Abstract
The skin and adnexa can be difficult to interpret because they change dramatically with the hair cycle throughout life. However, a variety of methods are commonly available to collect skin and perform assays that can be useful for figuring out morphological and molecular changes. This overview provides information on basic approaches to evaluate skin and its molecular phenotype, with references for more detail, and interpretation of results on the skin and adnexa in the mouse. These approaches range from mouse genetic nomenclature, setting up a cutaneous phenotyping study, skin grafts, hair follicle reconstitution, wax stripping, electron microscopy, and Köbner reaction to very specific approaches such as lipid and protein analyses on a large scale.
The skin in all species is considered by mass to be the largest of the intermediate-sized organs. 24 In addition to providing a barrier to the outside world, skin has a variety of functions including acquired and innate immunity, vitamin D metabolism, mechanics (stretch and compression), wound healing, sensory and autonomic functions, sociosexual communication, and thermoregulation.15,83
The laboratory mouse is the predominant animal model in biomedical research. 51 The most obvious cutaneous difference between many of the inbred strains of mice is coat color, 71 which determines the skin color. 96 However, each strain and even substrain may have major or minor differences in the skin and adnexa, such as AKR/J mice, which are all homozygous for a mutation in sterol O-acyltransferase 1 (Soat1ald/ald) that results in defects in the medulla, changing the appearance of the hair.22,108 In addition, each inbred strain has a predilection for developing specific diseases, sometimes totally distinctive to that strain, due to a specific mutation(s) or strain-specific background modifier genes,3,79,101 making it imperative to properly designate the full strain designation and any known genetic mutations it carries. 94 However, despite the use of relatively routine protocols for genetic engineering and chemical mutagenesis on a large-scale basis,7,8 determining the effects of a single gene mutation on the skin and adnexa in an inbred strain can still be a daunting task. We present here a relatively straightforward approach for working up new skin mutants with examples of specific state-of-the-art technologies to characterize pathological and molecular details. These approaches work well regardless of the cause of the mutation (genetic engineering, mutagenesis, and so on), including the spontaneous mutant mice that continue to arise in colonies worldwide.
Nomenclature
Pathologists are trained to use specific terminologies, including synonyms, for description of lesions and final diagnoses. Many consensus meetings have been set up to standardize mouse pathology nomenclature with the expressed goal of coordinating terms with those used for human diseases. 19 However, it is surprising how few pathologists and mouse biomedical researchers understand the highly standardized mouse genetic nomenclature. This is critical to ensure that investigators are working with the mice they think they have in their studies. Investigators often name the spontaneous mutations themselves, not infrequently using names or symbols already assigned to unrelated genes and allelic mutations. For example, a search for P120 yields 10 different genes, P38 yields 13, and Scd1 yields 4 (Scd1-4) (http://www.informatics.jax.org/marker/summary, 5 September 2022). The same is true for strain names. Many investigators use the abbreviation B6 for any C57BL/6 mouse strain. This designation is specifically restricted for C57BL/6J. Other substrains have additional letters added, such as B6NJ for C57BL/6NJ. These two strains, B6 and B6NJ, have a number of spontaneous mutations that make them different and can affect interpretation of results.36,72
The International Committee on Standardized Genetic Nomenclature for Mice (http://www.informatics.jax.org/nomen/) is currently run through the Mouse Genome Informatics Database at The Jackson Laboratory. It is important to contact the nomenclature committee directly soon after creating a new mutation to have a specific designation assigned so that the name is used in all subsequent works to avoid mislabeling. This way, the official designation can be used in all publications.
The basic designation is that inbred strains are all capital letters before the slash and capital and lower case after the slash, with a holder designation at the end. None of the letters are in italics. For C3H/HeJ mice, C3H is the strain, He (Dr. Heston developed the substrain), and J for the holder, The Jackson Laboratory. For the spontaneous mutation in the toll-like receptor 4 gene, the gene is designated Tlr4, and the allele defective lipopolysaccharide response is Lps-d, all in italics with the allele designation being superscripted (Tlr4Lps-d). The human gene would be listed in all capital letters and italics (TLR4). The protein is listed as TLR4, all in capital letters not in italics for both mice and humans. In this example, the entire mouse strain with this allelic mutation is designated C3H/HeJ-Tlr4Lps-d. There are many different designations for induced mutations that indicate how they were made. Examples are shown in Table 1 for the Tlr4 gene. There are many other specific genetic tools (such as consomic strains, where one chromosome has been replaced with that of another strain) available as described in detail elsewhere. 78
Examples of various nomenclatures for mutations in the mouse toll-like receptor 4 gene.
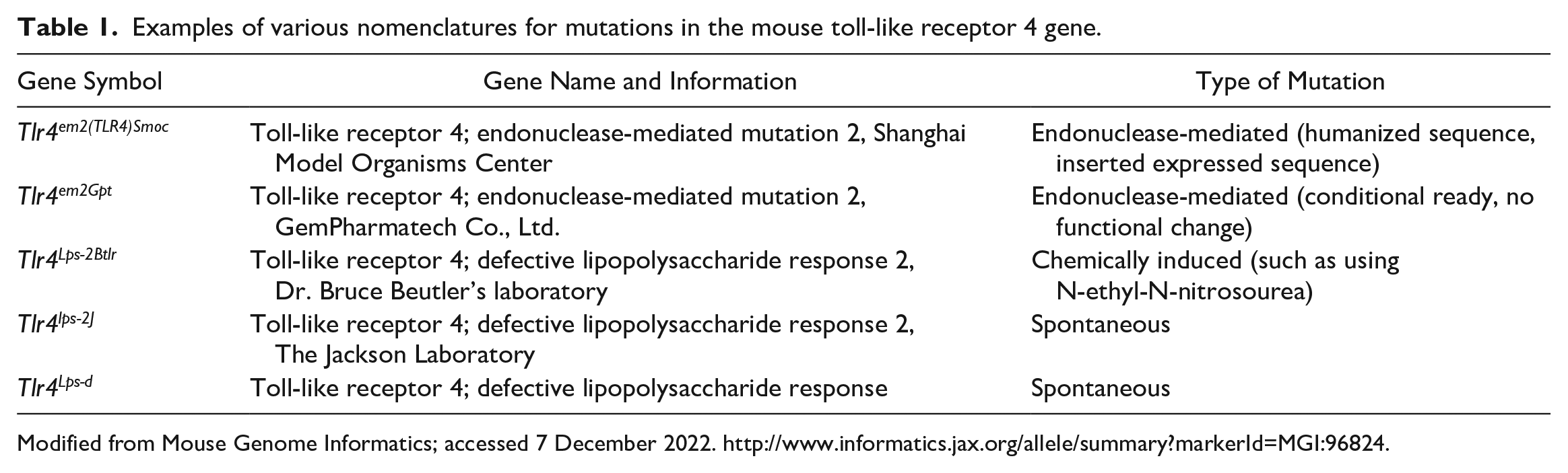

Modified from Mouse Genome Informatics; accessed 7 December 2022. http://www.informatics.jax.org/allele/summary?markerId=MGI:96824.
As a specific example of how to use nomenclature, creating white (albino) mice from pigmented mice can illustrate its importance. Pigment, regardless of type, can interfere with interpretation of phenotypes or specific assays, such as immunohistochemistry using dark-brown chromogens (diaminobenzidine) or looking for fine details with hair shafts. To circumvent this difficulty, it is possible to make a pigmented mouse white simply by crossing it with an albino mouse with a mutation in the tyrosinase gene. It is important to match the inbred strain background. For example, C57BL/6J wildtype mice, that are black, can be crossed with C57BL/6J-Tyr2J or C57BL/6NJ with either C57BL/6NJ-Tyrem31/GrsrJ or B6(Cg)-Tyrc-2J. Although this seems like a minor issue, C57BL/6J and C57BL/6NJ have a number of different polymorphisms (mutations) 72 that affect the frequency of B6 alopecia and dermatitis, 101 the presence or absence of retinal disease, 36 and a variety of other confounding phenotypes.
