Abstract
Objective/Background
Craniofacial growth and development is influenced by growth hormone, which is in turn facilitated by the growth hormone receptor (GHR). GHR gene polymorphisms have been found to be associated with mandibular morphology and skeletal facial profiles in various populations. The aim of this study was to determine the association of single nucleotide polymorphisms (SNP) of GHR gene rs6180, rs6182, and rs6184 with maxillomandibular parameters in the Indian population.
Setting and Sample Population
A cross sectional study was conducted on 174 male and female Indian subjects in the age range of 20 to 32 years who reported for orthodontic treatment.
Material and Methods
Venous blood and lateral cephalogram were collected from each participant for genetic and cephalometric analysis. Genotypes of each SNP were identified by polymerase chain reaction—restriction fragment length polymorphism. Independent t test and Analysis of Variance (ANOVA) were done for intra and inter group comparisons with SPSS software.
Results
SNPs rs6180, and rs6182 were found to be associated with sagittal position and length of maxilla and mandible, anterior facial height, and growth pattern. SNP rs6184 did not have any significant association with the craniofacial parameters.
Conclusion
The results suggest that the GHR gene might be a candidate gene for maxillomandibular morphology in the Indian population.
Keywords
Introduction
Craniofacial development is a complex process which is controlled by multiple genetic and environmental factors and it is fundamental to the origin of vertebrates. 1 Within the purview of orthodontics, it is essential to comprehend the etiology of maxillomandibular discrepancies which have various functional implications ranging from difficult airway, defects in speech and mastication, hearing loss, diminished growth and development, and esthetics to psychosocial issues. 2
Recent advancements have enhanced our knowledge of the genetic course of maxillomandibular development, variations in which may cause morphological and functional disturbances. Genetic variations include mutations and single nucleotide polymorphisms (SNPs). SNPs are caused when there is a change in a single nucleotide along the DNA sequence, which occurs once in every 100 to 300 bases and hence an individual can have millions of SNPs. These genetic variations may cause or modify disease risks or may lead to a natural phenotype. 3
Genes such as insulin like growth factor, growth hormone receptor (GHR), vascular endothelial growth factor, and transforming growth factor govern the maxillomandibular development. 4 Growth hormone (GH) is a significant contributing factor to longitudinal growth and development in the early post-natal years. 5 GH functions by regulation of cellular metabolism and differentiation and controls lipid, carbohydrate, and mineral metabolism.
The most important binding protein which facilitates the action of GH is the GHR. 6 Human GHR gene is present in chromosome 5p13.1-p12. Godowski et al 7 reported that the GHR gene has 9 exons that encode the receptor and several additional exons in the 5-prime untranslated region. The coding exons span at least 87 kilo base of chromosome 5. Exon 2 encodes the signal peptide, exons 3 through 7 encode the extracellular domain, exon 8 encodes the transmembrane domain, and exon 9 and part of exon 10 encode the intracellular domain. A range of factors such as development, nutrition, hormones including GH, and tissue specific factors control GHR gene expression.
Studies done on many regional populations, predominantly Asians, have shown significant associations between GHR gene polymorphisms and maxillomandibular parameters. Mandibular ramus length was found to be associated with SNPs rs6184 and rs6182 in the Japanese population8–11 and with SNP rs6180 in Indian population, 12 whereas SNP rs6180 was linked to ramus height in the Han Chinese population. 13 SNP rs6184 was found to be associated with mandibular length and lower facial height in Turkish subjects 14 , and to ramus length and lower facial height in Iranians, 15 while it was linked to only mandibular body length in Columbians. 16 Study by Adel et al did not find any association between GHR polymorphisms and mandibular morphology. 17 SNP rs2973015 was associated with mandibular prognathism in Brazilian patients in a study by Maranon-Vasqex et al. 18 Three dimensional mandibular morphology was studied by Nakwaki et al on Japanese subjects and found a correlation between GHR gene variant rs6180 and distance between the coronoid processes. 18
Information about the genetic makeup and the inherent growth potential of the patient will assist in determining the optimal time and mode of treatment. Growth curves can predict the expected growth pattern to a certain extent, but genetic information about the individual will strengthen this process. This personalized prediction can be used to evaluate the prognosis of treatment.
Indians constitute 17.7% of the world population and represent the largest diaspora in the world, with around 18 million of its citizens living in other countries. 19 To the best of our knowledge there is no study on Indian population assessing the effect of GHR gene variants rs6180, rs6182, and rs6184 with craniofacial morphology.
