Abstract
Companion animal cancer diagnostic reports are text-based documents containing essential information on tumor classification and diagnosis. Establishing an animal cancer registry requires integrating and extracting structured data from diverse report formats across multiple providers. This study presents the development of an object-oriented programming approach to standardize and automate cancer data collection for canine and feline patients, enabling the creation of the Australian Companion Animal Registry of Cancers (ACARCinom); Australia’s first national registry of cat and dog cancers. An object-oriented programming approach was developed using the C# language for data processing, tested on sample data from 6 data providers. The initial programming phase focused on designing a parser that identified report sections using regular expressions based on standardized headings. The text was then cleaned to remove unnecessary formatting and HTML tags. Data dictionaries containing preferred terms and synonyms were used to extract key information such as diagnosis, topography, grade, and metastasis, improving consistency and accuracy. A coordinate map of extracted terms was generated to analyze spatial relationships within the report, allowing prioritization of diagnoses. The system also logged parsing decisions and potential issues for expert review. Markup using HTML tags enabled clear visualization of parsed content within the original reports. Extracted data and patient metadata were stored in an intermediary database table, allowing veterinary pathology experts to review and refine entries before final import. This automated solution streamlines data extraction and standardization from diverse sources, enabling the efficient analysis of cancer records and enhancing research and surveillance capacity in veterinary oncology.
Cancer data and tumor registries provide a structured and systematic way to collect, store, and analyze information on cancer cases across a defined population. 6 These registries offer critical insights into the incidence, prevalence, and trends of different types of cancer, allowing healthcare professionals and researchers to identify patterns, risk factors, and potential causes. 23
As cancer remains a leading cause of illness and death among companion animals, there is a need for comprehensive data to inform clinical practice, guide research, and provide valuable epidemiological insights. Unlike human oncology, where population-based tumor registries are well-established, 34 veterinary cancer registries have traditionally been limited in scope, often relying on unstructured hospital- or pathology-based data.7,16,33 The development of more robust and sustainable veterinary cancer registries is essential to accurately monitor cancer incidence, identify risk factors, and evaluate treatment outcomes in animals. In many cases, the structure and content of veterinary pathology and clinical reports are not standardized and may exist in a variety of formats, including handwritten notes, free-text entries within electronic medical records, or scanned files. These unstructured formats are difficult to analyze using conventional data extraction methods, which are often designed for structured, text-based information. 3 The lack of standardization and uniformity means that essential data can be buried in narrative descriptions, making it challenging to systematically capture and analyze relevant information. Extracting cancer data from unstructured pathology or clinical records presents a significant challenge in both human and veterinary medicine.42,48 This problem is further exacerbated by the inconsistencies in data language and vocabulary. 15 For example, different pathologists might describe the same tumor characteristics using different terminologies or abbreviations, leading to variability in the data that are captured. In addition, veterinary pathology or clinical reports often vary significantly between institutions and practitioners, reflecting differences in training, resources, and reporting practices. Inconsistency in terminology not only makes it difficult to compile comprehensive datasets but also introduces the risk of data loss or misinterpretation, which can negatively impact the accuracy of cancer registries.
To address these challenges, innovative approaches are needed to extract meaningful data from unstructured and free-text pathology reports. One potential solution is the development of advanced natural language processing (NLP) algorithms capable of interpreting and standardizing free-text descriptions in pathology reports.4,25,26,46 NLP techniques, such as named entity recognition, information extraction, and text classification, can be particularly useful in automating the identification of key pathological features, diagnostic terms, and relevant clinical data. 47 Recent advancements in machine learning and NLP have made it possible to design models that can process free-text pathology reports with high accuracy. Additionally, there is a growing interest in creating standardized templates and reporting guidelines for pathology reports, which could help reduce variability and improve the consistency of data captured. 28 The adoption of standardized reporting practices, coupled with advanced data extraction tools, could significantly enhance the ability to compile accurate and comprehensive cancer datasets from pathology reports, ultimately improving cancer surveillance, research, and treatment outcomes.
A comparable effort to standardize veterinary cancer data using rule-based methods has been undertaken in the United Kingdom, where a national dataset of canine cancer pathology reports was curated from free-text records using a structured, rule-based approach. 41 This initiative shares similar goals with the Australian Companion Animal Registry of Cancer (ACARCinom) project, highlighting a growing international focus on the development of automated systems to transform unstructured veterinary pathology data into structured cancer registries.
The ACARCinom was created to provide a national repository of cancer data in companion animals and address the need for implementation of an automated solution for importing heterogeneous diagnosis reports for cancer patients and managing vast amounts of data efficiently. The ACARCinom project faced the challenge of importing a large volume of records from various organizations, including private pathology laboratories and university diagnostic pathology laboratories, each with differing data formats. In this study, we report the methodological steps to the development of an innovative and flexible NLP solution to allow efficient and automated extraction of critical diagnostic and patient data from semi-structured text.
