Abstract
There is a need to standardize pathologic endpoints in animal models of SARS-CoV-2 infection to help benchmark study quality, improve cross-institutional comparison of data, and assess therapeutic efficacy so that potential drugs and vaccines for SARS-CoV-2 can rapidly advance. The Syrian hamster model is a tractable small animal model for COVID-19 that models clinical disease in humans. Using the hamster model, the authors used traditional pathologic assessment with quantitative image analysis to assess disease outcomes in hamsters administered polyclonal immune sera from previously challenged rhesus macaques. The authors then used quantitative image analysis to assess pathologic endpoints across studies performed at different institutions using different tissue processing protocols. The authors detail pathological features of SARS-CoV-2 infection longitudinally and use immunohistochemistry to quantify myeloid cells and T lymphocyte infiltrates during SARS-CoV-2 infection. High-dose immune sera protected hamsters from weight loss and diminished viral replication in tissues and reduced lung lesions. Cumulative pathology scoring correlated with weight loss and was robust in distinguishing IgG efficacy. In formalin-infused lungs, quantitative measurement of percent area affected also correlated with weight loss but was less robust in non-formalin-infused lungs. Longitudinal immunohistochemical assessment of interstitial macrophage infiltrates showed that peak infiltration corresponded to weight loss, yet quantitative assessment of macrophage, neutrophil, and CD3+ T lymphocyte numbers did not distinguish IgG treatment effects. Here, the authors show that quantitative image analysis was a useful adjunct tool for assessing SARS-CoV-2 treatment outcomes in the hamster model.
Formalin fixation and pathological assessment of tissues remains a cost-effective and gold standard approach for assessing the protective efficacy of vaccines, preventive measures, and therapeutic regimens for infectious diseases including SARS-CoV-2. Yet, these assessments can be limiting due to the time involved and individual variation introduced in situations where studies need to be evaluated and experimental regimens triaged quickly, as became necessary during the height of the SARS-CoV-2 pandemic, and as are currently needed to assess the pathogenicity of novel variants of concern including the highly mutated Omicron variant.6,19 These limitations are confounded by interindividual variation when evaluating for subtle lesions. Historically, this has been addressed by the gold standard of 2 pathologists performing the scoring independently, but this also increases the amount of time required and is often neither feasible nor practical in situations where expedited review is requested. Finally, cross-institutional assessments can be hindered by variation in sample preparation. Although it is well accepted that formalin infusion of the lungs is necessary for accurate evaluation of pulmonary lesions, this is often not possible when large studies are being conducted with many animals, or where tissues are divided for various assays such as reverse transcription polymerase chain reaction (RT-PCR) that are not compatible with formalin fixation. Here, we demonstrate the use of quantitative tools to reproducibly compare histological findings between studies and potentially help expedite a first-pass analysis of tissues in the hamster model of SARS-CoV-2 infection.
The Syrian hamster (Mesocricetus auratus) has become a preferred small animal model for assessing the efficacy of vaccines and other therapeutics for COVID-19.4,16,21,33,34 Numerous studies have demonstrated the efficacy of immunization with SARS-CoV-2 spike protein either via mRNA administration or adenoviral vector delivery.34,35 Studies performed in hamsters assessing the efficacy of the Johnson & Johnson, Ad26-S adenoviral-vectored construct have demonstrated strong correlation of induction of neutralizing anti-spike IgG antibodies to protective efficacy against maximum weight loss and viral load (both genomic and subgenomic RNA) by nasal swab q-PCR.34,35 We aimed to explore whether a standardized, histopathological assessment coupled with quantitative image analysis could provide an early read-out on efficacy in the context of the Syrian hamster SARS-CoV-2 model. The use of digital pathology is expanding in human and veterinary medicine.3,7,15,37 It is becoming increasingly recognized that image analysis and machine learning algorithms can be trained to detect lesions broadly and in some cases triage normal from abnormal to better utilize pathologist’s time.1,2,3,5,15,36,37,40 However, the use of these applications may be limited if collection and processing of tissues varies across institutions. Here, we assess the therapeutic efficacy of purified IgG from SARS-CoV-2-challenged rhesus macaques to prevent SARS-CoV-2 disease in challenged hamsters, using a combination of quantitative image analysis and a semiquantitative scoring rubric. The objective was to highlight how and when these tools may be used to aid or expedite routine pathologic assessments with an emphasis on how tissue processing, sample amount, and time points evaluated may affect study results.
Materials and Methods
Animal Study Design
Convalescent non-human primates
To understand the immunity of the SARS-CoV-2 virus, 9 rhesus macaques were initially infected with SARS-CoV-2 WA1/2020 at 3 different doses—1.1 × 106 PFU, 1.1 × 105 PFU, and 1.1 × 104 PFU (n = 3/dose)—via the intratracheal route (IT) and intranasal route (IN) (1 ml in each route). Following infection, the animals were observed for clinical disease and were assessed by various virologic and immunologic assays. On day 35 following infection, the animals were re-challenged with the same virus at the same infection dose. Two weeks after re-challenge, a detailed necropsy was performed to observe the pathologic characteristics of the disease. Large-volume plasma from each animal was collected to purify polyclonal IgG, to look at the protective efficacy of the IgG in an adoptive transfer model. All animal studies were performed in accordance with all relevant local, state, and federal regulations and were approved by BIOQUAL’s Institutional Animal Care and Use Committee.
Hamsters
Golden Syrian hamsters (Envigo), male and female at a 50:50 sex ratio, 9 weeks of age, were randomly allocated to groups (n = 10 per group for high-, mid-, and low-dose IgG, and n = 20 for sham-treated animals). For IgG transfer, the indicated mass of purified IgG or buffer control was administered via intraperitoneal injection in 2.5 ml of volume. One day after IgG transfer, hamsters were challenged with 2.4 × 104 PFU WA1/2020 SARS-CoV-2 (Bioqual/Lot #061620-1000) administered by the intranasal route in a total volume of 100 µl (50 µl/nostril). Following challenge, body weights were assessed daily and weight loss that exceeded 20% was established as a humane endpoint requiring euthanasia. Sham-treated hamsters were sacrificed at days 2, 4, 7 (±1), 10, 14 (±1), and 16 (n = 3) following challenge; IgG-treated hamsters were sacrificed at day 4 (n = 3 per group) and day 14 (n = 6 per group). At indicated time points, peripheral blood was collected via the retro-orbital route to isolate serum for immunologic assays. Immunological and virological assays were performed blinded. IgG studies were conducted in compliance with all relevant local, state, and federal regulations and were approved by the BIOQUAL Institutional Animal Care and Use Committee. Two pilot studies were performed at the Tufts Regional Biosafety Laboratory to evaluate the effects of tissue processing on quantitative image analysis results. These studies were performed in accordance with approval of the Tufts Institutional Biosafety Committee. Hamsters were similarly 9 weeks of age, 50:50 male and female, and acquired from the same source (Envigo). Hamster infections performed at Tufts used 2019-nCoV/USA/WA1/2020 (BEI Resources, NR522-81) following a parallel protocol of intranasal delivery of 1 × 105 TCID50 in 100 µl (50 µl/nostril). Animals were evaluated for detailed histopathology at days 2 (n = 2), 4 (n = 4), 7 (n = 4), and 14 (n = 2) following challenge.
Necropsy and Histopathology
For studies performed at BIOQUAL, Inc., the left lung lobe was collected and placed free-floating in 10% neutral buffered formalin (NBF, formalin). For studies performed at Tufts University, hamsters were euthanized with an overdose of isoflurane. The thorax was exposed to the level of the trachea, the trachea incised, and an 18G, 50 mm plastic feeding needle for PO gavage with the bulb removed was inserted. The lungs and heart were removed en bloc by dissecting along the dorsal aspect of the thoracic cavity, followed by transection of the thoracic esophagus at the level of the diaphragm and liver. The accessory lobe was identified, clamped at the level of the bronchus with John’s Hopkins bulldog clamps, and removed for weighing, and frozen in RNAlater for tissue viral load. The heart was removed with care to not lacerate airways or pulmonary tissue and was weighed. The remaining lungs were infused slowly (approximately 30 seconds) with 1.5–2 ml of 10% NBF until the surface of the caudal lung lobes showed no folds, followed by clamping of the trachea with a second bull-dog clamp to maintain formalin instillation. The entire pluck and other tissues were transferred to 50 ml conical tubes with at least 30 ml 10% NBF. Lungs were fixed for 24–48 hours in 10% NBF and then transferred to 70% ethanol for processing without lobe disassociation.