Hence, the aim of this study was to determine the association between 3 GHR gene polymorphisms rs6180, rs6182, and rs6184 and specific craniofacial morphology (maxillary and mandibular parameters, sagittal maxillomandibular relations, vertical craniofacial parameters) in Indian population.
Methodology
Sample Size and Study Design
A cross-sectional observational study was conducted on 174 subjects (81 males and 93 females) aged between 20 years and 32 years, who reported to the Department of Orthodontics and Dentofacial Orthopaedics between September 2018 and January 2020. The study was approved by the institutional ethical committee with reference number “MAIDS/Ethical Committee/ 08,” and written informed consent was obtained from all the study participants. The sample size was calculated based on a previous study 16 and the power of this study was kept at 80%, alpha error 5%, and beta error 20% with 95% confidence interval.
Subjects requiring/undergoing fixed orthodontic treatment who were in Cervical Vertebrae Maturation Index completion stage 20 with no history of chronic diseases were included in the study. The exclusion criteria were presence of congenital anomalies, jaw tumors and cysts, any systemic disease, and previous history of facial trauma and facial surgeries.
Genotyping
Peripheral venous blood was collected to extract genomic DNA by conventional phenol chloroform method 21 which was used to determine the gene polymorphisms of GHR gene—rs6180, rs6182, and rs6184, using specific primer Polymerase Chain Reaction (PCR).
Primer 5.0 was used to design primers for each SNP.
rs6180 (I562L) rs6180- F 5’ -ACT TCT GTG AGG CAG ATG CCA AAA AGG GC-3’ rs6180-R 5′ -TCC CAG CAG CAG TGG TAA GGC TTT CTG T-3′ rs6182 and rs6184 (P561T and C422F) rs6182 and rs6184 F 5′- GGG AAG CAG ATC TCT TATG C-3′ rs6182 and rs6184 R 5′-TAT AGT CTG GGA CAG GCA TCT-3′
The PCR was done in a sterile 0.5 ml tube. The reaction mix was made with 30 ng genomic DNA, 200 µM of each dNTP, 0.25U Ex Taq, 0.1 µM of each primer. The PCR conditions were 40 cycles at 95°C for 1 min, 94°C for 30 s, 66°C for 45 s, 72°C for 45 s, and 72°C for 5 min.
PCR—Restriction Fragment Length Polymorphism (PCR-RFLP)
The polymorphic sites were determined by standardized PCR-restriction fragment length polymorphism (PCR-RFLP) method based on previous studies.9, 10, 13, 16 The sizes of the expected fragments before enzymatic digestion were 1037 bp for SNPs rs6184 and rs6182, and 672 bp for SNP rs6180. PCR products from SNPS rs6184, rs6182, and rs6180 polymorphic sites were respectively digested with Eco147I, Cac8l (Thermo Scientific®, Barrington, IL, USA), and HpyCH4V (New England Biolabs®, Ipswich, MA, USA) restriction enzymes at codon 561, 526, and 422.14, 16 The components were incubated at 37°C for 3 to 16 h. The RFLP products were resolved by gel electrophoresis. The sizes of the gel products were 397, 317, 275, and 80 base pairs (bp) for SNP rs6180; and 1037,808 and 229 bp for SNPs rs6182 and rs6184.
Determination of Craniofacial Morphology
Digital lateral cephalogram was obtained from each participant under standardized conditions (80 kvp, 15 ma). The radiographs were taken with the participants standing with Frankfort horizontal plane parallel to the floor and lips in repose. The cephalograms had a ruler for calibration of the linear measurements. The cephalometric tracing was done by a single examiner in a standardized manner on an 8 × 10″ acetate paper to avoid inter examiner variability. Cephalometric tracing was repeated after 6 weeks for 20 randomly selected samples and intra rater agreement was measured using kappa statistics.
The following parameters were measured in the lateral cephalogram:
Maxillary parameters: SNA, NAꞱPt A and Co-Pt A Mandibular parameters: SNB, NAꞱPog, Co-Gn, Go-Gn and Ar-Go Maxillomandibular relationship: ANB, Wits appraisal and Beta angle Growth pattern: Go-Gn to SN and FMA Anterior Facial Height: N-Me
Beta angle was used to divide the subjects into 3 groups as Beta angle has been proven as a significant indicator of anteroposterior jaw discrepancy. 22 Group I consisted of patients with Beta angle between 27° and 35°, group II with Beta angle less than 27°, and Group III with Beta angle more than 35°.