Materials and Methods
Definition of Minimum Registry Requirements and Inclusion Criteria
The International Agency for Research on Cancer (IARC) establishes minimum data requirements for cancer registries, ensuring accurate, standardized, and comprehensive data collection. These requirements generally encompass basic demographic information (eg, gender, age, and place of residence), cancer diagnosis specifics (eg., primary site, histology, and behavior), and vital status (eg, last follow-up date and date of death). 6 In the context of the ACARCinom project, key data elements included reference number, patient ID, consultation date, species, breed, sex, neuter status, age or date of birth, diagnosis (tumor name), tumor topography, metastatic site topography, grade, diagnostic certainty, and morphology, as per the Vet-ICD-O-canine-1 coding system. 39 Inclusion criteria for the cases to be imported in the ACARCinom database were cancer cases only, feline and canine species, and cancer cases confirmed by histology. Excluded cases included non-cancer diagnoses, cancer cases in other species, those diagnosed by cytology or gross post-mortem examination alone, and cases with uncertain diagnoses (reported as differential diagnoses often written as vs—versus, ddx, or dd in free-text fields).
Creation of Data Vocabularies
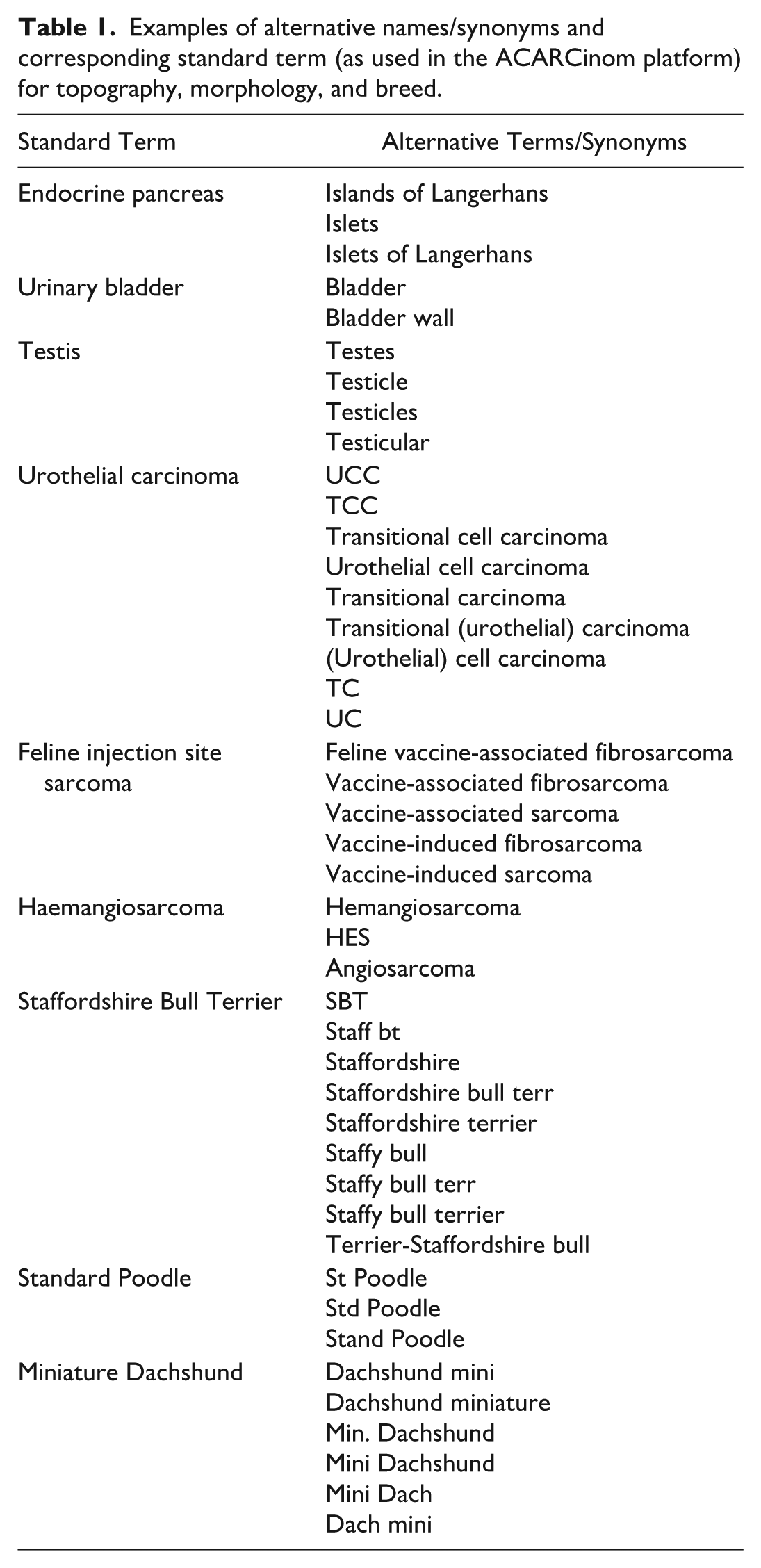
A standardized data vocabulary is critical for consistency in cancer registry data collection and analysis. The ACARCinom project developed and implemented its vocabulary based on the International Classification of Diseases for Oncology (ICD-O-3.2) and the veterinary coding system. 39 This vocabulary included terms for diagnoses, tumor topography, grading, certainty levels, and both feline and canine breeds, ensuring uniformity across data points. For tumor names, topography, and breed classifications, the ACARCinom vocabulary system was designed to include both a preferred or standard term for each category and a set of synonyms or alternative terms, as per examples provided in Table 1. This dual approach ensured that data from various pathology reports could be efficiently mapped to the preferred vocabulary. By incorporating synonyms, the system allowed for the extraction of information from pathology reports using a range of terminology, ensuring consistent mapping to the standard terms in the ACARCinom database. This method streamlined the extraction process and improved the accuracy of data harmonization across multiple institutions, regardless of the specific terminologies used.
Examples of alternative names/synonyms and corresponding standard term (as used in the ACARCinom platform) for topography, morphology, and breed.

Certainty levels were categorized as confirmed, high, moderate, low, and inconclusive based on the diagnostic modifiers in the unstructured text (Table 2).
Categories of certainty levels and corresponding terms/diagnostic modifiers as extracted from pathology reports of the ACARCinom data providers.

The ACARCinom vocabulary adopted the following grading schema:
Grade 1, grade 2, and grade 3 for canine mammary tumors, 37 feline mammary tumors,9,14,29 soft tissue tumors, 12 and lymphomas 45
