Abstract
Background
Non-invasive liquid biopsies of circulating tumor DNA (ctDNA) is a rapidly growing field in the research of non-small cell lung cancer (NSCLC). In this study, factors affecting the concordance of mutations in paired plasma and tissue and the detection rate of ctDNA in real-world Chinese patients with NSCLC were identified.
Methods
Peripheral blood and paired formalin-fixed paraffin-embedded tumor tissue samples from 125 NSCLC patients were collected and analyzed by sequencing 15 genes. Serological biomarkers were tested by immunoassay.
Results
The overall concordance between tumor and plasma samples and the detection rate of somatic mutations in ctDNA was 69.2% and 78.4%, respectively. The concordance and detection rate raised with clinical stage were stage I: 14.3%, 14.3%; stage II: 53.3%, 60.0%; stage III: 71.4%, 78.1%; stage IV: 74.1%, 85.2%. With increased tumor diameter, the concordance and detection rate raised from 33.33% to 71.64% and 33.33% to 80.8%, respectively. For patients with partial response, stable disease, progressive disease, and who were treatment-naïve, the concordance and detection rates were 0.0%, 62.7%, 75.2, 73.6%, and 16.7%, 61.9%, 83.3%, 86.5%, respectively. Serological markers: CEA, CA125, NSE, and CYFRA21-1 were significantly higher for patients with detectable somatic alterations in ctDNA than in those who were ctDNA negative (17.08 ng/mL vs. 3.95 ng/mL, 21.63 U/mL vs. 18.27 U/mL, 17.68 U/mL vs. 14.14 U/mL, and 6.55 U/mL vs. 3.81 U/mL, respectively).
Conclusion
Advanced-stage, treatment naïve or poor therapy outcome, and large tumor size were associated with a high concordance and detection rate. Patients with detectable mutations in ctDNA had a higher level of carcinoembryonic antigen, CA125, NSE, and CYFRA21-1.
Introduction
Lung cancer is the leading cause of cancer incidence and mortality worldwide. 1 Non-small cell lung cancer (NSCLC) is the most common type, accounting for approximately 85% of all lung cancer cases. 1 Targeted therapy 2 and immunotherapy 3 have shown remarkable effectiveness in increasing the survival of patients with NSCLC. Accurate identification of predictive somatic alterations is crucial for screening targeted drugs and advancing the understanding of treatment-induced tumor evolution. Tumor tissue is recommended for pathologic diagnosis and molecular testing. 4 However, there are approximately 30% of NSCLC patients could not obtain sufficient tissue for subsequent molecular analyses. 5 Testing circulating tumor DNA (ctDNA) isolated from peripheral blood has been shown as a complement to routine tissue-based diagnostic and a feasible means of identifying acquired resistance mechanisms. It could identify actionable tumor mutations at the time of diagnosis or recurrence of patients with advanced lung adenocarcinoma. Therefore, the non-invasive option, also known as liquid biopsy, is recommended in the College of American Pathologists (CAP) and the International Association for the Study of Lung Cancer (IASLC) guidelines. 6 However, the sensitivity of ctDNA detection and concordance to tumor tissue are so compromised to be restricted to use in clinical practice. It has been reported that the false-negative rate of a ctDNA test was up to 30%. 6 Therefore, it is essential to screen out the patients who would benefit from liquid biopsy.
Protein markers in serum, such as carcinoembryonic antigen (CEA), carbohydrate antigen 19-9 (CA19-9), carbohydrate antigen (CA125), cytokeratin 19 fragment (CYFRA21-1), and neuron-specific enolase (NSE) have been used conventionally in predicting the therapeutic effect and prognosis of NSCLC.7,8 Many studies have explored the integration of genomic mutations and protein markers in peripheral blood that could improve the NSCLC diagnosis.9,10 However, there are few reports about their correlations. In this real-world study, the genomic profile of patients with NSCLC and the concordance of variants in paired plasma and tissue samples was identified, and the relationship between clinical characteristics and the genomic variants in ctDNA was analyzed.
Patients and methods
Study participants
A total of 125 NSCLC patients from Peking University Cancer Hospital & Institute were enrolled from March 2017 to September 2020. Clinical features including sex, age, smoking history, tumor size, onset location, pathological subtype, and clinical stage were documented. Written informed consent was obtained from the participants in the study. This study was approved by the medical ethics committee of the Peking University Cancer Hospital & Institute (2016XJS01) and was conducted according to the Declaration of Helsinki Principles.