Intra and inter group comparisons were done for the genotype frequencies of each SNP with craniofacial parameters.
Statistical Analysis
Data was analyzed using SPSS software V16.0. The SNPs were expressed as percentages while craniofacial parameters were expressed as means and standard deviation. Chi-square test was used to analyze the difference in proportion of patients having GHR gene polymorphisms and independent t-test/Analysis of Variance (ANOVA) was done for craniofacial parameters.
Results
The genotypes observed were rs6180 – AA, AC and CC; rs6182 – GG and GT; rs6184 – CC and AC, as shown in Figures 1 and 2. Homozygous mutant genotype was not observed in SNPs rs6182 and rs6184.
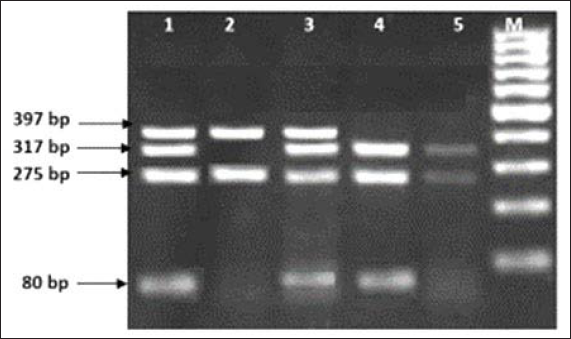
Lane M: 100 bp Marker
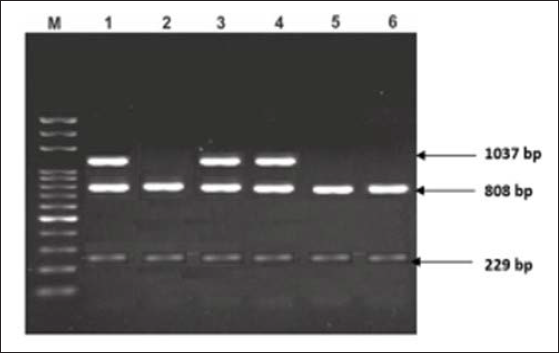
Lane M: 100 bp DNA marker
Intra rater agreement for SNPs with craniofacial parameters was not statistically significant as measured by the Kappa statistics.
The difference in proportion and association of the genotypes of rs6180, rs6182, and rs6184 polymorphism with each group was analyzed using chi-square test (Table 1). Patients in Group I had a higher proportion of AA genotype of rs6180, patients in Group II had a higher proportion of AC genotype, while patients in group III had a higher proportion of AC genotype of SNP rs6180. Higher proportion of patients in all 3 groups had GG genotype of SNP rs6182 and CC genotype of SNP rs6184. The association between the groups and the polymorphisms, however, was not statistically significant.
Chi-Square Test for Inter Group Comparison of Genotype Frequencies of Each SNP.
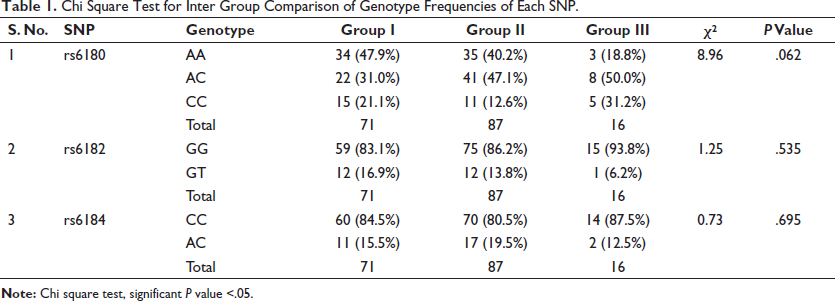
Inter group comparisons for each SNP with the 3 groups showed no statistically significant association.
One way ANOVA and independent t test were performed to compare the mean values of the craniofacial parameters with the genotypes of each SNPs (Table 2). Subjects with CC genotype of rs6180 had a retrognathic maxilla in comparison to the AA and AC genotypes, with a P value of .038 as depicted by N ⊥ Pt A. But on post-hoc analysis, there was no significant variation within the genotypes.
One Way ANOVA Test for Association of Genotypes of SNP rs6180, and Independent T Test for rs6182 and rs6184 with Cephalometric Parameters.