I/low grade, I/high grade, II/low grade, II/high grade, III/low grade, and III/high grade for canine mast cell tumors24,36 or either low grade/high grade or I/III if only one system was reported.
Definition of Recording and Reporting Specific Data Items
Guidelines were established for recording and reporting multiple primary cancers in the same individual, following the ICD-O-3.2 framework that defines multiple primaries as:
Two or more separate neoplasms in different topographic sites.
Certain conditions characterized by multiple tumors.
Lymphomas involving multiple lymph nodes or organs.
Two or more neoplasms of a different morphology arising in the same site.
A single neoplasm involving multiple sites, whose precise origin cannot be determined.
Multiple primaries, by definition, exclude neoplastic lesions that result from metastasis or recurrence of a primary cancer. 22 In the ACARCinom system, metastases were documented at the time of sample submission, either in conjunction with the primary tumor or as a distinct diagnosis. Further classification and differentiation of recurrent and metastatic lesions were scheduled for the data analysis phase to enhance accuracy and consistency.
Data Sources, Analysis of Reports, and IT Business Analysis
The ACARCinom project utilized anonymized pathology reports from 4 academic institutions (The University of Queensland, Murdoch University, University of Adelaide, University of Sydney) and 2 private laboratories (Gribbles Veterinary Pathology, IDEXX Laboratories). These organizations provided patient and case information, including diagnostic results, in text-based reports. While these reports were often semi-structured, they posed significant difficulties in extracting the relevant data needed for the project. Manually reviewing and processing each report would have been impractical and time-consuming, making automation essential. Sample diagnostic reports were analyzed to identify variations in terminology, structure, and completeness across different institutions. This analysis informed the development of a robust data management system that standardized report formats, ensuring consistency in how diagnostic information was captured. Most reports adhered to some level of standardization, often featuring headings for key sections, which aided in the extraction of critical data items. Depending on the specific data source, the name of the tumor or diagnosis, the site of the tumor or topography, the grade of the tumor, and the certainty level (diagnostic modifiers) were reported under or after specific headings, as listed in Table 3. Supplemental File S1 shows an example of a submission.
Specific headings or sections of the reports containing the name of the tumor (diagnosis), site of the tumor (topography), grade of the tumor, and certainty level.

Parser Development
The parser was developed using C# language for data processing (Application Programming Interface—API), Microsoft SQL Server database for storage, and Angular (Google) for the presentation layer (user interface, UI). All the details about the software tools used for the development of the ACARCinom database are included in Supplemental Table S1.
