Abstract
Novel goose parvovirus (NGPV) is continuously threatening the global duck industry, as it causes short beak and dwarfism syndrome among different duck breeds. In this study, we investigated the viral pathogenesis in the tongue of affected ducks, as a new approach for deeper understanding of the syndrome. Seventy-three, 14- to 60-day-old commercial Pekin ducks were clinically examined. Thirty tissue pools of intestine and tongue (15 per tissue) were submitted for molecular identification. Clinical signs in the examined ducks were suggestive of parvovirus infection. All examined ducks had short beaks. Necrotic, swollen, and congested protruding tongues were recorded in adult ducks (37/73, 51%). Tongue protrusion without any marked congestion or swelling was observed in 20-day-old ducklings (13/73, 18%), and no tongue protrusion was observed in 15-day-old ducklings (23/73, 32%). Microscopically, the protruding tongues of adult ducks showed necrosis of the superficial epithelial layer with vacuolar degeneration. Glossitis was present in the nonprotruding tongues of young ducks, which was characterized by multifocal lymphoplasmacytic aggregates and edema in the propria submucosa. Immunohistochemical examination displayed parvovirus immunolabeling, mainly in the tongue propria submucosa. Based on polymerase chain reaction, goose parvovirus was detected in 9 out of 15 tongue sample pools (60%). Next-generation sequencing confirmed the presence of a variant goose parvovirus that is globally named NGPV and closely related to Chinese NGPV isolates. Novel insights are being gained from the study of NGPV pathogenesis in the tongue based on molecular and immunohistochemical identification.
Waterfowl parvoviruses (WFPVs) are cross-species pathogens of waterfowls, which cause a highly contagious enteric disease known as Derzsy’s disease. The disease is characterized by its high morbidity and mortality rates in highly susceptible, antibody-free goslings and Muscovy ducklings. Birds that survive the infection persistently suffer from stunted growth and abnormal feathering.11,20 Classical goose parvovirus (GPV) infection occurs not only in geese but also in Muscovy ducks, whereas Muscovy duck parvovirus (MDPV) only causes disease in Muscovy ducks. 15 Since the first isolation of GPV in the 1960s in Hungary, and MDPV in the late 1980s in China, both viruses have continued to cause economic losses in the duck industry.17,28
Short beak and dwarfism syndrome (SBDS) in ducks was first reported in France in 1970 and in Poland in 1995. 29 It was suggested that this syndrome was caused by GPV, based on its serological identification, but the causative agent was neither isolated nor molecularly identified until 2006 when a study on SBDS in France declared that the incriminated pathogen was successfully isolated and identified by phylogenetic analysis as a variant GPV strain. 29 Following several SBDS outbreaks among different duck breeds, including Pekin, mule, and Muscovy ducks in China since 2015, the causative agent of SBDS has been globally known as a novel goose parvovirus (NGPV).6,14 In Egypt, NGPV was first detected and isolated from SBDS outbreaks in 2020 from Pekin and mule ducks. 32 Since then, SBDS outbreaks have surged in different Egyptian governorates.
The genomes of WFPVs are approximately 5100 nucleotides. The viruses have long, linear, and single-stranded DNA with an icosahedral symmetry and a diameter of 20–22 nm. The viruses have two main open reading frames. The 5′ open reading frame encodes the nonstructural proteins that are involved in viral replication, transcription, and assembly. The 3′ open reading frame encodes the capsid viral proteins, including structural proteins VP1, VP2, and VP3, which play significant roles in tissue tropism, host range, pathogenesis, pathogenicity, and immune responses. WFPVs are divided into two groups: the GPV-related group and the MDPV-related group. Irrespective of their high genomic similarity, both groups differ in their host ranges, immunogenicity, and nucleotide sequences.3,6,14,17,28,41
Since 2015, phylogenetic analysis, based on complete genome sequences, has revealed that WFPVs are divided into three branches, namely the classical GPV, NGPV, and MDPV. These three strains belong to different lineages within the species Anseriform dependoparvovirus in the genus Dependoparvovirus of the subfamily Parvovirinae of the family Parvoviridae. Moreover, they share similarities in their morphologies, physicochemical properties, culture characteristics, and genomic structures but differ in their pathogenesis, pathogenicity, and immunogenicity.10,38
WFPVs can be diagnosed using a variety of serological methods, including the agar gel precipitation test, virus neutralization test, and enzyme-linked immunosorbent assay. These traditional methods have been shown to be laborious and time-consuming and are only used as indirect diagnostic tools when the molecular detection is not available. 39 Furthermore, it is difficult to make an accurate differential diagnosis between NGPV, classical GPV, and MDPV based on serological assays because of the high genetic similarity and antigenic cross-reactivity between MDPVs and GPVs.6,8 Polymerase chain reaction (PCR) techniques can be differentiated between GPVs and MDPVs using highly accurate primers. 36 Consequently, sequencing techniques have been widely administrated for the confirmatory detection and differentiation between classical GPVs and NGPVs.21,25
Several studies inspected the negative impact of NGPV on the growth performance of ducks based on lesion scoring, serological identification, molecular detection, and histological examination of affected tissues including the liver, intestines, kidneys, muscles, and heart.5,25,26,37 Surprisingly, no previous work investigated the NGPV pathogenesis in the tongue and its drastic role in growth retardation despite the degenerative effect of NGPV infection on the tongue tissue.
The aim of the current study was to identify the implicated pathogen that is associated with SBDS in the tongues of naturally infected Pekin flocks using molecular, histopathological, and immunohistochemical methods.
Materials and Methods
Ethics Statement
Animal handling and sample collection were reviewed and approved by the Zagazig University Institutional Animal Care and Use Committee at Zagazig University in Egypt, with the approval number: ZUIACUC/2/F/176/2022.