Samples collection and targeted sequencing
The peripheral blood samples and paired formalin-fixed paraffin-embedded (FFPE) tumor tissues from 125 patients were collected. Venous blood in 10 mL ethylenediaminetetraacetic acid (EDTA) tubes were centrifuged twice at 1600 g 4°C for 10 min and 16,000 g 4°C for 10 min sequentially for ctDNA testing. Serum was isolated from another tube of peripheral blood to identify biomarker levels. FFPE samples were de-paraffinized with xylene followed by genomic DNA extraction using QIAamp DNA FFPE Tissue Kit (Qiagen). Genomic DNA from white blood cells and cell-free DNA were extracted using the DNeasy Blood & Tissue Kit (Qiagen) and the QIAamp Circulating Nucleic Acid Kit (Qiagen), respectively, according to the manufacturers’ instructions. Sequencing libraries were prepared following the protocols as previously reported. 11 Hybridization-based target enrichment was carried out with Nanjing Geneseeq Technolog Inc. lung cancer panel covering all coding regions and splice sites of the 15 NSCLC-associated genes, including AKT1, ALK, BRAF, EGFR, ERBB2, KRAS, MAP2K1, MET, NF1, NRAS, PIK3CA, PTEN, RET, ROS1, and TP53. All samples were tested in a Clinical Laboratory Improvement Amendments (CLIA)—CAP-certified genomic testing facility (Nanjing Geneseeq Technolog Inc., Nanjing, China). Somatic variants, including single nucleotide variant, indel, and fusion, were called using an internally validated bioinformatics analysis pipeline. 12 Sequencing data were demultiplexed by bcl2fastq (v2.19), analyzed by Trimmomatic to remove low-quality (quality<15) or N bases. SNPs and indels were called by VarScan2 and HaplotypeCaller/UnifiedGenotyper in GATK, with the mutant allele frequency (MAF) cutoff as 0.5% for tissue samples, 0.1% for liquid biopsy samples, and a minimum of three unique mutant reads. Common variants were removed using dbSNP and the 1000 Genome project. Germline mutations were filtered out by comparison with each patient's whole blood controls. The resulting somatic variants were further filtered through an in-house list of recurrent sequencing errors that was generated from over 10,000 normal control samples on the same sequencing platform.
Tumor biomarker testing
The Cobas E602 electrochemiluminescence immunoassay (Roche Diagnostics, Indianapolis, IN, USA) was used to determine the concentration of protein markers in serum, including CEA, CA19-9, CA125, CYFRA21-1, and NES. In addition, the chemiluminescence immune analyzer ARCHITECT i2000sr (Abbott Diagnostics, Abbott Park, IL, USA) was used to detect the biomarker SCC.
Statistical analysis