Mouse Colonies
Mouse colonies should be maintained as clean as possible behind limited-access barriers in environmentally controlled facilities. Small colonies consisting of a few breeding pairs may be sufficient for a small phenotyping program. Mapping studies for gene identification should be done using 2 inbred lines, one carrying the mutation under investigation, and the other being wildtype. Each hybrid generation should be observed for the abnormal phenotype as well as any change in disease severity, lesions identified, onset of disease, life span, or other phenotypes, all of which can point to the effects of major or minor modifying genes. If the gene is known and a molecular genotyping assay is available, it can be relatively simple to maintain a small colony if mutant mice are viable. If the mutation is recessive, then only mutant and control mice are needed for phenotyping. If the mutation is semidominant, then homozygous mutant, heterozygous mutant, and wildtype mice will be needed. Wildtype (Gene+/+) mice should be collected from the same colony at the same time as mutant mice for an accurate comparison. Gene mapping has changed over the decades from linkage with coat color genes, stable integration of retroviruses with Southern blots, 103 and PCR using microsatellite markers 82 to single-nucleotide polymorphisms now done on large arrays. 33
Numbers and Ages of Mice
A common question is how many mice of which sexes and ages should be evaluated for a minimum workup? Statisticians say a minimum of 3 mice are needed for each group (of inbred mice, not outbred stocks), but more is better. Traditional approaches suggested that a single animal with pronounced clinical signs is adequate to determine the primary phenotypic abnormalities. While this can be a starting point, it is always good to evaluate more than 1 mouse to confirm that changes seen are reproducible. However, the initial question is what other changes occur besides those in the organ of interest, namely the skin and adnexa. A comprehensive initial evaluation can be performed with a minimum of 2 female and 2 male mice, mutants and controls (8 total), at several ages (Fig. 1). While a disease (a lesion) is usually thought about in 3 dimensions (height, width, and depth), time is a fourth dimension that should be included in the evaluation of a new mutant. 25 Major changes in any animal’s life provide pivotal points for evaluation: birth (day 0), weaning (3 weeks), puberty (6 weeks), adulthood (15 weeks), and if the mice live long enough, geriatric age (12 months or older). 69 This has been the approach taken by the International Knockout Mouse project as a first histological screen of mice at 15 weeks of age. 68 Aging studies require larger numbers of mice, such as groups of 15, often just to allow for sufficient numbers to reach 2 years of age.79,80 Ages at which mice are weaned and go through puberty are about the same for most strains. The age when mice stop breeding varies, sometimes dramatically, by inbred strain, 21 so one may need to modify the ages for some strains if reduced fecundity is the definitive goal. By standardizing the ages, using the same age groups for each mutant mouse line evaluated, the results become more consistent and comparable across studies. This approach, when supported by detailed systemic histopathology, provides good insight into all organ systems and will define all future work with these mice.
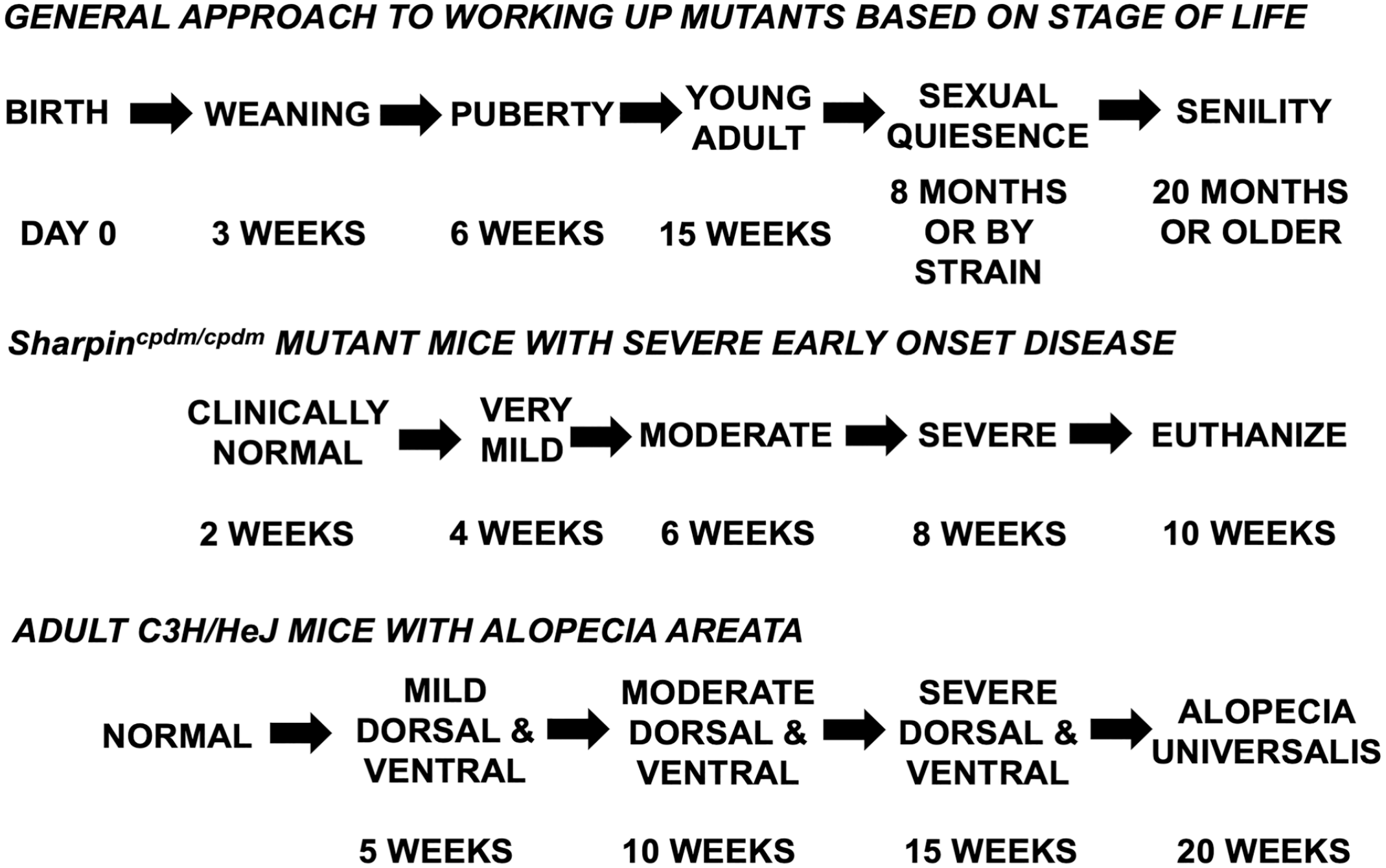
Several options are available for times or ages to collect samples for evaluation based on age or progression of the phenotype.