Clinical and Postmortem Examinations
Seventy-three, 14- to 60-day-old diseased Pekin ducks, representing 15 duck flocks, were collected from different localities at Sharkia governorate in Egypt over 1 year (2021–2022), which included 8 adult Pekin flocks (37 diseased ducks) that were more than 1 month old and 7 young Pekin flocks (36 diseased ducks) that were less than 1 month old. All 73 birds were submitted for clinical and postmortem examinations. Although all examined ducks were hatched from parvovirus-vaccinated breeders, it was anticipated that the collected ducks had parvovirus infection because they all exhibited beak dwarfism, growth retardation, and whitish diarrhea. In addition, some birds were suffering from general weakness and abnormal feathering (feather loss on the neck and back regions).
The growth retardation observed in each flock was assessed by measuring the body weight of all diseased ducks introduced to the clinic and calculating the average body weight of those diseased ducks representative of each infected flock (3–7 diseased ducks per flock). Next, we compared the mean body weight of the diseased ducks with that of the healthy ducks in the same flock. It is a part of descriptive data routinely collected from the farm owners and recorded in the case report file.
Sampling
Thirty tissue pools (3–5 tissue specimens/tissue pool) were collected under aseptic conditions from the intestine and tongue separately (15 tissue pools/organ), as each intestinal and tongue pool represented individual flocks (15 total flocks). All 30 tissue pools were transmitted at approximately 4°C in leak-proof containers with ice and preserved at −80°C for molecular detection (quantitative TaqMan real-time PCR and conventional PCR).
Nucleic Acid Extraction
DNA was extracted from the 15 intestinal and 15 tongue tissue pools according to the manufacturer’s instructions of the GeneJET Genomic DNA Purification Kit (Thermo Fisher Scientific, Baltics UAB, Lithuania, Catalog No: K0721). DNA extracts were quantified using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
qPCR and Conventional PCR
Oligonucleotide primers and probe (TIB MOLBIOL, Berlin, Germany) that were used for qPCR reactions are shown in Table 1. A 25-μL PCR reaction mix was composed of 12.5 μL of 2X Brilliant Multiplex qPCR Master Mix (Agilent technologies, Santa Clara, CA, USA, Catalog No: 600553), 0.5 μL of each primer (10 μmol/L each), 1 μL of probe (10 μmol/L), 3 μL of DNA template, and nuclease-free water in an amount to adjust the total reaction volume. The thermal cycle profile was set as 1 cycle of 95°C for 10 minutes, followed by 40 cycles of 95°C for 15 seconds, 58°C for 1 minute, and 72°C for 30 seconds using the AriaMx Real-time PCR System (Agilent Technologies, Santa Clara, CA, USA).
Oligonucleotide primers and probe used in the current study.
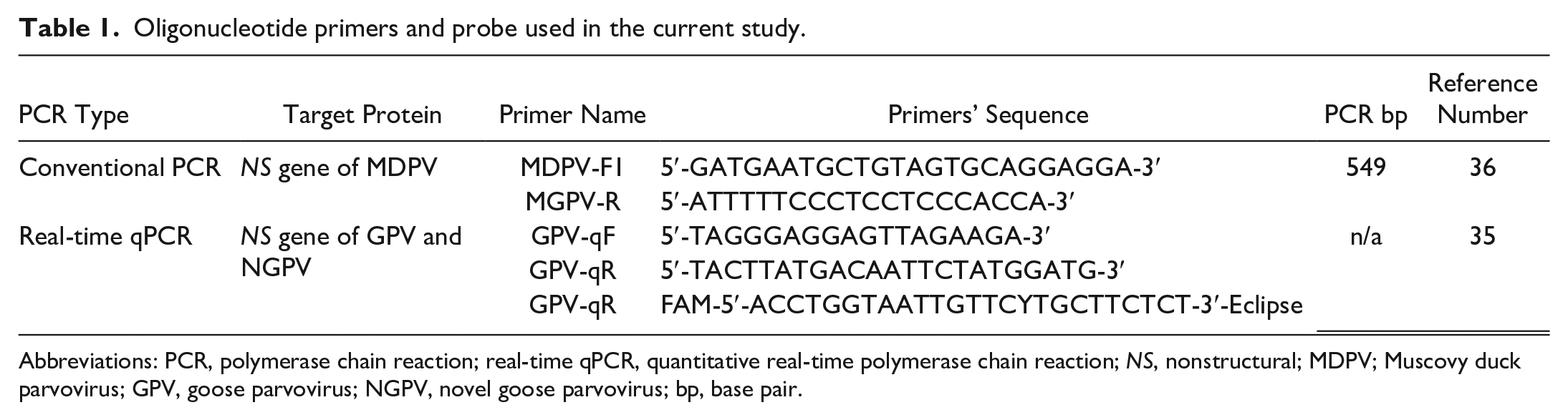
Abbreviations: PCR, polymerase chain reaction; real-time qPCR, quantitative real-time polymerase chain reaction; NS, nonstructural; MDPV; Muscovy duck parvovirus; GPV, goose parvovirus; NGPV, novel goose parvovirus; bp, base pair.
The 30 real-time qPCR samples were also resubmitted for a discriminative conventional PCR to detect MDPV. A total of 50 μL for each PCR reaction was optimized as follows: 25 µL of 2X DreamTaq Green PCR Master Mix (Thermo Fisher Scientific, Baltics UAB, Lithuania, Catalog No: K1071), 1 µL of forward primer, 1 µL of reverse primer (10 μmol/L each), 5 µL of DNA template, and 18 µL nuclease-free water. The thermal cycling conditions were adjusted as follows: one cycle of 95°C for 1 minute followed by 40 cycles of 95°C for 30 seconds, 60°C for 30 seconds, 72°C for 1 minute, and a final extension step at 72°C for 10 minutes. The amplification products were subjected to 1% agarose gel electrophoresis using the ready-to-use GeneRuler 1 kb DNA Ladder (Thermo Fisher Scientific, Baltics UAB, Lithuania, Catalog No: SM0313).
Histopathology and Immunohistochemistry
Seventy-three tongue specimens were collected from all examined Pekin ducks, preserved in neutral-buffered formalin, routinely processed, sectioned, and stained with hematoxylin and eosin. 1 Tissue sections were examined using a Leica DM4 B light microscope. Images for examined tissue sections were captured using a Leica DMC 4500 digital camera linked to LAS-X software (Leica, Wetzlar, Germany).