To calculate the concordance in FFPE and plasma ctDNA matched samples, gene variations in FFPE samples were used as the reference compared to ctDNA. The sensitivity, specificity, positive predictive value (PPV), and concordance were calculated as in the previous report. 13 To analyze the relationship of clinical features and the top two mutated genes, patients were divided into two groups according to whether TP53 or EGFR gene variations were detected in either FFPE or ctDNA samples. The differences in clinicopathological features were assessed by a two-sided Chi-square test or Fisher's exact test. Continuous variables were expressed in median and interquartile range (IQR) and analyzed by the Mann-Whitney U test. All analyses were performed using R 3.4.0 and SPSS version 22 (IBM). Differences with a P <0.05 were regarded as statistically significant.
Results
Demographics and clinical characteristics
The cohort included 78 males and 47 females, with a median age of 62 years (mean 62.7 years, IQR 57−70.8 years). Nearly half of the patients (53.6%, 67/125) had a smoking history, and 97% were male. Adenocarcinoma was the most common pathologic subtype (76.8%), followed by SCC (16.0%). Of the enrolled patients, 64.8% were at advanced stages, while stage I–II was less than 10% (Figure 1). The median interval between tissue and peripheral blood samples collection was 12 days (IQR 8–29 days).
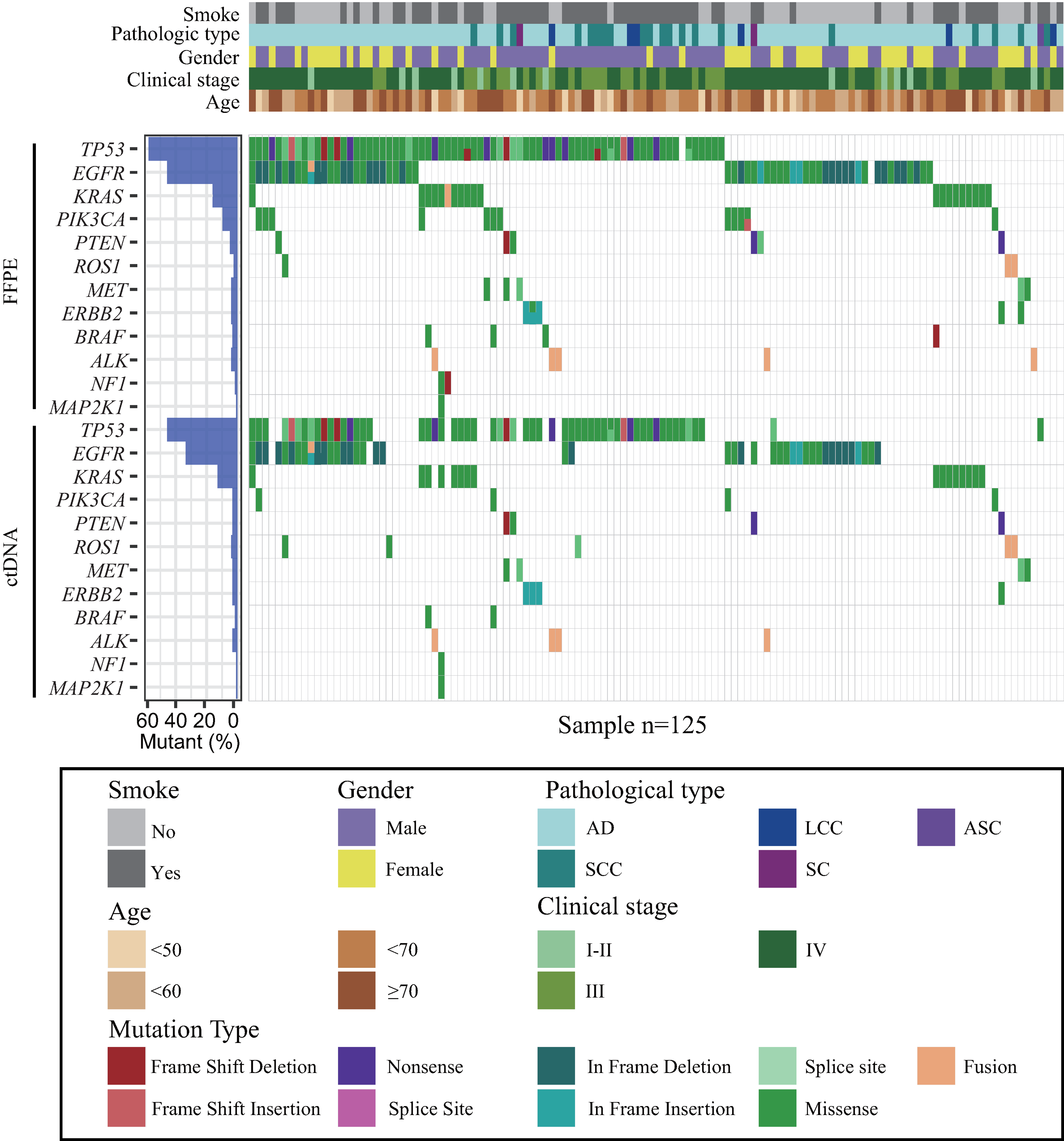
Summary of patient characteristics and mutation landscape in matched FFPE and plasma ctDNA sample pairs. Samples were classified by smoking history, gender, age range, pathological diagnosis, and clinical stage. Mutation type per sample in FFPE is compared to those in plasma ctDNA.
Mutation concordance in plasma and tissue sample pairs
Among the 125 patients with matched plasma and tissue samples, 122 (98.4%) carried 228 somatic mutations in either tissue or plasma. The other 3 cases had no gene variations in either tissue or plasma (Figure 2(a)). Patients were divided into three groups according to the distribution of gene variants in paired samples: 58.4% complete consistency (73/125), 20.0% partial consistency (25/125), and 21.6% absolute difference (27/125) (Figure 2(b)). There were 118 concordant mutations of 70 patients in both tissue and plasma. Twenty-five patients with 50 consistent variants contained additional discordant variants in either tissue (28 variants) or plasma (5 variants). One patient had a variant only in plasma, and 24 patients had 43 variants only in the tissue. The other two patients had 6 variants in both tissue and plasma, but those variants were discordant in different sample types (Figure 2(b)).