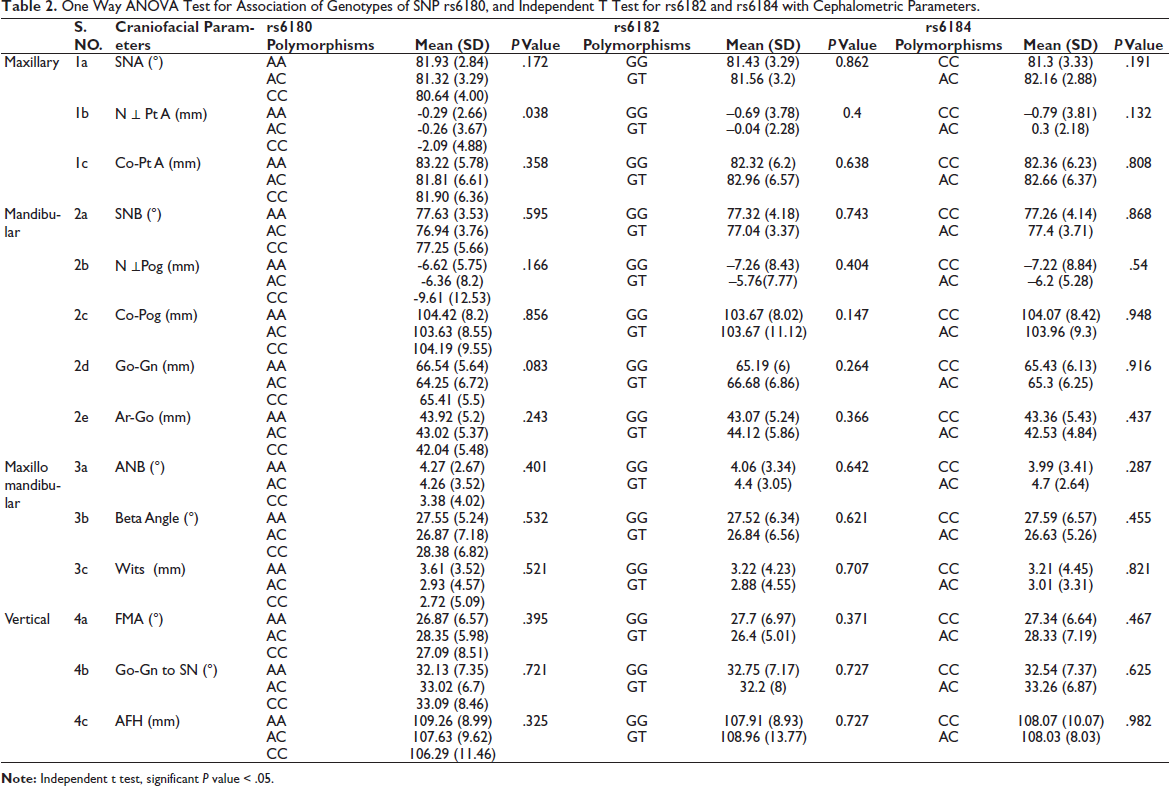
There was no statistically significant difference in any craniofacial parameter between the genotypes of SNPs rs6182 and rs6184.
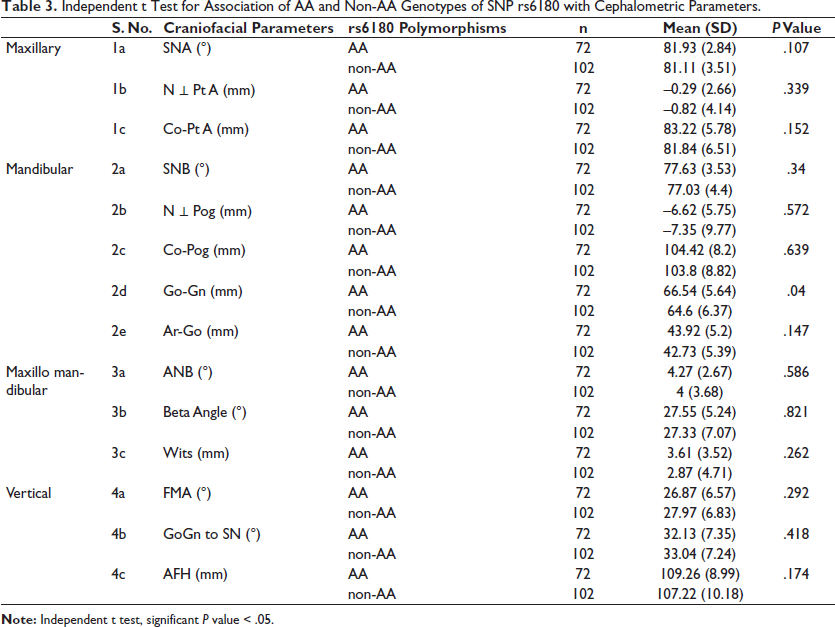
Craniofacial parameters in subjects with AA genotype of rs6180 compared with those with non-AA genotypes depicted that the mandibular length (Go-Gn) was significantly reduced in non -AA genotype, with a P value of .04 (Table 3).