Immunohistochemistry
Paraffin blocks were sectioned at 5 µm thickness. Slides were baked for 30–60 minutes at 65°F and then deparaffinized in xylene and rehydrated through a series of graded ethanol to distilled water. Heat-induced epitope retrieval (HIER) was performed using a pressure cooker on steam setting for 25 minutes in citrate buffer (Thermo; AP-9003-500) followed by treatment with 3% hydrogen peroxide. Slides were then rinsed in distilled water and protein blocked (BioCare, BE965H) for 15 minutes followed by rinses in 1× phosphate-buffered saline. The following primary antibodies were used and diluted in DaVinci Green Diluent (BioCare, PD900M): rabbit anti-Iba-1 antibody (Wako; 019-19741) diluted 1:4000; rabbit anti-myeloperoxidase antibody (DAKO, A0398) diluted 1:1500; rabbit anti-SARS-nucleocapsid protein antibody (Sino Biological clone R040, 40143-R040) diluted 1:1500, and rabbit anti-CD3 antibody (DAKO, A0452) diluted 1:300. All antibodies were incubated for 60 minutes at room temperature, followed by rabbit Mach-2 HRP-Polymer (BioCare; RHRP520L) for 30 minutes and then counterstained with hematoxylin followed by bluing using 0.25% ammonia water. Labeling was performed on a Biocare IntelliPATH autostainer. Tissue lesions were independently assessed by 2 board-certified veterinary pathologists (AJM, CP-M).
Digital Image Analysis
Quantitative image analysis was performed using HALO software (v3.0.311.405; Indica Labs) on at least one cross-section of lung lobe from each animal. In cases where more than one cross-section was available, data from all lung lobes were quantified as an individual data point. For SARS-N, Iba-1, MPO, and CD3 positivity by immunohistochemistry (IHC), the Indica Labs—Multiplex IHC algorithm (v3.1.4) was used for quantitation. In all instances, manual inspection of all images was performed on each sample to ensure that the annotations were accurate. Random Forest-based machine learning tissue classifier in HALO was trained by a board-certified pathologist (CP-M) to differentiate areas of parenchymal consolidation in the lung (inflammation), unaffected lung including alveoli, airways (bronchi and bronchioles), airway epithelium, and vasculature, using healthy hamster lung tissue and selected samples of SARS-CoV-2-infected hamster lung tissues. As with quantitative analysis, manual inspection of all images was performed on each sample to ensure that the classifications were accurate.
IgG purification
Polyclonal IgG was purified from the plasma of convalescent rhesus infected with SARS-COV-2 (WA1/2020) using the NAb™ Protein G Spin Column (Thermo Scientific). Plasma was incubated for 1 hour in Protein G columns preconditioned with Protein G IgG Binding Buffer (Thermo Scientific) to capture maximum IgG. Columns were washed 6 times, and IgG was eluted using Protein G IgG Elution Buffer (Thermo Scientific). The eluted IgG was buffer exchanged with 1× DPBS, analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), and quantified using spectrophotometry, and the concentration was normalized to 10 mg/ml.
RT-PCR
SARS-CoV-2 E gene (sgRNA) and N gene (gRNA) levels were quantified by RT-PCR. Standards were made by synthesizing a gene fragment of the subgenomic E or genomic N gene. The gene fragments were then cloned into pcDNA3.1+ expression plasmid using restriction site cloning (Integrated DNA Technologies). The inserts were subsequently transcribed in vitro to RNA using the AmpliCap-Max T7 High Yield Message Maker Kit (CELLSCRIPT). The standards were prepared in log dilutions for RT-PCR assays from 1 × 1010 to 1 × 10−1 copies/ml. Viral loads were determined from lung tissue. The RNA was extracted as follows: Total RNA was extracted on a QIAcube HT instrument using the RNeasy 96 QIAcube HT Kit according to the manufacturer’s specifications (QIAGEN). Extracted total RNA and standard dilutions were reverse-transcribed using SuperScript VILO Master Mix (Invitrogen) according to the manufacturer’s specifications. A TaqMan custom gene expression assay (Thermo Fisher Scientific) was designed using the sequences targeting the E gene sgRNA. The sequences for the custom assay were as follows: forward primer, sgLeadCoV2.Fwd: CGATCTCTTGTAGATCTGTT CTC; E_Sarbeco_R: ATATTGCAGCAGTACGCACACA; and E_Sarbeco_P1 (probe): VIC-ACACTAGCCATCCTTACTG CGCTTCG-MGB. For the genomic N assays, the sequences for the forward (F) and reverse (R) primes and probe (P) were as follows: 2019-nCoV_N1-F: 5′-GACCCCAAAATCAGCG AAAT-3′; 2019-nCoV_N1-R: 5′-TCTGGTTACTGCCAGTTG AATCTG-3′; and 2019-nCoV_N1-P: 5′-FAM-ACCCCGCATT ACGTTTGGTGGACC-BHQ1-3′. Samples and standards were run in duplicate reactions on the QuantStudio 6 and 7 Flex Real-Time PCR Systems (Applied Biosystems). Thermal cycling conditions were as follows: initial denaturation at 95°C for 20 seconds, then 45 cycles of 95°C for 1 second, and 60°C for 20 seconds. Standard curves were used to calculate sgRNA copies, and copy number was normalized to the input weight of lung tissue (copies per gram); the quantitative assay sensitivity was 100 copies/g.
ELISA
RBD-specific binding antibodies were assessed by ELISA. Briefly, plates were coated with 0.5 μg/ml of SARS-CoV-2 RBD protein (Schmidt Lab, Ragon Institute), diluted in 1× PBS, and incubated at 4°C overnight. After incubation, plates were washed once with a wash buffer (0.05% TWEEN-20 in 1× PBS) and blocked with 350 μl of casein block per well. The block solution was discarded after 2–3 hours of incubation at room temperature and plates were blotted dry. Three-fold serial dilutions of hamster serum in casein block were added to wells and plates were incubated for 1 hour at room temperature.
Plates were then washed 3 times and goat anti-hamster IgG HRP (Southern Biotech) diluted to 0.1 µg/ml in casein block, or anti-rhesus IgG HRP (NHP Reagent Resource) diluted to 1 µg/ml in casein block, was added to wells and incubated at room temperature in the dark. After 1 hour, plates were washed 3 times, and 100 μl of SeraCare KPL TMB SureBlue Start solution was added to each well. Development was halted with the addition of 100 μl of SeraCare KPL TMB Stop solution per well. The absorbance at 450 nm was recorded using a VersaMax microplate reader. The raw OD values were transferred into GraphPad Prism for analysis. A standard curve was interpolated using a sigmoidal 4-parameter logistic (4PL) fit. To quantify the endpoint titer, the interpolation function was used to calculate the dilution at which the OD value would be equal to a value of 0.1.
Results
Pretreatment With Purified Rhesus Macaque IgG Resulted in Dose-Dependent Induction of Anti-NHP Antibodies in Syrian Hamsters
Hamsters were administered 25, 2.5, or 0.25 mg of purified IgG derived from plasma of SARS-CoV-2-challenged rhesus macaques, or sham-treated (buffer control), 1 day prior to challenge of the hamsters with SARS-CoV-2 (Fig. 1). Non-human primate (NHP) anti-spike protein receptor-binding-domain (RBD) antibodies were measured by ELISA to quantify transferred antibodies at days 0, 3, 6, 9, and 14 postchallenge (dpc) (Fig. 2a–d). Robust levels were detected in all hamsters receiving IgG in a dose-dependent manner at both days 0 and 3. Animals that received higher doses (25 and 2.5 mg) had detectable IgG titers for an extended period and all animals showed titer reduction over the 14-day period (Fig. 2a, b). Levels of hamster anti-spike protein RBD antibodies were like-wise measured by ELISA at days 0, 3, 6, 9, and 14 postchallenge (dpc) (Fig. 2e–h). No hamster anti-RBD was detectable at either 0 or 3 dpc in all hamsters, including sham-treated animals, consistent with lack of an endogenous response to the virus at these early time-points. Anti-hamster RBD antibody responses were consistently detected in hamsters in a dose-dependent manner beginning at day 6 in hamsters receiving 2.5 and 0.25 mg IgG doses (Fig. 2f, g), while only one animal had detectable anti-hamster RBD in the 25 mg dose group (Fig. 2e). All sham-treated hamsters showed high anti-RBD titers at day 6 following challenge (Fig. 2h). Animals receiving the highest IgG dose (25 mg) had significantly higher binding anti-NHP RBD titers at 12 dpc (Fig. 2i) and significantly lower anti-hamster RBD binding and neutralizing titers at 12 dpc (Fig. 2j, k) as compared to sham-treated hamsters, indicative of decreased viral replication in animals receiving the highest IgG dose.