An alternative approach to evaluate time as a part of the phenotype is once the clinical phenotype is known, from initial onset of lesions to progression to severe disease, and then mice can be collected just before lesions appear and at regular intervals before the time of death (natural or euthanasia due to disease severity). For example, chronic proliferative dermatitis mutant mice (Sharpincpdm/cpdm) appear to be normal at 2 weeks of age but have to be euthanized by 10 weeks of age.35,53 Therefore, studying them at 2, 4, 6, 8, and 10 weeks of age provided useful intervals to help work out the molecular pathogenesis of the disease. Another example is alopecia areata in C3H/HeJ mice, a cell-mediated autoimmune disease that causes hair loss that naturally waxes and wanes (Fig. 1). 84 However, in the skin graft model of alopecia areata, 41 the disease progresses in a predictable manner, and mice can be followed up at 5-week intervals after skin engraftment (Fig. 1). 12 Such systematic approaches provide insight into how to grade progression of disease, which can be applied to therapeutic response. 97
As the hair cycles throughout life and mice are altricial, not fully developed at birth, careful evaluation at multiple time points is essential. In addition to in-depth analyses at the ages or disease stages discussed above, the hair cycle provides additional challenges to interpretation. Mice with darkly pigmented hair (black, agouti, or gray) can have their backs shaved and followed up over time. Hair shafts in anagen are pigmented, which can be visualized and followed up over time in the shaved areas at the gross level (Fig. 2a) with comparison to histologic staging (Fig. 2b, c).96,104,105 Gray scale cards can be prepared to provide scoring systems for the new hair growth. 14 No two littermates are in quite the same stage of the hair cycle if necropsied at the same time. Therefore, groups of 3 or more are needed to generate mean numbers for comparison between mutant and control mice. If mice are euthanized every 3 days from birth through the embryonic (18 days postpartum) and first adult hair cycles (up to 40 days of age), it is possible to graph the hair cycle by measuring hair follicle length). This is complicated by the fact that there are 4 different types of hair follicles producing 4 different types of hair shafts in the mouse. However, the percentage of hair follicles can be estimated based on anatomical criteria into anagen (score: 3), catagen (score: 2), or telogen (score: 1) and graphed using a computerized spreadsheet (Fig. 2b, c). 96 Graphic presentations of these data can be created with this approach or by using detailed scoring systems for the mouse hair cycle. 47 The phenotype can be dramatically different from late anagen where a diagnostic feature is evident to catagen when severe follicular dystrophy is the prominent feature, as is the case with the various allelic mutations of the desmoglein 4 (Dsg4) gene.43,81
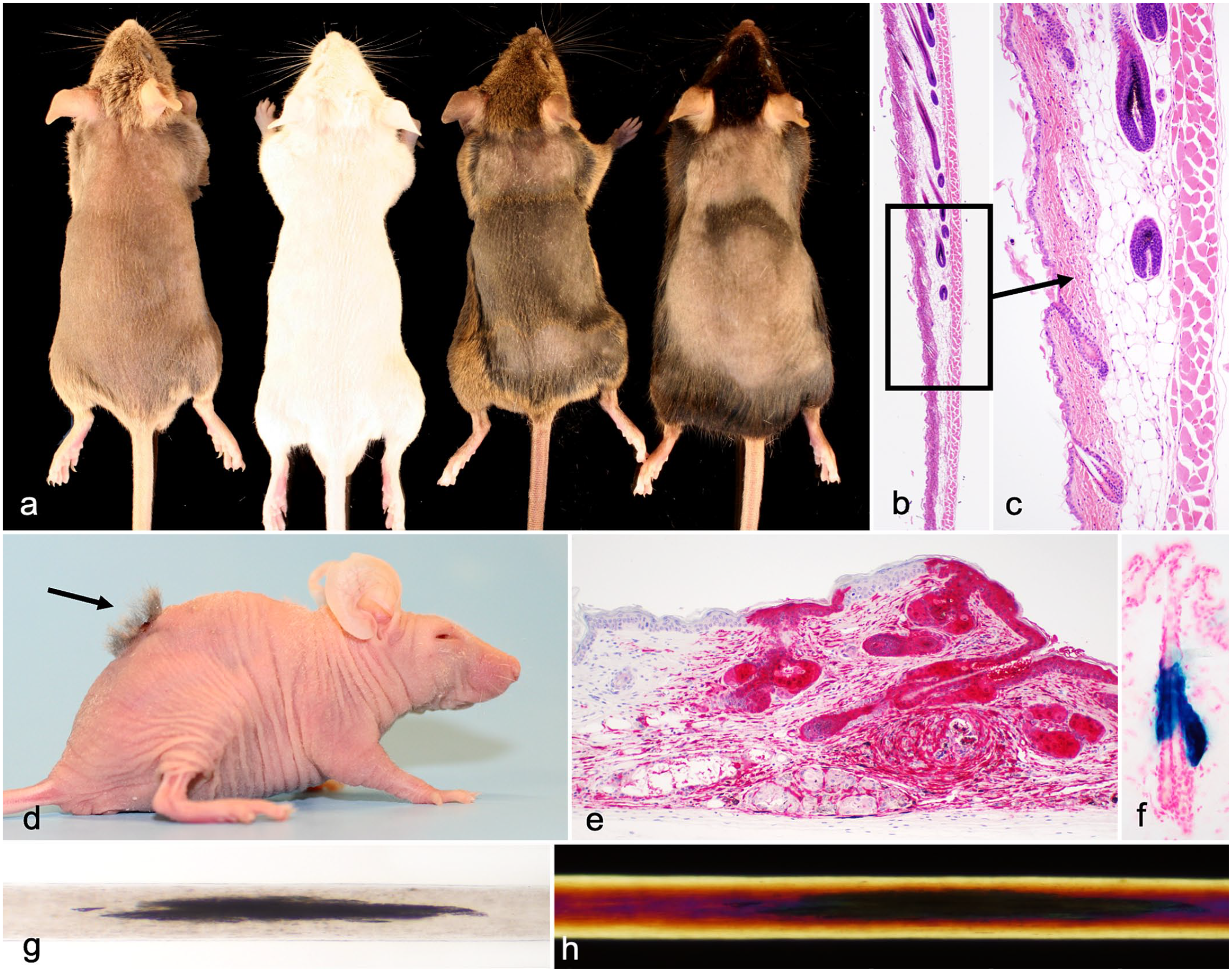
(a) Four different mouse strains shaved to show the grossly evident hair wave. Only the heavily pigmented strains (C3H/HeJ and C57BL/6J, right) have dark areas indicating actively growing hair. Left to right: DBA/2J, BALB/cByJ, C3H/HeJ, and C57BL/6J. (b) Histology of the junction of a hair wave to illustrate transition from telogen to anagen. Skin, mouse, C57BL/6J. Hematoxylin and eosin. (c) Higher magnification of Fig. 2b. (d) Hair follicle reconstitution (arrow) using cells from a C57BL⁄6-Tg(UBC-GFP)30Scha⁄J mouse (that expresses green fluorescent protein [GFP] in all cells) grafted onto an albino nude mouse (CByJ.Cg-Foxn1nu/nu) that lacks T cells. (e) Immunohistochemical detection of GFP in donor cells to differentiate them from the host cells from the mouse in Fig. 2d. Immunohistochemistry using a rabbit anti-GFP antibody. (f) Far2tm1(KOMP)Wtsi/2J mouse (normal mouse with a lacZ reporter gene) expressed LacZ protein only in sebaceous glands in the skin where fatty acyl CoA reductase 2 (FAR2) is expressed. (g) X-gal staining of frozen sections. Light and dark bands in a human hair with pili annulate. Whole mounted human hair evaluated using white light. (h) Red and dark bands in a human hair with pili annulate. Same human hair as in Fig. 2 but evaluated with polarized light.
