Abstract
Millions of children have multiple nutritional deficiencies, threatening their optimal growth, development, and quality of life. Revealing the magnitude and underlying biology of malnutrition from a greatly expanded set of practical biomarkers will be critical for developing appropriately targeted and evaluated interventions. However, our abilities to reveal and quantify the many forms of malnutrition, other than by anthropometry and occasional use of biochemical indicators, remain limited. Plasma proteomics holds great promise as a basis for developing novel biomarkers to facilitate assessment of growth, micronutrient status, inflammation, and other health status of populations while also providing biological insight into causes and adverse consequences of malnutrition. Discovery-driven plasma proteomics has been shown to reveal functional biomarkers of nutritional and health status, identifying clusters of protein biomarkers from which field-friendly, comprehensive, and low-cost methods could be developed for assessing populations. In this brief review, we summarize several key discoveries to date and discuss potential public health applications of proteomics-based biomarkers in reporting the extent and metabolic features of undernutrition in low-resource settings.
Introduction
Progress toward reducing child stunting, wasting, and multiple micronutrient deficiencies remains unacceptably slow, jeopardizing normal growth and development, health, and survival of children in low-income countries. Although nutritional deficits likely disturb multiple organs, tissues, and body systems, 1 our abilities to reveal underlying biology of malnutrition at a population level are limited with existing public health indicators that generally offer minimal biological insight. Biomarkers that are sensitive to shifts in population nutriture, responsive to underlying causes, and predictive of long-term health outcomes will improve our ability to assess and prevent nutritional deficiencies and their related health consequences. Proteins found in the plasma have great potential as functional biomarkers, as they mediate every biological process: transporting nutrients and hormones, executing biochemical reactions, fighting against exogenous deleterious agents, supporting structure of tissues, and traveling short or long distances to mediate a myriad of cellular responses. 2 A recent revolution in high-throughput proteomics technologies enables simultaneous screening of thousands of proteins present in the plasma in an unbiased way, opening new opportunities to develop novel biomarkers of child nutrition and health status.
Over the past 15 years, we have conducted discovery-driven plasma proteomics to identify proteins associated with different aspects of child nutrition and health. This work was driven by a vision to detect and develop biomarkers for a field-ready, multiprotein assay that can both offer deep biological insights and provide more complete information on nutritional and health status of populations than is currently available with conventional biomarkers. We hypothesized that networks of plasma proteins that are involved in regulatory biological pathways of micronutrient metabolism, inflammation, child growth, cognition, and other metabolic functions and disease processes could be detected and quantified. We posited that clusters of proteins possess sufficient variability to predict micronutrient status of populations and reflect diverse facets of nutrition-related health status, with an expectation of predictive proteins being quantifiable on inexpensive multiplex systems in the future. Encouraged by preliminary 2-dimensional gel electrophoresis findings of nutrient–protein associations in Nepalese pregnant women, 3,4 we tested our hypothesis in a cohort sample of 500 school-aged Nepalese children born to mothers from an earlier antenatal micronutrient trial. 5 In the child cohort, nutritional and health status was extensively characterized using conventional biochemical and anthropometric indicators. 6,7 Using state-of-the-art tandem mass spectrometric, statistical, and bioinformatics methods, 8 we profiled nearly 1000 proteins that formed the basis for our discoveries. 9 Here, we provide a brief summary of proteins associated with a variety of indicators of nutritional and health status and discuss the potential public health use of plasma protein biomarkers in low-resource settings.
Proteins and Micronutrients
Plasma proteomes indeed do exist with conventional, individual micronutrient status indicators, with initial evidence of “proof of concept” emerging with predictive associations between vitamins A, D, and E, copper, and selenium and their major carrier proteins (9-11). The largest proteome was observed with copper (>200 proteins); proteomes with ∼50 to 160 correlates exist for vitamins A, E, B6, and β-cryptoxanthin, and smaller proteomes (<50) have been shown for other nutrient indicators, including plasma ferritin, vitamin K, β-carotene, and selenium (Figure 1). 9 -13 Importantly, small-sized clusters may not be “weak” proteomes. For example, selenoprotein P, a circulating selenium transporter and one of three proteins associated with selenium status, explains 63% of the variability in plasma selenium concentrations. 11 Proteomes of other micronutrients have not, to date, been found predictive of micronutrient status but do reflect complex direct to systemic interplay between nutrients across various biological systems. It is worthy to note that proteomes of ferritin, copper, vitamin A, vitamin B6, β-cryptoxanthin, and β-carotene highly overlap with the proteome of inflammation (Figure 1), 14 suggestive of altered regulation of these nutrients during inflammation.