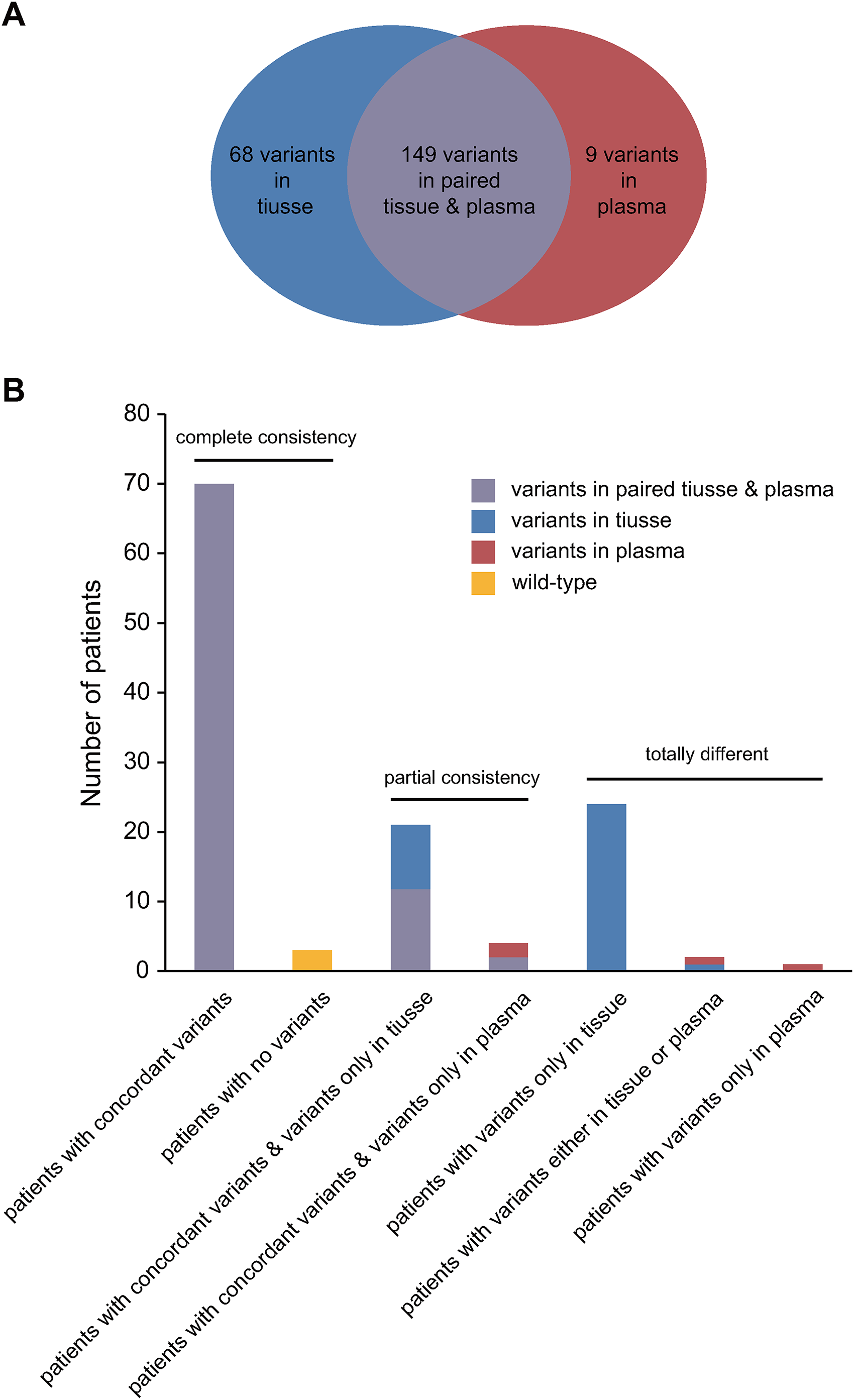
The concordance of variants in paired tissue and plasma ctDNA. (a) All 226 somatic variants were divided into three groups: Only in tissue (blue), only in plasma (red), and in both samples (purple). (b) Patients with NSCLC were divided into complete consistency, partial consistency, and total difference groups according to the distribution of gene variants in paired samples. The height of the bar indicated the number of patients. The bars were colored to indicate the proportion of the different types of variations.
The overall concordance between FFPE and matched plasma ctDNA was 69.2%, with a plasma sensitivity of 70.8% (95% CI: 61.6%–78.6%), specificity of 42.9% (95% CI: 11.8%–79.8%), and a PPV of 95.4% (95% CI: 88.1%–98.5%). The concordance was 14.3%, 53.3%, 71.4%, and 74.1%, ranging from stage I to IV (P = 0.012), and were significantly associated with tumor diameter: with 33.33%, 69.57%, and 71.64% for diameter <1 cm, ≥1 cm and <3 cm, and ≥3 cm, respectively (P < 0.001).
Response to treatment was another impact factor on the concordance (P = 0.001). For all the six patients with partial response (PR), somatic variants in tissue were undetected in ctDNA. For patients evaluated as stable disease (SD) or progress disease (PD) the concordance between ctDNA and tissue was 62.7% and 73.6%. Treatment-naïve patients had a higher concordance than those with treatment history (75.2% vs. 60.5%)
The relationship between clinical characteristics and the detection rate of gene variations in plasma ctDNA
For consistency analysis, the results of tissue testing were regarded as the gold standard. However, due to intratumoral, intermetastatic, and intrametastatic heterogeneity, a single tissue specimen could not completely contain all tumor subclones. After excluding the experimental and sequencing errors, variants that are present in plasma but not in tissue should be real. Therefore, the relationship between the genomic features of plasma ctDNA and clinical characteristics was subsequently analyzed.
The overall detection rate of somatic mutations in plasma ctDNA was 78.4%. The detection rate of ctDNA for patients with advanced-stage, poor treatment outcome, and ≥1 cm tumor diameter was significantly higher than in those with early-stage, good treatment outcome, and <1 cm tumor diameter. However, this was not affected by sex, age at diagnosis, smoking, sampling site, and pathological subtype (Table 1).
The relationship between clinical characteristics and the presence of gene variations in plasma ctDNA.
