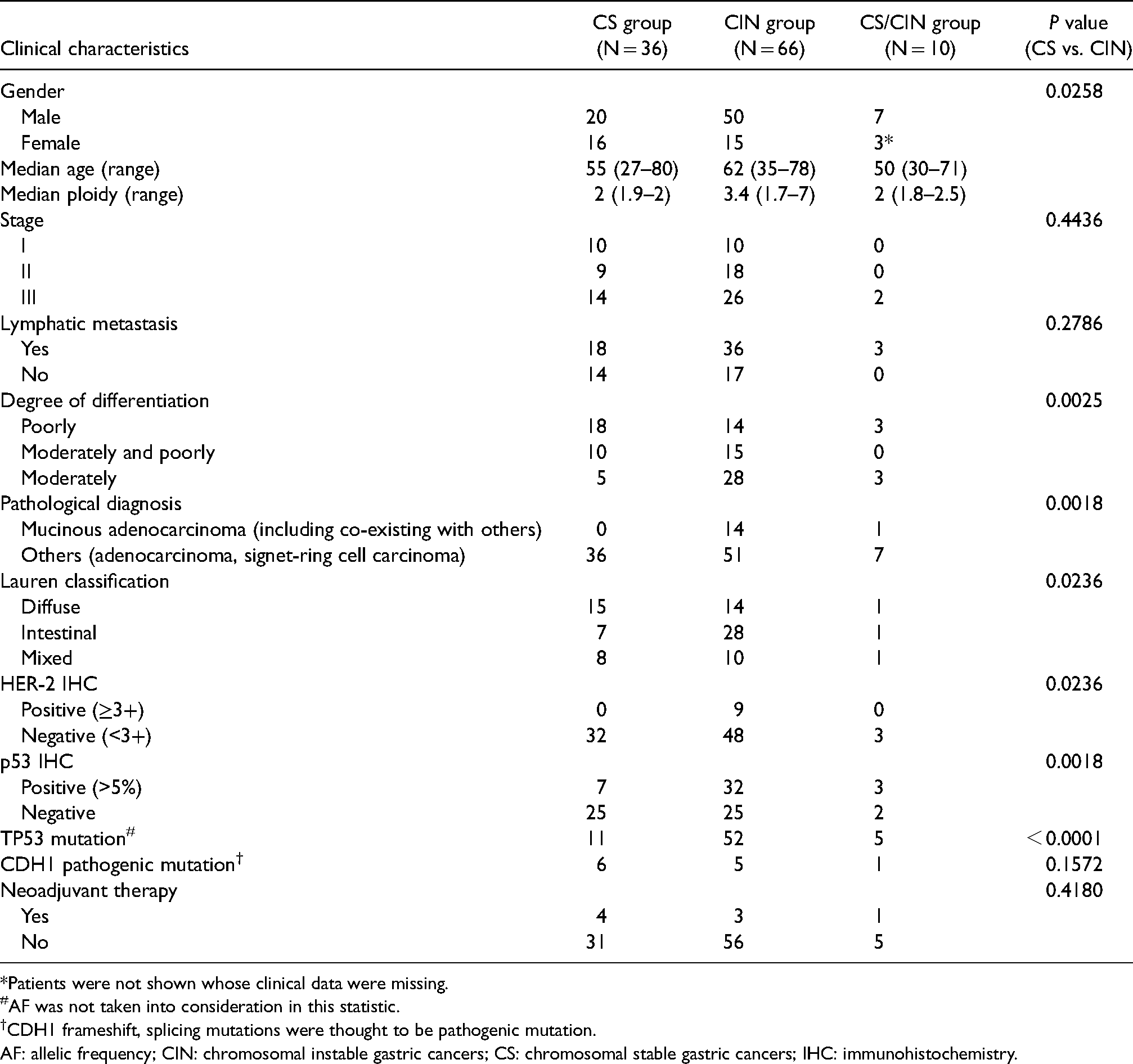
*Patients were not shown whose clinical data were missing.
#AF was not taken into consideration in this statistic.
†CDH1 frameshift, splicing mutations were thought to be pathogenic mutation.
AF: allelic frequency; CIN: chromosomal instable gastric cancers; CS: chromosomal stable gastric cancers; IHC: immunohistochemistry.
DNA-sequencing methods and variant calling
Genome DNA were extracted and subjected to exome or targeted sequencing, captured respectively by Agilent SureSelect Human All Exon V6 Enrichment Kit (Agilent, Santa Clara, CA, USA) or Agilent SureSelect XT custom 2.9 M probes, and sequenced by Illumina HiSeq platform (Illumina, San Diego, CA, USA). TrimGalore 0.6.6. was employed to pre-process the raw sequence data. Alignment utilized the BWA-MEM algorithm based on the hg38 reference genome. Marking duplicates and base recalibration were conducted following the Genome Analysis Toolkit (GATK4) best practices. Somatic mutations and short insertions/deletions were called and post-filtered using MuTech2, annotated with GATK Funcotator. Somatic copy number alterations were identified by the CNVkit comparing sequence coverage of targeted regions in the tumor relative to tumor-adjacent tissues. Tumor mutation burden (TMB) values were calculated as the ratio of mutations per megabase of the panel.