Use of rhesus macaque immune sera to prevent SARS-CoV-2 infection in hamsters. Study schema; serum isolated from SARS-CoV-2-challenged and recovered rhesus macaques was purified and administered to Syrian hamsters 1 day prior to SARS-CoV-2 intranasal challenge.
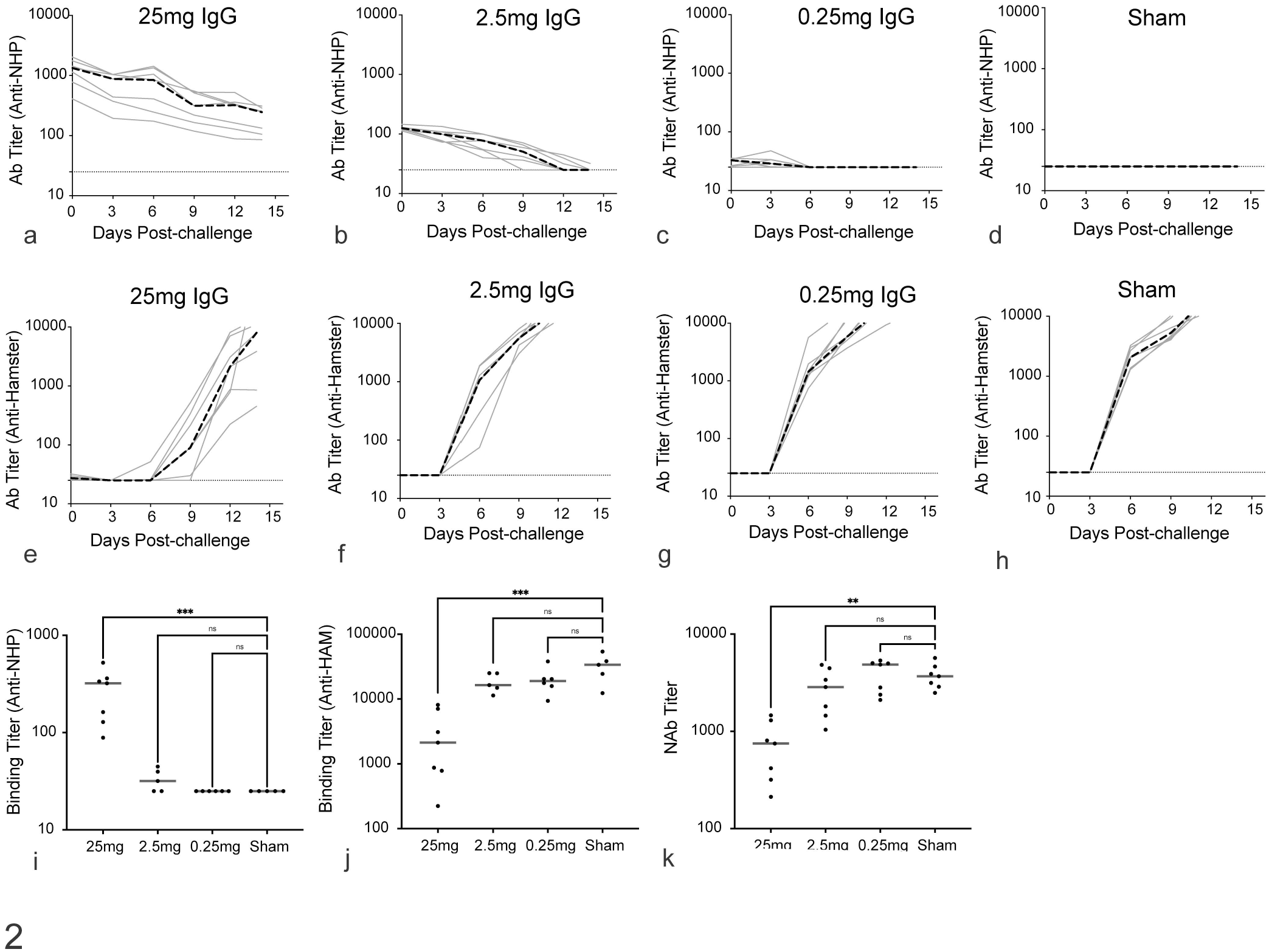
Antibody responses in SARS-CoV-2-challenged Syrian hamsters receiving prophylactic immune serum from rhesus macaques. Longitudinal ELISA binding anti-rhesus macaque spike RBD antibody responses in SARS-CoV-2-challenged hamsters receiving (a) 25 mg, (b) 2.5 mg, (c) 0.25 mg, and (d) sham-treated, and longitudinal ELISA binding endogenous anti-hamster spike RBD antibody responses in challenged hamsters receiving (e) 25 mg, (f) 2.5 mg, (g) 0.25 mg, and (h) sham-treated on days 0, 3, 6, 9, 12, and 14 days following challenge. Hamster anti-NHP (i), anti-hamster (j), and neutralizing anti-spike RBD antibodies (k) in hamsters 12 days following SARS-CoV-2 challenge. Kruskal-Wallis 1-way ANOVA, ***P < .001, **P < .01. Black dashed line = median (a–h); gray line = median (i–k). n = 10 per group for high-, mid-, and low-dose IgG, and n = 20 for sham-treated animals.
High-Dose SARS-CoV-2 Immune IgG Protected Syrian Hamsters From Weight Loss During SARS-CoV-2 Challenge
Hamsters were intranasally challenged with 2.4 × 104 PFU of 2019-nCoV/USA/WA1/2020 (BEI Resources, NR-52281) and monitored daily for weight and viral replication (based on nasal swabs). Maximum weight loss was typically observed between 4 and 6 days postchallenge with complete recovery by day 14 as previously reported (Fig. 3a). 34 Despite significant levels of anti-NHP RBD generated in response to IgG treatment, hamsters receiving the highest dose of immune IgG (25 mg) were completely protected from weight loss following SARS-CoV-2 challenge, and animals receiving a mid-level dose were partially protected. Animals receiving low dose of immune NHP IgG demonstrated weight loss comparable to sham-treated animals. Viral burden across multiple tissue compartments was accessed by RT-PCR for subgenomic E gene (Fig. 3b). Viral RNA was detected in the lungs of all challenged animals. Notably, animals that received a high dose of immune IgG had approximately 1 log lower sgRNA in lung as compared to mid and low IgG-treated and sham-treated animals. High-dose IgG also significantly reduced viral burden in the trachea and heart. High-dose IgG protected against dissemination of virus to the intestine, kidney, spleen, and liver.