Hyperimmune serum against WFPVs was prepared in four New Zealand rabbits by using an inactivated oil emulsion vaccine (Ceva-phylaxia, Budapest, Hungary, Batch No: 001KG1D) through a series of 0.5-ml scheduled injections. 30 The blood samples were collected and centrifuged for serum separation. Purification of serum was conducted via precipitation using ammonium sulfate. 27 Purified IgY was used as the primary antibody at a dilution of 1:500 in phosphate-buffered saline.
Fifteen tongue tissue sections, representative from all examined Pekin duck flocks, were submitted for immunohistochemistry (IHC). The IHC protocol was as follows. 24 Heat-induced antigen retrieval was conducted in a microwave oven for 15 minutes using the Tris-EDTA buffer. Tissue sections were washed with phosphate-buffered saline. Endogenous peroxidase blocking step was performed by adding three drops of 3% H2O2 to tissue sections and incubating for 10 minutes. Tissue sections were incubated with the primary antibodies (rabbit anti-WFPV, 1:500 in phosphate-buffered saline) for 2 hours in a humidity chamber, followed by washing with phosphate-buffered saline and incubation with horseradish peroxidase–labeled secondary antibodies for 30 minutes at room temperature (Bio-SB, Santa Barbara, CA, USA). Finally, a diaminobenzidine substrate mixture was used for substrate detection (Bio-SB, Santa Barbara, CA, USA). 4 For negative control tissue slides, the primary antibody was replaced by negative serum on WFPV-infected tongue tissue specimens. In addition, normal noninfected Pekin duck tongue tissues were incubated with the primary anti-WFPV antibodies. For positive control tissue slides, primary antibodies with proven reactivity to avian tissues were obtained, including anti-smooth muscle actin (Clone 1A4 [Ready-To-Use], ScyTek, USA), and anti-BCL2 (26593-1-AP, Proteintech, Germany), and were used on tissue sections from various duck organs, including tongues and intestines.
Next-Generation Sequencing
Two isolates from one flock, extracted from the tongue and intestinal tissues, were selected for sequence analysis. The DNA library was prepared by viral genome DNA fragmentation followed by ligation of adapters to the end of DNA fragments. The sequences of these adapters were recognized by the Illumina sequencing platform and allowed for the amplification of the DNA fragments using the Nextera XT DNA Library Preparation Kit (Illumina, San Deigo, CA, USA, Catalog No: FC-131-1024). The prepared DNA library was then loaded onto the Illumina MiSeq sequencer (Illumina, San Deigo, CA, USA, Serial No: SY-410-1003). The fluorescent signals generated during this process were detected by the sequencer and were used to determine the sequence of each DNA fragment.
The raw sequencing data were processed and analyzed using the Bacterial and Viral Bioinformatics Resource Center (BV-BRC) (bv-brc.org) and Molecular Evolutionary Genetics Analysis Version.11 (MEGA.11) software. Data were submitted to the National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/). The phylogenetic tree was designed using the MEGA.11 software and based on the blast analysis of both whole genome–sequenced isolates in the current study and up to 20 NGPV complete genome sequences that were homologous to the study isolates. The NCBI reference GPV virulent B strain (accession no: NC_001701) was blasted with both identified strains and inserted within the whole genome phylogenetic tree to determine the nucleotide diversity among their genomes (on the whole genome, VP, and nonstructural levels). The viral genome was annotated and interpreted using the Vgas viral genome annotation system (http://guolab.whu.edu.cn/vgas).
All procedures for molecular identification were applied according to the biosafety measures in the Biological Prevention Department, Main Chemical Labs, Chemical Warfare, Ministry of Defense, Egypt.
Results
Clinical and Postmortem Findings
Among examined duck flocks, the morbidity rate of most flocks ranged from 2.25% to 11.4%, but one flock had a morbidity rate of 33.3% and another two flocks reached 50%. Only 3 flocks had mortalities, which did not exceed 1.5%. The examined ducks exhibited variable clinical signs including growth retardation and whitish diarrhea in all 73 examined ducks (100%), abnormal feather loss and skin redness on the neck and back areas of 35 out of the 73 diseased ducks (48%), and general weakness in 21 out of the 73 diseased ducks (29%). All examined ducks had short beaks either with protruding tongues in 50 out of 73 examined ducks (68%) or nonprotruding tongues in 23 out of 73 examined ducks (32%).
The gross lesions of examined tongues were variable in relation to the age of examined ducks. Necrotic, swollen, and congested protruding tongues were present in all adult ducks greater than 30 days of age (37 out of total 73 examined ducks, 51%). Tongue protrusion without any congestion or swelling was observed in 20-day-old ducklings (13 out of 73 examined ducks, 18%), while the apparently normal (nonprotruded) tongues were recorded in 15-day-old ducklings (23 out of 73 examined ducks, 32%) (Fig. 1a–d). Necrosis was mainly restricted to keratinized lingual apex (known as a lingual nail) and keratinized lateral barbs located in both dorsolateral sides of tongue, which appeared as blackish discoloration compared to the apparently normal structures in the nonprotruding tongues of diseased young ducks (Fig. 1b, c). Mild intestinal congestion was identified in 57 out of 73 (78%) ducks, while only 16 out of 73 examined ducks had apparently normal intestines (22%).