Chromosomal integrity analysis
R package Sequenza 3.0.0 was used for quantifying ploidy and copy number alterations. Chromosomal integrity was classified as CS, CIN, and intermediate state (CIN/CS) based on Sequenza results with default parameters, modified with allelic imbalance and scarHRD (R package scarHRD 0.1.1) results. CS represents >95% available segments along the genome having two copies of the total copy number; CIN represents >25% available segments that do not have two copies; and otherwise CS/CIN. Sex chromosomes were excluded from the analysis. Visualization of copy number alterations was performed via the copy number 1.30.0 R package. Mutational analysis of DNA damage repair (DDR)- and cell cycle regulation (CCR) custom pathways-related genes (Table S3) was performed by maftools 2.6.5 R packages.
Immunohistochemical assay
Sample fixation for immunohistochemical assay was performed with 10% neutral buffered formalin and the sections (3 microns in thickness) were cut from paraffin embedded tissue. The primary antibodies targeting ERBB2 proteins (Clone: SP3, zsbio, Beijing, China) and TP53 proteins (Clone: DO-7, zsbio, Beijing, China) were used for immunostaining.
Statistical analysis
In correlation analysis, we calculated P-values via Fisher's exact test or the Chi-square test using GraphPad Prism 9.0 and calculated the correlation coefficient (Cramer's V) between chromosomal integrity and various clinical characteristics using SPSS 20.0. Because both of these variables are categorical with two or more possible values per variable, we chose Cramer's V value to indicate the strength of the association. Cramer's V ranges from 0 to 1: 0 indicates no relationship and 1 indicates complete correlation. The P-value was used to evaluate the authenticity of the association. A P-value less than or equal to 0.05 was considered statistically significant.
Results
Two states of chromosomal integrity in gastric cancer
We performed whole-exome sequencing (WES) for gastric carcinoma tissue and adjacent normal tissue. When investigating CNVs across the entire genome, we observed two different states of chromosomal integrity of gastric cancer: CS and CIN (Figure 1(a) and (b)). CS preserved intact diploid genome, while CIN manifested aneuploid genome characterized by widespread breakpoints inside chromosomes and the consequent CNVs, including deletions and amplifications. We described the chromosomal integrity from the angle of DNA double-strand break (DSB) in which CS means little DSBs and CIN means numerous DSBs. In the pilot project of 12 cancers, 5 were divided into the CS group, 6 into the CIN group, and the remaining 1 (sample 6) represented a transient state of chromosomal integrity (Figure 1(d)). Our results showed consistency with previous findings that somatic copy number alterations were changed at significant frequency across multiple cancer types. 8
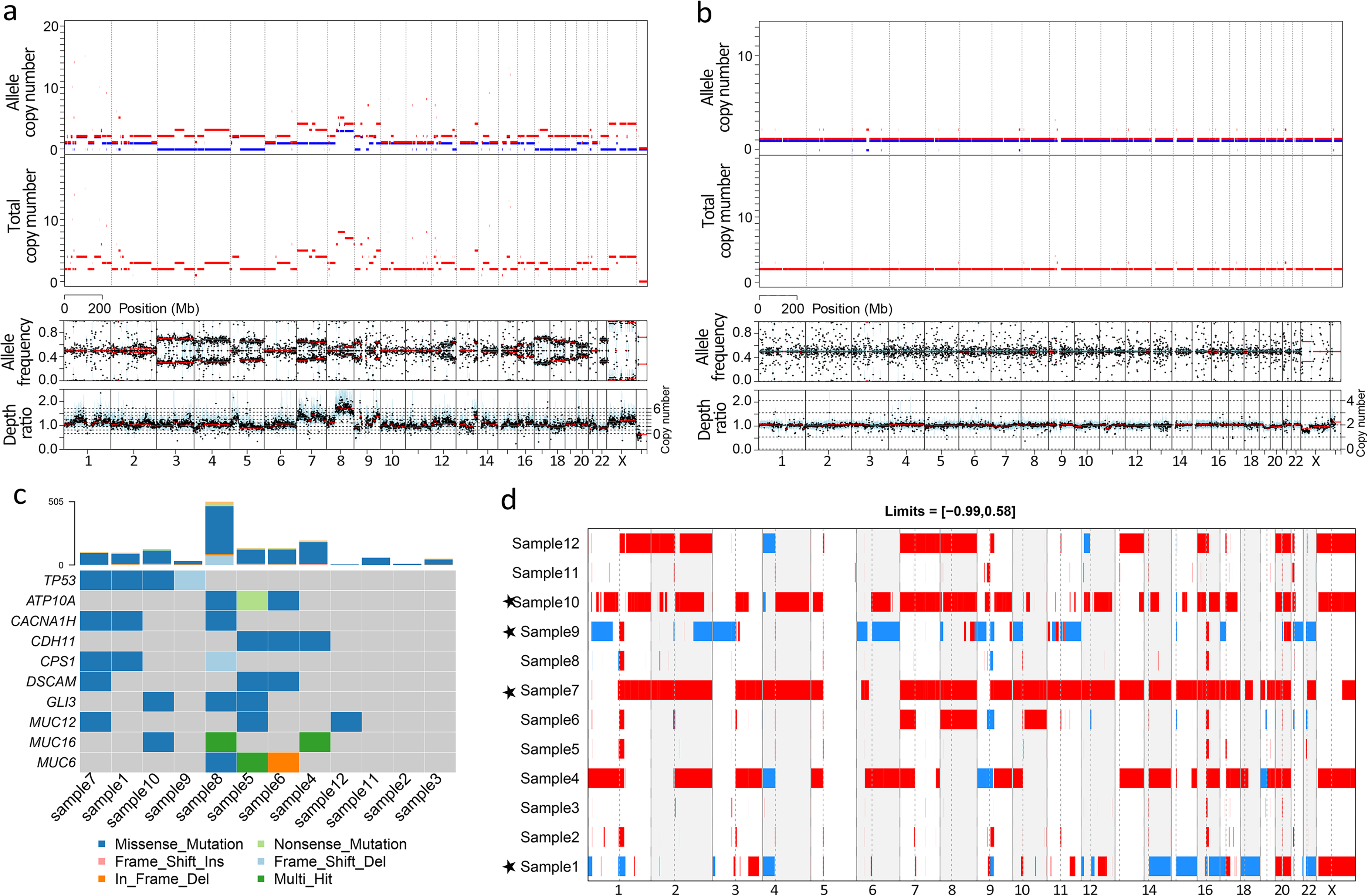
