Abstract
Background
Epidemiological studies have reported a comorbid relationship between migraine and thyroid dysfunction.
Methods
We investigated the genetic relationship between migraine and thyroid function traits using genome-wide association study (GWAS) data.
Results
We found a significant genetic correlation (rg) with migraine for hypothyroidism (rg = 0.0608), secondary hypothyroidism (rg = 0.195), free thyroxine (fT4) (rg = 0.0772), and hyperthyroidism (rg = –0.1046), but not thyroid stimulating hormone (TSH). Pairwise GWAS analysis revealed two shared loci with TSH and 11 shared loci with fT4. Cross-trait GWAS meta-analysis of migraine identified novel genome-wide significant loci: 17 with hypothyroidism, one with hyperthyroidism, five with secondary hypothyroidism, eight with TSH, and 15 with fT4. Of the genes at these loci, six (RERE, TGFB2, APLF, SLC9B1, SGTB, BTBD16; migraine + hypothyroidism), three (GADD45A, PFDN1, RSPH6A; migraine + TSH), and three (SSBP3, BRD3, TEF; migraine + fT4) were significant in our gene-based analysis (pFisher’s combined P-value < 2.04 × 10−6). In addition, causal analyses suggested a negative causal relationship between migraine and hyperthyroidism (p = 8.90 × 10−3) and a positive causal relationship between migraine and secondary hypothyroidism (p = 1.30 × 10−3).
Conclusion
These findings provide strong evidence for genetic correlation and suggest complex causal relationships between migraine and thyroid traits.
Keywords
Introduction
Migraine has been linked with several comorbidities including neurological, psychiatric, cardiovascular, and endocrine disorders. These comorbid conditions are known to complicate the clinical presentation and treatment of migraine (1). Previous epidemiological and case-control studies investigating the association of migraine with thyroid dysfunction have indicated that thyroid dysfunction causes migraine and vice versa (2–5). The increased prevalence and risk of thyroid dysfunction (2 to 6% globally) has consequences on quality of life (6). Studies have reported that hyperthyroidism is associated with an increased risk of cardiovascular diseases (CVD) (hazard ratio [HR] = 3.39) (7) and breast cancer (HR = 2.04) (8). Hypothyroidism has been reported to be associated with an increased risk of diabetes mellitus (HR = 1.58) and also CVD (HR = 1.20) (8,9).
The most common thyroid dysfunction traits are hypothyroidism and hyperthyroidism. Both these traits are differentiated based on varying levels of thyroid stimulating hormone (TSH) and free thyroxine (fT4) and share a complex inverse relationship. Pathological processes within the thyroid gland result in hypothyroidism (high TSH and low fT4) and hyperthyroidism (low TSH and high fT4), however rarely, processes arising from the hypothalamus or pituitary result in secondary hypothyroidism (low fT4 with inappropriately low TSH) (6,10). These thyroid dysfunction traits are suggested to have health consequences among migraine cases. Studies have reported a relationship between hypothyroidism and migraine (2,11,12). For example, a case-control study reported hypothyroidism incidence to be 10.8% in the migraine population and 6.2% in the general population with a hazard ratio of 1.411 (95% CI = 1.01–1.97). The study also found migraine cases to have a 41% increased risk of developing hypothyroidism (12).
Despite evidence from observational epidemiological studies, a clear interpretation of their increased co-occurrence is lacking, including whether genetic factors are involved in their comorbidity. Thus in the present study, we utilise genome-wide association studies (GWAS) summary statistics to investigate the genetic overlap between migraine and the thyroid traits of hypothyroidism, hyperthyroidism, secondary hypothyroidism, TSH, and fT4, at the genome-wide and regional locus level. We also used genetic approaches including cross-trait meta-analysis and gene-level genetic overlap, to identify shared genetic components and related pathophysiology between migraine and thyroid traits. Finally, we use Mendelian randomisation (MR) to look for causal relationships between migraine and thyroid traits.
Materials and methods
GWAS summary statistics for migraine and thyroid traits
We utilised the GWAS summary statistics from the latest migraine GWAS by Hautakangas et al. 2022 study (13). The GWAS summary statistics for thyroid traits were obtained from PANUK Biobank for hypothyroidism, hyperthyroidism, and secondary hypothyroidism, while TSH and fT4 were obtained from publicly available data from the Teumer et al. 2018 study (14).
Female- and male-specific GWAS summary statistics for migraine were obtained from the Anttila et al. 2013 study (15), while for TSH and fT4 were obtained from the Teumer et al. study (14). Sex-specific GWAS summary statistics were not available for hypothyroidism, hyperthyroidism, and secondary hypothyroidism. The details of cases and controls for all traits are provided in the online Supplementary Table 1. All participants in the present study were of European ancestry.
Genetic correlation
To estimate single nucleotide polymorphism (SNP)-based heritability between migraine and thyroid traits and to assess genetic correlation (rg) between them, we performed cross-trait linkage-disequilibrium (LD) score regression (LDSC) (https://github.com/bulik/ldsc) (16). LDSC analyses were performed on the liability scale and utilised the software’s pre-calculated European LD scores from the 1000 Genomes Project EUR reference sample. Because of overlapping UK Biobank samples between migraine and hypothyroidism, hyperthyroidism, and secondary hypothyroidism, we performed both constrained and unconstrained (genetic covariance intercept) genetic correlation analyses.