Independent t Test for Association of AA and Non-AA Genotypes of SNP rs6180 with Cephalometric Parameters.
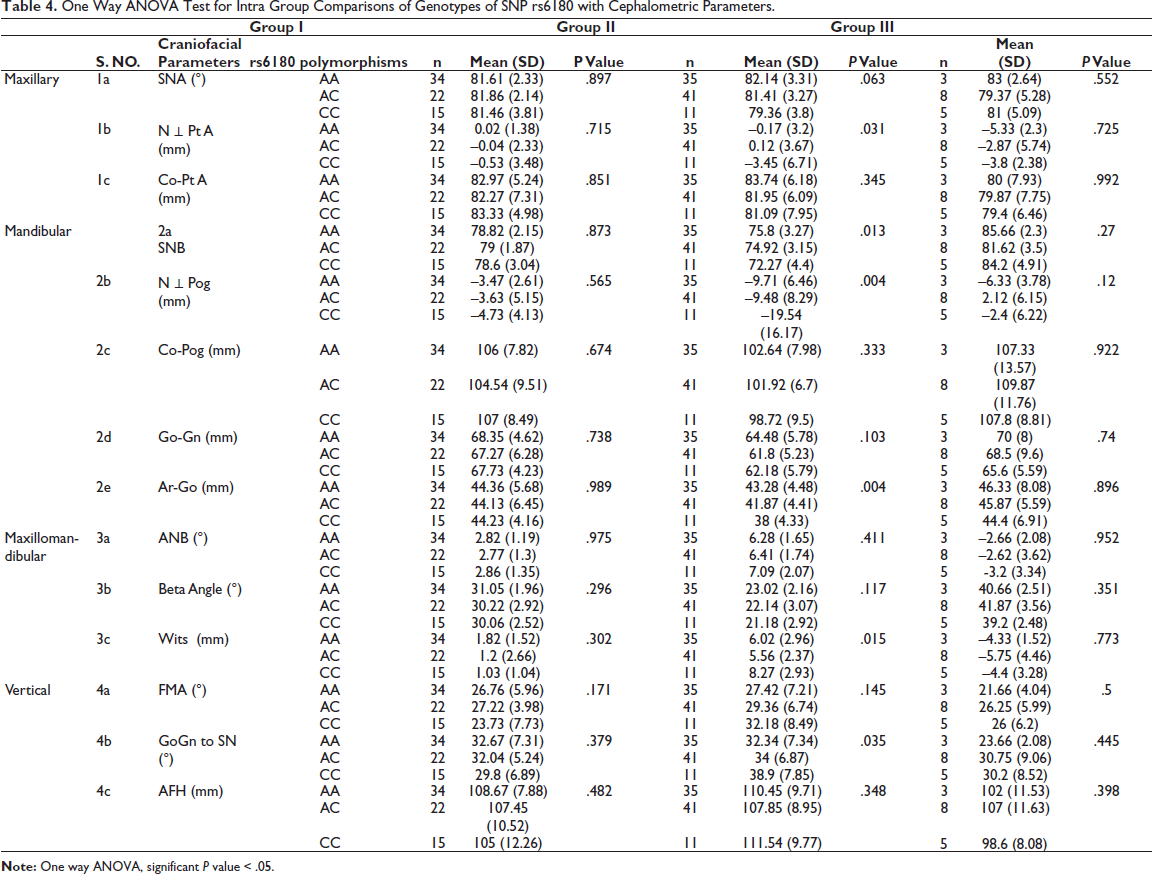
Intra group comparisons of the genotypes of SNPs rs6180, rs6182, and rs6184 with the craniofacial parameters are illustrated in Tables 4, 5, and 6.
One Way ANOVA Test for Intra Group Comparisons of Genotypes of SNP rs6180 with Cephalometric Parameters.