The data parsing process for converting “raw” data into the ACARCinom system involved several key steps, outlined below. The “raw” data consisted of Microsoft Excel spreadsheets, each provided by different data contributors and formatted with varying columns and data types. While these spreadsheets typically included some straightforward columns, such as dates, species types, and sex, the real challenge lay in the free-form report text columns. These text fields represented the core data that the project aimed to analyze, and they required careful extraction and processing to ensure accuracy and consistency.
Finding the Relevant Part(s) of the Report Text to Begin Analysis
The primary objective of the parser was to extract critical information such as diagnosis, topography, grades, certainty levels, and metastasis details from the report text. To achieve this, the parser utilized data dictionaries created specifically for the project. Many dictionary items consisted of straightforward text terms, such as cancer names (eg, “round cell tumour” or “haemangioma”—in British English as per the original database), which the parser searched for within the report text. Other dictionary items could be regular expressions. Regular expressions are formal patterns used to describe and identify specific sequences of characters within text. They function as a declarative language for text processing, enabling precise matching, extraction, and transformation of data based on syntactic structure. At their core, regular expression patterns are built from a compact syntax that encodes rules about character types, repetition, positioning, and grouping. 8 Regular expression dictionary terms greatly extended the matching power within the text, particularly when matching topography, where many different terms, including acronyms, may be used to describe the location of a cancer. Examples included skin, integument, right, rhs, left, lhs, axilla, overlying, lateral, tarsus, etc.
Each data dictionary contained “preferred” terms, along with supplementary data utilized by the parser. For instance, in the case of diagnosis terms, the dictionary included morphology codes and preference settings such as display colors. Furthermore, dictionary terms could have multiple synonyms to account for common misspellings, name changes, or colloquial names. When a synonym was identified in the report text, the parser mapped it to the corresponding preferred term.
Using the headings of each report, along with subsequent headings that indicated the end of the previous section, we constructed a series of regular expressions to find the text blocks of interest. Each provider’s data file had its own converter rules for finding sections within the reports. In cases where preferred headings were not found, we employed a fallback strategy using less common heading keywords. Ultimately, if no specific terms were identified, the parser extracted the entire report text. Once the relevant sections were harvested, they underwent a “cleaning” process to remove unnecessary formatting and undesirable HTML tags, if present.
An example of the regular expression rules is provided in Supplemental File S2.
Parsing the Report Text
The parser began by constructing a coordinate map for all identified dictionary terms within the report. This map facilitated an in-depth positional analysis of the terms, allowing us to:
Identify relationships between all the diagnoses, topography, grade, and metastasis items.
Prioritize diagnoses based on their positions within the text.
Consolidate diagnoses into unique sets, addressing instances where multiple potential cancer types are suggested or where a cancer name is mentioned repeatedly.
Evaluate additional properties related to the terms, such as positive or negative sentiment, references to differential diagnoses, and any reference systems employed in the text.
Present the parsing results in a clear and distinguishable format for ease of understanding.
Internal to the parser, the coordinate map is an array containing each captured statement, its positional coordinates, the length of the statement, and an internal identifier.
As the parser analyses the report text, it also compiled a ranked list of issues, along with the rationale behind parsing decisions and any other relevant information for users reviewing the results. For instance, the parser noted if a diagnosis or topography term was not found. It also indicated if a specific diagnosis was excluded due to terms like “ruled out” or “benign,” or if a differential diagnosis was presented. Additionally, the parser cross-referenced previous diagnostic records for the animal, if available, to identify inconsistencies, often stemming from simple data entry errors, such as mislabeling species type, sex, or age. These issues were assigned an impact value, ranging from informative to critical. Any parsed record with a critical issue could not be processed further (ie, saved to the database) without user review. Critical issues typically involved missing diagnostic data, inability to determine species, detection of multiple valid diagnoses without a clear preferred option, or the absence of any diagnoses.
Evaluation of Parser Performance
Each time the parser processed a set of records, it logged the batch start time, end time, and the total number of records processed. This information was then sent back to the user as a notification upon completion.
The parsing component operated as part of a broader application running on a single server. Parser performance could fluctuate based on several factors, including the number of active users, the number of parsing batches in progress, and the size of the free-form report text within each record. The parser performance exceeded our expectations and easily processed the historical data. For example, the parser on average was capable of processing 1000 records in 75.96867 seconds (including the time used by the database operation to save the records).
The processing statistics per 1000 records are shown in Supplemental Table S2.