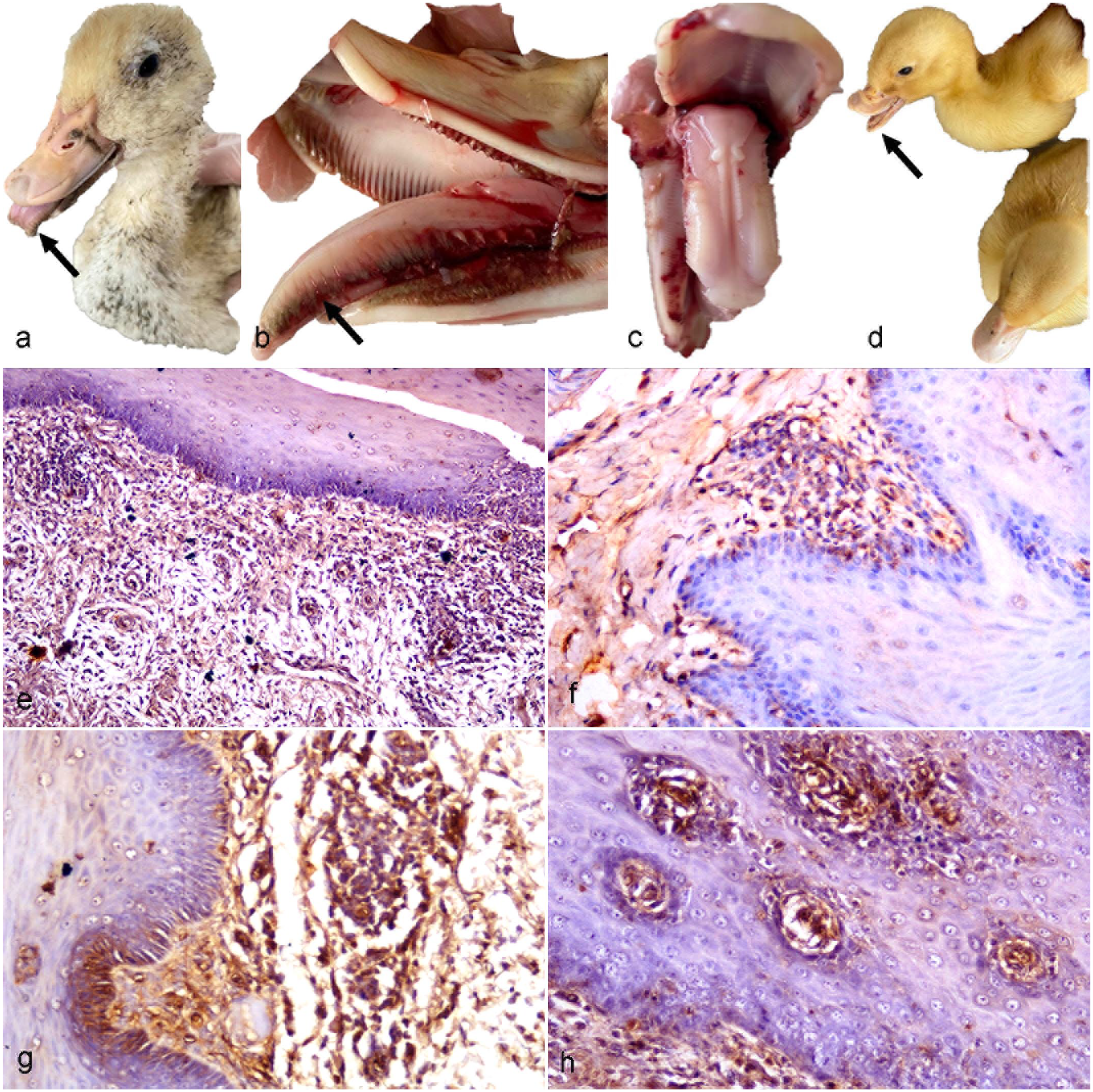
Clinical and postmortem findings of goose parvovirus infection in tongues of Pekin ducks. Diseased adult ducks are characterized by (a) congested protruding tongue (arrow) and (b) necrosis (arrow) of the dorsolateral border of the protruding tongue. Diseased young ducks are characterized by (c) an apparently normal structure of nonprotruding tongue and (d) beak atrophy (arrow) comparing to the normal duck (bottom right). (e–h) Goose parvovirus immunohistochemistry, tongue, Pekin ducks. (e) Immunolabeling is distributed in the propria submucosa of a young duck, (f) the propria submucosa of an adult duck, (g) the mucosa and propria submucosa of a young duck, and (h) the stratified epithelium of mucosa of a young duck.
qPCR and Conventional PCR Findings
Among the examined flocks, 9 out of 15 tongue tissue pools were GPV-positive (60%). Four GPV-positive pools were harvested from adult flocks with tongue protrusion, and 5 GPV-positive pools were harvested from young Pekin flocks without tongue protrusion. The 6 GPV-negative tongue tissue pools were from flocks with tongue protrusion, suggesting there is no correlation between tongue protrusion and PCR positivity. However, all 15 intestinal pooled samples were GPV-positive (100%). The 30 tissue pooled samples were resubmitted for a discriminative conventional PCR assay and confirmed the absence of MDPV.
Histopathological and Immunohistochemical Findings
Forty-three of the 73 tongue tissue sections that were collected from the 9 PCR-positive flocks (including 20 adult and 23 young diseased ducks) showed histologic lesions. Meanwhile, 30 tongue tissue sections, collected from the 6 PCR-negative flocks (including 17 adult and 13 young diseased ducks), were histologically abnormal, and 6 of these tongue tissues, representing each flock, subjected to immunolabeling did not have an identifiable viral antigen. Nine tongue tissue sections harvested from PCR-positive flocks were subjected to immunolabeling, and all exhibited the viral antigen.
Microscopical examination of the tongue revealed various histopathological alterations that varied among the adult and young Pekin duck flocks. The affected adult flocks had necrosis of the superficial epithelial layer of the protruding tongue, which was frequently associated with vacuolar degeneration. Focal lymphoplasmacytic inflammatory cell infiltrates were observed in the subepithelial surface, which contained numerous congested blood vessels. Marked glossitis was detected in young flocks and was characterized by multifocal lymphoplasmacytic aggregates in the propria submucosa in association with congested blood vessels and marked edema (Fig. 2).