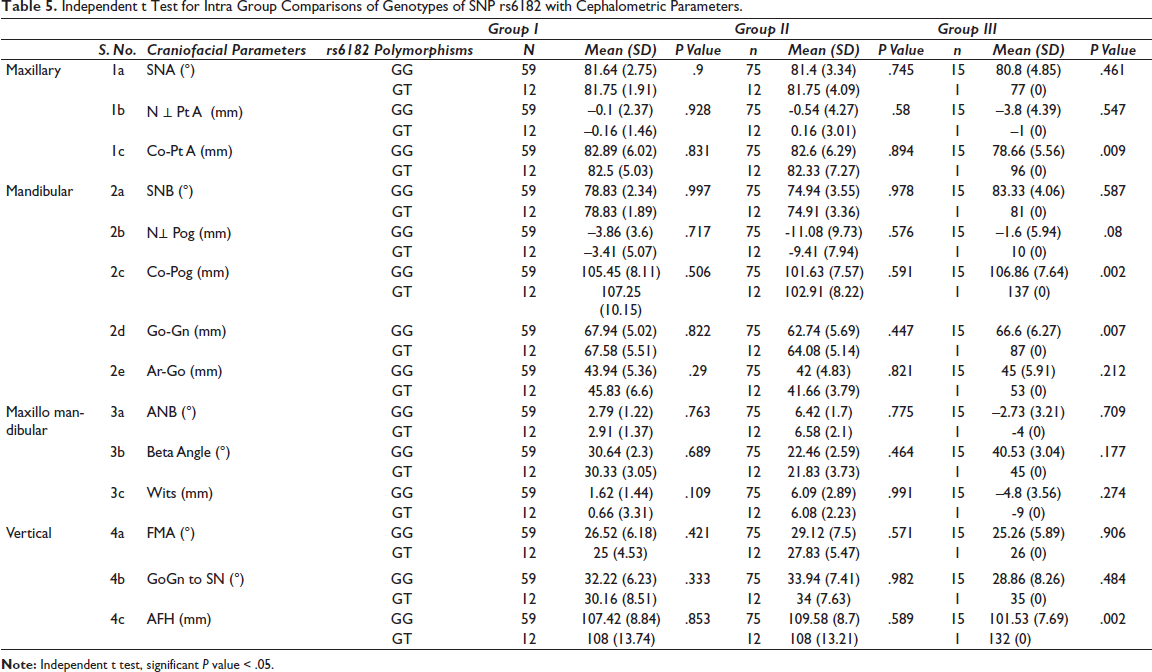
Independent t Test for Intra Group Comparisons of Genotypes of SNP rs6182 with Cephalometric Parameters.
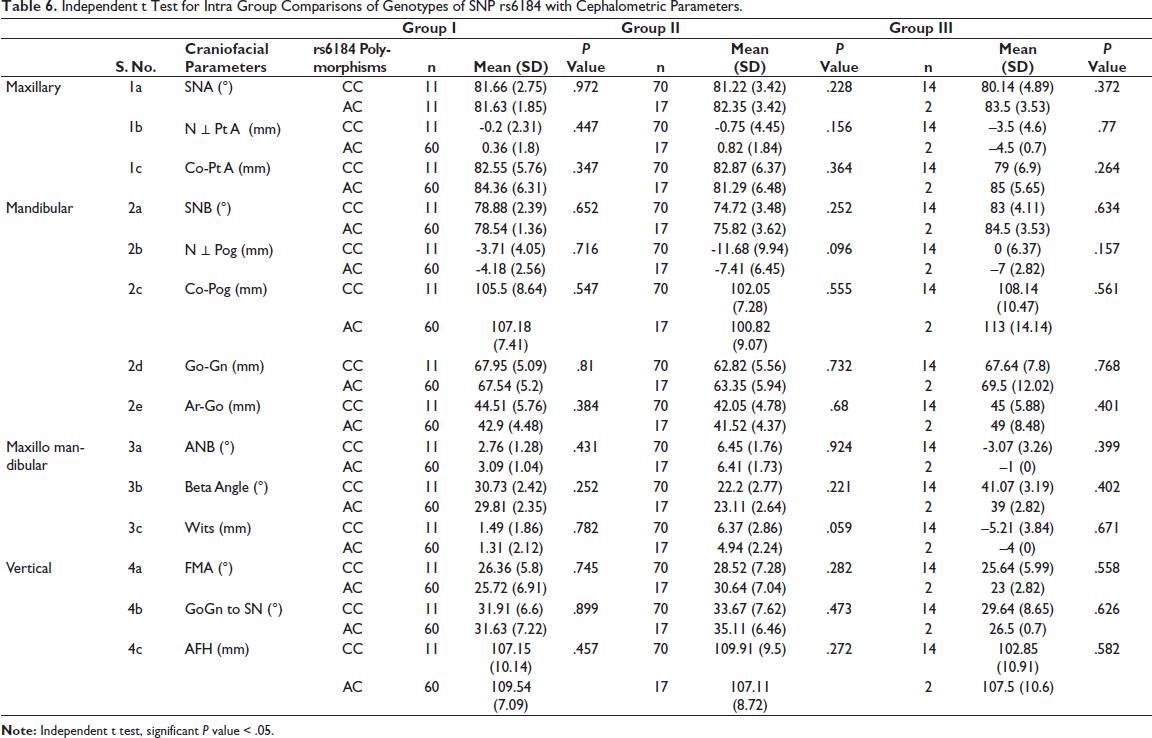
Independent t Test for Intra Group Comparisons of Genotypes of SNP rs6184 with Cephalometric Parameters.