Quantitative Measurements
Many mice carrying mutations that affect the skin have no obvious morphologic abnormality as the change may only be in the hair cycle, such as with a null mutation in the fibroblast growth factor 5 gene (Fgf5go, angora mice). 93 Measuring a variety of cutaneous parameters can be useful to help define a mutant phenotype, such as the full epidermal thickness (or Malpighian layer and stratum corneum), dermis, hypodermal fat layer, sebaceous glands, and length of hair follicles. Epidermal thickness, including Malpighian layer versus the stratum corneum, can be useful when studying scaly skin disease models, especially when using cre-recombinase transgenics to delete the expression of a particular gene in different tissue types to define the response. 92 Several of these anatomic structures, notably the hypodermal fat layer, sebaceous glands, and hair follicles, vary in size with the hair cycle, making them another useful way to define the hair cycle. It is important to have samples that represent all the phases of the hair cycle for this reason. Sometimes, the phenotype is best seen at specific hair cycle stages. For example, subtle abnormalities in hypodermal fat can appear normal in telogen and require skin with anagen hairs to identify the lesion. 74
To obtain reproducible results and to minimize variability, it is critical to use an internal histologic standard and not randomly make measurements. The hair follicle provides such a reference point. Measurements should only be taken in areas of sections where all or most of the hair follicles are observed. Digital cameras have manual measurement options that allow fairly rapid manual collection of this type of data. 53
Epidermal proliferation rates can be quantified using bromodeoxyuridine (BRDU) incorporation. Age- and sex-matched control and mutant mice are injected intraperitoneally with BRDU at 50 µg/g of body weight to label cells entering the “S” phase of the cell cycle. After 1 hour (mice 12 days of age and older) or 2 hours (newborn mice), mice are euthanized, and tissues are collected and fixed for routine histology. Immunohistochemistry is performed using an anti-BRDU antibody. Positive nuclei are counted per 1000 cells or per millimeter if the skin is collected and processed flat. Alternatives that use existing paraffin sections include labeling with antibodies directed at proliferating cell nuclear antigen or Ki67. Mitotic rates can be calculated in a similar manner in routine hematoxylin and eosin–stained sections or with only hematoxylin. 73
Skin Grafts and Hair Follicle Reconstitution
Skin grafting was a method used to work out the concept of histocompatibility 31 and later for genetic quality control. 1 It is also a useful tool to investigate fundamental mechanisms of a mutant mouse phenotype. A major inflammatory response in mouse skin poses a problem as to how to determine if the fundamental underlying issue is in the mouse’s immune system or a normal immune response to something abnormal in the skin. Full-thickness skin graft is one approach to answer this question. Skin can be grafted from the mutant mouse to a histocompatible littermate or to an immunodeficient mouse. The severe combined immunodeficiency mutation (Prkdcscid) has been moved onto a number of congenic strains making it a useful recipient. C3H/HeJ mice develop alopecia areata, a cell-mediated autoimmune disease, as they age. 84 After grafting skin from these mice onto young C3H/HeJ-Prkdcscid/scid mice that are histocompatible, do not have clinical alopecia areata, and are immunodeficient, the hair regrew in the graft. However, when affected skin was grafted onto clinically normal young C3H/HeJ mice with a normal immune system that are histocompatible, the graft did not regrow hair, but the mouse developed patchy alopecia that progressed over 20 weeks to alopecia universalis, total baldness. 41 The methods are described in detail elsewhere. 70 Similar studies have been carried out with other mutant mice with a scaly skin disease. 85
Another grafting approach used to understand cellular interactions of hair follicles and fibroblasts is hair follicle reconstitution (Fig. 2d). A number of different methods can be used with mice that express a reporter gene in all cells (such as one that expressed green fluorescent protein [GFP] [C57BL⁄6-Tg(UBC-GFP)30Scha⁄J] 67 ) that can be visualized by fluorescence microscopy or immunohistochemistry, 106 to differentiate donor from recipient cells (Fig. 2e). Alternatively, albino recipients (such as the immunodeficient CByJ.Cg-Foxn1nu/nu mice) receiving cells from pigmented donors (such as C57BL/6J) form follicles that are pigmented and represent successful reconstitution. The reconstituted hair follicles in this case are more obvious not just because the donor cells are pigmented but also because the recipient nude mice have severe follicular dystrophy. 34 The reciprocal approach can also be taken as BALB/cByJ wildtype are albino with a normal immune system while B6.Cg-Foxn1nu/numice are pigmented and immunodeficient.
Reporter Genes for Localizing Gene Expression
Reporter genes are used to track the cellular and subcellular localization of a specific gene or protein or monitor cellular activity. Escherichia coli beta-galactosidase (commonly called lacZ) is one such gene that can be included in a gene construct to visualize where a protein is expressed (Fig. 2f). Many are linked in cre-recombinase constructs, and image databases provide insight into their utilization (http://www.informatics.jax.org/downloads/reports/MGI_Recombinase_Full.html). 26 Other reporter gene systems are available including GFP, enhanced GFP, and other gene constructs in a variety of other colors. Antibodies are available to detect a number of these fluorescent proteins; antibodies using IHC provide stronger signals and stable slides that can be archived. 106 Luciferase-based reporters are also available as reported elsewhere.16,17,37,49 These can help define where a cre-recombinase construct is actually expressed rather than just where one expects it to be expressed 92 or where a gene is actually normally expressed. 95 This information can aid with interpretation, especially to separate strain-specific lesions from those associated with the mutated gene under investigation.
Wax Stripping/Depilation to Synchronize the Mouse Hair Cycle
Wax stripping is commonly used to synchronize the adult mouse hair cycle. This method results in mild transitory acanthosis in response to the mild injury to the barrier, which is not seen in unmanipulated normal mouse skin of animals over 2 weeks of age. The hair cycle can be followed up in a reproducible manner using this approach; 96 in adult mice, one can see the progressive changes in hair follicle structure, size, and pigmentation.
Mice are anesthetized with isoflurane (or whatever method the institutional animal care and use committee approves). Puralube, (Pharmaderm Animal Health, Melville, NY) is applied to each eye. Surgi-wax hair remover (American International Industries, Los Angeles, CA) is heated to a liquid using a microwave or a professional wax warmer (Satin Smooth Deluxe Series SSW4C; BaBylissPRO, Glendale, AZ) and allowed to cool to a semiliquid state. When its temperature, measured using a thermometer, reaches 100°F and is confirmed by testing against the wrist of the technician applying it, the wax is then applied to the dorsal hair of the mice from the neck to the base of the tail. Once dried, the matted hair is rolled off from the base to the neck, thereby removing all hair. The mouse hair cycles in a predictable manner following wax stripping, which enables defining lesions at specific time points in the cycle.44,81 This method only works when the hair shaft is strong enough to be pulled out as a unit. If there is a follicular dystrophy such that only the shaft above the surface is removed, such as with nude mice (Foxn1nu/nu), little to no synchronization will occur.
Manual Hair Plucking
Mouse hairs can be plucked manually using the thumb and forefinger to examine hair shafts for structural abnormalities or for ectoparasites. Forceps may damage hair shafts. Gently plucking hairs will easily remove hair shafts from follicles in the telogen (exogen) stage of the hair cycle. Anagen follicles are actively growing, and hair shafts are firmly attached within follicles. Plucking hairs from follicles in anagen may result in damage or induction of abnormalities in hair shafts already weakened by structural defects. Plucking hair shafts from telogen follicles allows for examination of the whole structure from root (club) to tip. Hair should be stored in a clean cryopreservation tube (Nalge Nunc International, Denmark). When studying mutant mice, standardized collection techniques for skin and hair are as important as for any other organ. Hair should be collected from the same area on every mouse in a study such as from the right lateral skin surface from shoulder to the pelvic region. Avoid areas where full-thickness skin will be collected for histologic examination. Plucking hairs will artifactually distort the inner root sheath and other structures in the hair follicle as discussed with wax stripping. If the vibrissae (very long straight hair fibers around the eyes, muzzle, and lower limbs) appear abnormal, samples of these should be plucked as well, from the same side as where the body hair was plucked. The vibrissae should be stored in a separate Nunc tube. Other specialized hair fiber and follicle types can be collected and examined if necessary.
Hair plucking works well with adult mice, as most hair follicles are in prolonged telogen, so they can be easily removed without damage. This is in sharp contrast to human hairs that are predominantly in anagen for long periods of time and are therefore difficult to remove without causing damage. Many physicians question the quality of samples collected in this manner. To avoid this unjustified concern, 1-cm2 sections of skin can be removed, and hair can be examined directly.