Pairwise GWAS analysis
We utilised a Bayesian pleiotropy association test implemented in the pairwise-GWAS (GWAS-PW) (https://github.com/joepickrell/gwas-pw) software to identify shared genomic regions influencing migraine and thyroid traits (17). It estimates local (regional) genetic correlations and posterior probabilities of association (PPA) of genomic regions shared across migraine and thyroid traits under four defined models: (i) association with migraine only (PPA1); (ii) association with thyroid trait only (PPA2); (iii) shared association with both migraine and thyroid trait via a SNP (PPA3); shared association with both migraine and thyroid trait but via two distinct SNPs (PPA4). Further information is provided in the online Supplementary Methods. To estimate the genetic overlap, we considered PPA3 > 0.5 to identify pleiotropic regions across migraine and thyroid traits via a shared SNP.
Cross-disorder GWAS meta-analysis and its characterisation
We conducted a cross-trait GWAS meta-analysis of migraine and thyroid traits to identify shared loci using the METASOFT software (http://genetics.cs.ucla.edu/meta) inverse variance-weighted (IVW) fixed effect (FE) model (18). To allow for heterogeneity effects across traits, the METASOFT Han and Eskin’s random-effects (RE2) model was also utilised (18). A SNP p-value < 5 × 10−8 was considered to be genome-wide significant (GWS), while p-value < 0.05 was considered to be nominally significant. For SNPs and loci to be considered novel, they should have a meta-analysed GWS p-value (pmeta-analysed < 5 × 10−8) but should not be GWS in either of the two traits (5 × 10−8 < pmigraine/thyroid traits < 0.05).
Meta-analysis results were annotated using the FUMA software (https://fuma.ctglab.nl/) to characterise the SNPs and loci associated with migraine and thyroid traits (19). The meta-analysed SNPs were first filtered based on their association with migraine and thyroid traits, i.e., with p-values for both traits in the range of 5 × 10−8 < pmigraine/thyroid traits < 0.05. The remaining GWS SNPs were used as input to FUMA. FUMA then performed LD clumping to identify regions containing SNPs that are correlated (in LD) with each other at r2 < 0.6, and independent lead SNPs that are not in LD with each other at r2 < 0.1. A comprehensive description of the steps followed to identify the novel loci are in online Supplementary Methods.
Mendelian randomisation between migraine and thyroid traits
To explore the causal relationship between migraine and thyroid traits, we performed two-sample Mendelian randomisation (2SMR) analysis using the R statistical package (https://mrcieu.github.io/TwoSampleMR/) (20). First, we tested for a causal effect of each thyroid trait (exposure) on migraine (outcome). Second, we performed reverse MR analyses to test for a causal relationship between genetic risk for migraine (exposure) and each thyroid trait (outcome). For 2SMR analysis, LD-independent GWS SNPs at p-value < 5 × 10−8 (r2 < 0.1) were utilised as instrumental variables (IVs). We performed 2SMR analyses using the IVW, the weighted median, and the MR-Egger method (21,22). Additionally, to test for horizontal pleiotropy and possible outliers, we conducted Mendelian randomisation pleiotropy residual sum and outlier (MR-PRESSO) analysis (23). To further validate our results, we performed Cochran’s Q statistics for heterogeneity test, Wald ratio for single SNP MR analysis and IVW for ‘leave-one-out’ analysis.
Another MR analysis using a bi-directional Generalised Summary-data based Mendelian randomisation (GSMR) was performed using GSMR software (24). This analysis estimates the effect of LD-independent GWS SNPs on summary statistics of a thyroid trait (bzx) and migraine (bzy) to estimate the causal association (bxy) between a thyroid trait (exposure) and migraine (outcome), and vice-versa. To exclude putative pleiotropic SNPs from the analysis, the GSMR package also uses the HEIDI outlier method (HEIDI-outlier p-value < 0.01) (24). The LD-independent GWS SNPs for migraine and thyroid traits were selected at a p-value < 5 × 10−8 (r2 < 0.1), except for secondary hypothyroidism for which the threshold was lowered to a p-value < 1 × 10−5 (r2 < 0.1) due to the small number of GWS SNPs.
Also, to test for a causal relationship at the genome-wide level we performed latent causal variable (LCV) analysis which estimates the genetic causality proportion (gcp) of each thyroid trait on migraine (25). It is based on a latent variable mediating the genetic correlation between the two traits, where a gcp of zero would mean no genetic causality and a gcp of one would mean full genetic causality. The LCV analysis was performed on the GWAS summary statistics of each trait and pre-calculated LD scores from 1000G accompanying the LDSC software.
Gene-based association study
To further confirm our SNP-level genetic overlap analysis and identify shared genes across migraine and thyroid traits, we performed a gene-based association analysis. Our gene-based analysis was conducted using the GATES test (26) implemented in the Fast ASsociation Tests (FAST) (27) package. The common SNPs between migraine and thyroid traits were assigned to 34,212 genes from NCBI. More details are provided in online Supplementary Methods. The output provides us with the best significant SNP assigned to a gene with their respective p-values. We note that neighbouring genes may have correlated results due to LD between the topmost significant SNP assigned to each gene.
Independent gene-based test
We performed independent gene-based analysis using the ‘genetic type I error calculator’ GEC software (28) as implemented in previous studies (29,30) to estimate the effective number of independent genes across migraine and thyroid traits. The best SNPs assigned to genes across migraine and thyroid traits GWAS were used as input for GEC analysis. GEC was utilised (i) to overcome the potential for correlation across neighbouring gene-based association results; and (ii) to generate unbiased data for assessing the gene-level genetic overlap between migraine and thyroid traits.