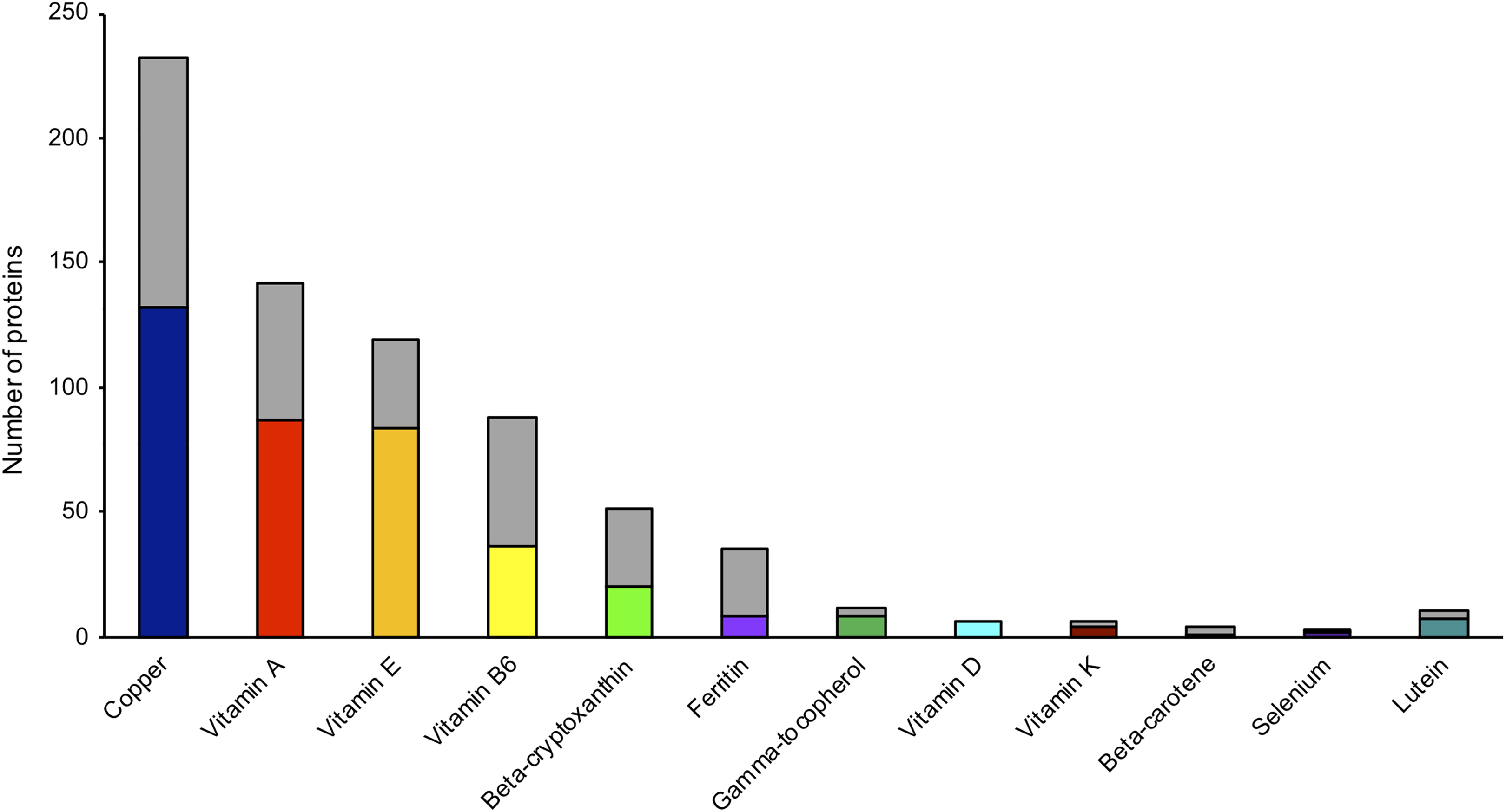
Proteomics-based plasma proteins associated with micronutrients in school-aged Nepalese children. Proteome of each micronutrient is defined by proteins quantified by mass spectrometry and associated with plasma concentrations of copper, retinol (vitamin A), α-tocopherol (vitamin E), pyridoxal-5’-phosphate (vitamin B6), β-cryptoxanthin, ferritin, γ-tocopherol, 25(OH)D (vitamin D), protein induced in vitamin K absence-II (vitamin K), β-carotene, selenium, and lutein, controlling a false discovery rate (q < 0.10). Stacked gray bars represent proteins commonly associated with each micronutrient and inflammation, indicated by α-1-acid glycoprotein, controlling a family-wise error rate of 0.1% (P < 1.0 × 10−6).