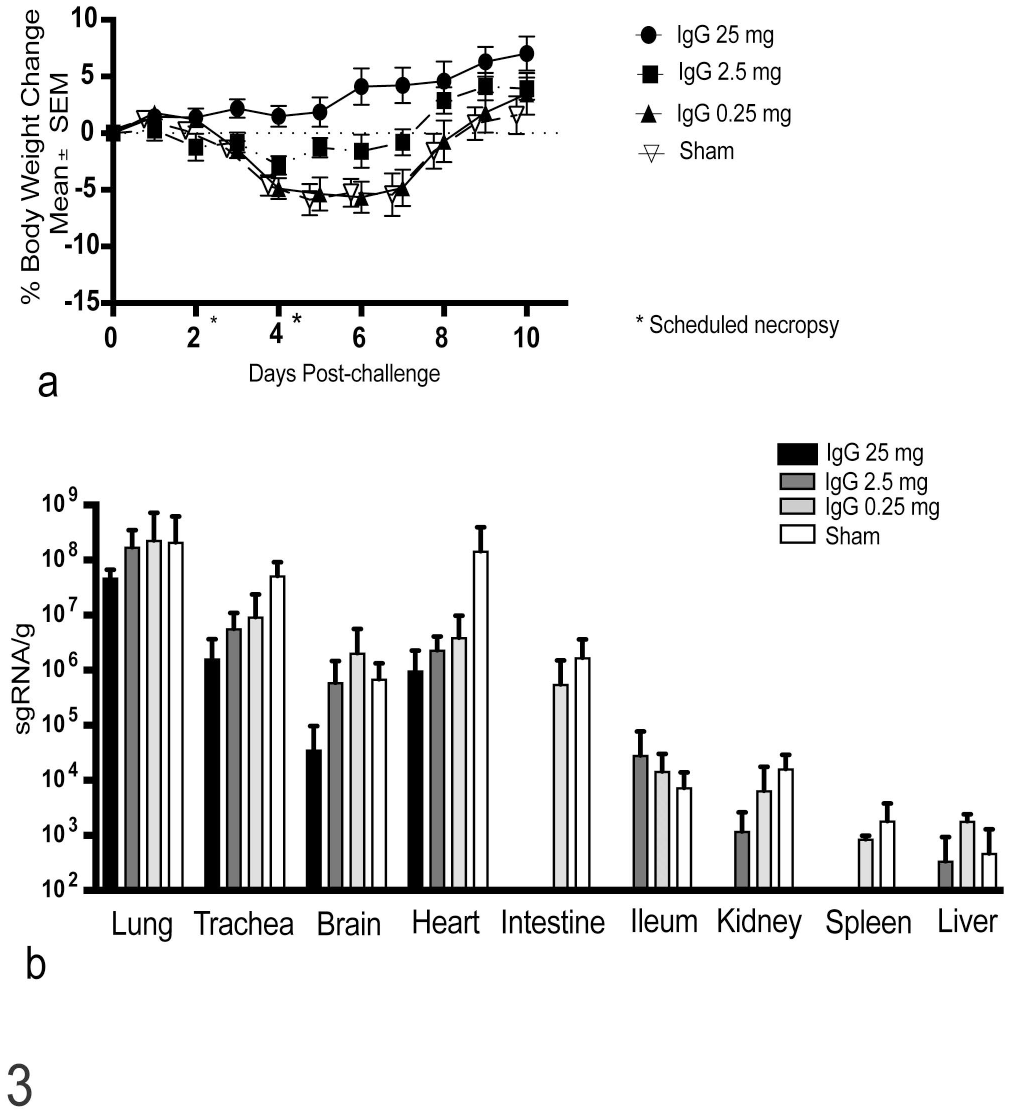
Therapeutic efficacy of macaque immune IgG in SARS-CoV-2-challenged Syrian hamsters. Syrian hamsters were intranasally challenged with 2.4 × 104 PFU SARS-CoV-2 WA1/2020 and weight was monitored daily for 14 days following challenge. (a) Median weight loss over time in animals receiving 25 mg, 2.5 mg, 0.25 mg, or sham IgG from previously SARS-CoV-2-challenged rhesus macaques. (b) RT-PCR for subgenomic SARS-CoV-2 RNA in tissues collected.
A Regional Scoring System Adequately Predicted Disease Severity and Response to Treatment
Semiquantitative scoring systems have been widely used for evaluating SARS-CoV-2 lesions in hamsters. 25 We adopted a 6-feature cumulative scoring system that encompasses the major pathologic features of SARS-CoV-2 disease for assessing vaccine efficacy in hamsters 35 which greatly assisted visualization of pathology data for the broader scientific audience. Standard assessment of (1) airways (bronchiolar epithelium), (2) interstitium, (3) vasculature (vessels), and (4) alveolar spaces was performed with an additional score assigned for (5) edema and (6) alveolar epithelial regeneration (type II pneumocyte hyperplasia) as a marker for previous diffuse alveolar damage (DAD). Each feature was assigned a score of 0 (no lesions), 1 (mild), 2 (moderate), and 3 (severe), the details of which have been previously reported (Figs. 4 and 5). 19 Here, we used this scoring system to evaluate for SARS-CoV-2 lesions in IgG- and sham-treated hamsters. 35 Cumulative pathology scores were highest at 7 days following challenge (Fig. 6a). Bronchiolar epithelial damage occurred early postchallenge and peak scores were assigned at 2 days following challenge (Fig. 6b), which coincided with massive viral replication in this compartment in the hamster model. 34 Vascular changes and associated leakage and edema were observed as early as 4 days following challenge and resolved by 10 days following challenge (Fig. 6c and d). Immune cell infiltrates increased from days 4 to 7 and could be broadly scored by percent lung consolidation (interstitium) and alveolar infiltrates (Fig. 6e, f). Regeneration, observed as type II pneumocyte hyperplasia, peaked at 7 days following challenge (Fig. 6g).
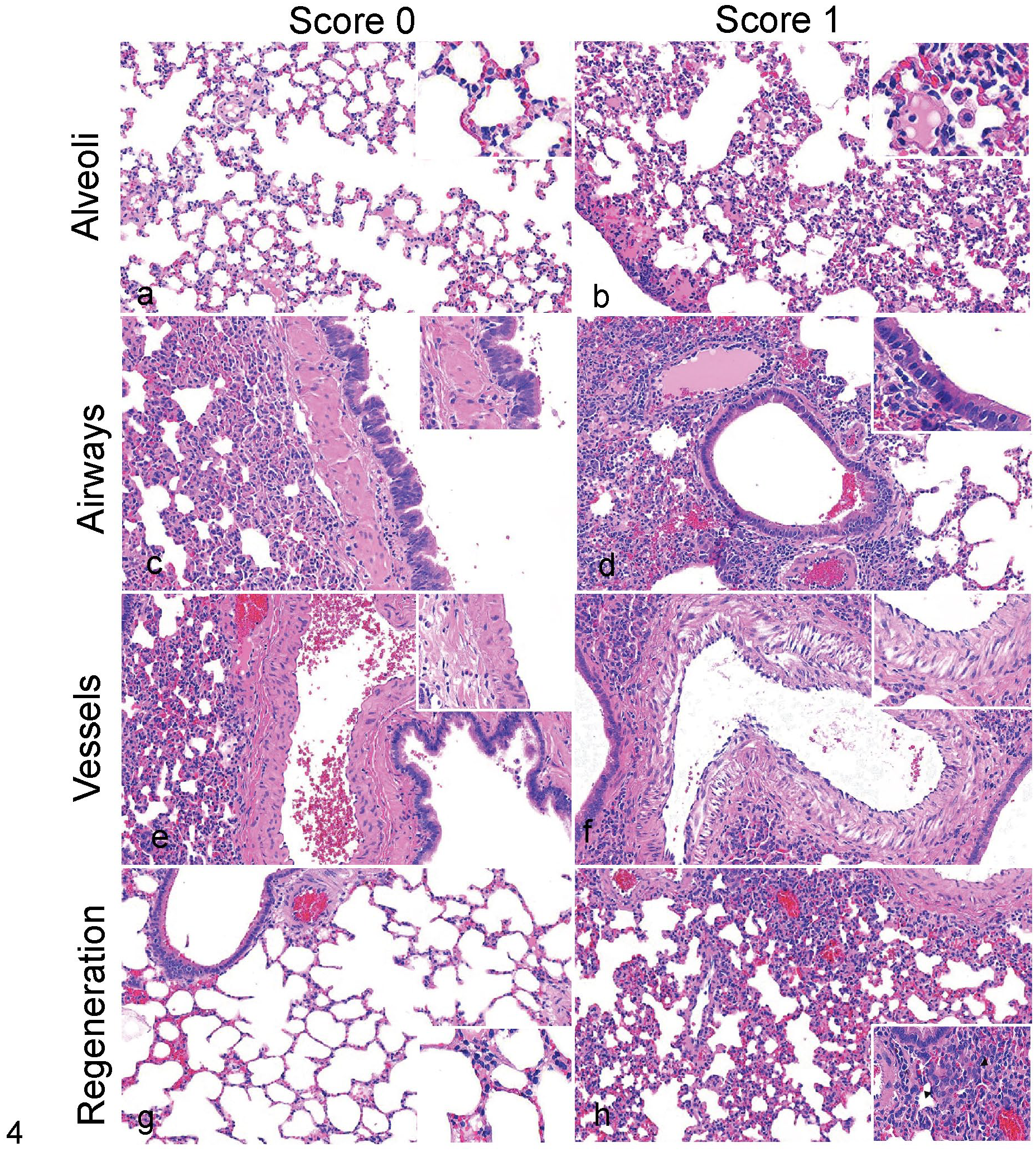
Semiquantitative scoring system of normal to mild histopathological pulmonary features in SARS-CoV-2-infected Syrian hamsters. Representative lesions and severity scores for 0 (normal) and 1 (mild) lesions for: airspaces (alveoli), bronchi and bronchioles (airways), arteries and veins (vessels), and type II pneumocyte hyperplasia (regeneration) in lungs of SARS-CoV-2-challenged Syrian hamsters. Hematoxylin and eosin. (a) Normal alveoli. (b) Mild, multifocal to locally extensive filling of the alveoli with macrophages and neutrophils, with mild to moderate expansion of the interstitium with patchy distribution (affecting <50% of the parenchyma). (c) Normal bronchial epithelium with columnar ciliated epithelium. (d) Mild and multifocal attenuation of the bronchiolar epithelium with multifocal loss of cilia. (e) Normal pulmonary artery with flat and inconspicuous endothelium and no perivascular inflammation. (f) Reactive endothelium (mildly plump cells and prominent nuclei) with rare subendothelial inflammation (endothelialitis) and occasional mild perivascular inflammation. (g) Normal alveoli lined by flat and inconspicuous type I pneumocytes. (h) Rare foci of type II pneumocyte hyperplasia, typically around bronchioles.