Gene-level genetic overlap
Analysing the effective number of independent genes obtained using independent gene-based association results, we assessed whether the proportions of genes overlapping migraine and thyroid traits, at three nominal p-value thresholds (p < 0.1, p < 0.05, p < 0.01), were more than expected by chance (see online Supplementary Methods).
A significant binomial test p-value indicates that the observed number of overlapping genes was more than expected by chance.
To identify individual genes associated with migraine and thyroid traits at a p < 0.05, we combined the gene-based p-value for the respective traits using the Fisher’s combined P-value method (FCP) (29,30). We then assessed the shared genes for both migraine and thyroid traits reaching genome-wide significance based on our FCP results.
Pathway analysis
To identify potential biological mechanisms and pathways related to the overlapping genes of migraine and thyroid traits, we performed functional enrichment analysis using ‘g:GOst’ tool implemented in the g:Profiler software (http://biit.cs.ut.ee/gprofiler/) (31). The tool covers data sources including Gene Ontology, Reactome, WikiPathways, Kyoto Encyclopedia of Genes (KEGG), Human Protein Atlas, CORUM, and Human Phenotype Ontology, updated on a regular basis (31). In the present study to identify shared pathways and mechanisms, we utilised genes overlapping migraine and thyroid traits at p < 0.05 (FCP < 1 × 10−4). We applied the recommended ‘g:SCS algorithm’ to adjust for multiple testing and restricted the term size of the pathways to 5 and 350. In case the enriched pathways were biased due to the presence of LD across the neighbouring genes, we ensured that the top SNPs of the genes in any respective pathway were not in LD (r2 < 0.1).
Results
Univariate SNP-based heritability across traits
The SNP-based heritability (h2SNP) estimates from LDSC analysis of the migraine and thyroid traits GWAS summary statistics are reported in online Supplementary Table 1. Migraine and all five thyroid traits were found to have significant (p < 0.05) h2SNP and were therefore utilised in subsequent cross-trait genetic analyses.
Genome-wide genetic overlap between migraine and thyroid traits
Using LDSC, we found a significant positive genetic correlation between migraine and hypothyroidism (rg = 0.0608, p = 5 × 10−4), migraine and secondary hypothyroidism (rg = 0.195, p = 3.8 × 10−3), and migraine and fT4 (rg = 0.0772, p = 3 × 10−4). We also found a significant negative genetic correlation between migraine and hyperthyroidism (rg = –0.1046, p = 2.8 × 10−3). However, we did not find a significant correlation between migraine and TSH (rg = –0.0104, p = 0.6801). LDSC indicated a small amount of sample overlap for migraine with hypothyroidism, hyperthyroidism, and secondary hypothyroidism, although the genecov intercept was very close to zero (–0.0032, –0.0072, and –0.0091, respectively) and when constrained to zero had a negligible impact on the results. Detailed SNP-based genetic correlation results are provided in Table 1.
Genetic correlation between migraine and thyroid traits.

h2, heritability of the traits; valid SNPs, number of SNPs used in estimating genetic correlation; rg, genetic correlation; SE, standard error; Z, Z-score; p, p-value; Intercept t1(SE)/t2(SE), LD score regression intercept for trait 1 (SE) and trait 2 (SE); Intercept (genecov), LD score regression intercept on genetic covariance scale.
LDSC analysis of the female- and male-specific datasets did not find a significant genetic correlation between migraine and TSH, or migraine and fT4 (online Supplementary Table 2).
Pleiotropic variants between migraine and thyroid traits
We used the GWAS-PW approach to identify pleiotropic loci associated with both the migraine and thyroid traits GWAS. A pleiotropic locus in a region identifies SNPs associated with both traits regardless of the SNPs’ direction of effect. In contrast to the LDSC genome-wide genetic correlation results, applying GWAS-PW analysis to trait combinations of migraine with hypothyroidism, hyperthyroidism, and secondary hypothyroidism, all 1703 of the tested genomic regions had a PPA4 > 0.9 and PPA3 < 0.5, indicating no genomic region contained a SNP that was strongly associated with both migraine and the thyroid traits.
When applying the GWAS-PW approach to migraine and TSH, although the LDSC genome-wide genetic correlation was not significant, we identified significant local genetic correlation at two loci (PPA3 > 0.5) (Table 2): one on chromosome 9 at 135.3–137 Mb (lead SNP: rs8176645; associated with migraine risk and high TSH levels) and one on chromosome 2 at 43.3–44.3 Mb (lead SNP: rs12712881; associated with migraine risk and lower TSH levels). For migraine and fT4, we identified GWS local genetic correlation at 11 loci across ten chromosomes (PPA3 > 0.5) (Table 3). When examining the direction of the effect allele of these 11 index SNPs at the pleiotropic loci in migraine and fT4, the effect allele for nine index SNPs were associated with increased fT4 levels being associated with an increased risk of migraine (Table 3). Tables 2 and 3 list the SNPs that are significant for migraine and TSH, and migraine and fT4 within the implicated genomic regions.
Pleiotropic loci with top significant SNPs influencing migraine and TSH identified by GWAS-PW.