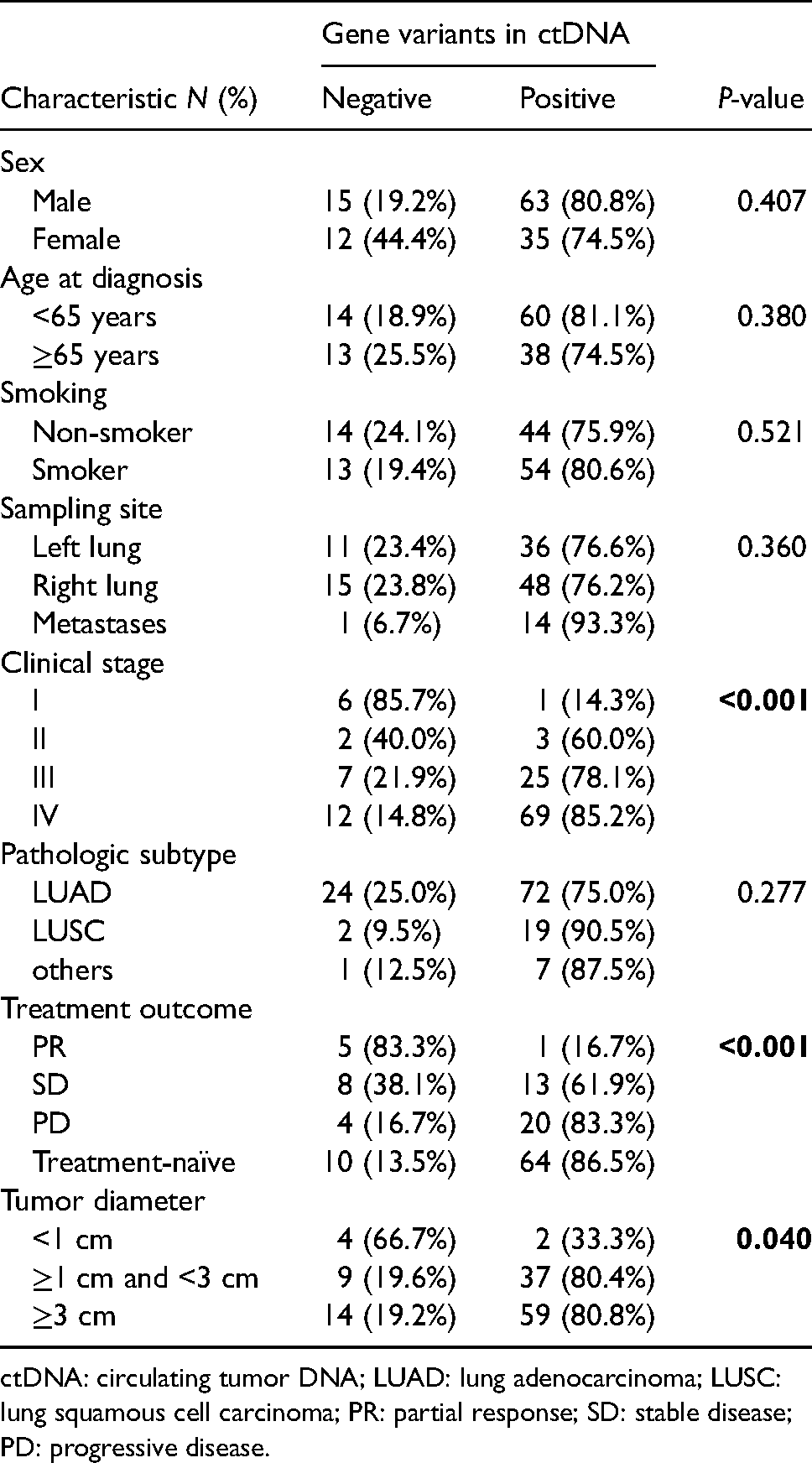
ctDNA: circulating tumor DNA; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; PR: partial response; SD: stable disease; PD: progressive disease.
Patients with mutations detected in ctDNA had significantly higher levels of CEA, CA125, NSE, and CYFRA21-1 in serum than those negative. The median values between the two groups of CEA, CA125, CA199, NSE, CYFRA21-1, and SCC were 17.08 ng/mL versus 3.95 ng/mL, 53.75 U/mL versus 21.96 U/mL, 21.63 U/mL versus 18.27 U/mL, 17.68 U/mL versus 14.14 U/mL, 6.55 U/mL versus 3.81 U/mL, and 1.0 U/mL versus 0.8 U/mL, respectively (Figure 3).