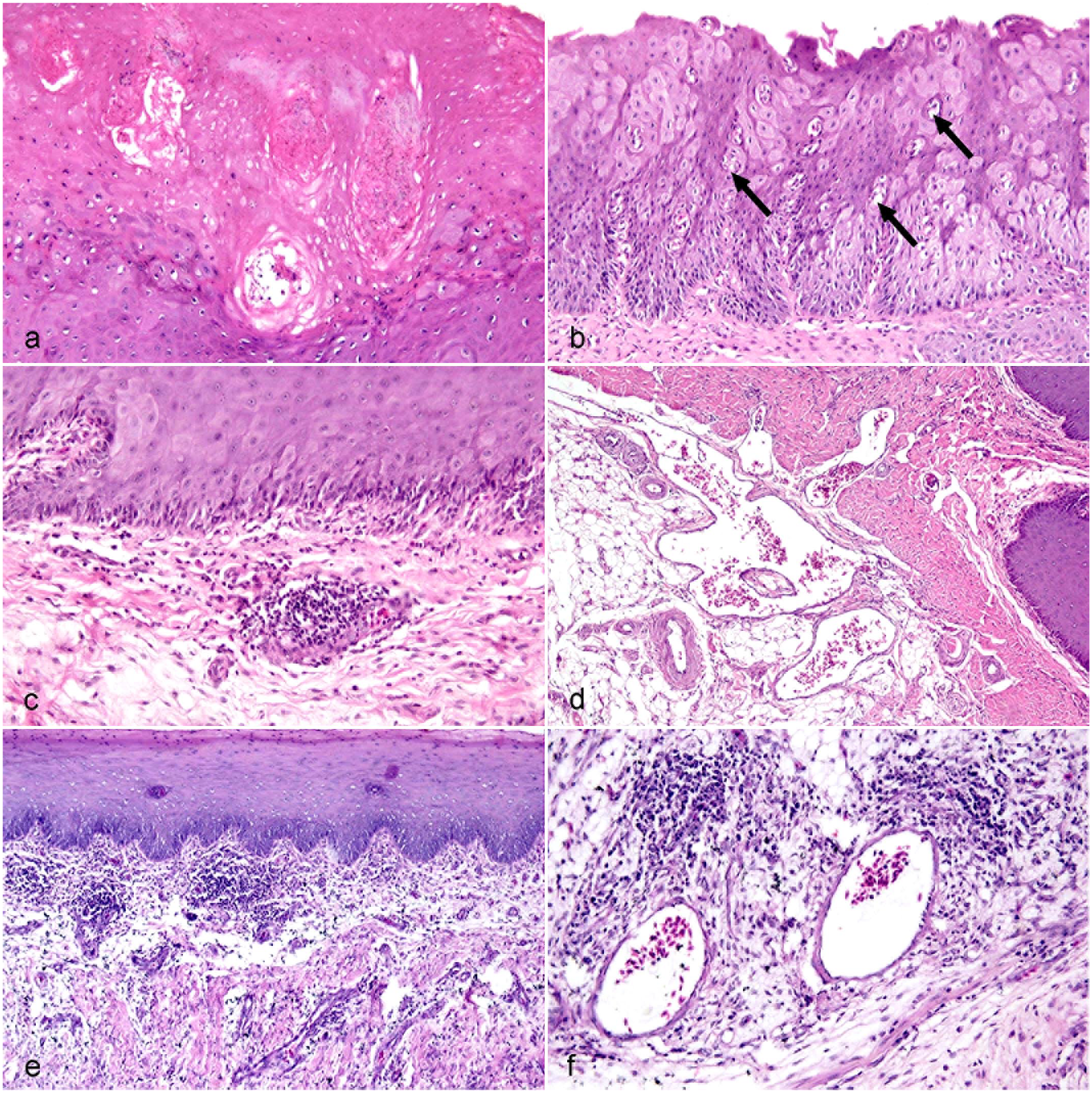
Goose parvovirus infection in the tongue of Pekin ducks. Hematoxylin and eosin. (a) Necrosis of the superficial epithelial surface in an adult duck. (b) Vacuolar degeneration (arrows) of stratified epithelial cells in an adult duck. (c) Moderate perivascular and mild, diffuse, subepithelial mononuclear infiltrate in the propria submucosa in an adult duck. (d) Numerous dilated blood vessels in the propria submucosa in an adult duck. (e) Moderate inflammation in the superficial propria submucosa in a young duck. (f) Multifocal lymphocytes and plasma cells and edema in the propria submucosa in a young duck.
Positive immunolabeling of the viral antigen was mainly detected in the propria submucosa layer in both PCR-positive adult and young Pekin ducks. However, higher immunolabeling was observed in the young flocks with the intense inflammatory reaction. Viral antigen was detected in the epithelial cells of the mucosal surface in some flocks (Fig. 1e–h).
A normal histological structure was represented in examined noninfected ducks as compared to the histopathological alterations observed in infected ones. There was no viral antigen immunolabeling in noninfected flocks. Similarly, there was no immunolabeling in infected flocks in sections without inclusion of the primary antibody (Supplemental Figure S1).
Next-Generation Sequencing
Two strains in the current study, isolated from the tongue and intestine of the same flock, namely GPV-EgyArmy -ZU-237-2022 and GPV-EgyArmy-ZU-246-2022 were submitted for next-generation sequencing, which confirmed that the whole genome lengths were 5104 bp and 5102 bp, respectively. Sequence data were submitted to GenBank under the accession numbers of OR416221 and OR416219, respectively. Both strains are classified as a variant GPV (later named NGPV), as they showed up to 98.86% similarity with each other and were closely related to recent Chinese NGPV isolates identified since 2015. Twenty NGPV strains, which had the highest nucleotide identity with strains in the present study (98%–99%), were selected to draw the whole genome phylogenetic tree (Figs. 3 and 4).