Two states of chromosomal integrity in gastric cancer identified by WES. (a) and (b) Representative genomes for chromosomal stability gastric cancers and chromosomal instability gastric cancers were selected to show copy numbers in sample 10 and sample 11 patients, respectively, and were analyzed by the Sequenza R algorithm package. (c) Mutational landscape of the 12 samples. The top bar plot shows the somatic mutation count for each sample. The bottom heat map shows the top 10 mutated genes. (d) Overview of copy number variation in the 12 samples. The Y-coordinate indicates samples and the X-coordinate indicates chromosomes. The colors red and blue indicate gains and losses, respectively. Gains and losses are defined by the estimated copy number (log2CN −1) above 0.58 or below the −0.99 threshold. The star (★) shows the identified TP53 mutation with ≥10% allele frequency.
Subsequently, we enlarged the scale of the sample size to confirm the finding. A total of 112 gastric cancer samples with stage IA–IIIC were collected, most of which were treatment-naïve (Table 1; Table S1). Instead of WES, targeted sequencing covering 2.9M genomic regions was employed to investigate chromosomal integrity. Our results showed that regions based on a panel of 612 genes could be an approximate reflection on genome-wide CNVs, which can be used to distinguish the two states of chromosomal integrity (Figure S1(a) and (b)). More than half of the patients (59%, 66/112) were the CIN state; CS only accounted for around 32% and CIN/CS for 9% (Figure S1(c)). The ploidies in CIN were highly variable while CS maintained an almost constant diploid (Figure S1(d)). The proportion was unexpected as all of those patients were locally or regionally primary gastric cancer, in contrast to the reported 37% whole-genome duplications in TCGA samples. 21 Neoadjuvant chemotherapy was performed for eight patients with different regimens. Given that four belonged to the CS group and three to the CIN group, it seemed that preoperative treatment did not affect genome integrity and chromosomal integrity (P = 0.418). On the whole, patients with gastric cancer in our study were stratified into two cohorts based on the states of chromosomal integrity.
TP53 mutation contributes to CIN
We next explored what was driving the transformations of chromosomal integrity starting from the diploid genome to chromosome breakage, to the gain or loss of some fragments, to the high disorder of the aneuploid genome. We investigated whether there was any association between genetic mutations and chromosomal instability. Indeed, we identified the most frequent mutations in the genome integrity pathway, including TP53 and ataxia telangiectasia mutated (ATM) (Figure 2(a)). Additionally, considering that the genes involved in the DNA damage response pathway were frequently mutated in cancer and homologous recombination deficiency could lead to DNA double-strand breaks, we speculated patients in the CIN group would harbor genetic mutations associated with DDR. However, we did not find the causal correlation between those gene variations and the states of chromosomal integrity: neither in well-characterized homologous recombination repair genes like BRCA1/2, PARP1 nor in non-homologous end-joining genes like MRE11/NBS1, Ku70/Ku80 (Figure 2(a) and (b); Table S3).