Parser Result Storage and Processing
The parser results, along with associated data such as patient information, were stored in a temporary holding table within the database. Each newly imported record was tagged as “machine parsed,” and its readiness status was set based on whether the record, including the parsed report, met all predefined rules. A human expert (specialist veterinary pathologist, CP) then reviewed the reports flagged by the system, making adjustments as needed. If a human examiner updated a record, it was marked as “human examined” and underwent a validity check. If all rules were satisfied, the record status was updated to “ready.” If the human examiner identified any missing terms in the parser results, they could easily modify the dictionaries and re-upload the dataset for parsing. Any existing records in the holding table were then updated with the latest parsing results.
For a record to be marked as “passed” and ready for saving to the database by the parser, it had to meet a minimum set of criteria such as an explicit diagnosis and topological location. The parser checked for both under- and over-interpretation of these rules. For example, it was possible for the parser to interpret more than one diagnosis, particularly when the author had outlined differential diagnoses. In such instances, these were flagged for human confirmation.
To minimize the risk of incorrect records being processed without a flag, a proportion of parser-passed (ie, “machine-passed”) records were randomly sampled for manual review. In addition, the final data saving step included independent rule-based validation checks outside the parser’s logic. In the unlikely event that an unfit record was marked as ready, these external checks failed, preventing it from entering the database. Such issues were logged and reported to the user in the import completion email. Finally, any records marked as “ready,” whether machine or human reviewed, could be fully imported into the main database.
Performance Evaluation of Cancer Parsing
All cancer registries, in particular population-based cancer registries but transferable to different types of registration systems, should be able to give some objective indication of the quality of the collected data. Two dimensions of quality are validity (proportion of cases with specific characteristics that actually have that attribute—indicator of accuracy of recording) and completeness (the extent to which all cancer cases in a defined population are recorded). 21 To assess the accuracy of the parsing system, a board-certified veterinary pathologist (CP) reviewed random samples of 300 submissions (220 single tumors, 80 multiple tumors), including 54 metastatic lesions in both single and multiple tumors group, parsed by the system and “machine ready” (all rules met—confirmed cancer cases). This review aimed to verify accuracy and validity. An additional 300 submissions flagged by the system as “not machine ready” were also assessed to quantify the amount of missing information in the record (Fig. 1). Completeness can typically be evaluated when at least 2 independent data sources are available, such as comparing histopathology data with cases identified through screening programs or treatment records using capture–recapture methods. However, this approach is not currently feasible for veterinary cancer registries, which generally rely on a single data source—histopathology submissions. Consequently, completeness could not be quantitatively assessed in our study.
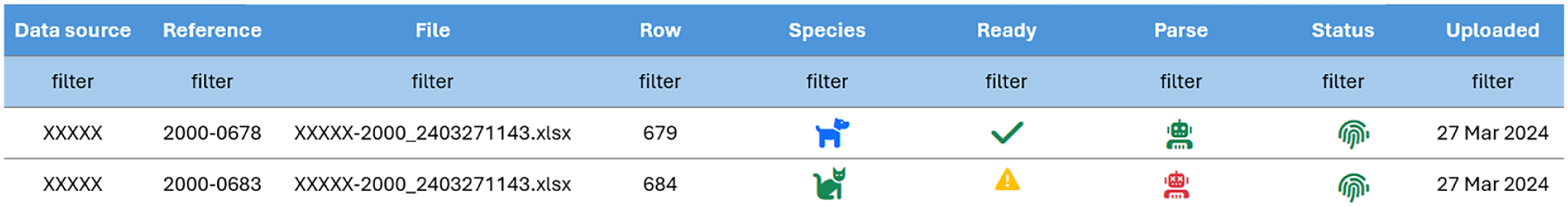
Example of submissions that are “machine ready” (confirmed cancer case, first row) and “not machine ready” (record flagged by the system for incompleteness, second row). The green check mark represents “parsed record—machine ready,” the yellow triangle means “parsed record—to be checked,” the green robot means “passed parsing,” the red robot identifies “failed parsed records,” and the green fingerprint represents “unique cases” (correct patient ID, patient ID found).
Results
Parsing of Cancer Diagnoses
Once the parsing of the report text was complete, the resulting coordinate map was utilized to mark up the report text with HTML tags that delineate the diagnosis sections and each individual data type, such as diagnosis, topography, grade, and metastasis, identified within the text. These tags are standard HTML elements applied to format and structure the visual display of the parsed report in the user interface. An example segment of the parsed report text is illustrated in Fig. 2a, while Fig. 2b shows the raw text of the report marked-up with HTML tags, including items flagged with a minus sign (to be removed). Figure 2c is a closer view of the marked-up report text.