For many years, the standard approach to evaluate plucked mouse hair was to place double-sided sticky tape on a glass slide, attach the hairs to the tape, and then look at them under a microscope. Hairs would not be in a single plane, making photomicroscopy difficult to nearly impossible. This can be easily corrected by placing the plucked hairs on a clean microscope slide, gently separating them, placing a drop of mounting medium on the slide (Permount mounting media; Fisher Scientific, Fair Lawn, NJ), and dropping a coverslip over the sample. This forces all the hairs to lay in the same plane and creates a permanent sample that can easily be stored. Slides are examined in a standard compound microscope using white, polarized, or dark-field illumination (Figs. 2g, h, and 3a–e).
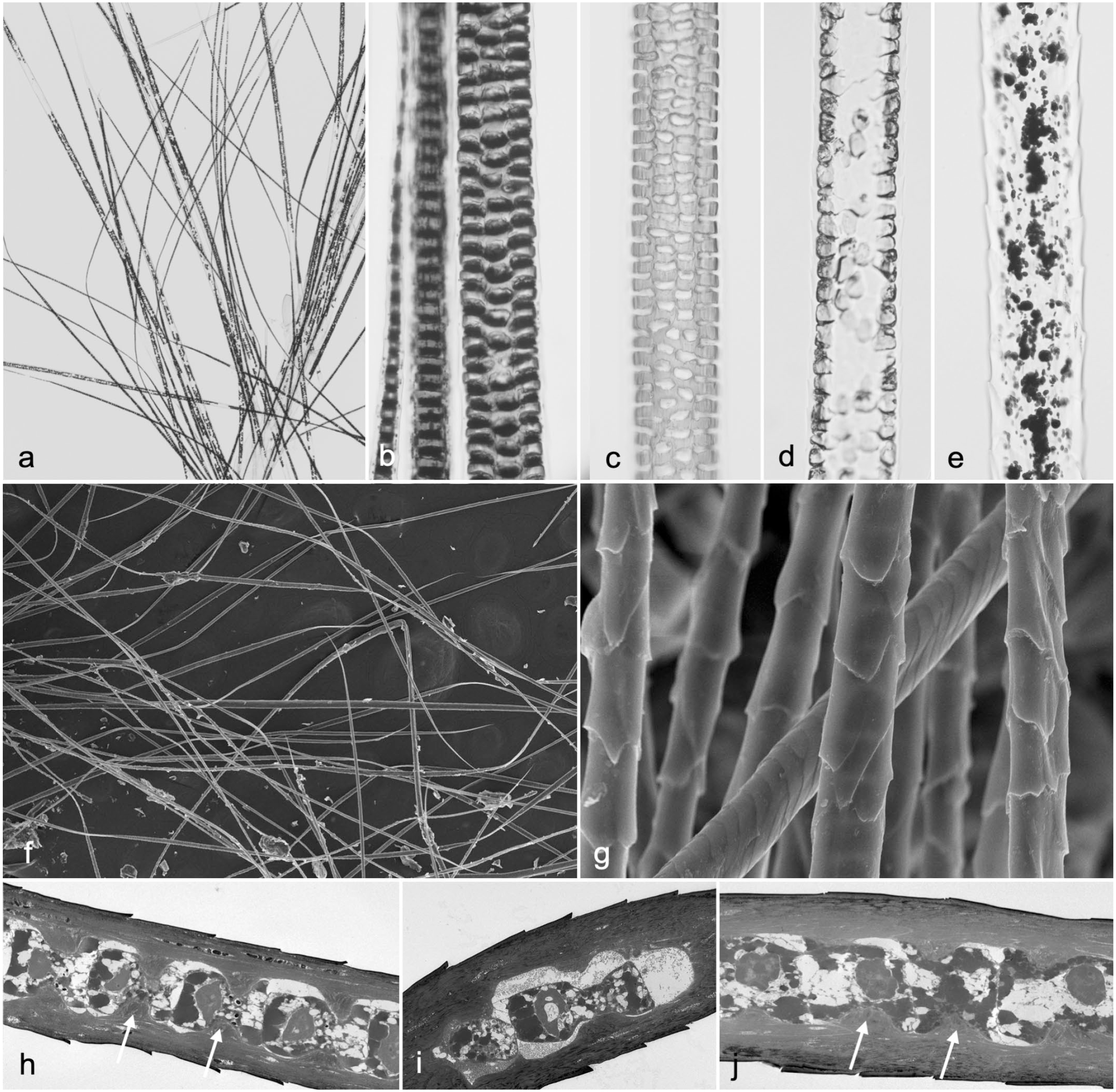

(a–e) Microscopic aspect of manually plucked hair placed on a glass slide and covered with a coverslip using a drop of mounting medium. (a) AKR/J-SOAT1ald/ald mouse. (b) Pigmented C3H/HeJ+/+ mouse. Normal zigzag (septate pattern) and guard hairs (septulate pattern). (c) Albino BALB/cByJ+/+ mouse. Normal guard hair. Note the septulation pattern of the medulla. (d) AKR/J-SOAT1ald/ald mouse. Abnormal guard hair medulla with no septulation. (e) Beige mutant mouse (SBL/LeJ-Foxq1sa/sa Lystbg/bg). Abnormal guard hair. Note the clumped melanin granules (beige phenotype) and lack of septulation of the medulla (satin phenotype). (f–j) Plucked hair. Transmission electron microscopy. (f) B6N(Cg)-Far2tm2b(KOMP)Wtsi/2J mutant mouse with normal hair. (g) Higher magnification of Fig. 3f. Note the variation in cuticle patterns representing different hair types or region of the hair shaft. (h) LP/J pigmented mouse, pretreated hair. Longitudinal sections of a hair. Note the projections (white arrows) that fix the locations of the medulla cells. (i) Albino AKR/J-SOAT1ald/ald mouse, pretreated hair. Longitudinal sections of a hair. Note the projections in the LP/J mouse hair are relatively lacking in the AKR/J hair, which is deficient in trichohyalin. (j) FVB/NJ (albino) mice, pretreated hair. Note the projections are prominent (arrows) similar to those in the LP/J hair in Fig. 3h.
Microbial Screening
It is always important when working up a new mutant mouse with a skin phenotype to routinely run a microbiological screen, using standard bacterial and mycological cultures. Hair pluck evaluation as well as routine histopathology should identify ectoparasites, a common cause of alopecia in mice. It is surprising how many institutions have colonies heavily infected with ectoparasites. Some viral infections will present with classical cytopathological effects, such as koilocytosis and intranuclear inclusions for mouse papillomavirus infection, 27 or epithelial hyperplasia, ballooning degeneration, and prominent intracytoplasmic inclusions with mouse pox (ectromelia) infection. 13 Routine special stains may also identify infectious agents. It is better to have the data on hand when presenting a new model for a scaly skin disease, such as psoriasis, even if the physicians dismiss it as dermatophytosis.
KÖbner Reaction
The Köbner reaction describes exacerbation of a skin lesion by relatively mild trauma, such as a psoriasis flare following tape stripping. 107 Mutant mice with a scaly skin disease can develop a Köbner reaction following trauma, such as a biopsy.48,77 To test for this phenomenon, remove the hair using an electric shaver to obtain a clean site. Take single-sided sticky tape, stick it on the skin, and gently remove. Repeat several times until there is a slight oozing of serum at the skin surface. Observe the mice for several days watching for exacerbation of the lesions at the site tested.
Collecting and Trimming Skin for Histologic Evaluation
The skin superficially appears to be homogeneous, but when carefully examined, it is clearly not. Different hair types are found on the truncal skin (guard hairs, auchene, zigzag, and awl), eyelids (truncal hairs, vibrissae, and cilia), muzzle (truncal hairs and vibrissae), tail skin (tail hairs), and around the anus (truncal hairs and perianal hairs). It is often overlooked that the distal limbs have vibrissae hairs as well as truncal hairs. 87
Transcriptome studies of the skin can be so different between normal anatomic sites in an individual to suggest they are separate organs.12,99 Therefore, it is important to collect skin from more than one anatomic site. Dorsal and ventral skin can be different in some mutant phenotypes; however, because normal skin can be hard to tell apart, placing dorsal skin samples along with ear (auricular cartilage) and tail (thick epidermis) but ventral skin with muzzle and eyelid skin provides a way to distinguish these sites, as these other tissues have distinct anatomic features.