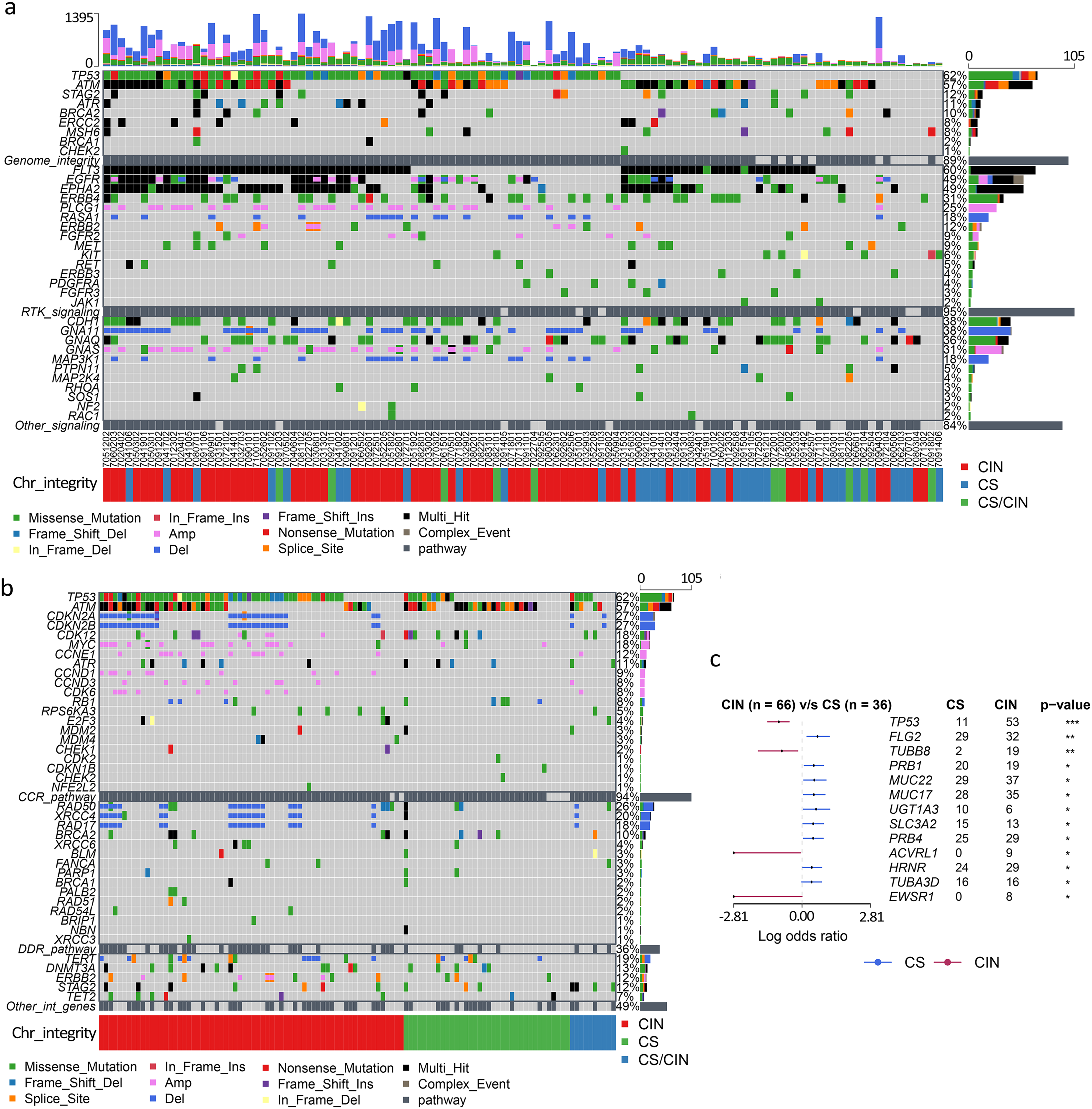

Significantly mutated genes in gastric cancer with different chromosomal integrity. (a) The top three mutated pathways in the 112 gastric cancer samples identified by targeted sequencing. The top panel shows the number of somatic variations in each sample. The color represents the type of variation. The middle heatmap shows the mutational landscape of gastric cancer. Each column denotes a sample and each row denotes a gene or pathway. The bottom shows the state ofchromosomal integrity of each sample. (b) DDR and CCR custom-pathways-related genes mutational landscape in the samples identified by targeted sequencing. The samples are sorted by clinical features. (c) The forest plot for differences of mutated genes between the two cohorts (P-value is set as 0.05). CNV in the genes meta-analysis is not included.