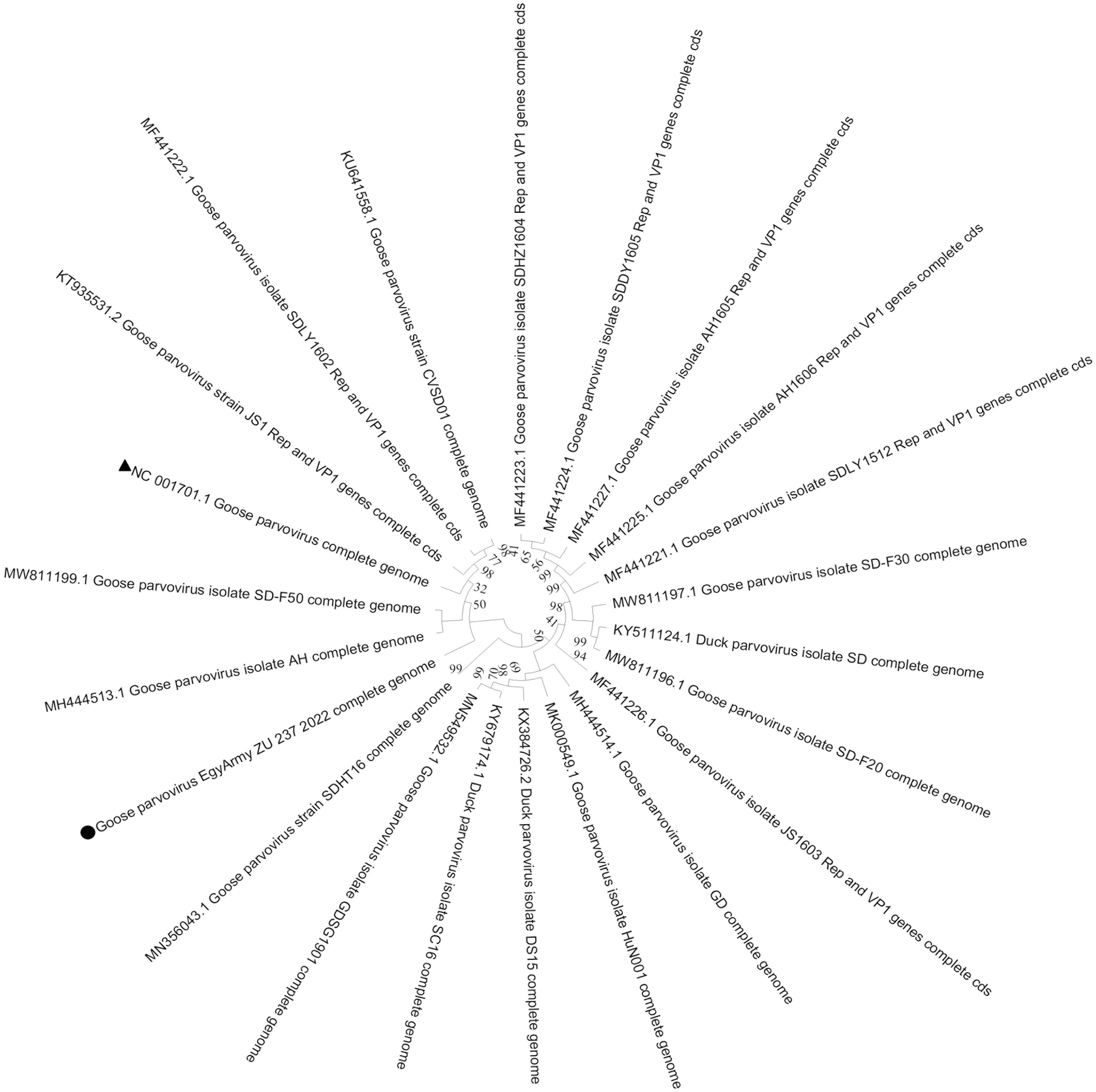
Bootstrap phylogenetic trees based on the full genome sequences of 20 novel goose parvovirus strains closely related to the GPV-EgyArmy-ZU-237-2022 strain; 500 replicates. The circle refers to the strain in current study, and the triangle refers to classical goose parvovirus strain B.
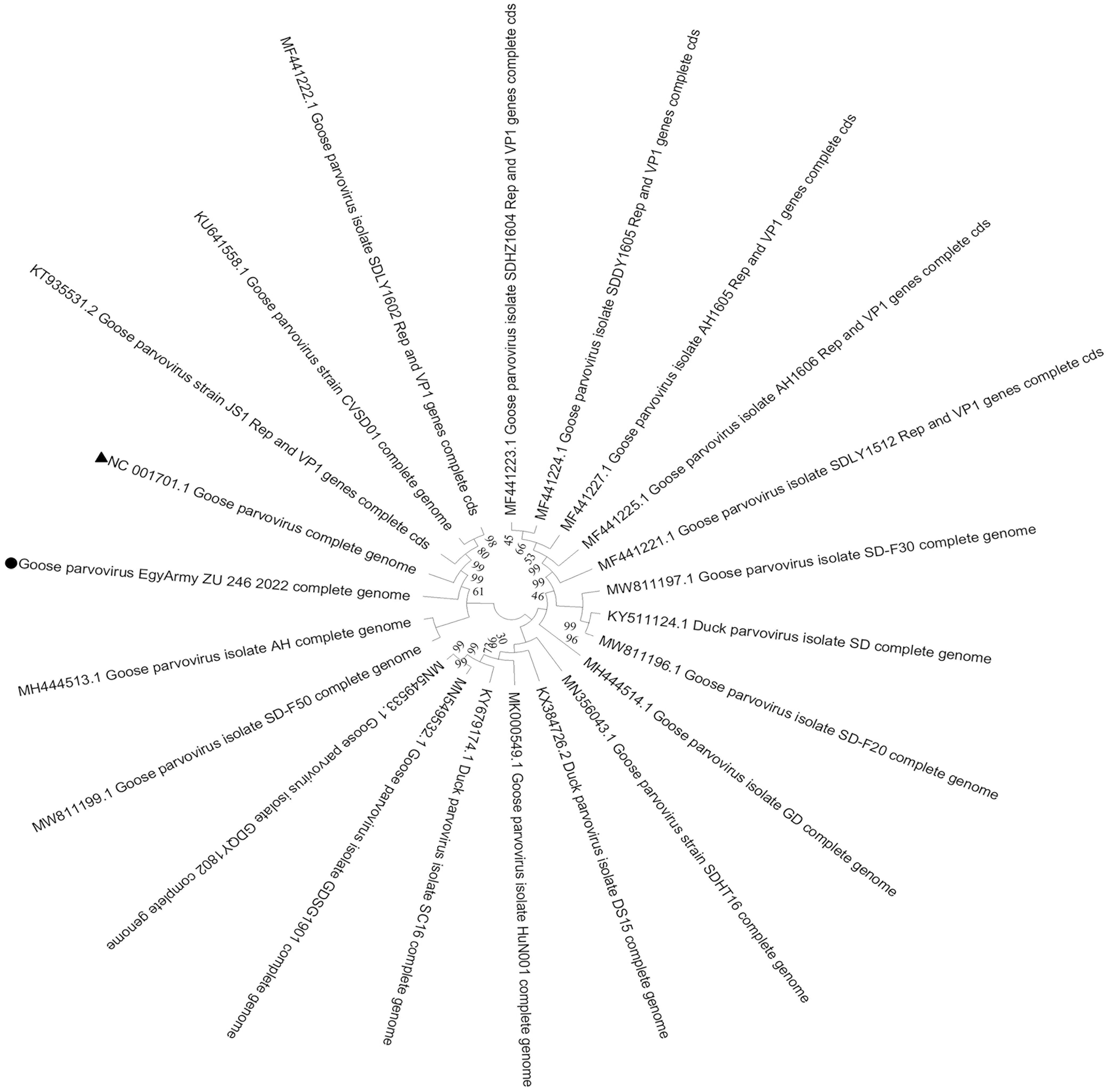
Bootstrap phylogenetic trees based on the full genome sequences of 20 novel goose parvovirus strains closely related to the GPV-EgyArmy-ZU-246-2022 strain; 500 replicates. The circle refers to strain in current study, and the triangle refers to classical goose parvovirus strain B.
Gene annotation of the GPV-EgyArmy-ZU-237-2022 and GPV-EgyArmy-ZU-246-2022 strains as well as the nucleotide identity of the classical GPV B strain with the two strains is shown in Table 2.
Genome annotation and the nucleotide identity with classical goose parvovirus B strain of the two full genome–sequenced strains under the present study.
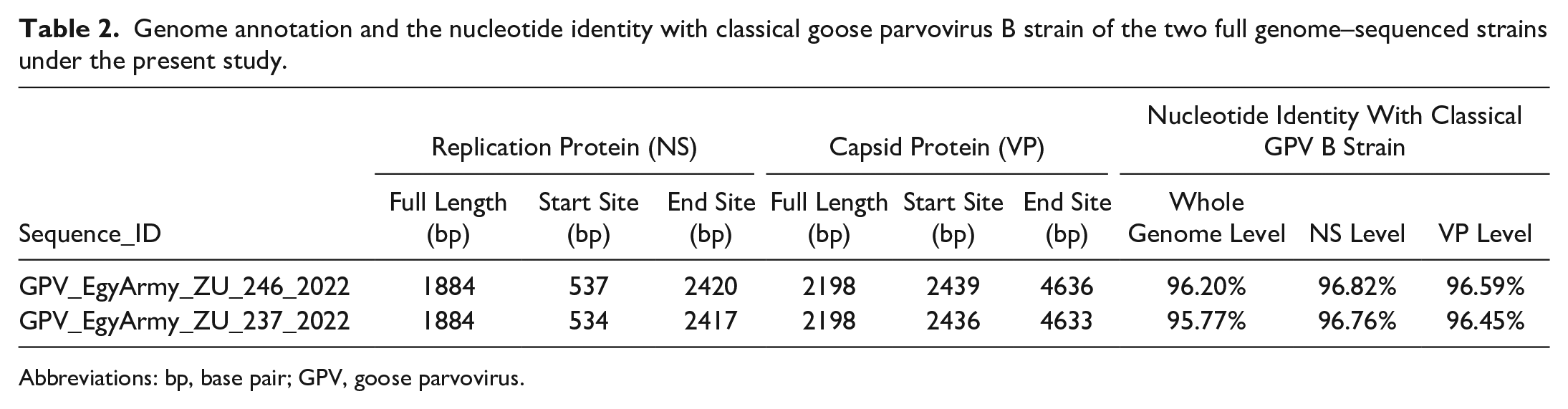
Abbreviations: bp, base pair; GPV, goose parvovirus.
Discussion
Since detailed studies about NGPV pathogenesis are still not sufficient enough to understand the full scope of the disease, 17 the current study provides a new perspective on the viral pathogenesis and demonstrates the drastic impact the virus has on the tongues of NGPV-affected ducks.