Dorsal and ventral skin should be fixed on a card or foil, and anterior-posterior aspects are marked on the card. The samples are trimmed longitudinally in the direction of the hair growth, anterior-posterior to optimize chances of getting full hair follicles in vertical sections. Horizontal sections, looking straight down on the sample, can be prepared using a piece of skin measuring approximately 0.7 × 0.7 cm2, haired side down, sectioned from the skin surface down. Tail skin can be removed from the underlying connective tissue and bone at the time of necropsy and trimmed in the same orientation as the dorsal and ventral skin. Alternatively, the skin can be left intact on the tail/tail bone, decalcified, and trimmed in cross and longitudinal orientations. Eyelids can be removed with all the skin of the head as one unit. Eyelids can be cut bisecting both lids then making a second cut just posterior or anterior to the corners of the eyelids so that the upper and lower eyelids remain attached to one another. Muzzle skin can be trimmed by making a cut approximately 3–4 mm from the front edge of the muzzle. The ear can be cut in half lengthwise, and then one of the halves can be cut completely off from the scalp. Lay this half flat on the cutting surface and make a second cut parallel to the first to obtain a section similar in size and shape to that of the dorsal and ventral skin sections. Foot pads and nails are collected by amputating the distal limb, followed by fixing, decalcification, and trimming down the midline. This nail unit should always be processed separately (as is the case with all tissues containing bone) in separate cassettes from other soft tissues (skin samples).
Scanning Electron Microscopy and X-ray Element Analysis
Skin excised surgically at the time of necropsy, skin biopsy punches, plucked hairs (Fig. 3f, g), and amputated distal digits (nails and foot pads) can be examined using scanning electron microscopy (SEM) to get a 3-dimensional perspective of phenotypic deviations. Samples of mouse skin (1.5–2.0 cm2) can be removed from mutant and control mice, placed connective tissue side down on dry nylon mesh, and immersed in cold 2.5% glutaraldehyde in 0.1-M cacodylate buffer. A front and rear foot can be amputated at the carpus/tarsus and prepared in a similar manner. After overnight fixation at 4°C, samples are washed twice with 0.1-M cacodylate buffer and postfixed in 0.5% osmium tetroxide in 0.1-M cacodylate buffer. Samples are then dehydrated in a series of graded ethanols to a final series of 3 changes in 100% ethanol. Samples are critical point dried, by gently flushing specimens 4 times with CO2 for 5 minutes while gradually increasing the temperature in a critical point dryer (Balzers, Union, FL) to 41°C. Over a period of 30–40 minutes, pressure is released slowly to allow CO2 to evaporate while allowing the sample to return to room temperature. The samples are then attached to a clean aluminum SEM stub using double-sided tape and sputter-coated with 4 nm of gold. 2 Element analysis can be performed on hair and nail samples using an X-ray microanalysis system (EDAX Inc., Mahwah, NJ) attached to the SEM, especially for investigating trichothiodystrophy (see Supplemental Data). 42
Transmission Electron Microscopy
Samples of mouse hair (several milligrams) can be collected from a euthanized mouse by gently plucking as most are in telogen in adult animals. Ordinarily, untreated hair is nearly impervious to embedding medium, resulting in sectioning artifacts. However, the shaft can be permeabilized by partial reduction of protein disulfides through incubation in sodium dodecyl sulfate (SDS) plus dithiothreitol (DTT) at room temperature for 1–3 hours. 56 It is then immersed in Karnovsky fixative, postfixed with osmium tetroxide, and embedded in a reduced viscosity epoxy-resin combination (Araldite 6005 and EMbed-812, Thermo Fisher Scientific, Waltham, MA). Ultrathin sections (80 nm) are stained with uranyl acetate and lead citrate and examined by transmission electron microscopy. As shown in Fig. 3h–j, the mild pretreatment before fixation preserves the fine structure well enough to permit visualization of the physical basis for the hair interior defect phenotype. 62 A more thorough treatment with this detergent under reducing conditions, dissolving the keratins and solubilizing 80% of the total protein, yields a limp hair shaft consisting only of isopeptide cross-linked material. 63 Not much protein appears to be extracted from the normal cuticle because of its high content of isopeptide bonding, but the cuticle of hair from the matted mutant (Tmem79ma) shows defects with mild reduction and completely disintegrates with more vigorous reduction in SDS. 64
Digits are amputated at the time of necropsy and fixed in 6% glutaraldehyde in 0.05-M phosphate buffer and decalcified in ethylenediaminetetraacetic acid for 4 weeks. Following decalcification, small tissue blocks are postfixed with osmium tetroxide and embedded in an epoxy-resin combination. Semithin serial sections can be obtained from these blocks stained with toluidine blue for evaluation and selection of regions for ultrastructural evaluation. 29 Ultrathin sections can be made from selected blocks, contrasted with 1% uranyl acetate and 1% lead citrate, and examined by transmission electron microscopy.2,42,55 Figures and descriptions of normal mouse skin and adnexa are available from multiple sources.46,83,88,90
Tissue Collection for Other Special Methods
Skin can be frozen for use in various biochemical, molecular, or immunofluorescence studies. Frozen sections lack detail but maintain epitopes for immunohistochemical studies. Lipids can be detected in frozen sections using oil red O, Sudan black, osmium, and other stains. 38 A 1.5 × 2.0-cm section of skin can be removed from the dorsal surface of the mouse at a standardized location, such as the dorsal interscapular region. This skin is trimmed to approximately 0.3 × 1.5 cm to form strips that run lengthwise from the head to the tail to properly orient hair follicles. The strips of skin are placed on a piece of aluminum foil, and a bead of optimal cutting temperature (OCT) solution (Miles Inc., Elkhart, IN) is run along one edge. The foil is then dipped into liquid nitrogen to instantly freeze the samples. Once frozen, samples are quickly removed from the foil and placed on edge in a clear plastic mold (HistoPrep disposable base molds; Fisher Scientific, Pittsburgh, PA) filled with the OCT solution. The mold is then placed back into the liquid nitrogen and allowed to freeze. Frozen molds are wrapped in foil and stored at −80°C until needed. Both the molds and the foil are labeled, with the accession number used for the animal being necropsied.
In situ hybridization techniques can use tissues collected in fixatives routinely used for histopathology. Currently, commercially available kits can be used for paraffin or frozen sections. Most are now optimized to use freshly collected samples fixed in neutral buffered 10% formalin fixation. However, the sections need to be processed within a few days of collection as the kits usually do not work on archival material.
For molecular studies (gene mapping), spleen, liver, and kidneys are routinely removed at necropsy, snap-frozen with liquid nitrogen in screw-capped plastic vials (NUNC tubes; Nalgen Nunc International, Denmark), and stored at −80°C for subsequent DNA extraction. Skin can be collected in the same manner for transcriptome studies. Collecting multiple tissues, especially storing them in separate freezers, provides backup should one set be destroyed or not be useful for whatever reason. Large-scale single-nucleotide polymorphism arrays can be used for gene mapping (GigaMUGA arrays; https://www.neogen.com/search/?searchString=gigamuga+arrays). 45 These samples can be processed for RNA sequencing and other molecular assays.