Another commonly mutated pathway, CCR, was implicated in maintaining genome integrity. Approximately 94% of samples had at least one variation in the CCR pathway (Figure 2(b)). TP53 is a pivotal gene involved in multiple aspects of the cell cycle, whose variations occurred in 60.7% (68/112), the most frequent alteration in our samples. Previous studies reported that TP53, RB1, and PTEN played preeminent roles in regulating cell proliferation and had been validated in the establishment of CIN.22–24 Our results showed a strong relationship between TP53 dysfunction and CIN (Cramer's ϕ = 0.645, P < 0.0001; Figure 2(c)), but not RB1 or PTEN. In the DNA damage checkpoint genes, ATM was the second most common gene mutation in the pathway; however, there was no significant difference between the CIN and CS groups (P = 0.09), which contrasts with a previous study. 25
We performed a clinical enrichment analysis and, once again, found that TP53 mutations were enriched in the CIN subtype (P = 1.48e-6, fdr = 0.001). More importantly, TP53 mutational frequency was of even greater concern for the interpretation of mutational effects. The median of allelic frequency (AF) was 23.95% (95% CI 17.3–27.6%), 0% (95% CI 0–0%) and 3.95% (95% CI 0–16.4%) in patients with CIN, CS, and CS/CIN (Figure 3(b)). About 60% (9/16) of patients with AF less than 10% reserved stable chromatin structures. The proportion was approximately 4% (2/52) in the cohort of AF above 10%. The two mutations were p.L194F with AF 11.6% and p.R282W with AF 17.4%. We identified a total of 56 variants in the TP53 gene via Mutech2 analysis (Figure 3(a)). Most of these were missense mutations located in the CpG dinucleotides of the DNA-binding domain, as well as five splicing mutations, and two single copy deletions. As immunostaining of ≥10% was considered positive for TP53 immunohistochemistry (IHC), TP53 mutations were in concordance with the immunohistochemical results in 95.7% of patients (Table S1; Figure 3(c)).
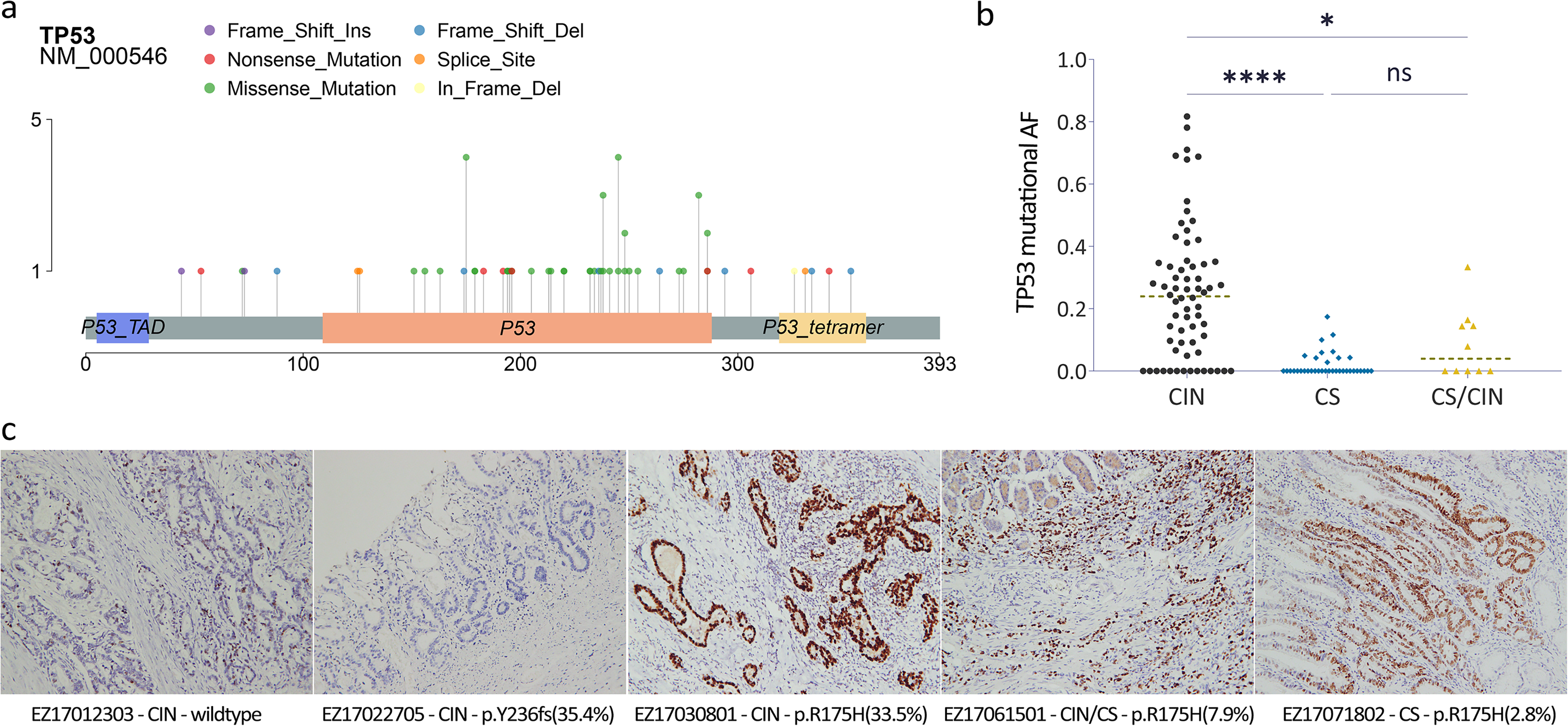
Characteristics of TP53 mutations in gastric cancer. (a) A lollipop plot of amino acid changes of TP53 protein identified from the 112 samples is shown. The number in the Y-axis shows recurrent occurrence, and the number in the X-axis indicates the amino acid sequence. (b) The distribution of TP53 mutational frequency across CIN, CS, and CS/CIN. (c) IHC assessment of distinct TP53 mutation status. The number in brackets denotes the allelic frequency of TP53 mutation.