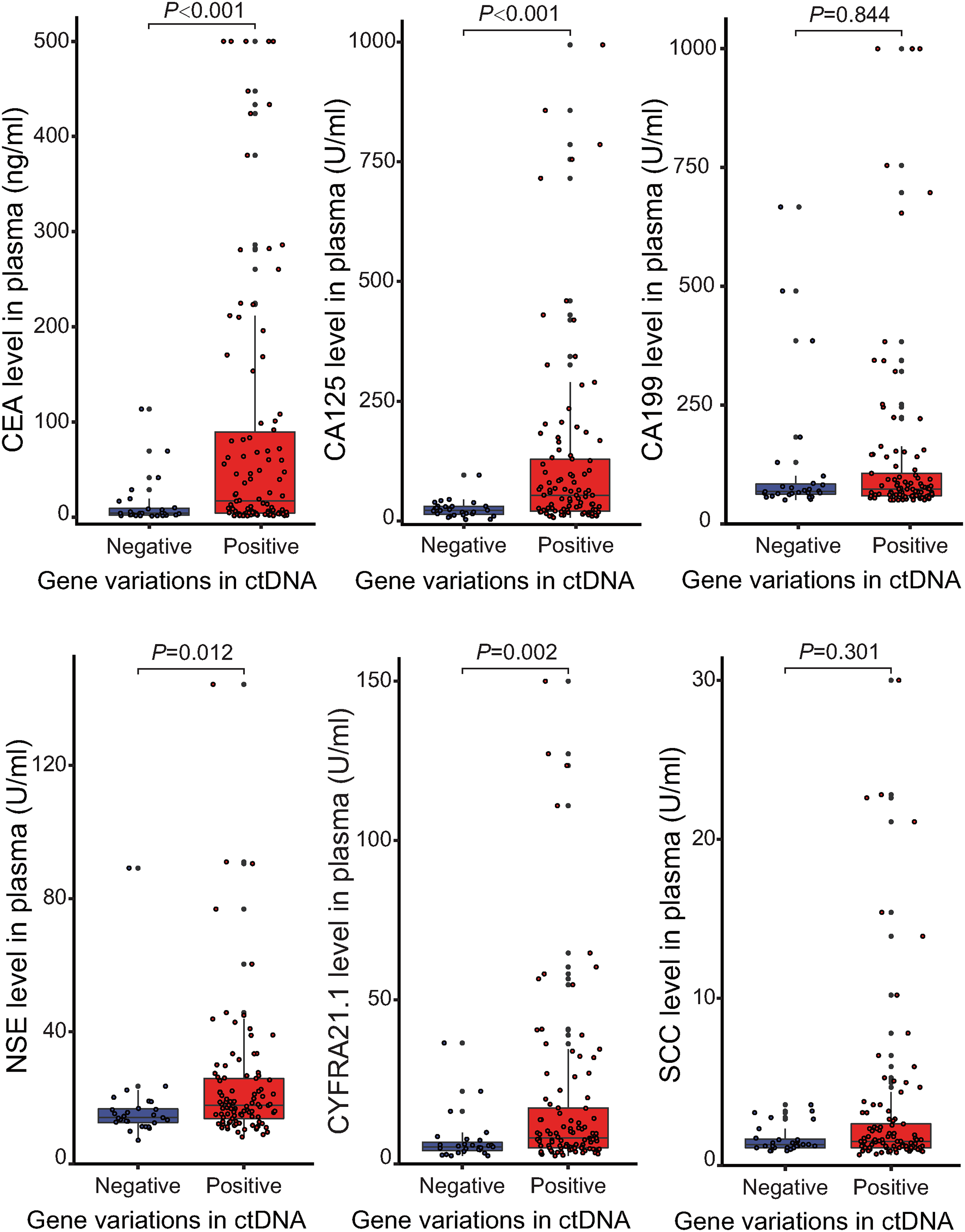
Relationship between tumor diameter and levels of serological biomarkers and the presence of gene variations in plasma ctDNA.
Somatic alterations distribution in tissue and matched plasma samples
The majority of variations identified in FFPE and plasma were missense (64.8% and 63.1%). Insertions and deletions accounted for 19.2% of FFPE and 19.4% of ctDNA variations. For FFPE samples, the top five most frequent somatic alterations were in the following genes: TP53 (58.4%), EGFR (46.4%), KRAS (15.2%), PIK3CA (10.4%), and PTEN (5.6%). The top five most frequently mutated genes in plasma samples were TP53 (45.6%), EGFR (33.6%), KRAS (13.6%), ROS1 (4.8%), and PTEN (3.2%) (Figure 1). Except for ROS1, the detection rate of all other genes in tissues was higher than that in plasma.
Somatic mutations of EGFR were enriched in exon 18–21. The most common EGFR alterations were L858R (35.7%) and the in-frame deletions in exon 19 (35.7%), followed by T790 M (10.0%) (Figure 4(a)). The mutations located in the TP53 most commonly occurred in its DNA-binding domain between amino-acid residues 106–285. C242, R248, R273, and D281 were the most common point mutations in TP53, accounting for 21.1% of all variants (Figure 4(b)). Mutations in codon 12 are the predominant somatic alterations in KRAS (77.8%), and the G12C accounted for 38.9% of all KRAS mutations.