Chunk, ID representing the LD region; NSNP, number of SNP in the LD region; chr, chromosome; st, start position of the LD region; sp, stop position of the LD region; PPA_1, estimated posterior probability of model 1 (locus affecting only migraine); PPA_2, estimated posterior probability of model 2 (locus affecting only TSH); PPA_3, estimated posterior probability of model 3 (locus affecting both migraine and TSH via a single SNP); PPA_4, estimated posterior probability of model 4 (locus affecting both migraine and TSH via two different SNPs); SNP, single nucleotide polymorphism; EA, effect allele; NEA, non-effect allele; Beta, effect of association; p, p-value; LD, linkage disequilibrium.
Pleiotropic loci with top significant SNPs influencing migraine and fT4 identified by GWAS-PW.

Chunk, ID representing the LD region; NSNP, number of SNP in the LD region; chr, chromosome; st, start position of the LD region; sp, stop position of the LD region; PPA_1, estimated posterior probability of model 1 (locus affecting only migraine); PPA_2, estimated posterior probability of model 2 (locus affecting only fT4); PPA_3, estimated posterior probability of model 3 (locus affecting both migraine and fT4 via a single SNP); PPA_4, estimated posterior probability of model 4 (locus affecting both migraine and fT4 via two different SNPs); SNP, single nucleotide polymorphism; EA, effect allele; NEA, non-effect allele; Beta, effect size of EA; p, p-value; LD, linkage disequilibrium.
Shared loci between migraine and thyroid traits
We conducted genome-wide cross-trait meta-analyses to identify loci that may share association with migraine and each thyroid trait using the METASOFT software. We selected SNPs with a meta-analysed p-value < 5 × 10−8 and trait-specific 5 × 10−8 < pmigraine, thyroid trait < 0.05.
Applying the approach detailed in online Supplementary Methods identified 17, one, five, eight, and 15 novel GWS loci from the meta-analysis of migraine and hypothyroidism, migraine and hyperthyroidism, migraine and secondary hypothyroidism, migraine and TSH, and migraine and fT4, respectively (Table 4).
Genome-wide significant SNPs from cross-trait meta-analysis between migraine and thyroid traits.

SNP, single nucleotide polymorphism; EA, effect allele; NEA, non-effect allele; chr, chromosome number; pos, position of SNP; p, p-value; Beta, effect size of EA; SE, standard error of beta; HPO, hypothyroidism; HPER, hyperthyroidism, SHPO, secondary hypothyroidism; TSH, thyroid stimulating hormone; fT4, free thyroxine.
Overall, among the novel migraine loci, rs3795310 on chromosome 1 had the strongest significance for hypothyroidism (pmeta = 2.42 × 10−10). The second strongest overall signal was rs5017041 at chromosome 10 with hypothyroidism (pmeta = 1.67 × 10−9). We found one overlapping gene, PFDN1 at chromosome 5, associated with migraine and TSH (lead SNP: rs6870103, pmeta = 4.55 × 10−10) as well as with migraine and fT4 (lead SNP: rs6897346, pmeta = 4.19 × 10−8).
Causal effect of migraine on thyroid traits
We conducted 2SMR, GSMR and LCV analyses to investigate potential causal relationships between migraine and each thyroid trait.
When testing for a causal effect of each thyroid trait on migraine, we utilised 211, 21, 28, 56, and 24 independent GWS SNPs as IVs for hypothyroidism, hyperthyroidism, secondary hypothyroidism, TSH, and fT4, respectively. 2SMR analysis only found some weak nominally significant evidence for a causal effect of hyperthyroidism on migraine with MR-Egger (β = –0.04, p = 3.30 × 10−2) and GSMR found weak evidence for a causal effect of fT4 on migraine (β = 0.04, p = 4.50 × 10−2) (Table 5a and 5b).