Genotype CC of rs6180 exhibited significant difference in NA ⊥Pt A (P value – .031), SNB (P value – .013), NA ⊥ Pog (P value – .004), Ar-Go (P value – .004), Wits (P value – .015), and Go-Gn to SN (P value – .035) amongst patients in group II, when compared to the other genotypes. Those subjects who had the CC genotype were likely to have a more retrognathic maxilla, retrognathic mandible, a smaller ramal length, and a more vertical growth pattern.
There was significant difference between GG and GT genotypes of SNP rs6182 and Co - Pt. A (P value – .009), Co-Pg (P value – .002), Go-Gn (P value – .007), AFH (P value – .002) in Group III subjects as assessed by independent t-test. Subjects with GG genotype had a more retrognathic maxilla, and those with GT had a longer effective mandibular length and a longer mandibular body length, and a longer anterior facial height.
Significant associations were not revealed between SNPrs6184 and craniofacial parameters amongst subjects in groups I, II, and III.
Discussion
Craniofacial growth is controlled by a multitude of genetic, hormonal, and environmental factors. Bone homeostasis is regulated by local growth factors, cytokines, and hormones, and GH has an evident role in it. GH, a peptide, is secreted by the cells of anterior pituitary which is under the control of the hypothalamus and is also controlled by other hormones like thyroid hormone, sex steroids, and glucocorticoids. The hormone stimulates periosteal apposition through osteoblasts and through forces exerted by muscles on bones. It majorly affects endochondral ossification such as that occurring in the mandibular condyle and cranial base. 23
Individuals who have GH deficiency have smaller craniofacial measurements—small facial height and width, small head circumference, protruding frontal bone, saddle nose and a convex profile, small mandibular and maxillary length, and smaller cranial base length. Cranial base angle, gonial angle, and the angle between the maxillary and mandibular planes are increased.23, 24
The functions of GH are exerted through its gene expression. 24 GH action is activated by binding to its receptor—the GHR, and its encoding gene is located on the proximal short arm of chromosome 5. GHR is present in almost every cell of the body.
Any change in the receptor may have serious implications. Such individuals may have short stature, decreased bone mineral density and concentration, decreased muscle strength, thin skin and hair, delayed puberty, and increased adiposity. Mutation in the extracellular domain of the GHR gene may cause loss of function, resulting in GH insensitivity. Studies have shown that individuals with polymorphisms in GHR gene show varied response to GH. 25
Many studies have been conducted to determine the effect of GHR polymorphisms on craniofacial morphology. These studies were conducted in Asian, Turkish, and Columbian subjects. We investigated the effects of GHR SNPs rs6180, rs6182, and rs6184 which were common across the above-mentioned studies, in Indian population. While most of these studies predominantly studied only mandibular morphology, our study assessed both maxillary and mandibular morphology.
We used venous blood for our study as blood sample is the gold standard for DNA analysis, and has many advantages including minimal chances of contamination, fast and quick procedure, and a significantly higher percentage of amplifiable human DNA when compared with saliva sample. 26
Maxillary parameters
The effect of GH on maxillary growth has been debated. Some studies showed increased maxillary length with GH administration, whereas others found no significant effect. Studies have also shown that GH stimulates periosteal bone formation and muscle mass enhancement which in turn affects the skeletal tissues associated with it. 27
Research on GH deficient children has shown that maxillary length is shorter in such patients. GH treatment in these patients brought about an increase in maxillary length. Cranial base ossifies by endochondral ossification and change in cranial base parameters may influence the position of maxilla as well as mandible. 27 Hence, GH may affect the growth of maxilla indirectly.