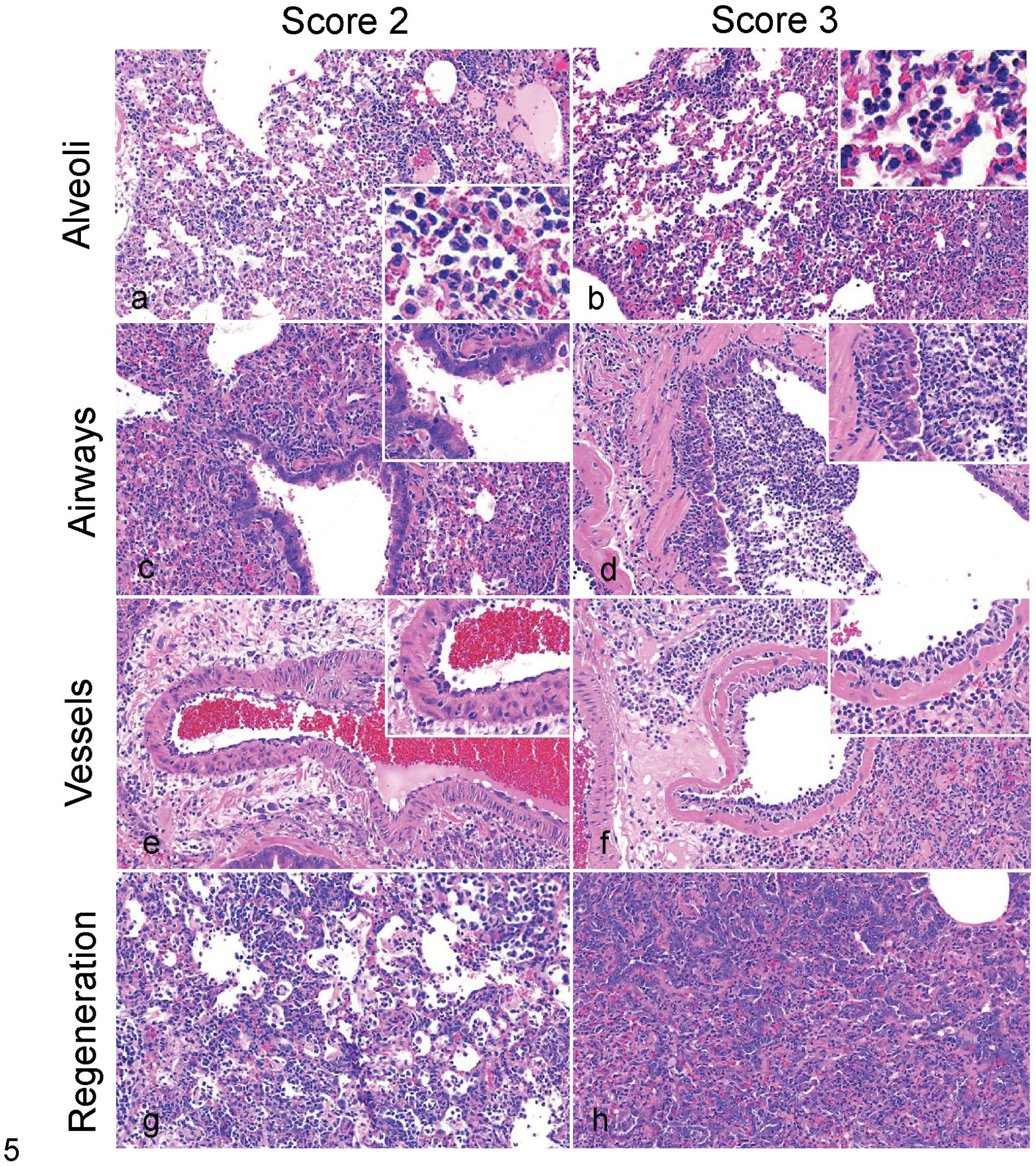
Semiquantitative scoring of moderate to severe histopathological pulmonary features in SARS-CoV-2-infected Syrian hamsters. Representative lesions and severity scores for 2 (moderate) and 3 (severe) lesions for: airspaces (alveoli), bronchi and bronchioles (airways), arteries and veins (vessels), and type II pneumocyte hyperplasia (regeneration) in lungs of SARS-CoV-2-challenged Syrian hamsters. Hematoxylin and eosin. (a) Moderate filling of the alveoli with macrophages and neutrophils and mild to moderate expansion of the interstitium involving >50% of the parenchyma. (b) Marked filling of alveoli with neutrophils, macrophages, and cellular debris and moderate to marked expansion of the interstitium of >50% of the parenchyma. (c) Moderate to marked attenuation of the bronchiolar epithelium with multifocal loss of cilia, frequent sloughing of epithelial cells, occasional syncytia formation, and mild to moderate inflammation (neutrophils, macrophages, and lymphocytes). (d) Marked and diffuse attenuation of the epithelium, loss of cilia, degeneration and necrosis of epithelial cells with frequent sloughing, and marked predominantly neutrophilic inflammation. (e) Reactive endothelium with mild to moderate endothelialitis, mild to moderate perivascular inflammation and edema, and infrequent intramural inflammation. (f) Marked and diffuse endothelialitis with markedly reactive endothelium, prominent perivascular inflammation and edema, and frequent intramural inflammation. (g) Multifocal areas of alveoli lined by hyperplastic type II pneumocytes with some nuclear pleomorphism. (h) Locally extensive areas of type II pneumocyte hyperplasia that pack alveolar spaces and frequently show karyomegaly and nuclear pleomorphism.