Example of parsing of the report text from of the ACARCinom data provider. (a) Example segment of the parsed report text. The text highlighted in yellow indicates the diagnosis identified by the parser. (b) Marked-up results: Identification of the site of the tumor, diagnosis, and grade through HTML tags. Note the items marked-up with a minus sign; this indicates the parser excluded these from consideration, in preference for the found diagnosis statement and related site. (c) A closer view of the marked-up report text where the parser found a result. Each color represents the HTML tag corresponding to a specific term.
Performance of Cancer Parsing
For reports containing a single tumor diagnosis as well as multiple tumors, all the cases were successfully recorded. In cases involving metastatic lesions, the accuracy rate was 88.9%, with 48 successful results out of 54. All failed records originated from retrospective necropsy reports, which were exclusively in free-text format without structured subheadings.
The following issues were evidenced in the confirmed submissions:
Record of the generic term “tumor/tumors”: Reports were recorded as tumor cases whenever the word tumor appeared in any section. This necessitated an update to the diagnosis dictionary, removing tumor/tumor as a synonym for Tumor cells, uncertain whether benign or malignant (8001/1) (in British English as per original database)
Duplicate entries: Some cases contained multiple entries for the same tumor when reports included synonyms in brackets (eg, Perivascular myoma [angioleiomyoma]).
The submissions flagged by the system contained the following issues:
Misspellings: Errors in breed names (eg, Alaskan Melemute instead of Alaskan Malamute), tumor site names (eg, aritenois instead of arytenoid), and diagnoses (eg, haenangioma instead of haemangioma). When identified, such misspellings, acronyms, and synonyms were added to the dictionaries to improve recognition in subsequent iterations.
Unrecognized acronyms: Use of abbreviations for breed and tumor site not present in the data dictionaries (eg, Wlfh for Wolfhound, FLB for forelimb)
Unspecified tumor site: Reports either lacked tumor site information (eg, unlabeled sample) or mentioned the site in a different section
Missing Data: absence of essential information such as postcode or patient ID
Non-cancerous cases: Submissions without a confirmed cancer diagnosis
Erroneous tumor classification: As previously mentioned, reports were flagged when the word tumor (in British English as per the original database) triggered incorrect classifications.
Data Source Validation and Error Detection Procedures
To ensure consistency and data integrity across sources, each partner institution is required to notify the registry of any changes in data formatting or report layout prior to submitting new datasets. In addition to this communication protocol, the registry portal includes an automated verification system that checks the structure of uploaded files. If a discrepancy in format is detected, the system halts the upload and returns an error message indicating that the file format does not match the expected data source. This double-check mechanism ensures that any modifications in data layout are promptly identified and addressed before data integration.
Final Structure of the Database
The final ACARCinom database (available at: acarcinom.org.au 32 ) consists of the following columns:
Data source: Data provider or data source
Reference: Number of the pathology report (reference number assigned by the submitting laboratory)
Consultation: Date the tumor was submitted to the laboratory
Species: Canine or feline
Patient: Anonymous number assigned to the patient by the data provider/clinician
Age (in years) or extrapolated from the date of birth and the consultation date
Sources: Original Excel file containing the specific extracted data
Samples: Number of tumors recorded per patient
Breed: 545 canine breeds and 136 feline breeds
Sex: Sex of the cat or dog including castration status, when known
Postcode: Practice where the sample was taken or laboratory where the sample was processed
Diagnosis: Specific name of the tumor using the preferred names, as per vocabulary (N = 900)
Primary site: Specific site of the tumor using the preferred names, as per vocabulary (N = 557)
Metastasis: Site of metastasis, if any
Grade: Using a 2- or 3-tier system, as per literature 2
Certainty: Using the 5 certainty levels
As of October 8, 2025, a total of 11,169 records have been parsed and processed into the ACARCinom database. This number is continually updated as new data are submitted by participating laboratories, reflecting the dynamic and evolving nature of the resource.
Discussion
The ACARCinom platform represents a landmark achievement to support the management and transformation of veterinary oncology data, leveraging cutting-edge NLP technology to automate the extraction of critical information from unstructured pathology reports. Historically, cancer data collection for epidemiological research has been hindered by challenges such as inconsistent report formats, varied terminologies, and documentation discrepancies.20,43 At the core of the ACARCinom platform is an extendable base parser, which has been developed and performance tested to convert complex or unstructured veterinary oncological diagnostic text into a readable format. ACARCinom’s flexible and extendable NLP parser effectively circumvents data interpretability issues, delivering enhanced accuracy, consistency, and efficiency in data extraction, being capable of interpreting companion animal data across diverse report formats. This innovation not only minimizes the time and resources required for manual data entry but also paves the way for more standardized and robust veterinary cancer registries.