Proteomic Analysis
An important issue to address when mouse skin or adnexa look different from normal is the basis for the abnormal appearance. Proteomic analysis offers a good way to assess differences in protein content that may help elucidate the pathogenesis of skin diseases.18,60 In general, differences in appearance are reflected in readily distinguishable protein profiles. Cornified features such as the hair shaft, nail plate, and epidermal stratum corneum are readily accessible by snipping or tape stripping without invasive treatment, but an analysis requires careful sample preparation. Extended reduction of disulfide bonds in the presence of strong denaturants has proved satisfactory in obtaining high protein yields. Although isopeptide bonds in corneocytes will prevent the use of certain peptides in database searching, this impediment involves only a minority of peptides (15%−20% of lysine residues), and thus only a small reduction in protein coverage. This conclusion applies to the detergent-solubilized proteins and to the transglutaminase cross-linked envelope structures that remain insoluble after disulfide reduction in strong denaturants.56,59
Early studies analyzed milligram amounts of cornified sample, but more recent reports exploit the high sensitivity of mass spectrometry using much smaller protein amounts (eg, 2 cm of a single hair shaft). 23 For this purpose, one must exercise great care to avoid exogenous contamination, especially from ubiquitous ambient dust containing fragments of skin and hair. Because mass spectrometry is incompatible with SDS-containing samples, the detergent is removed before analysis, generally before tryptic digestion. For this purpose, precipitation of the protein with ethanol is satisfactory for milligram amounts of sample, but to improve yields for low protein amounts, sodium dodecanoate is used instead of SDS during reduction and alkylation and is removed by acidification and ethyl acetate extraction before digestion. 109
Newborn mouse epidermis can be analyzed after separation from dermis 39 or using tape circles measuring 1 or 2.2 cm in diameter (CuDerm, Dallas, TX) to collect layers of stratum corneum. In the latter case, corneocytes are eluted from the tape by overnight immersion in a solution of 2% SDS in 0.1 M sodium phosphate buffer. 58 After rinsing several times in the buffer to remove adventitious material, the corneocyte proteins are reduced in a mixture of 2% SDS, 50-mM DTT, and 0.1-M ammonium bicarbonate buffer and then alkylated with iodoacetamide. Reducing hair samples at elevated temperatures 109 results in approximately 90% protein solubilization by the digestion. Yields of keratin-associated proteins are higher with lower-temperature treatment although total protein solubilization by digestion is lower. 23 Mouse nail plate is isolated by dissection from digits after boiling in a solution of 2% SDS in 0.1-M phosphate buffer (pH 7) with careful removal of any attached tissue. The nail is heated in a mixture of 2% sodium dodecanoate, 50-mM ammonium bicarbonate, and 50-mM DTT for 5 minutes in a boiling water bath and then incubated overnight in a 70°C oven before alkylation with iodoacetamide, detergent removal, and tryptic digestion. 86
Mass spectrometric analysis of digests and database searching provide a protein profile of hundreds or even thousands of protein constituents in a sample. Compilations of these proteins in software such as Scaffold (https://www.proteomesoftware.com/products/scaffold-5) offer convenient platforms to compare relative amounts of a given protein in parallel samples using (weighted) spectral counts. Estimates of the amount of protein by label-free quantification permit comparisons of relative amounts of different proteins. 59 In either case, one must be aware that identical peptides can be generated from different proteins. This phenomenon is encountered particularly in families of highly homologous proteins such as the keratins. Thus, quantitative estimates are sometimes limited to clusters of such proteins. Some examples where this approach has been used are given below.
Epidermis
Often large differences in appearance are reflected in large differences in protein profile. For example, mice lacking the suprabasal epidermal AP1 transcription factor function exhibit a striking keratoderma phenotype. Befitting the important role of AP1-mediated transcription in epidermal differentiation, the proteomic analysis revealed dramatic differences compared with control mice as a result of the gene targeting in levels of keratins and other filament proteins, junctional proteins, metabolic enzymes, protease inhibitors, and proteins serving as transglutaminase substrates. 66 Figure 4a–c illustrates examples in several of these categories. Proteins expressed at lower-than-normal levels indicate suppression of suprabasal differentiation, and those at higher-than-normal levels appear to reflect intracellular stress. Cross-linked envelopes in the stratum corneum are known to be comprised of a broad spectrum of proteins, a major fraction of which are keratins.30,59 This finding confirms by proteomic analysis that mice with ablation of type 1 or 2 keratins have defective cross-linked envelopes and numerous barrier defects. 32 By contrast, a proteomic analysis can also reveal compensatory changes that mask expected functional or morphological defects. In the loricrin (Lortm1Der) knockout mouse, for instance, numerous other proteins that can serve as transglutaminase substrates are induced to yield cross-linked envelopes that function normally, helping rationalize that the adult skin is indistinguishable from that normal in appearance. 59
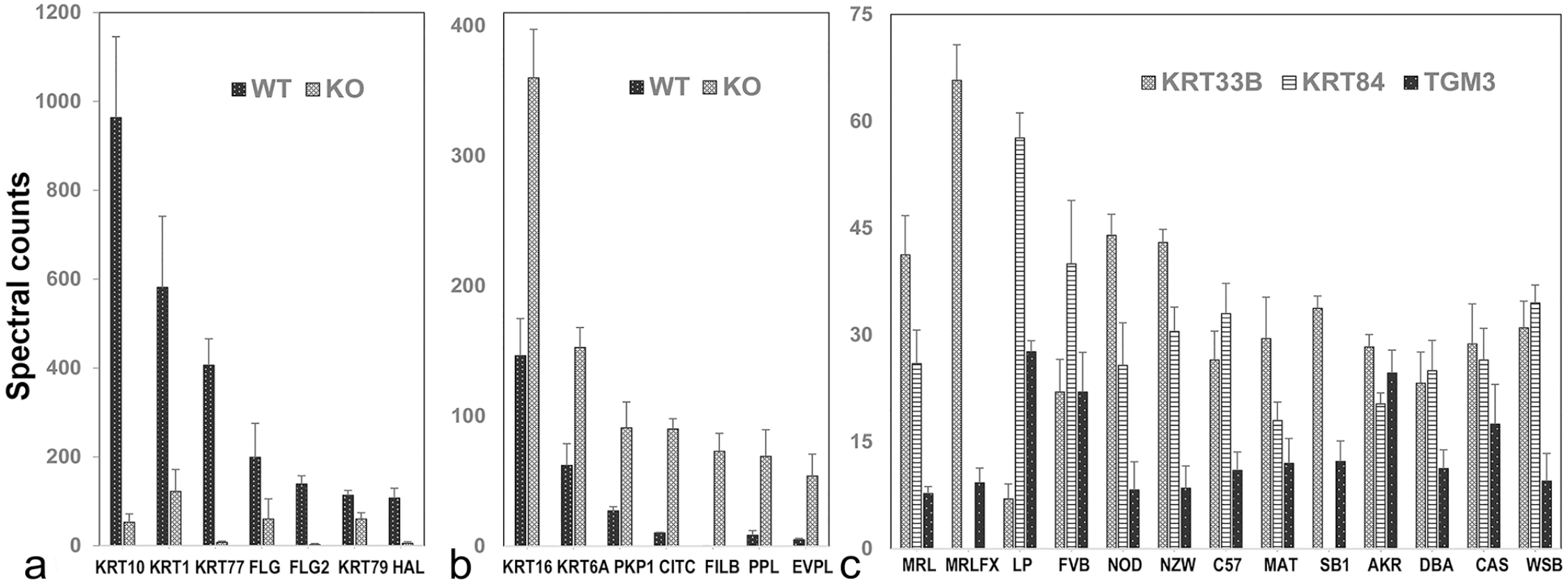
Protein profiling of mouse epidermis and hair shaft. Epidermal samples from mice with defective suprabasal AP1 transcription factor function showed (a) greatly suppressed levels of several major keratins, filaggrin (FLG), filaggrin family member 2 (FLG2), and histidine ammonia lyase (HAL) and (b) considerably higher levels of keratins 6A and 16, the junctional proteins plakophilin 1 (PKP1), periplakin (PPL), envoplakin (EVPL), as well as filamin beta and clathrin heavy-chain linker domain containing 1. (c) A survey of pelage hair from 13 mouse strains revealed highly variable levels of numerous proteins, including keratin 33B (KRT33B), keratin 84 (KRT84), and transglutaminase-3, E polypeptide (TGM3). KO, knockout; WT, wildtype.