A Plasma Inflammasome
Inflammatory conditions in a population are varied and complex. To date, the nutrition community has pegged the definition of inflammation to common, circulating chronic and acute-phase markers of infection, α-1-acid glycoprotein (AGP) and C-reactive protein. Among identified proteomes, the plasma proteome of inflammation, characterized with plasma AGP, has been the largest and most comprehensive in terms of the involvement of multiple biological systems, numbering ∼100 proteins suggesting potent, systemic alterations in plasma protein composition during inflammation. 14 Proteins positively associated with AGP are largely involved in host defense mechanisms in response to stress, including proteolytic attack processes, transport of pro-oxidative metabolites, and immune activation. Interestingly, proteins negatively associated with AGP include numerous extracellular matrix proteins that were found to be associated with mid-upper arm muscle area, an anthropometric indicator of lean body mass, as well as nutrient and hormone transporters, plausibly indicating degradation of muscle tissues and redistribution of nutrients and hormones during phases of inflammation. A similar, though less extensive, proteome was associated with C-reactive protein. 14 Substantial alteration in the plasma proteome composition during inflammation may illustrate the body’s metabolic priorities to first-line host defense mechanisms at the expense of suppressed nutrient transport and anabolic processes that may be critical for child growth.
Cognition and Inflammation
Health effects of chronic inflammation may extend to cognitive impairment. 15 Proteins that were found to be positively and negatively associated with intellectual function, evaluated by a measure of nonverbal general intelligence a year after blood sample collection, are known to be involved in diverse biological processes, including lipid/cholesterol transport (apolipoproteins), insulin-like growth factor 1 (IGF-1) regulation (IGF-1 binding proteins), thyroid hormone transport (transthyretin), and inflammatory response (acute phase proteins and complement factors). After adjusting for known risk factors for impaired development in poor societies (eg, child nutritional status, household wealth and maternal education), only negative associations with proteins largely involved in inflammation remained significant. Although we were unable to detect direct mediators of child cognitive function, the identified proteins imply that low-grade, prolonged systemic inflammation may disturb cognitive developmental processes of children.
Attained Child Growth and Body Composition