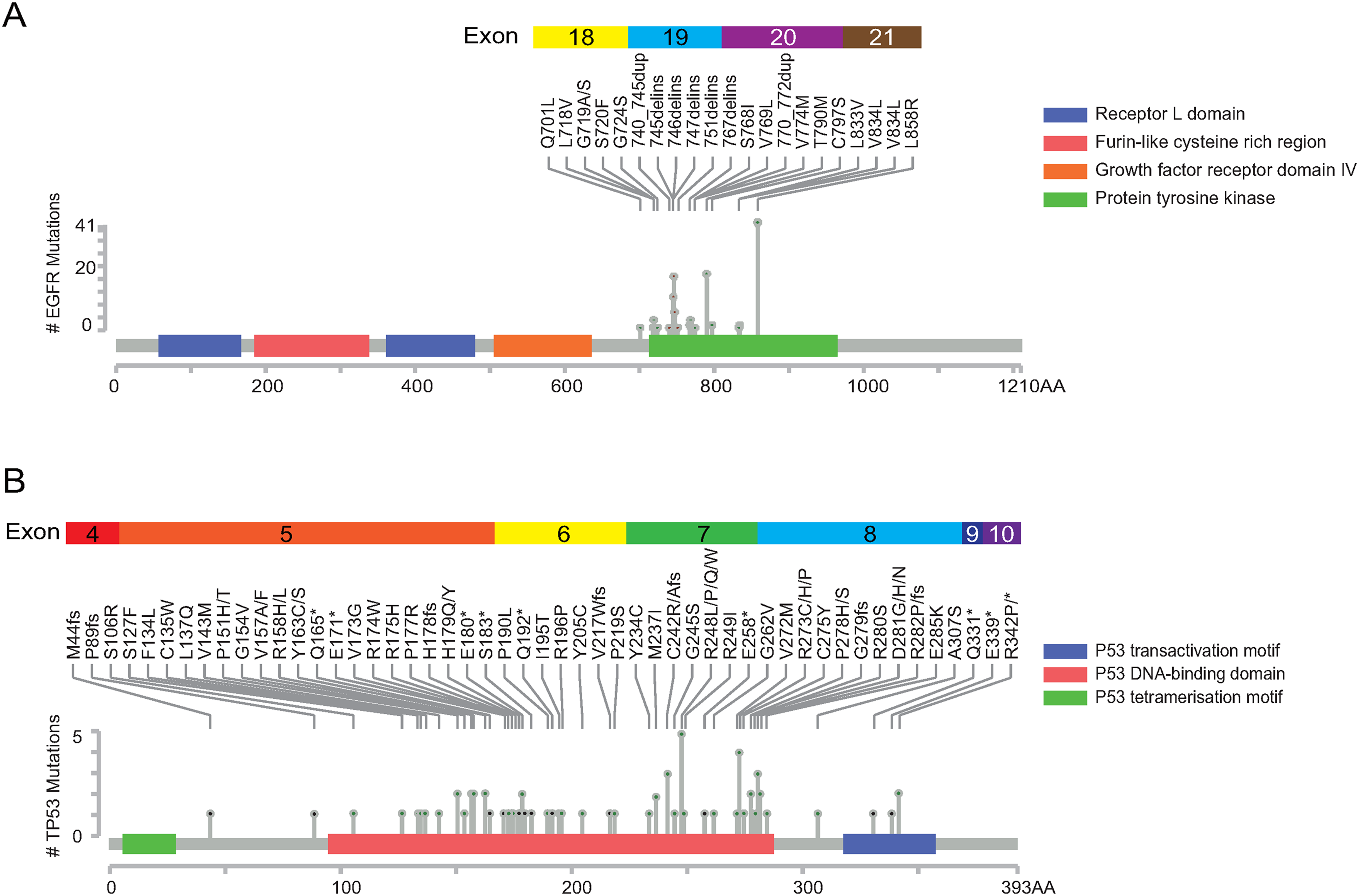
Distribution and localization of somatic SNVs and indels targeting EGFR (a) and TP53 (b) in NSCLC patients. Colorful boxes above the pictures represent exons of EGFR or TP53. The grave bar and colored boxes on it represented the protein structure and domains. The height of the lollipop in the middle of the figure represents the gene mutation frequency. Most of the mutations were localized in the protein tyrosine kinase domain of EGFR (a) and the DNA-binding domain of TP53 (b).
Except for females, non-smokers, and adenocarcinoma—which are well-known factors related to EGFR mutated-type—EGFR mutations were more common in patients with advanced-stage, smaller tumor size, poor treatment outcome, higher CEA level, and lower CYFRA21-1 and SCC levels. Additionally, EGFR variants are more easily detected in the primary site than in metastases (Supplementary Table 1). In contrast, TP53 mutations were enriched in males, smokers, and patients with higher CYFRA21-1 levels.