To the best of our knowledge, none of the previous studies have shown any association between GHR gene polymorphisms and maxillary parameters. Our study is the first to find such an association. CC genotype of rs6180 SNP was found to be significantly associated with N ⊥ Pt A. Subjects with CC genotype had a retrognathic maxilla when compared to those with other genotypes. Intra group comparisons revealed that Group II individuals with AC genotype of SNP rs6180 had a more prognathic maxilla and Group III individuals with the GG genotype of rs6182 SNP had a smaller maxillary length (Co-Pt A). Genotype rs6184 did not have any significant association with the maxillary parameters.
Mandibular Parameters
There is evidence for presence of GHR in the chondroprogenitor and chondroblast layers of mandibular condyle. 23 This may explain the effect of GHR gene polymorphisms on mandibular morphology. In concordance with previous studies done on GHR gene polymorphisms, our study also found significant associations of the SNPs with mandibular size.
In our study the AA genotype, which was the wild homozygous type of rs6180, was identified to be significantly associated with increased mandibular body length (Go-Gn) when compared with AC and CC genotypes clubbed together, ie, subjects who had a change in base pair from the normal wild type had a smaller mandible.
Within Group II subjects, CC genotype was associated with a retrognathic mandible (N⊥Pog), smaller ramal length (Ar-Go), increased Wits appraisal, and a more vertical growth pattern (Go-Gn to SN). Hence it can be proposed that the CC genotype of rs6180 could lead to a more severe Skeletal Class II malocclusion.
This SNP was associated with skeletal malocclusion in only one other study, the study by Zhou et al 13 in Chinese population. In their study, the CC genotype was associated with increased ramal length.
Group III individuals with GT genotype of SNP rs6182 had an increased effective mandibular length (Co-Pog), increased mandibular body length (Go-Gn), and increased anterior facial height (N-Me). Whereas we found associations between GT genotype and mandibular length, Kang et al concluded in their study that GG genotype of SNP rs6182 was associated with greater mandibular height in Chinese population. 9
The association of SNP rs6184 with mandibular parameter was not statistically significant in the present study.
Our findings were in contrast to the findings of other studies,8, 10, 11, 14, 16 which showed that SNP rs6184 was related to the mandibular length. Tobón-Arroyave et al 16 showed that the CA genotype of the SNP rs6184 in Columbians was associated with a decreased ANB angle and increased mandibular length. Bayram et al 14 also concluded in their study that the CA genotype of SNP rs6184 was associated with greater effective mandibular length and lower facial height in Turkish subjects. Yamaguchi et al 8 differed in their study by showing an association between AA genotype of rs6184 SNP and increased ramus length, while Sasaki et al 11 elicited an association between CA genotype of SNP rs6184 and reduced mandibular growth in Japanese population. Tomoyasu et al 10 concluded in their study that the CC genotype of SNP rs6184 was associated with increased mandibular height in Chinese.
The differences in genetic association could be due to racial and ethnic variation. Research shows that genetic variation within population of the same continent is more than that between different continents. A total of 85% to 90% of genetic variation has been found within intracontinental population whereas only 10% to 15% variation is seen in intercontinental population. 19 This could possibly be the reason why this research does not replicate the results of studies conducted in other populations of Asia.
Genotype analysis could facilitate treatment planning especially for growth modulation by gauging the craniofacial growth potential and help determine the prognosis of orthopedic treatment.
Limitations and Future Scope
A larger sample size would permit the evaluation of association of the genotypic frequencies with craniofacial parameters based on gender. Previous studies have shown an association between maxillomandibular parameters and gender.28, 29
Conclusion
Our study suggests that the GHR gene might be a candidate gene for maxillomandibular morphology in the Indian population.
SNP rs6180 was found to be associated with sagittal maxillary position and SNP rs6182 with maxillary length. SNP rs6180 and rs6182 were associated with sagittal mandibular position and length, ramal length, anterior facial height, and growth pattern. No significant association was found with the genotypes of SNP rs6184.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The study was approved by the institutional ethical committee with reference number ‘MAIDS/Ethical Committee/ 08’.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Informed Consent
The participant has consented to the submission of the article to the journal.