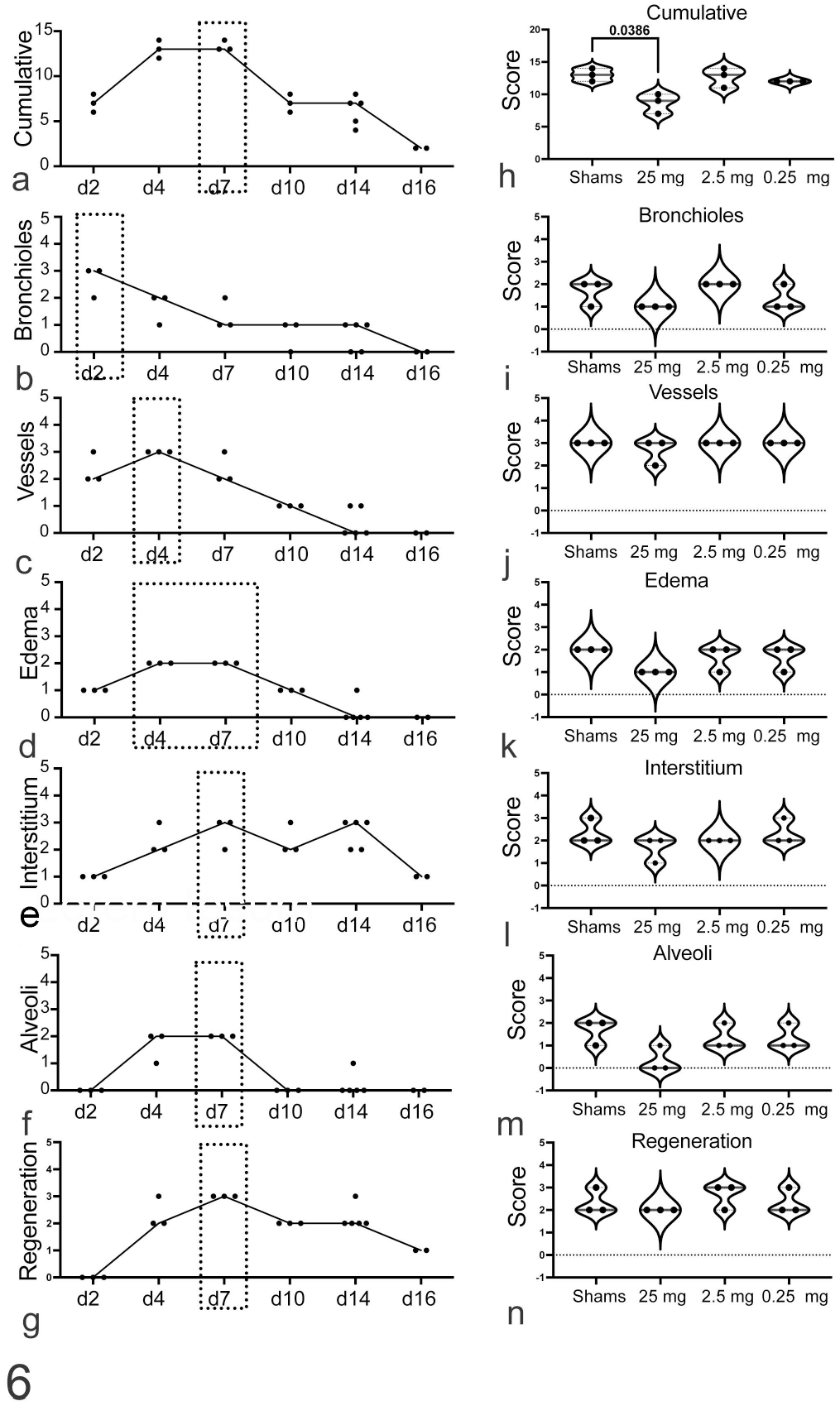
Effects of macaque immune sera on SARS-CoV-2 lesions in challenged hamsters. Semiquantitative (a) cumulative pathology scores, and individual scores for various anatomic compartments and pathologic features: (b) bronchioles, (c) vessels, (d) edema, (e) interstitium, (f) alveoli, and (g) regeneration in sham-treated animals at different times. Black dashed boxes indicate peak scores. Cumulative (h) and individual scores (i–n) for pathologic features in sham-treated compared to IgG-treated (25 mg, 2.5 mg, 0.25 mg) hamsters at 4 days post challenge (dpc); Black line = median. One-way ANOVA, Kruskal-Wallis test post hoc analysis. Day 2 (n = 3), Day 4 (n = 3), Day 7 (n = 3), Day 10 (n = 3), Day 14 (n = 5), Day 16 (n = 2).
Cumulative pathology scores at day 4 reflected trends in weight loss and animals that received the highest dose IgG (25 mg) had significantly lower cumulative scores than sham-treated animals (Fig. 6h). Assessment of bronchiolar epithelial lesions (airways) consistently mirrored the cumulative pathology score at day 4 (Fig. 6h, i) even in non-formalin-infused lungs. Few differences were detected in the vascular, interstitium, and regeneration scores between treated and sham animals, while edema and alveolar scores trended lower in IgG-treated animals (Fig. 6j–n).
Image Analysis Algorithms for Inflammation and Consolidation Correlated With Cumulative Pathology Scores and Maximum Weight Loss
There is significant interest in incorporating machine learning, artificial intelligence, and quantitative image analysis into pathologic assessments to increase pathologist efficiency and to minimize the number of slides requiring detailed manual assessment and semiquantitative scoring in both veterinary and human pathology.1,3,5,7,15,36,37,40 We asked whether quantitative image analysis for lung inflammation and consolidation could be similarly used for SARS-CoV-2 research in a time-limited setting. Furthermore, we wanted to understand whether image analysis could be more sensitive than clinical measures (maximum weight loss) or semiquantitative lesion scoring to detect subtle differences in treatment groups. We also wanted to explore the robustness of these methodologies in assessing tissues across institutions. We evaluated lungs from IgG-treated and sham-treated and challenged hamsters that were formalin-instilled (Fig. 7a) and compared these results to non-formalin-instilled lungs (Fig. 7b). Using the tissue classifier module in HALO, 15 regions of consolidation/inflammation (yellow), vasculature and smooth muscle (cyan), and uninvolved lung (purple) were classified by a board-certified veterinary pathologist (CP-M). The module was trained on instilled lung specimens and then applied to 2 independent studies performed at 2 different institutions to quantitate the percentage of affected lung (Fig. 7c). By day 7 following challenge, approximately 20% of the lung was affected when assessed in formalin-instilled lungs representing the peak of lung inflammation. Interstitial inflammation and consolidation had completely resolved by 14 days postchallenge and no significant difference between infected and uninfected animals was detected. Consolidation data generated from machine-learning modules corresponded to maximum weight loss (Fig. 3) and cumulative pathology scores (Fig. 6a) but was less sensitive when used in non-instilled lungs from IgG-treated and sham-treated animals (Fig. 7d). Visual inspection of algorithm outputs revealed that regions of lung consolidation were misinterpreted as vasculature in non-instilled regions of lung. Therefore, machine-learning methods to measure lung inflammation will likely be most useful in expanded lung specimens.