Hair Shaft
A study of the hair interior defect mouse, where cells of the medulla have a scattered distribution, showed a deficiency in the trichohyalin level. This finding helps rationalize the lack of microscopic projections from the cortex cells into medulla cells that normally hold the latter in place, thus preventing their stabilization in a regular array. 62 Distinct differences in protein profiles are observed in hair shafts from different mouse strains where the appearance is strain-specific. However, hair shafts from different mouse strains where the hair shafts do not appear abnormal also differ. 57 The protein profile of pelage hair can distinguish mouse strains in pairwise comparisons (Fig. 4c). The conclusion from such proteomic analysis, that such differences in expression level have primarily a genetic basis, has been confirmed in studies of hair shafts from human monozygotic twin pairs. 109
Nail Plate
Ablation of the autophagy-related 7 (Atg7tm1Tchi) in keratin 14 (Tg(KRT14-cre)1Amc)-positive mouse epithelia results in a substantial increase in the number of proteins, mostly enzymes and nonstructural proteins, that survive cornification in the nail plate. 28 A naturally occurring frameshift mutation in a mouse keratin produced degenerative changes in the nail plate, including altered protein profiles, suggesting this mouse mutation could provide a good model for laminitis of the horse hoof. 86 Analysis of hair and nail proteins, other than keratins, has only rarely been amenable to traditional biochemical methods. Thus, the advent of proteomic analysis of nail and hair by mass spectrometry after trypsin digestion has permitted identification of proteins present at substantial levels, which are sparsely studied. For example, the protein V-set and immunoglobulin domain containing 8 (VSIG8) is a membrane-bound protein of unknown function present at moderately high levels in differentiating cells of hair shaft, nail plate, and oral cavity, but not epidermis.61,65 Exploring the functions of such novel proteins in mutant mouse strains may deepen our understanding of normal and disease states in nail and hair.
Skin Surface Lipid Analysis
Skin surface lipids can be systematically collected relatively easily from laboratory mice and other species. In humans, there are large areas of skin without a high density of long, terminal hair shafts covering the skin, such as the forehead, upon which a cotton swab containing acetone can be wiped, whereas mice are covered with a very dense hair coat. Mice should be euthanized or anesthetized (such as with tribromoethanol or isofluorane, for short-term anesthesia), 4 and the hair shaved with electric clippers. Mice with scratches and cuts in the shaved area should be avoided because contamination from the blood and subcutaneous fat could skew the results. The mice are held to stretch the skin in a metal round tart mold (8–8.5 cm in top diameter, 5.5 cm in bottom diameter, 2.5 cm in depth; available at cookware stores online). A hole (2 cm in diameter) is cut in the bottom and sanded to remove rough edges. A mouse is placed in a mold to make a relatively tight seal and expose the dorsal lumbar skin from the bottom hole. The exposed skin is immersed with swirling in 5 ml of acetone in a glass petri dish (10 cm in diameter) for 5 seconds, and then the acetone extract is transferred into a glass tube (13 × 100 mm). The petri dish is rinsed with 5 ml of acetone and combined with the initial acetone extract. The acetone extracts are dried using nitrogen gas. Nitrogen at low flow is sent through the tubes to dry the acetone, leaving only the lipids at the base. Multiple samples can be dried using a manifold such as the Reacti-Vap Evaporator (Thermo Scientific, Waltham, MA) or N-EVAP Nitrogen Evaporator (Organomation Associates, Berlin, MA). The samples can be sealed and stored indefinitely at this point.
New Technology for Molecular Analysis of Mutant Mice
An amazing diversity and range of new technologies, building on existing methods or totally new approaches, are becoming adopted for phenotyping mutant mice. Emerging methods are in transition from organ-based analysis of transcriptomics, metabolomics, lipidomics, and proteomics to single cell–based methods including imaging.11,18,40 It is now possible to do these types of analyses of single cells in serial tissue sections. These developing technologies will move research from analysis of all layers of the skin to regions and now each cell type within a section, thereby defining each cell’s role in the disease state.
Comparative Pathology
Skin and adnexal diseases are commonly clustered based on major types of lesions such as diseases of glands, scaly skin diseases with (psoriasiform) or without (ichthyosiform) inflammation, blistering diseases, nail diseases, and many others.50,88–90 These broad classifications provide a useful starting structure for more detailed evaluations, as defined in both human and veterinary medicine, to obtain a definitive diagnosis. A good place to start is several of the large, multivolume, human dermatopathology textbooks, many of which have an animal models (primarily mouse) section.5,6,10 However, more detailed approaches are now available using dermatology ontologies that can be integrated into diagnostic medical records software programs.20,75,100
Comparing a mutant mouse phenotype at the gross (clinical) and histologic levels to similar diseases in humans and other species can identify the general class of disease if not the exact condition. This can be fortuitous, when the mouse gene affected is known, and a corresponding disease is identified in humans (or other species) where the homologous gene is mutated. When multiple species are similarly affected, this is a phylogenetically conserved disease and likely to be a very good model. An example is what the investigator called the “sore ear mutant mouse” that turned out to be a hypomorphic mutation of laminin gamma 2 (Lamc2jeb), making it a very good model for junctional epidermolysis bullosa. 9 Similarly, the histologic features of alopecia areata in C3H/HeJ mice are quite similar to those of human alopecia areata. 84 However, nothing was known at the time about the complex genetics of alopecia areata until studies in both mice82,98 and later humans 52 began to demonstrate overlap at the molecular level.54,91
Collaborating with expert dermatologists or dermatopathologists, particularly those with expertise on the disease under investigation or similar diseases, can provide great insight into the criteria used to definitively diagnose the human disease by systematic demonstration of features in the mouse model. The catch, however, is that while mice are often maintained as inbred strains in a controlled environment, humans are not. Most humans are very outbred, which creates tremendous heterogeneity among patients, making direct matches to homogeneous mouse models difficult, if not impossible.76,102 The key is to know what one is looking for; hence, the value of collaborating with experts on the human disease and finding the best match. This may require moving the allele onto a different inbred background that takes advantage of strain-specific background modifier genes. Alternatively, a phylogenetically conserved genetic model for a disease that is found in many different species, including humans, and has the same primary gene mutated can validate many of the basic mechanisms of disease pathogenesis.
Summary
The laboratory mouse has proven itself to be an extraordinary model organism for many diseases that are orthologs of human diseases. Careful workup of a mouse model, especially complete necropsy, can define lesions in organs beyond the primary interest that are often overlooked by physicians in their patients. Integrating modern molecular technologies to better understand modifier genes, molecular pathways, and ultimately therapeutic approaches outlined here will carry these models from novelties to heavily used tools to define and ultimately treat diseases in humans and other species.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231170329 – Supplemental material for Phenotyping mice with skin, hair, or nail abnormalities: A systematic approach and methodologies from simple to complex
Supplemental material, sj-pdf-1-vet-10.1177_03009858231170329 for Phenotyping mice with skin, hair, or nail abnormalities: A systematic approach and methodologies from simple to complex by John P. Sundberg and Robert H. Rice in Veterinary Pathology
Footnotes
Acknowledgements
The authors thank Nicholas Gott for his assistance with the histology and immunohistochemistry and Lesley Bechtold for the scanning electron micrographs.
Supplemental Material for this article is available online.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the United States Department of Agriculture (National Institute of Food and Agriculture)/University of California Agricultural Experiment Station project CA-D-ETX-2152-H and MorphoPHEN ERASMUS-EDU-2022-PEX-EMJM-MOB-101082155.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