Discussion
Liquid biopsy is a rapidly growing field in lung cancer study, offering an alternative to standard procedures when tumor tissue is not available and providing a rapid and dynamic assessment of emergent resistance mechanisms that can be used to guide treatment decisions.14,15 However, an excellent molecular ctDNA test should retain an acceptable sensitivity, specificity, and concordance to molecular testing in the tumor tissue. Previous studies have shown that there are many impact factors on concordance. Chen et al. 16 reported a gradual increase in concordance according to the clinical stage: 57.9%, 66.7%, and 90.0% for stages I, II, and III, respectively, which was consistent with our study. Tumor volume was also reported to be strongly correlated with ctDNA level per plasma mL.17–19 A mathematical model predicts a median ctDNA detectable tumor size of 0.83 cm for informed monthly cancer relapse testing. 20 In this study, NSCLC patients with tumor diameter <1.0 cm had a notable lower mutation concordance. In contrast, patients with a tumor diameter ≥1.0 cm had an acceptable concordance. With the increase in tumor size, the concordance rate raised slowly. The tumor diameter and clinical stage are indicators for the volume and burden of the tumor, which both reflect the number of the malignant cells and the total amount of ctDNA released into peripheral blood.
The ctDNA level in plasma was also associated with cell proliferation. 20 In this study, patients’ response to treatment was another factor impacting the concordance: the better the treatment outcome, the lower the concordance. Patients with advancing disease had the highest concordance. According to Response Evaluation Criteria in Solid Tumors (RECIST) guidelines, the shrinkage or increase of target lesions and the development of a new lesion or not are important indicators for response evaluation. 21 The poor outcome represents the resistance to current treatment and the strong proliferative activity of tumor cells.
In general, there are two inconsistent aspects between the results from plasma and tissue: false positive and false negative. False positive is defined as gene variants detected in plasma but not in tissue, revealing tumor heterogeneity. 22 The limitation of ctDNA applied in clinical was mainly attributed to its higher false-negative rate. Besides the advanced clinical stage, poor treatment outcome, and larger tumor size mentioned above, patients with higher levels of CEA, CA125, NSE, and CYFRA21-1 were more likely to have a positive ctDNA test result. These serological biomarkers are generally accepted as useful prognostic factors, 23 as predictors of efficacy for chemotherapy, 24 targeted therapy, 25 and immunotherapy, 7 and as markers of postoperative recurrence and metastasis.26,27 It has also been reported that the expression levels of CEA, CYFRA21-1, and NSE significantly correlated with cfDNA concentration. 28 However, the relationship between the levels of CA199 and SCC, and the presence of gene variants in ctDNA needs further study.
Different from EGFR, TP53 mutations favor males and smokers. This phenomenon has been reported extensively and is attributed to preferential DNA adduct formation by carcinogenic polycyclic aromatic hydrocarbons (PAH) found in cigarette smoke. 29 Benzopyrene-diol-epoxide (BPDE), which is metabolized from the PAH is implicated as a major contributor to TP53 G > T transversions in smokers. 30 Other more common TP53 mutations could be explained as positions of methylated CpG transitions due to the mutation of 5′-methylated cytosine. 31
There are some limitations in this study. First, it was subjected to a limited number of patients and potential sample selection bias due to its retrospective nature. Second, only single nucleotide variants, indel, and fusion in 15 genes were detected, limiting the generalization of the findings. Therefore, future efforts should be directed toward prospective trials with a larger sample size, improved techniques and comprehensive evaluation.
In conclusion, tissue and plasma could provide complementary information of somatic variations for Chinese NSCLC patients. Early-stage, good treatment outcome, and small tumor size were significantly associated with low concordance between matched tissue DNA and plasma ctDNA. Patients with advanced stage, non-treatment history, larger tumor diameter, and a higher level of CEA, CA125, NSE, and CYFRA21-1 in serum had a higher detection rate of ctDNA.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155221099036 - Supplemental material for Comparison of the somatic mutations between circulating tumor DNA and tissue DNA in Chinese patients with non-small cell lung cancer
Supplemental material, sj-docx-1-jbm-10.1177_03936155221099036 for Comparison of the somatic mutations between circulating tumor DNA and tissue DNA in Chinese patients with non-small cell lung cancer by Meng Zhang, Yi Feng, Changda Qu, Meizhu Meng, Wenmei Li, Meiying Ye, Sisi Li, Shaolei Li, Yuanyuan Ma, Nan Wu and Shuqin Jia in The International Journal of Biological Markers
Footnotes
Acknowledgments
We thank all the patients who participated in this study, and the suggestions from Dr. Yuyan Wang.
Author contributions
M.Z., Y.F., N.W., and S.J. conceived the idea of the study; M.Z., C.Q., M.M., S.L., and M.Y., performed the research; M.Z. and Y.M. analyzed data; M.Z., S.L., and Y.F. interpreted the results; M.Z. and S.J. wrote the paper; all authors discussed the results and revised the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
The study has been performed in accordance with the Declaration of Helsinki and has been approved by the Beijing Cancer Hospital Medical Ethics Committee (2016XJS01).
Written informed consent to participate in the study was obtained from participants. A copy of the consent form is available.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Key R&D Program of China, (grant number 2018YFC0910700), Capital’s funds for health improvement and research (grant number 2018-2-1023), and National Natural Science Foundation of China (grant number 82073312).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