Associations with height-for-age (HAZ) and weight-for-age (WAZ) z-scores in Nepalese children derived from the World Health Organization (WHO) reference, as well as arm muscle area, reveal that circulating proteins may reflect underlying biological processes of somatic growth. 16 The biological functions in which the identified proteins are mainly involved are summarized in Table 1. The most remarkable finding is that IGF-1 and its binding proteins were commonly and strongly associated with HAZ, WAZ, and arm muscle area. The result likely reflects the essential roles of the IGF-1 axis in child bone and muscle growth and supports an ability of plasma proteomics to detect biologically insightful correlates of lean mass growth. For example, proteins uniquely associated with HAZ suggest that specific biological pathways such as nutrient transport (ie, afamin [AFM]), bone mineralization (tetranectin [CLEC3B]), and vascular development (vasorin [VASN]) may sustainably contribute to bone elongation, while subclinical activation of the immune system (represented by S100A12) may delay skeletal growth. A distinctive network of structural or binding components of extracellular matrix represents the proteome of arm muscle mass. The extracellular matrix is a major structural framework of connective tissue and a body protein reservoir that plays numerous roles in muscle tissue remodeling and host response to injury. 17
Proteomics-Based Plasma Proteins Associated Anthropometric Indicators and Their Representative Biological Functions in School-Aged Nepalese Childrena,b
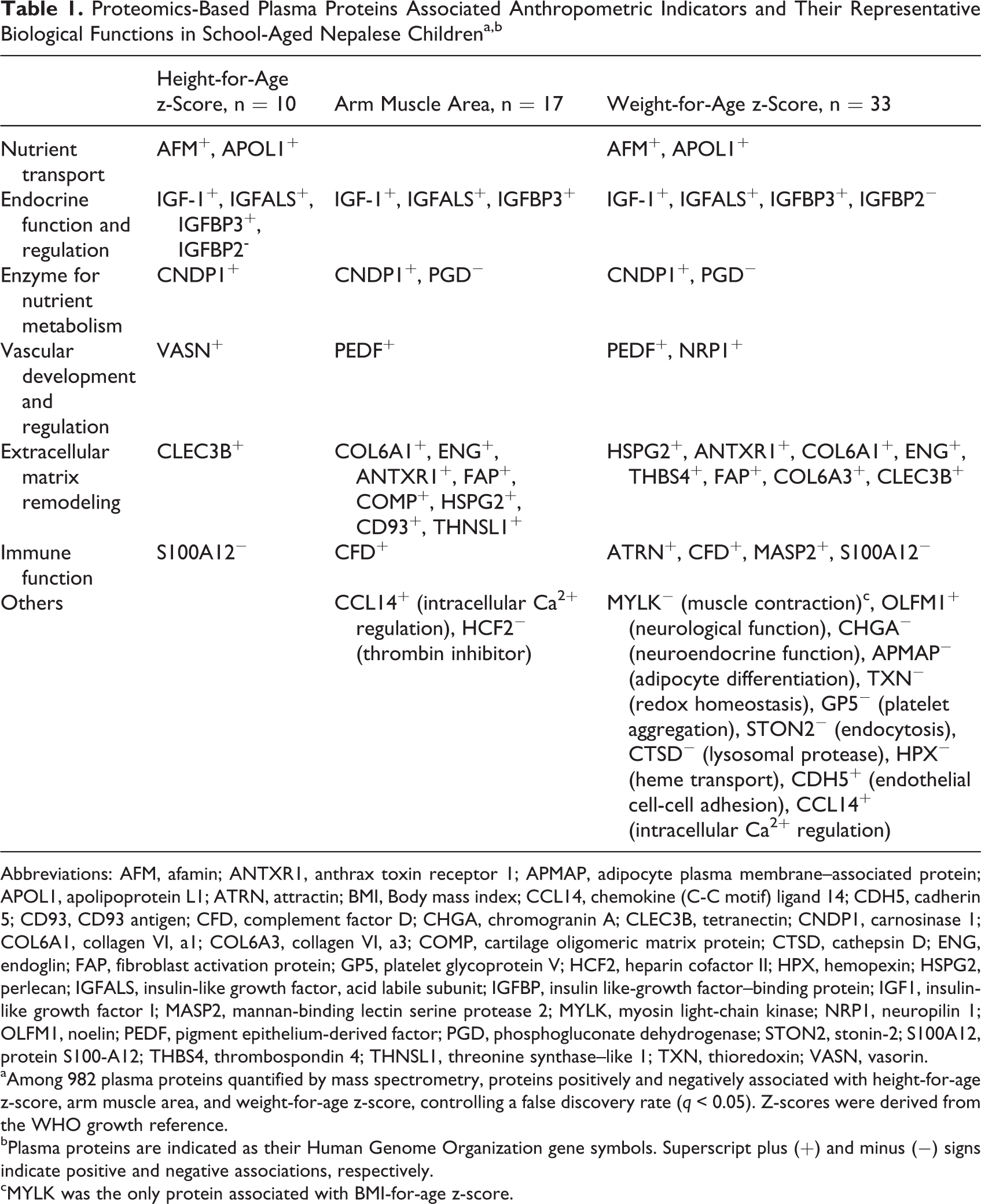
Abbreviations: AFM, afamin; ANTXR1, anthrax toxin receptor 1; APMAP, adipocyte plasma membrane–associated protein; APOL1, apolipoprotein L1; ATRN, attractin; BMI, Body mass index; CCL14, chemokine (C-C motif) ligand 14; CDH5, cadherin 5; CD93, CD93 antigen; CFD, complement factor D; CHGA, chromogranin A; CLEC3B, tetranectin; CNDP1, carnosinase 1; COL6A1, collagen VI, a1; COL6A3, collagen VI, a3; COMP, cartilage oligomeric matrix protein; CTSD, cathepsin D; ENG, endoglin; FAP, fibroblast activation protein; GP5, platelet glycoprotein V; HCF2, heparin cofactor II; HPX, hemopexin; HSPG2, perlecan; IGFALS, insulin-like growth factor, acid labile subunit; IGFBP, insulin like-growth factor–binding protein; IGF1, insulin-like growth factor I; MASP2, mannan-binding lectin serine protease 2; MYLK, myosin light-chain kinase; NRP1, neuropilin 1; OLFM1, noelin; PEDF, pigment epithelium-derived factor; PGD, phosphogluconate dehydrogenase; STON2, stonin-2; S100A12, protein S100-A12; THBS4, thrombospondin 4; THNSL1, threonine synthase–like 1; TXN, thioredoxin; VASN, vasorin.
aAmong 982 plasma proteins quantified by mass spectrometry, proteins positively and negatively associated with height-for-age z-score, arm muscle area, and weight-for-age z-score, controlling a false discovery rate (q < 0.05). Z-scores were derived from the WHO growth reference.
bPlasma proteins are indicated as their Human Genome Organization gene symbols. Superscript plus (+) and minus (−) signs indicate positive and negative associations, respectively.
cMYLK was the only protein associated with BMI-for-age z-score.