MR results for migraine and thyroid traits.
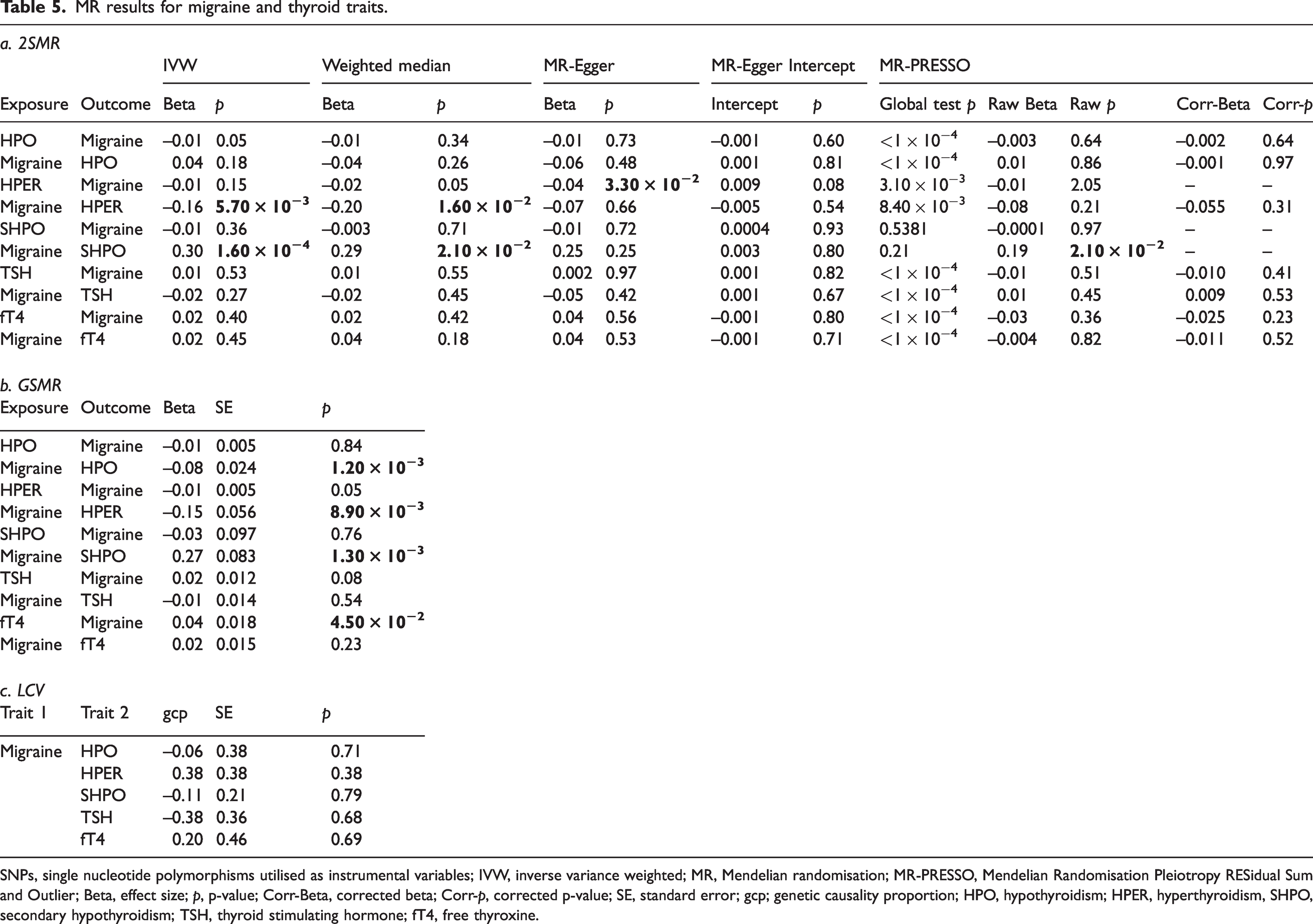
SNPs, single nucleotide polymorphisms utilised as instrumental variables; IVW, inverse variance weighted; MR, Mendelian randomisation; MR-PRESSO, Mendelian Randomisation Pleiotropy RESidual Sum and Outlier; Beta, effect size; p, p-value; Corr-Beta, corrected beta; Corr-p, corrected p-value; SE, standard error; gcp; genetic causality proportion; HPO, hypothyroidism; HPER, hyperthyroidism, SHPO, secondary hypothyroidism; TSH, thyroid stimulating hormone; fT4, free thyroxine.
In contrast, reverse analyses testing for a causal relationship between genetic risk for migraine and each thyroid trait, we utilised 158 independent GWS SNPs as IVs for migraine. 2SMR found a significant causal effect of migraine on hyperthyroidism with IVW (β = –0.16, p = 5.70 × 10−3) and weighted median (β = –0.20, p = 1.6 × 10−2), but not with MR-Egger (β = –0.07, p = 0.66) and MR-PRESSO (β = –0.07, p = 0.21). We also found a significant causal effect of migraine on secondary hypothyroidism with IVW (β = 0.30, p = 1.60 × 10−4), weighted median (β = 0.29, p = 2.1 × 10−2), and MR-PRESSO (β = 0.19, p = 2.10 × 10−2); but not with MR-Egger (β = 0.25, p = 0.25). The MR-Egger intercepts for both migraine on hyperthyroidism (intercept = –0.005, p = 0.54) and migraine on secondary hypothyroidism (intercept = 0.003, p = 0.80) did not deviate significantly from zero, suggesting no evidence for pleiotropy. Similarly, reverse GSMR analyses found a significant negative causal relationship between migraine and hyperthyroidism (β = –0.146, p = 8.90 × 10−3) and a significant positive causal relationship between migraine and secondary hypothyroidism (β = 0.27, p = 1.30 × 10−3). Interestingly, the reverse GSMR analyses also found a significant negative causal relationship between migraine and hypothyroidism (β = –0.080, p = 1.20 × 10−3), but found no evidence for a causal effect of migraine on TSH or fT4 (Table 5a and b).
LCV analyses found no evidence for genome-wide causality between migraine and any thyroid trait (Table 5c).
Gene-level genetic overlap
We performed gene-level genetic overlap analysis to identify and assess the proportion of associated genes overlapping the migraine and each thyroid trait GWAS. To determine if the proportion of overlapping genes was more than expected by chance, we performed binomial tests for genes associated at three p-value thresholds (Table 6, online Supplementary Tables 3–6). For instance, when analyzing migraine with hypothyroidism at pgene < 0.05, the observed proportion (0.245) of overlapping associated genes was significantly higher (pbinomial = 1.68 × 10−8) than the expected proportion (0.175) of the overlapping genes. A similar pattern of results was observed at the two other thresholds (pgene < 0.01 and pgene < 0.1).
Independent gene-based association analysis and gene-based genetic overlap between migraine and hypothyroidism.

1Migraine dataset obtained from Hautakangas et al. (13), 2Hypothyroidism dataset obtained from PANUK Biobank Neale lab, 3Raw number of genes (total number of genes obtained in the gene-based association analysis using GATES software), 4Effective number of independent genes (the total number of independent genes obtained in the independent gene-based test using the ‘genetic type 1 error calculator’ method), 5Proportion of the total effective number of independent genes.