Among the examined Pekin ducks, high morbidity rates, low mortalities, and beak atrophy with or without tongue protrusion were presumptively indicative for NGPV infection during the field diagnosis. Accordingly, the same clinical signs were previously recorded in several studies investigating naturally and experimentally occurring SBDS.7,8,29 The low mortality and beak atrophy are discriminating between fully susceptible, maternal antibody-free, classical, and novel GPV-infected ducks. Since classical GPV has mild pathogenicity in mule and Pekin ducks, NGPV, a variant GPV strain, is not completely adapted to cause severe disease in such duck breeds, which might be the reason for the low mortality caused by this strain.18,26,29
The tongues of adult Pekin ducks were necrotic, swollen, congested, and protruding, compared to the apparently normal nonprotruding tongues (31.5%) in 15-day-old ducklings. With increasing age, affected ducks had smaller beaks with increased tongue protrusion and more severe inflammation. The harmful impact of such tongue lesions on duck growth performance is seen as major economic losses among affected flocks. Supporting the presence of NGPV infection in the tongue prior to inflammatory changes, PCR results revealed the positive NGPV detection in protruding tongue pools of 4 adult flocks and nonprotruding tongue pools of 5 young flocks. The urgent question was whether NGPV is the sole infectious agent or there are other factors that could be the cause of such inflammatory reactions among protruding tongues, such as bacterial coinfections, oxidative stress, or adverse environmental conditions.