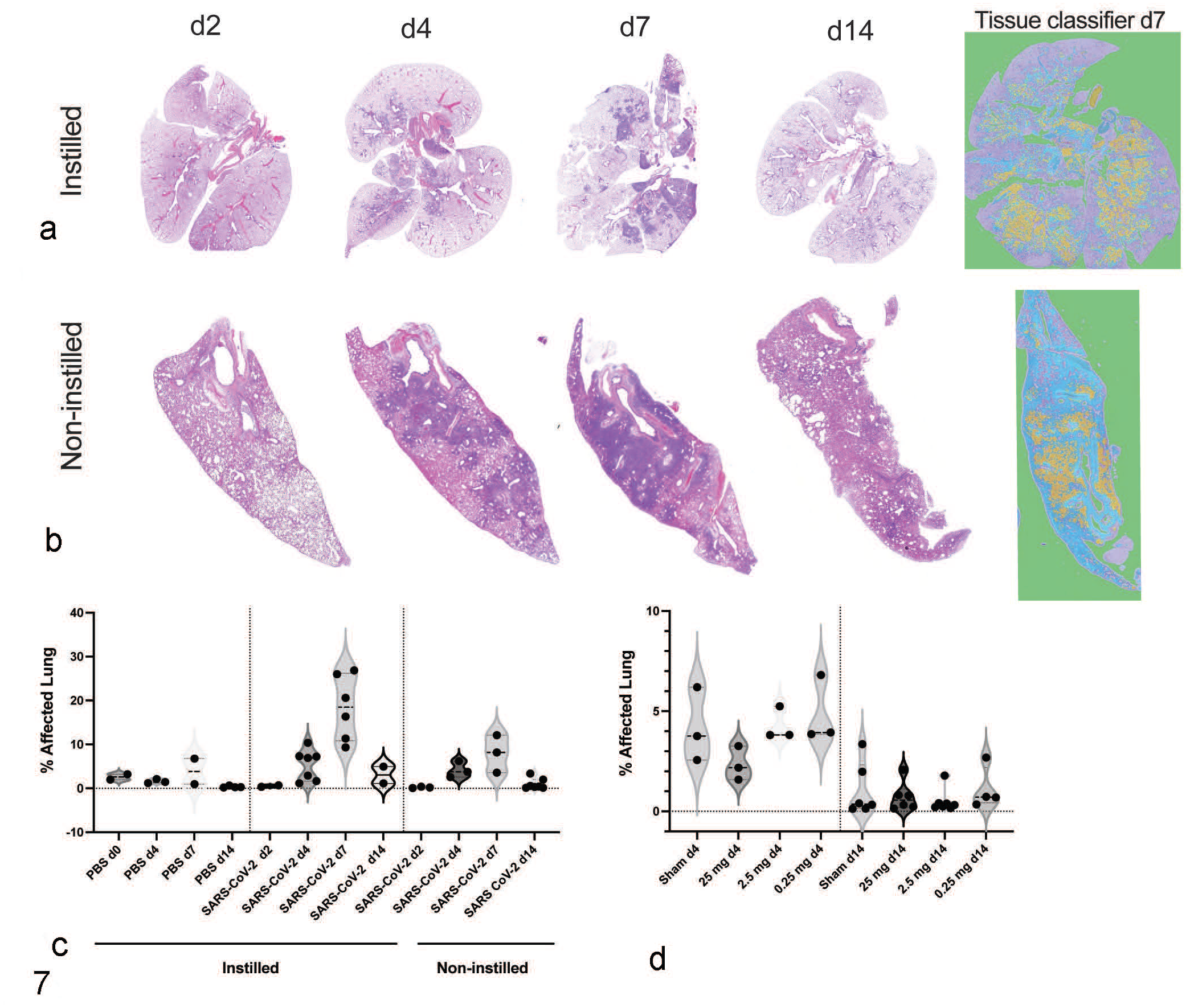
Quantitative image analysis using machine learning, of lung inflammation induced by SARS-CoV-2 infection, in formalin-infused and non-infused lungs from different institutions. Whole-slide digital scans of lung from SARS-CoV-2-infected hamsters at days 2, 4, 7, and 14 following challenge in (a) formalin-instilled lungs at institution 1 and (b) non-instilled lungs at institution 2. HALO Indica Labs tissue classifier artificial intelligence module was trained to recognize lung consolidation (yellow), connective tissue (cyan), and normal/unaffected parenchyma (purple). (c) Longitudinal quantitation of consolidation was reported as percent affected lung in SARS-CoV-2-challenged hamsters, showing peak inflammation at day 7 following challenge. (d) Percent lung affected in IgG-treated and sham-treated SARS-CoV-2-infected hamsters at days 4 and 14 following challenge at institution 2 where tissues were not instilled with formalin. n = 2–7 animals per time point.
Both Macrophage and T-cell Infiltration Peaked by 7 Days Post-infection as Measured by Quantitative Assessment of Immunohistochemistry
During the race to develop vaccines and therapeutics for COVID-19, many institutions and investigators were conducting large hamster studies where tissue lesions were important, but not necessarily the most critical endpoint for product advancement. We asked whether one or more specific measures of pulmonary inflammation in hamster lungs post-challenge could serve as an orthogonal endpoint for overall treatment response using quantitative image analysis. We were specifically interested in understanding which inflammatory infiltrates correlated best with clinical parameters and histopathologic scoring. Using previously validated antibodies for immunohistochemistry in hamsters 34 with minor alterations in antibody concentration to optimize signal-to-noise ratio with quantitative image analysis, we assessed degree of SARS-CoV-2 nucleocapsid protein (Fig. 8a), macrophage (ionized calcium binding protein, Iba-1) (Fig. 8b), neutrophil (myeloperoxidase, MPO) (Fig. 8c), and T-cell (CD3) infiltration (Fig. 8d) in hamster lungs post-SARS-CoV-2 challenge with quantitative image analysis (Fig. 8e–k). SARS-CoV-2 positivity was highest at day 2 following challenge (Fig. 8e). Macrophage, neutrophil, and T cell inflammation peaked by 7 days following challenge as assessed by number of chromogen-positive cells/tissue area (Fig. 8e–h) consistent with previously published data9,34 and correlated with peak scores for interstitium, alveolar spaces, regeneration, and cumulative pathology scores (Fig. 6a and e–h). However, no significant differences were detected in IgG-treated hamsters with assessment of these markers at day 4 following challenge in non-formalin-instilled lungs (Fig. 8i–k).
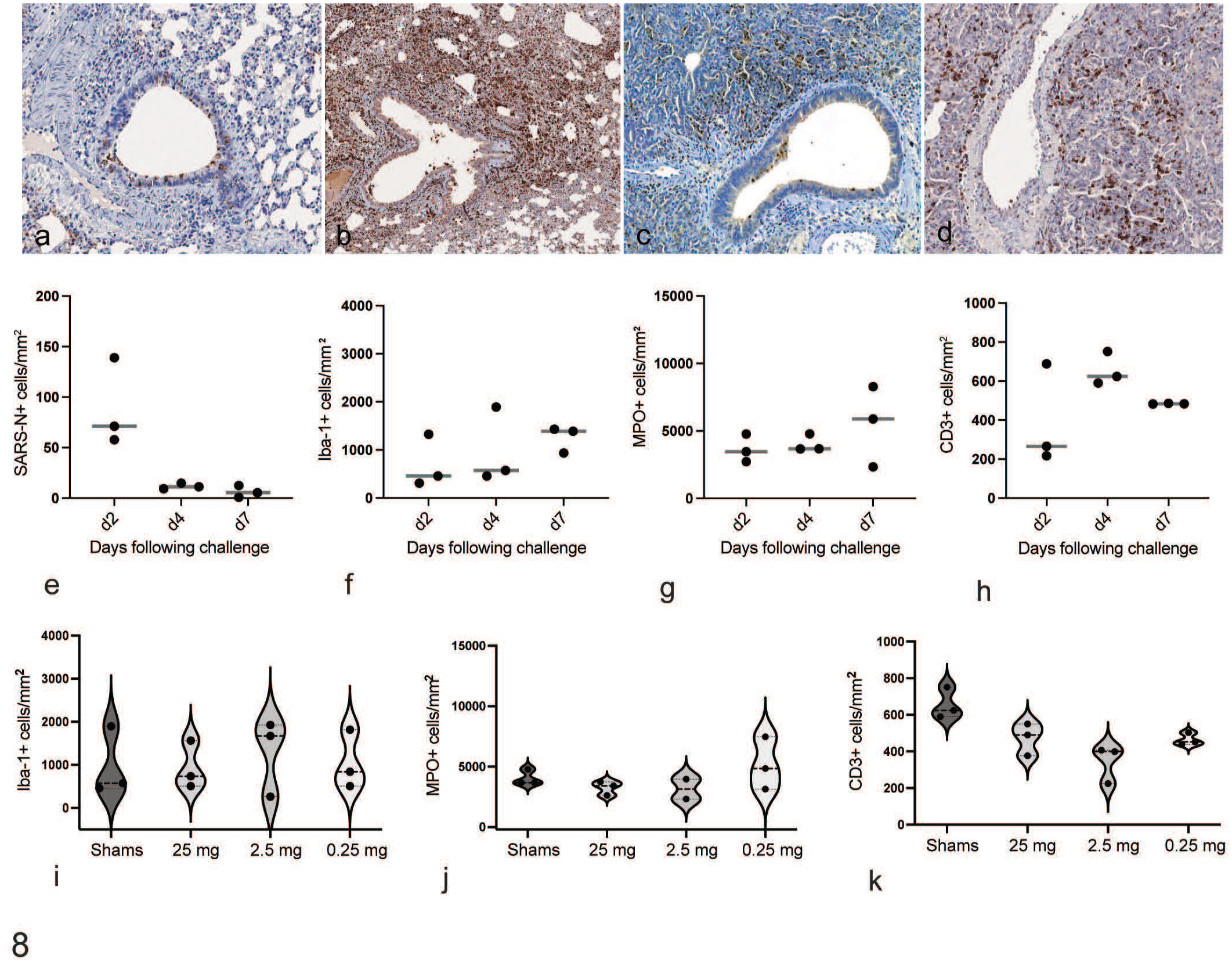
Quantitative image analysis of immunohistochemical markers of lung inflammation in IgG-treated and SARS-CoV-2-challenged hamsters. Lungs were immunolabeled for (a) SARS-N, (b) Iba-1, (c) myeloperoxidase (MPO), and (d) CD3, and HALO Multiplex IHC module (Indicalabs) was used to quantify the number of (e) SARS-N+, (f) Iba-1+, (g) MPO+, and (h) CD3+ cells per tissue area at days 2, 4, and 7 following SARS-CoV-2 challenge. (i–k) Quantitative image analysis of (i) macrophages (Iba-1+), (j) neutrophils (MPO+) cells, and (k) T lymphocytes (CD3+) per unit area in sham and IgG-treated hamsters at 4 days following SARS-CoV-2 challenge. Horizontal line = median.