The accuracy of the parsing system, when compared with manual review by veterinary pathologists, shows promising results, particularly in cases of single tumor diagnoses, with all cancer cases successfully recorded. However, slightly lower accuracy rates in cases involving metastatic lesions or multiple tumors compared to single tumor diagnoses highlight areas for further development. From a diagnostic pathology perspective, the frequency of metastatic disease or multiple tumors depends largely on the type of submission. Metastatic disease is more commonly encountered in post-mortem (necropsy) cases, where widespread dissemination of tumors can be observed, especially with tumors with well-known metastatic potential (eg, lymphoma, hemangiosarcoma, and osteosarcoma). 27 In contrast, biopsy submissions are typically focused on individual, discrete tumors, with fewer cases involving multiple lesions. As a result, while the current system performs well for biopsy-based datasets, improving its ability to accurately capture complex cases seen in necropsy reports would enhance its broader applicability, and ensure the completeness and reliability of cancer datasets. These complex scenarios require nuanced recognition and coding capabilities, suggesting potential improvements in the parser’s algorithm for identifying secondary lesions and concurrent diagnoses. For example, in cases where a primary neoplasm such as a mammary carcinoma metastasizes to regional lymph nodes or lungs, concurrent detection of both the primary tumor and metastatic lesions is crucial for accurate prognostication. 38 Current parsers may under-recognize secondary processes if they are not explicitly coded or described in the clinical narrative, leading to incomplete datasets and potential misclassification. Incorporating structured recognition of common metastatic patterns and associations between primary and secondary lesions could significantly enhance data quality and reliability. Nevertheless, the ability of the parser to accurately map diverse diagnostic terminologies to standardized codes, such as those in the Vet-ICD-O-canine-1 coding system, highlights its potential for broader application in veterinary registries globally. The Vet-ICD-O-canine-1 coding system is the first veterinary-specific adaptation of the ICD-O, which was developed to standardize the recording of tumor morphology and topography in canine cancer registries. 39 Originating from a collaborative initiative involving veterinary oncologists, pathologists, and epidemiologists, 40 the Vet-ICD-O-canine-1 enables consistent cancer registration practices and facilitate comparative oncology research across institutions and countries. 41 This coding system aligns veterinary oncology data with human oncology standards while addressing species-specific differences in tumor biology and presentation. 17 ACARCinom builds upon and directly links with the principles underpinning Vet-ICD-O-canine-1 by operationalizing standardized coding in routine diagnostic data through the automated parser.
Another key achievement was the creation of a standardized data vocabulary, which not only enhanced the consistency of the data but also allowed for more comprehensive analysis across different institutions. The importance of standardized vocabularies in cancer pathology has long been recognized in human medicine. Landmark efforts such as the development of ICD-O 17 and the Systematized Nomenclature of Medicine (SNOMED) 18 have enabled structured, comparable recording of tumor diagnoses worldwide, forming the backbone of cancer registries like SEER (Surveillance, Epidemiology, and End Results) in the United States and the European Network of Cancer Registries (ENCR).1,13 These systems provide standardized morphology and topography codes that support epidemiological research, healthcare planning, and the identification of emerging cancer trends. 35 In veterinary oncology, however, the lack of a unified data vocabulary has historically limited the ability to pool and compare cancer data across clinics, institutions, and countries. 31 Establishing a shared terminology is critical for accurate tracking, research collaboration, and ultimately for advancing comparative oncology initiatives that link animal and human health. The development of data dictionaries for cancer registries represents a pivotal step toward addressing these challenges. Historically, such initiatives have been driven by the need for standardized and comparable data to support effective cancer surveillance and research. In human oncology, organizations such as the International Association of Cancer Registries (IACR) have played a leading role in establishing global standards for data collection, coding, and presentation, thereby enabling comparability across regional and national registries. 5 Similarly, the North American Association of Central Cancer Registries (NAACCR) 30 has developed and maintained detailed data dictionaries that guide registry operations, inform data quality protocols, and evolve in response to new research needs and clinical practices.
The ACARCinom project has made significant strides in addressing this gap by developing a comprehensive veterinary cancer dictionary tailored to canine and feline cases. Building on the Vet-ICD-O-canine-1 framework, the ACARCinom dictionary standardizes diagnostic terminology across multiple fields, including tumor type, anatomical site, biological behavior, and breed. By harmonizing the often heterogeneous language found in veterinary pathology reports, the ACARCinom dictionary transforms disparate diagnostic entries into a coherent, codified framework. By integrating synonyms and aligning diverse diagnostic terminologies with standardized codes, the system reduces ambiguity, minimizes errors, and ensures that data from varied sources are both compatible and interoperable. This approach is essential for large-scale epidemiological studies and cancer surveillance, as it facilitates accurate cross-referencing and aggregation of data from diverse institutions and geographies.