Microscopical examination of the tongue revealed that the necrosis and vacuolar degeneration of the superficial epithelial layer of the protruding tongues were the main features in NGPV-affected adult ducks. Although the nonprotruding tongues of NGPV-positive young ducks were macroscopically normal, they had severe propria submucosal inflammation characterized by multifocal lymphoplasmacytic aggregates, congested blood vessels, and edema. Neither necrosis nor vacuolar degeneration was microscopically detected in young Pekins. No heterophilic infiltration was detected either in adult or young ducks, although it did not necessarily mean the absence of bacterial infection. Furthermore, the propria submucosa was more severely affected in the young than the adult. These histologic findings correlated with the IHC results in which viral antigen immunolabeling was mainly detected in the propria submucosa of tongues with a higher immunoreactivity observed in the young ducks than in adults. Histopathological examination of tongues of NGPV-suspected cases was only performed in one experimental study, in which affected protruding tongues were microscopically characterized by interstitial inflammation, loose connective tissue stroma, and edema. 5 Such results were comparable to our findings.
Intestinal tissue pools of all examined Pekin flocks were PCR-positive and were confirmed by next-generation sequencing as NGPV infection. The small intestine is the key marker for WFPV infection as a primary target organ of parvovirus replication that leads to significant damage in the intestinal structure and enteric clinical disease. 17 Parvovirus replication takes place in cells undergoing mitosis because the virus needs a polymerase to replicate efficiently. 34 It has been suggested that, following the infection, replication occurs in the intestinal wall. The virus enters the bloodstream and reaches its target organs. 28 The virus DNA detection in the different tissues of affected waterfowls is consistent with the observed clinical signs and pathological changes. 39
There are two proliferative growth zones in the beak frontonasal mass which are associated with the bone morphogenetic protein 4 activity. 40 The beak development is influenced greatly by the NGPV infection. The shortening of the beak is suggested to be a consequence of the virus’s effect on beak morphogenesis, as the bone morphogenetic protein 4 activity may be negatively affected during NGPN infection, which in consequence can result in the abnormal beak shape. 29
Conventional PCR confirmed the absence of MDPV infection in this study. Only two studies declared that SBDS may be also caused by MDPV, whether it was accompanied by tongue protrusion or not.13,22 PCR techniques cannot differentiate between GPVs and NGPVs due to the high genome similarity between these classical and variant strains. 35 Such differentiation based on PCR can discriminate only between GPVs (either classical or variant strains) and MDPVs. 36 As a result, all isolates in the current study could be identified as GPVs depending on PCR. As differentiation is not only mediated by PCR but also by sequencing techniques, next-generation sequencing and phylogenetic analysis of whole genome sequences of both isolates (GPV-EgyArmy-ZU-237-2022 and GPV-EgyArmy-ZU-246-2022) proved both as variants of GPV (later named NGPV), with up to 98.8% nucleotide similarity among each other, 98%–99% with other NGPV strains, and only 95.77%–96.20% with the classical GPV strain, on the basis of whole genome level.
Since the full NGPV molecular identification in 2015, a great genetic similarity (up to 99%–99.5%) between NGPVs has been recorded either in Mainland China2,9,19 or in other countries, such as Poland and Vietnam,12,23 compared to the lower similarity with classical GPV strains(up to 96.5%). In our study, next-generation sequencing confirmed the absence of any other viral infections that may be associated with enteritis and growth retardation such as duck circovirus and duck enteritis virus. Next-generation sequencing has allowed not only for the accurate detection of viral genomes affecting avian species but also the study of genomic diversity among them. 16
All examined Pekin flocks were collected from NGPV endemic areas. The high parvovirus resistance against adverse environmental conditions, and the difficult inactivation of parvovirus with several commercial disinfectants,31,33 allows the virus to persist in the environment for a long period, allowing the environment to be a consistent source of infection probably of one or more circulating strains.
Until now, there is no commercial vaccine against SBDS and no commercial vaccine containing NGPV strains. Based on the high identity between NGPV and classical GPV, it is more likely to employ inactivated and live-attenuated vaccines containing classical strains to prevent and control SBDS infection. As the cross-protective efficacy is not enough for SBDS prevention, it is urgent to develop matched vaccines to include recent isolates and effectively control SBDS.2,9 The genetic evolution of WFPVs consequently leads to vaccination failure because of outdated commercial vaccines and/or defects in the vaccination regimen itself. 21 Since the first SBDS detection in Egypt in 2020, advanced research has been carried out for investigating the current situation of NGPV in Egypt including pathogenicity, immunogenicity, and vaccine evaluation studies, which will give an in-depth knowledge for better SBDS prevention and control strategies.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858241249108 – Supplemental material for Short beak and dwarfism syndrome among Pekin ducks: First detection, full genome sequencing, and immunohistochemical signals of novel goose parvovirus in tongue tissue
Supplemental material, sj-pdf-1-vet-10.1177_03009858241249108 for Short beak and dwarfism syndrome among Pekin ducks: First detection, full genome sequencing, and immunohistochemical signals of novel goose parvovirus in tongue tissue by Amal A. M. Eid, Mohamed A. Lebdah, Sarah S. Helal, Mohamed G. Seadawy, Abdelgalil El-Gohary, Mohamed R. Mousa, Ayman H. El-Deeb, Fakry F. Mohamed and Reham M. ElBakrey in Veterinary Pathology
Footnotes
Author Contributions
SSH performed the practical work starting from the study design, clinical and postmortem examination, sample collection, nucleic acid extraction, PCR detection, manuscript writing and publishing process; AAME supervised the practical work starting from the design of work plan until the publishing process; MGS supervised the molecular work and performed the sequencing and phylogenetic analysis; MRM performed the histopathology, immunohistochemistry, and the design of figure plates; AE and MAL shared the supervision process; AHE contributed to the molecular detection and publishing process; FFM contributed to the manuscript language editing and publishing process; RME contributed to clinical and postmortem examination and publishing process.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