Clinical significance of chromosomal integrity
The 2010 WHO classification categorized gastric cancers into four major histologic subtypes: tubular, papillary, mucinous and poorly cohesive (including signet ring cell carcinoma). In our samples, 13.4% (15/112) was mucinous adenocarcinoma, or co-exist with other histologic patterns. All mucinous adenocarcinomas were restricted to CIN cohort (including one CS/CIN sample; P = 0.0018) and nine of them owned TP53 mutation. Alternatively, gastric cancers were divided into intestinal and diffuse histological subtypes according to the Lauren classification. We identified 30 diffuse patients, 36 intestinal patients and 19 patients with mixed type. Compared with patients with CIN, CS revealed enrichment of the diffuse subtype (P = 0.0236). The clinical features of patients were generally different between CS and CIN cohorts. While patients with CIN were prone to HER2 positive and mucinous adenocarcinoma, patients with CS had a poorly differentiated diffuse subtype (Table 1; Figure 4(d)). It was taken into consideration that CDH1 is generally accepted as a driver mutation for the hereditary diffuse gastric cancer syndrome, 26 we identified pathogenic CDH1 somatic mutations in 12 patients—all of which were the pure diffuse type with one exception of the mixed type (Table S1).
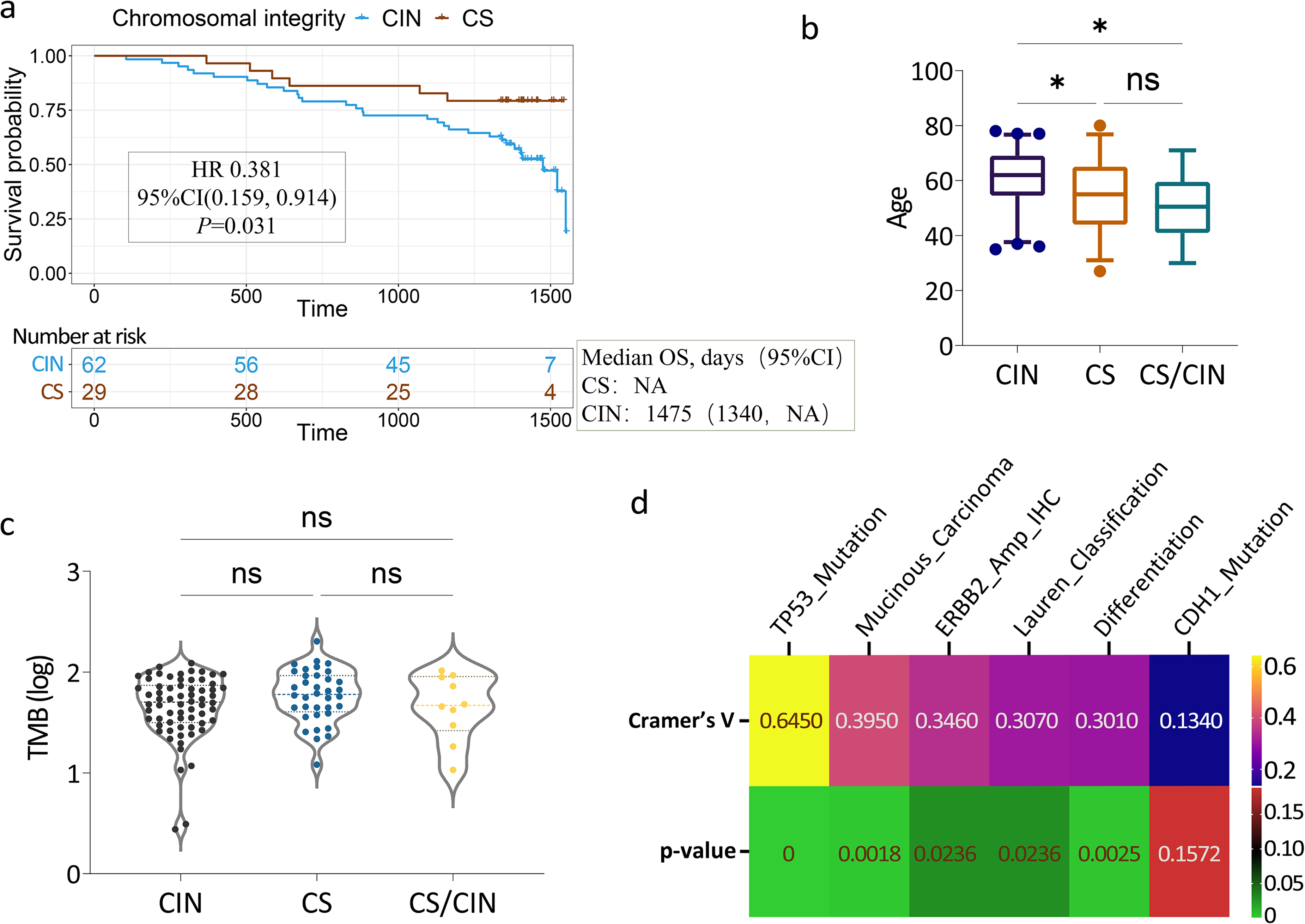
Clinical significance of chromosomal integrity. (a) Kaplan–Meier plots for OS by chromosomal integrity of gastric cancer patients. (b) The age distribution of patients with different chromosomal integrity. (c) The TMB distribution of patients with different chromosomal integrity. (d) Correlation analysis between chromosomal integrity and various clinical manifestations. Cramer's V indicates the strength of the association. The P-value indicates the statistical significance of the association.