While the parser algorithm currently developed has made significant strides, challenges remain, particularly in processing cases with ambiguous diagnoses or incomplete information. These issues necessitate a manual review process as a fail-safe mechanism to ensure data quality and reliability. While this hybrid approach maintains the integrity of the dataset, it underscores the need for further refinement of the parser’s capabilities in consultation with the existing data providers. The process of parser refinement involves close collaboration with pathology specialists working in diagnostic pathology laboratories, which in turn will enhance their routine data entry and report making. Enhancing its ability to handle differential diagnoses, recognizing patterns in complex pathology descriptions, and addressing missing data points could significantly reduce the need for human intervention.
To ensure the ACARCinom system remains robust and accurate over time, regular updates are planned to accommodate evolving terminology and structural changes in data from contributing laboratories. The rule-based components, including parsing logic and dictionaries, are modular and can be readily modified as needed. Maintenance is supported by automated logging of unparsed cases, periodic audits of “machine-passed” records, and direct feedback from users. Moreover, ongoing partnerships with pathology laboratories facilitate early identification of changes in reporting practices, enabling proactive system updates. In addition, any files or formats not recognized by the system are automatically flagged and reported in the notification email generated at the end of each upload, helping to quickly identify compatibility issues for review and correction.
While the current system is rule-based, the rapid progress of transformer-based and generative language models offers promising avenues for future development. 44 It is important to note that the ACARCinom project was initiated in 2021, prior to the significant advances in large language model capabilities that have since transformed NLP. At that time, rule-based methods provided a more reliable and computationally feasible solution for processing domain-specific veterinary pathology data. Although contemporary models such as Bidirectional Encoder Representations from Transformers (BERT) 19 and Generative Pre-Training Transformer (GPT) 47 architectures can achieve high performance on general NLP tasks, their effectiveness in veterinary contexts remains limited without substantial fine-tuning and retraining.10,11 Moreover, practical constraints related to data protection and privacy would necessitate local deployment rather than reliance on external APIs. Nevertheless, future iterations of ACARCinom could explore and compare LLM-based parsers to evaluate their accuracy, scalability, and cost-effectiveness relative to the current system.
Based on this foundation, the ACARCinom project has successfully demonstrated the feasibility and effectiveness of using an NLP-based approach to manage large-scale, unstructured cancer data in veterinary oncology. By developing a flexible parsing architecture and a standardized vocabulary system, the project has overcome many of the challenges associated with extracting meaningful data from diverse and semi-structured reports. This solution is capable of processing thousands of records per minute, minimizing the need for human intervention and review. Years of historical records across Australian diagnostic centers can now be processed and the data made available for research.
The parser’s high accuracy, particularly in single tumor cases, suggests that such an approach could be widely implemented in other veterinary cancer registries, facilitating better data collection and analysis on a national or even international scale. Furthermore, the ability to integrate new terms and terminologies ensures that the system is future-proof, capable of adapting to the evolving nature of veterinary pathology reporting. Together, this technology creates a robust system for tracking cancer cases, streamlining diagnosis, and supporting research in veterinary medicine.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858251413572 – Supplemental material for Development and implementation of a data parsing protocol for companion animal cancer data
Supplemental material, sj-pdf-1-vet-10.1177_03009858251413572 for Development and implementation of a data parsing protocol for companion animal cancer data by Chiara Palmieri, Matt Taylor, Mike Rickerby, Peter Bennett, Mieghan Bruce, Mark Krockenberger, Philippa McLaren, Thelma Meiring, Kerrie Mengersen, Gabriele Rossi, Anne Peaston, Attracta Roach, Andrew Stent and Ricardo J. Soares Magalhaes in Veterinary Pathology
Footnotes
Acknowledgements
We sincerely thank all data providers for their valuable contributions, which have been instrumental in the success of this project. We gratefully acknowledge the Australian Research Data Commons (ARDC) for their continued support of the ACARCinom project, which has significantly contributed to the development of the data infrastructure.
Author Contributions
CP, PB, MB, MK, PM, TM, KM, GR, AP, AR, and AS contributed to the design of the platform and conceptualization of the project; CP, MB, PB, MK, PM, TM, GR, AP, AR, and AS provided the data for the development of the parser; the manuscript was written by MT, MR, and CP with contribution and revision from all the other authors.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The ACARCinom project (![]() ) received research funding from the Australian Research Data Commons (ARDC). The ARDC is funded by the National Collaborative Research Infrastructure Strategy (NCRIS).
) received research funding from the Australian Research Data Commons (ARDC). The ARDC is funded by the National Collaborative Research Infrastructure Strategy (NCRIS).
Supplemental material for this article is available online.