A gene was considered to be GWS at FCP pgene < 2.04 × 10−6 adjusted for testing the estimated maximum number of independent gene-based tests (0.05/24,562). All genes overlapping migraine and each thyroid trait GWAS at FCP pgene < 2.04 × 10−6 are reported in online Supplementary Tables 7–11. Combining gene-based tests for association across migraine and each thyroid trait using the FCP method, we identified 851 unique genes showing evidence of being shared by both traits, of which 205 genes have previously been reported to be associated with migraine or the thyroid traits GWAS used in the present study.
These results provide evidence for a significant gene-level genetic overlap between migraine and each thyroid trait and identify novel genes associated with these genetically correlated traits.
Pathway analysis of overlapping genes
We performed pathway analysis of overlapping genes associated with migraine and each thyroid trait at a p < 0.05 threshold using the g:GOst tool implemented in g:Profiler software. After retaining the most significant gene within sets of correlated genes with their top significant SNPs in LD (r2 < 0.1), pathway analysis was performed for 655, 172, 69, 155, and 166 genes associated with pFCP < 1 × 10−4 overlapping migraine with hypothyroidism, hyperthyroidism, secondary hypothyroidism, TSH, and fT4, respectively. The pathway analyses identified 37, 55, 32, ten, and two biological pathways/processes enriched for the overlapping genes (Supplementary Tables 12–16). Some significantly enriched pathways include TAP/TAP1 binding, allograft rejection, Staphylococcus aureus infection, systemic lupus erythematosus (SLE), type 1 diabetes mellitus (T1D), rheumatoid arthritis (RA), and activation of complement component 3 (C3) and C5 proteins of the immune system.
Discussion
In the last decade, the relationship between migraine and thyroid traits has been studied using observational epidemiological and cross-sectional association studies. More recently GWAS have been performed to identify genetic factors associated with these traits. These studies have reported an increased co-occurrence of migraine and thyroid traits and identified SNPs and genes associated individually with migraine or thyroid traits. Here we perform the first known genetic study to investigate the genetic and causal relationship between migraine and thyroid traits. Utilising GWAS summary statistics, we found a significant genetic correlation between migraine and hypothyroidism, hyperthyroidism, secondary hypothyroidism, and fT4; and also shared pleiotropic regions between migraine and TSH, and migraine and fT4. Furthermore, cross-trait meta-analysis and gene-level overlap analysis identified a shared genetic basis underlying migraine and thyroid traits. Testing for a causal association, we found a significant causal relationship between migraine and hyperthyroidism, and migraine and secondary hypothyroidism.
LDSC tests for genome-wide genetic correlation by quantifying the average sharing of genetic effects between two traits (32). The positive genetic correlation observed between migraine and hypothyroidism is consistent with several previous cross-sectional case-control studies indicating similar conclusions (2,3,12). The observed positive genetic correlation between migraine and secondary hypothyroidism has not been studied previously, although it could relate to (be explained by) a recent finding which suggested that increasing loss of hypothalamic control (over the hypothalamo-limbic connection) can result in an increased susceptibility to a migraine attack (33). The positive genetic correlation observed between migraine and fT4, is not consistent with the existing clinical data, for example, a study suggested migraine to be associated with low thyroid hormone levels (34). Inconsistent with the inverse relationship between levels of TSH and fT4, the positive genetic correlation observed for hypothyroidism and fT4 with migraine risk suggests their relationship is driven by (poly)genetic factors across the genome. In contrast to our findings with hypothyroidism, the negative genetic correlation observed between migraine and TSH could be supported by a previous study that observed a low level of serum TSH was associated with prolonged migraine attacks, indicating an inverse relationship between TSH levels and migraine (4). However, both TSH and fT4 exhibit a complex relationship and their correlation with common conditions is frequently broken and sometimes inverted, as observed in our results (35). Our finding of a significant genetic correlation between migraine and thyroid traits suggests that this observed association between migraine and thyroid traits is due, in part, to shared genetic factors.
We next identified pleiotropic loci shared across migraine and thyroid traits and identified two pleiotropic regions across migraine and TSH, and 11 pleiotropic regions across migraine and fT4. The top three pleiotropic loci were found on chromosome 9 at 135.3–137 Mb (migraine and TSH), chromosome 6 at 31.0–31.6 Mb (migraine and fT4), and chromosome 17 at 15.0–16.4 Mb (migraine and fT4).
The novel loci identified in our cross-trait GWAS meta-analysis mapped to genes including RERE, BTBD16, HTT, TGFB2, PFDN1, ATXN2, BRD3, and SLC14A2. RERE (Arginine-Glutamic Acid Dipeptide Repeats) (lead SNP: rs3795310), a protein coding gene in the proximal 1p36 region, encodes for a nuclear receptor coregulator that positively regulates retinoic acid signalling (36). Although minimal data exists for RERE, its deficiency has been related to lesions of the corpus callosum and previously associated with migraine with aura and hyperthyroidism (37,38). BTBD16 (lead SNP: rs5017041) encodes a protein containing a BTB/POZ domain mediating protein-protein interactions. RERE mutations have been reported with major depressive disorder (MDD) and mood disorder (39), while BTBD16 mutations have been reported with bipolar disorder (40), which have been associated with both migraine and hypothyroidism. This evidence suggests that the novel variants rs3795310 and rs5017041 mapping to RERE and BTBD16 could have potential effects on migraine with hypothyroidism and hyperthyroidism, and migraine and hypothyroidism comorbidity, respectively. Also, PFDN1 on chromosome 5, was associated with both migraine and TSH (lead SNP: rs6870103) as well as with migraine and fT4 (lead SNP: rs6897346). This gene encodes a member of the prefoldin beta subunit family, which binds and stabilises newly synthesised polypeptides, allowing them to fold correctly. Although the exact role of these genes in migraine and thyroid traits is unknown, they are expressed in both brain and thyroid gland tissues.