Our classification, which was based on chromosomal integrity, predicted the survival outcomes of patients. The median OS in cohort CS was significantly longer than in cohort CIN (OS: not reached vs. 49.2 months, P = 0.025), which suggested a better prognosis in CS patients (HR = 0.381, 95% CI 0.159–0.914, P = 0.031) (Figure 4(a)). This longer OS appeared to be independent of other variables. However, CS patients were preferentially prone to be poorly differentiated (P = 0.0025), which was consistent with the histological subtype of diffuse adenocarcinomas. CIN was also related to the age of patients; namely, older people were more likely to be CIN (Figure 4(b)). However, TMB had no difference between two states (Figure 4(c)), in accordance with CIN results from CCR deficiency but not DDR deficiency, which leads to higher TMB. 27
Discussion
Most previous studies have reported DNA copy number alterations in advanced cancers or metastatic tumors.7,28 Herein, we identified two states of chromosomal integrity in locally or regionally primary gastric cancer. We observed aneuploidy and regional CIN in more than half of newly diagnosed cancer, which highlighted that an extensive chromosomal metabolic disorder exists in the development and spread of cancers. CIN did not seem to directly lead to carcinogenesis, per se, but it does provide more opportunities to break through limitations of cell proliferation and to acquire hallmarks of cancer gradually. 29
Although some significant CNVs were identified in our study (data not shown) and in various types of cancer elsewhere,21,30 they were probably the recurrent regions of amplification of oncogenes or the deletion of tumor suppressor genes. Indeed, regions containing ERBB2, EGFR, PLCG1, MYC, CCND1, CCND3, and CCNE1 oncogenes were frequently amplified, while tumor suppressor genes such as CDKN2A, CDKN2B, RAD17, and RAD50 were deleted (Figure 2(a)). Given CNVs were highly heterogeneous in specific cancers,31,32 they may be the result of a random accumulation of CIN. Therefore, stratifying patients based on the states of chromosomal integrity are significant both biologically and clinically.
We have shown that the underlying cause of CIN was loss-of-function of TP53. Strikingly, TP53 was the most significant gene in the mutational analysis and the pathway analysis, if not the only one. As somatic TP53 mutations occur in TCGA pan-cancer at 42% of samples, 33 it is significant to be expanded to a larger pancancercohort for the two states of chromosomal integrity. When explaining the impact of TP53 mutation on chromosomal integrity, we should pay more attention to mutational allelic frequency. Maybe low AF (<10%) of the gene mutation may still have a role play, or the functional effect was unable to be distinguished. However, in contrast, not all CIN patients harbored the TP53 mutation; cells appeared to exist in non-mutational p53 inactivating mechanisms, such as aberrant methylation of the promoter. 34 Other genes in the TP53 pathway may have altered in a complementary fashion. MDM2 and MDM4 have been stated to function as upstream regulators of TP53. However, we did not detect functional alterations in both these genes.
CIN was commonly identified in tumors but rarely used in oncology clinical practice. Our study initially took advantage of dichotomy of CS and CIN and showed some clinical relevance, which would promoteclinical transformation. Classification based on chromosomal integrity would help patient stratification, including the right choice of sensitive cases for chemotherapy or immunotherapy regimens. As our results implied that CIN was the consequence of cell cycle dysregulation, we speculated that CIN was sensitive to DNA-damaging drugs while CS was sensitive to cell-cycle disturbance according to the strategy of synthetic lethality in cancer therapy.35,36 Indeed, platinum-based adjuvant chemotherapies represent standard therapies in gastric cancer, and the addition of radiotherapy to adjuvant chemotherapy demonstrates improved disease-free survival for intestinal-subtype patients 37 ; however, the addition of docetaxel showed clinical benefit in undifferentiated patients. 38 Although CIN influenced clinical the therapeutic effect, whether chemoradiotherapies reshaped the genome was still inconclusive, since neoadjuvant chemotherapy with various regimens in our cohort did not seem to affect cancer genome stability and chromosomal integrity. For immunotherapy, CIN-derived copy number deletion at the DNA level may be the cause of immune escape through neoantigen deletion. 39 CIN may also provide a more evolutionary chance for cancer cells under the pressures of immunological surveillance and immunotherapies. 40 These studies suggested that CIN could be a possible biomarker for the response prediction to immunotherapies.