Discussion
Routine pathologic assessment of formalin-instilled lungs with hematoxylin and eosin staining and interpretation by a veterinary pathologist is important in triaging novel vaccines and treatment modalities, but there are some limitations to standard pathologic evaluation including pathologist-to-pathologist variation and time needed for analysis. Quantitative image analysis for assessing SARS-CoV-2 lesions in hamsters has been successfully used to evaluate lesions in studies of therapeutic efficacy, vaccine protection, and pathogenesis.4,8,9,34 Here, we highlight how incorporation of select image analysis tools may aid histologic assessment of SARS-CoV-2 lesions in large in vivo studies to assist study triage prior to complete pathologic assessment and scoring, in time- or resource-limited settings. However, we emphasize that some analysis tools may be less useful if tissue collection and processing is not standardized.
We evaluated the therapeutic efficacy of rhesus macaque immune sera on preventing SARS-CoV-2 viral replication, clinical disease, and lesions in hamsters. This study had 3 treatment groups and 1 sham group. Hamsters receiving the highest IgG dose of 25 mg were protected from weight loss. Notably, sham-treated hamsters in this study had less weight loss than anticipated. Using both semiquantitative pathologic scoring and quantitative digital image analysis, we evaluated lesions across IgG treatment groups. Use of a simple cumulative scoring system allowed detection of the therapeutic effect in the highest IgG dose group that mirrored virologic assessment and clinical parameters. Numerous studies have highlighted the effects of SARS-CoV-2 replication on the bronchiolar epithelium and disease progression in hamsters.13,14 We highlight that evaluation of key tissue compartments such as the bronchioles and vessels, and assessment of individual features such as regeneration, are good measures of therapeutic efficacy when time permits a detailed histopathologic evaluation.
It is widely accepted that lung inflation is critical for detailed pathologic evaluation in rodent models. This has been well recognized in lung physiology studies and in rodent models of acute lung injury and emphysema where lung morphometry is regularly used. It is also well known that fixation, inflation/instillation, and airway pressure must be standardized for proper assessment of treatment outcomes in these models. 29 In this study, the lack of lung expansion via air or fixative obscured semiquantitative measures of interstitial disease across IgG treatment groups.
We compared samples from this study to samples acquired with formalin instillation of the lung at our institution to test the robustness of digital pathology algorithms and machine learning/artificial intelligence software modules to show that quantitative image analysis platforms perform better and more consistently on formalin instilled compared to non-instilled lungs. Quantitative image analysis using machine-learning tools, such as the HALO Random Forest tissue classifier module (IndicaLabs), require up-front training by a pathologist or someone with histologic expertise. Once algorithms were adequately trained, we showed that a machine-learning platform could generate meaningful pathology data that correlates tightly with maximum weight loss in our hamster pilot studies. However, the HALO tissue classifier algorithm, trained to detect consolidation in SARS-CoV-2 challenged hamster lung, underperformed on samples from the immune sera treatment study, due to insufficient expansion of alveolar spaces. We also showed via whole-slide scanning of multiple lung lobes how variable the distribution of interstitial disease may be in hamsters infected with SARS-CoV-2. Standardized fixation and formalin instillation of lung, and attention to proper and consistent trimming and orientation of lung lobes during embedding, will further improve performance and consistency from image analysis algorithms.
We also evaluated immune cell infiltrates using an IHC image analysis tool (Multiplex IHC Module) to compare the time course of different immune cell infiltration with markers for macrophages, T lymphocytes, and neutrophils and assessed how well these markers correlated with clinical parameters and scoring for interstitial disease. Immunopositivity for Iba-1 (macrophages) by IHC provides a useful orthogonal measure of tissue lesions that correlates with overall clinical disease irrespective of lung expansion in hamsters.9,24,27,38 In this study, we did not detect robust differences in IgG treatment groups, likely related to samples being collected prior to the peak of macrophage inflammation. We did notice a trend toward decreased numbers of CD3+ T lymphocytes in the lungs of IgG-treated hamsters, but this did not correspond to maximum weight loss. Neutrophilic inflammation is an important component of SARS-CoV-2 lesions, but neutrophils are notoriously difficult to study ex vivo due to fragility and short-life span.11,23,26,39 Using quantitative assessment of IHC positivity and size exclusion, we evaluated numbers of MPO+ neutrophils in IgG-treated and untreated hamsters and did not detect any differences between groups.
There are advantages and disadvantages to the use of quantitative image analysis for SARS-CoV-2 pathology research. A major advantage is that assessment using image analysis can be performed on formalin-fixed paraffin-embedded tissue specimens. However, necropsies performed in biosafety level 3 containment introduce technical limitations that make pressure-controlled instillation of the lung difficult to execute. Likewise, instillation of formalin into the lungs may alter immune cell number and distribution in air spaces. Alternatively, suture ligation of the trachea prior to opening the thoracic cavity can maintain sufficient lung inflation for quality pathologic assessment. 30 Here, we show that regional atelectasis can make both scoring and quantitative image analysis less consistent, but the effects of over-distension on quantitative image analysis have not been explored.10,17,20,29
The development of standardized tissue collection, processing, and analysis techniques for SARS-CoV-2 studies using the hamster and mouse models will greatly improve comparison and interpretation of cross-institutional vaccine and therapeutic studies. With the rapid spread and reports of variable pathogenicity of the B.1.1.529 (Omicron) variants in the hamster model,6,22,28 we propose a method for combined virology assessment and formalin instillation of lung for improved detection of lesions and virus for SARS-CoV-2 research. For cross-institutional comparison of data and broad adoption of pathologic endpoints, scoring metrics and image analysis data alike will need further validation. Another important consideration is time of sampling. We show here that inflammation as measured by cumulative scoring, quantitation of percent lung affected, and immune cell infiltrates peaked between days 4 and 7. Our inability to detect differences in immune cell infiltrates in hamsters receiving high-dose IgG may have been the result of sampling at day 4 rather than day 7, or due to insufficient lung expansion for detection by image analysis algorithms. Correlation of pathologic endpoints to clinical parameters is one accepted means to validate image analysis endpoints.12,18,31,32
Immunohistochemistry and quantitative image analysis may be useful as adjunct measures of disease and inflammation that can be assessed initially to prioritize detailed pathological assessments and pathologist time. In settings where adequate formalin instillation of the lung can be achieved consistently, artificial intelligence modalities that can assess the percentage of affected lung provide an alternative measure of overall disease burden that may guide future study design and implementation while more detailed histopathological assessments are being completed. Combined use of traditional semiquantitative and quantitative image analysis for pathologic evaluation, as shown here for SARS-CoV-2 in the Syrian hamster model, does require an investment in instrumentation, namely a high-quality and fast digital slide scanner and image analysis software, a wide range of which are available in the market. However, given the increased accessibility of these tools across institutions, adoption of some of these tools not only provides an opportunity for pathologists to generate quantitative data, but also greatly improves accessibility and reproducibility of pathological assessment for animal models of human disease.
Footnotes
Acknowledgements
We thank Shannon O’Connor, Sarah Gardner, Julia Barrett, and Owen Sanborn for their technical assistance. The following reagent was deposited by Centers for Disease Control and Prevention and obtained through BEI Resources, NIAID, NIH: SARS-Related Coronavirus 2, Isolate USA-WA1/2020.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a COVID-19 Fast Grant, Mercatus Center, George Mason University to AJM and DHB.