Our gene-level analyses revealed a significant genetic overlap across migraine and the tested thyroid trait (pgene < 0.05). Of the genes mapping to novel loci identified from the cross-trait meta-analysis, six (RERE, TGFB2, APLF, SLC9B1, SGTB, and BTBD16; migraine and hypothyroidism), three (GADD45A, PFDN1, and RSPH6A; migraine and TSH) and three (SSBP3, BRD3, and TEF; migraine and fT4) were identified to be GWS in our gene-based analysis (pFCP < 2.04 × 10−6). TGFB2 (transforming growth factor beta 2) encoding for a secreted ligand of the TGFB family, is involved in the recruitment and activation of SMAD family transcription factors that regulate gene expression. Its signalling pathway is known to modulate repair processes and tissue development (41). Also, its receptor (TGFBR2) has been associated with vascular disease (42) and has been identified as a novel susceptibility gene in a clinic-based migraine GWAS (43). GADD45A (growth arrest and DNA-damage-inducible, alpha), encodes a protein that responds to environmental stress by activating p38/JNK pathways. p38/JNK pathways have been found to contribute towards initiation and maintenance of pain and inflammation (44), while GADD45A itself was recently prioritised as a functional candidate among the TSH GWAS loci (45). These shared genes reveal potential shared biological mechanisms for migraine and thyroid trait function.
Lastly, pathway analysis using overlapping genes for migraine and thyroid traits identified many biological pathways/processes. A large proportion of the overlapping genes in pathways across migraine and thyroid traits were related to the immune system, this suggests that both migraine and thyroid traits involve immune regulation. Recent reviews on the involvement of the immune system in migraine have noted clinical studies that have shown dysregulated immune systems among migraine cases (46) and that thyroid hormones and the immune system regulation are bidirectionally related (47). For example, a recent review supports a link between autoimmune/immunological diseases and an increased risk for migraine (48). Also, lowering of C3 with normal C4 and total complement activity has been reported among migraine cases thus supporting the involvement of the complement system (49).
A recent meta-analysis implied that the thyroid dysfunctional state of the thyroid gland and migraine pathogenesis could share a possible link (50), and looking at the genetic results in the current study using causal analysis, we found a stronger and significant negative causal relationship between migraine and hyperthyroidism and a significant positive causal relationship between migraine and secondary hypothyroidism suggesting that their association might be due to both shared molecular genetic mechanisms as well as causality. Although the underlying mechanism(s) remain unclear, a possible hypothesis is emerging for role of the hypothalamus in migraine and thyroid dysfunction comorbidity. Clinical studies have reported that the hypothalamus and limbic system are the attack-initiating brain structures during a migraine attack, where the induced migraine attack is associated with changes in hormonal status and menstrual cycle (51). A recent study supporting this hypothesis has shown that migraine cases are represented by neuro-endocrinological changes which lead to changes in the hypothalamically-regulated hormone status such as changes in levels of TSH, testosterone, and growth hormones. The pain transmitted to the hypothalamus during a migraine attack may contribute to the increase or decrease in the levels of TSH and fT4 (based on the individual set-point of the hypothalamus-pituitary-thyroid [HPT] axis) to cause thyroid dysfunction among migraine cases (51). This could relate to the mechanisms underlying the observed link between migraine and hyperthyroidism, and migraine and secondary hypothyroidism. The relationship between migraine and hypothyroidism could be consistent with biological pleiotropy rather than causality.
Studies evaluating the effect of normalising TSH levels have been reported to effectively reduce the severity and frequency of migraine attacks (52,53). The overall result of our genetic study suggests that in euthyroid, even a small difference or change in fT4 levels within the reference range is positively associated with migraine, however, in cases with established primary or secondary hypothyroidism with unphysiologically low fT4 below the reference range, or very high for hyperthyroidism, other mechanisms seem to be involved. As TSH is sensitive to minor changes in the fT4 levels, abnormal TSH levels are detected earlier than abnormal fT4 in thyroid dysfunction and our results indicate that fT4 levels have a significant relationship with migraine. Also because of the complex relationship observed with secondary hypothyroidism, fT4 levels should be measured in migraine patients to see if thyroid dysfunction influences or underlies their migraine attacks. Thus, testing of blood TSH and fT4 levels in migraine patients is encouraged, particularly when an increased frequency of migraine attacks co-occur with thyroid dysfunction symptoms— e.g., anxiety, irritability and nervousness, trouble sleeping, weight loss, muscle weakness, irregular (less frequent) menstrual periods, increased sensitivity to heat, and vision problems or eye irritation for hyperthyroidism; and fatigue, weight gain, forgetfulness, frequent and heavy menstrual periods, dry and coarse hair, hoarse voice, and reduced tolerance to cold temperatures for hypothyroidism (10). This is not currently recommended by the national or international guidelines to test for migraine and thyroid comorbidity.