In clinical application, TP53 mutation detection, but not immunohistochemistry (Figure 3(c)), would serve as a proxy for CIN. In view of TP53-mutant-associated cancer deaths of more than 500 million people today, 41 specific treatments for TP53 mutated tumors are required in this considerable population of unmet need. The functions of TP53 mutation have been heavily studied, but its role in cancer remains elusive. Overlooking functional versatility of TP53 protein, we focusing on the impact of TP53 dysfunction on cell cycle regulationand the consequential CIN would promote development of personalized therapy towards TP53-mutant tumors.
Several limitations exist in this study. First, whole-exome sequencing was conducted in 12 patients and targeted sequencing for cancer-associated genes was conducted in 112 patients. Because the samples were not available, we could not evaluate the concordance between the two sequencing strategies, although we used the same standards and parameters as in the chromosomal integrity analysis. Second, the mechanisms underlying CIN were complex as well as diverse. We only investigated the relationship between genetic mutation and chromosomal integrity, and found an excellent consistency of TP53 dysfunction and CIN. Gene expression patterns and epigenetic modifications should be included in future studies. Third, the follow-up and treatment courses after surgical operations were incomplete. To make CIN-based personalized cancer therapies into reality, more retrospective and prospective studies regarding the clinical significance of chromosomal integrity are warranted in future clinical trials.
Conclusions
CIN is prevalent in cancers but with an uncertain clinical relevance. We investigated CNV across the entire genome and divided gastric cancer into two cohorts based on the states of chromosomal integrity: chromosomal stability and chromosomal instability. Our results also demonstrated a strong relationship between chromosomal integrity status and TP53 mutation. Our study initially revealed some clinical significance of chromosomal integrity: that CIN patients were prone to HER2-positive and mucinous adenocarcinoma, as CS patients had a diffuse subtype and poorly differentiated but longer OS. We propose that classifying cancers based on chromosomal integrity would contribute to the development of more efficient treatment and would enable personalized therapy.
Supplemental Material
sj-xlsx-1-jbm-10.1177_03936155221106217 - Supplemental material for Clinical significance of chromosomal integrity in gastric cancers
Supplemental material, sj-xlsx-1-jbm-10.1177_03936155221106217 for Clinical significance of chromosomal integrity in gastric cancers by Rukui Zhang, Zhaorui Liu, Xusheng Chang, Yuan Gao, Huan Han, Xiaona Liu, Hui Cai, Qiqing Fu, Lei Liu and Kai Yin in The International Journal of Biological Markers
Supplemental Material
sj-xlsx-2-jbm-10.1177_03936155221106217 - Supplemental material for Clinical significance of chromosomal integrity in gastric cancers
Supplemental material, sj-xlsx-2-jbm-10.1177_03936155221106217 for Clinical significance of chromosomal integrity in gastric cancers by Rukui Zhang, Zhaorui Liu, Xusheng Chang, Yuan Gao, Huan Han, Xiaona Liu, Hui Cai, Qiqing Fu, Lei Liu and Kai Yin in The International Journal of Biological Markers
Supplemental Material
sj-xlsx-3-jbm-10.1177_03936155221106217 - Supplemental material for Clinical significance of chromosomal integrity in gastric cancers
Supplemental material, sj-xlsx-3-jbm-10.1177_03936155221106217 for Clinical significance of chromosomal integrity in gastric cancers by Rukui Zhang, Zhaorui Liu, Xusheng Chang, Yuan Gao, Huan Han, Xiaona Liu, Hui Cai, Qiqing Fu, Lei Liu and Kai Yin in The International Journal of Biological Markers
Footnotes
Acknowledgements
For technical advice, we thank the project of Shanghai's Double First-Class University Construction and Development of High-Level Local Universities: Intelligent Medicine Emerging Interdisciplinary Cultivation Project, and Medical Research Data Center of Fudan University. For clinical information collection, we thank the department of information, Changhai Hospital, Naval Military Medical University.
Author contributions
Rukui Zhang: data analysis, data interpretation, investigation, and draft writing. Zhaorui Liu: samples collection, clinical information collection, and data analysis. Xusheng Chang: clinical information collection. Yuan Gao: clinical follow-up. Huan Han: immunohistochemistry experiment. Xiaona Liu: data analysis. Hui Cai: clinical follow-up. Qiqing Fu: draft reviewing. Lei Liu: research design, supervision, and funding acquisition. Kai Yin: research design, supervision, and funding acquisition. Rukui Zhang, Zhaorui Liu, and Xusheng Chang contributed equally to this work.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Changhai Hospital. Informed consent was obtained from all patients for the study.
Data availability
The data used to support the findings of this study are available from the corresponding author upon reasonable request
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was supported by the National Key Research and Development Program of China (No. 2016YFC0901903), Medical Innovation Research Program of Science and Technology Commission of Shanghai Municipality (No. 21Y11913100) and Health Industry Clinical Research Project of Shanghai Municipal Health Commission (No. 20204Y0416).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