In these Nepalese children, who present with a lean phenotype, it is noteworthy that no proteins were associated with fat mass as indicated by mid-upper arm fat area and triceps and subscapular skinfolds. 16 The only protein associated with BMI-for-age z-score (BMIZ) was myosin light-chain kinase (MYLK), known to play a prominent role in muscle contraction. The negative association between MYLK and BMIZ suggests that this protein may leak from muscle tissue due to muscle wasting or muscle tissue damage. As a composite measurement, the proteome of child weight comprises the sum of proteins associated with child stature and muscle mass as well as proteins of diverse functions, such as neurological processes (noelin [OLFM1]) and cellular response to catabolic stress (thioredoxin [TXN]). Although we used indirect measurements of body dimension and composition, the identified proteomes of child linear growth and body composition reflect the complex interplay among multiple biological pathways and systems (nutritional, endocrine, immune, and musculoskeletal) that cannot be discerned from anthropometry.
Proteome Persistence: Small Head Size at Birth
As a birth cohort, we have been able to explore the existence of sustained proteomic patterns throughout early childhood that may reflect fetal growth, indicated by measurements of weight, length, and head circumference taken within ∼72 hours after birth. 5 A novel cluster of more than 20 cytoskeletal proteins were 8% to 20% less abundant in children with a small head size at birth (<−2 z-scores of the WHO growth standard 18 ) compared to children born with a normal head size (≥−2 z-scores), after adjusting for potential confounders. 19 Interestingly, no proteins differed by other indicators of poor fetal growth such as weight, length, or weight-for-length at birth. The clinical significance of reduced circulating abundance of proteins responsible for intracellular structure and cell motility in children with a restricted head size at birth is unclear. However, this result illustrates the potential for the childhood plasma proteome to reflect sustained consequences of restricted tissue growth or development during fetal life.
Way Forward: Transforming Biomarker Discovery to Public Health Application
A detectable plasma proteome has been shown to have the potential to estimate population plasma micronutrient and inflammatory status and reflect the pathobiology of poor growth and impaired cognition among Nepalese children. There is an urgent need to replicate and expand these findings in other life stages and population settings of varied nutritional and health status and assess the responsiveness of the plasma proteome to nutrition interventions. To be of public health service, however, proteins of predictive utility need to be migrated from mass spectrometry for discovery onto field-friendly, high-throughput, low cost, targeted instrumentation. Such a platform would allow for the concurrent assessment of multiple biomarkers that could comprehensively profile the nutritional and health state of populations. The platform would minimize plasma sample volume, technical requirements, assay costs, and delays in analyses. This transformation will require multiple phases, including more expansive discovery, verification and development of biomarkers, and continued transdisciplinary efforts across diverse stakeholders, including epidemiologists, trialists, the biotechnology community, and funders. Further, opportunities are abound for integrating multiple omics platforms to expand nutritional, metabolic, homeostatic, and disease risk profiles for public health application in the future. 20
Footnotes
Acknowledgments
The authors are grateful to core members of the Johns Hopkins Nutriproteomics Research Team, including Parul Christian, Robert Cole, Lauren DeVine, John Groopman, Shelly Herbrich, Bareng AS Nonyane, Robert O’Meally, Ingo Ruczinski, Lee Shu-Fune Wu and James Yager. The authors thank the Bill & Melinda Gates Foundation for its support of the original antenatal micronutrient trial and the 7-year postpartum follow-up study (GH614, Ellen Piwoz), and the nutriproteomics research reported here (OPP5241, Yiwu He). Our work has been further supported by the Sight and Life Global Nutrition Research Institute (Klaus Kraemer), Baltimore, MD, Royal DSM Ltd, Kaiseraugst, Switzerland, and through a Faculty Innovation Award from the Johns Hopkins Bloomberg School of Public Health, Baltimore, MD. Finally, the authors are thankful to the Hildegard Grunow Foundation for the Rainer Gross Award and the support for the publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