The present study has three possible limitations. First, since we utilised GWAS summary statistics from European ancestry our conclusions may not generalise to other ancestries. Second, our conclusions are limited to the general susceptibility of migraine and thyroid traits as the sex-specific GWAS datasets were either not available or underpowered. Third, although there is a slight sample overlap between the migraine and PANUK thyroid traits GWAS datasets, we do not expect it to affect our conclusions, indeed the LDSC genetic correlation analyses indicated a very small sample overlap with genecov intercept (gcov int) values very close to zero.
Conclusion
The present study provides important insight into the genetic and causal relationship between migraine and thyroid traits. Our results show a significant genetic correlation between migraine with hypothyroidism, hyperthyroidism, secondary hypothyroidism, and fT4. The loci, genes and pathways identified as being shared between migraine and the examined thyroid traits provide biological insight into their comorbid relationships. Also, the causal relationship observed between migraine and hyperthyroidism, and migraine and secondary hypothyroidism; and the shared genomic regions between migraine, TSH, and fT4 indicate the importance of measuring both fT4 and TSH to assess thyroid dysfunction in migraine patients.
Article highlights
Migraine risk is significantly correlated with hypothyroidism, hyperthyroidism, secondary hypothyroidism, and fT4. Migraine shares pleiotropic loci with TSH and fT4. A causal relationship exists between migraine and hyperthyroidism, and migraine and secondary hypothyroidism.
Supplemental Material
sj-pdf-1-cep-10.1177_03331024221139253 - Supplemental material for Shared genetics and causal relationships between migraine and thyroid function traits
Supplemental material, sj-pdf-1-cep-10.1177_03331024221139253 for Shared genetics and causal relationships between migraine and thyroid function traits by Sana Tasnim, Scott G Wilson, John P Walsh, Dale R Nyholt and The International Headache Genetics Consortium (IHGC): for the ICON study group in Cephalalgia
Supplemental Material
sj-pdf-2-cep-10.1177_03331024221139253 - Supplemental material for Shared genetics and causal relationships between migraine and thyroid function traits
Supplemental material, sj-pdf-2-cep-10.1177_03331024221139253 for Shared genetics and causal relationships between migraine and thyroid function traits by Sana Tasnim, Scott G Wilson, John P Walsh, Dale R Nyholt and The International Headache Genetics Consortium (IHGC): for the ICON study group in Cephalalgia
Footnotes
The International Headache Genetics Consortium
The members of the IHGC are Verneri Anttila, Ville Artto, Andrea C Belin, Anna Bjornsdottir, Gyda Bjornsdottir, Dorret I Boomsma, Sigrid Børte, Mona A Chalmer, Daniel I Chasman, Bru Cormand, Ester Cuenca-Leon, George Davey-Smith, Irene de Boer, Martin Dichgans, Tonu Esko, Tobias Freilinger, Padhraig Gormley, Lyn R Griffiths, Eija Hämäläinen, Thomas F Hansen, Aster V.E Harder, Heidi Hautakangas, Marjo Hiekkala, Maria G Hrafnsdottir, M Arfan Ikram, Marjo-Riitta Järvelin, Risto Kajanne, Mikko Kallela, Jaakko Kaprio, Mari Kaunisto, Lisette J.A Kogelman, Espen S Kristoffersen, Christian Kubisch, Mitja Kurki, Tobias Kurth, Lenore Launer, Terho Lehtimäki, Davor Lessel, Lannie Ligthart, Sigurdur H Magnusson, Rainer Malik, Bertram Müller-Myhsok, Carrie Northover, Dale R Nyholt, Jes Olesen, Aarno Palotie, Priit Palta, Linda M Pedersen, Nancy Pedersen, Matti Pirinen, Danielle Posthuma, Patricia Pozo-Rosich, Alice Pressman, Olli Raitakari, Caroline Ran, Gudrun R Sigurdardottir, Hreinn Stefansson, Kari Stefansson, Olafur A Sveinsson, Gisela M Terwindt, Thorgeir E Thorgeirsson, Arn MJM van den Maagdenberg, Cornelia van Duijn, Maija Wessman, Bendik S Winsvold, and John-Anker Zwart.
Acknowledgements
We would like to thank the research participants and employees of 23andMe, Inc. for making this work possible. The full GWAS summary statistics for the 23andMe discovery data set will be made available through 23andMe to qualified researchers under an agreement with 23andMe that protects the privacy of the 23andMe participants. Please visit ![]() for more information and to apply to access the data. In addition, we acknowledge the contributions of the PANUK Biobank, the International Headache Genetics Consortium (IHGC), and the ThyroidOmics consortium (Teumer et al. 2018) for sharing their data. The first author would like to thank the Queensland University of Technology (QUT) and the Australian Government's Research Training Program (RTP) for providing a scholarship.
for more information and to apply to access the data. In addition, we acknowledge the contributions of the PANUK Biobank, the International Headache Genetics Consortium (IHGC), and the ThyroidOmics consortium (Teumer et al. 2018) for sharing their data. The first author would like to thank the Queensland University of Technology (QUT) and the Australian Government's Research Training Program (RTP) for providing a scholarship.
Author contributions
ST and DRN conceived and designed the study; ST performed the analysis and interpreted the results with assistance from DRN.; IHGC provided and curated the data; ST wrote the manuscript with critical inputs from DRN, SGW, JPW, IHGC, and all authors reviewed, revised, and agreed on the final version of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
