Abstract
The emergence of the coronavirus disease 2019 (COVID-19) caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) inspired rapid research efforts targeting the host range, pathogenesis and transmission mechanisms, and the development of antiviral strategies. Genetically modified mice, rhesus macaques, ferrets, and Syrian golden hamsters have been frequently used in studies of pathogenesis and efficacy of antiviral compounds and vaccines. However, alternatives to in vivo experiments, such as immortalized cell lines, primary respiratory epithelial cells cultured at an air–liquid interface, stem/progenitor cell-derived organoids, or tissue explants, have also been used for isolation of SARS-CoV-2, investigation of cytopathic effects, and pathogen–host interactions. Moreover, initial proof-of-concept studies for testing therapeutic agents can be performed with these tools, showing that animal-sparing cell culture methods could significantly reduce the need for animal models in the future, following the 3R principles of replace, reduce, and refine. So far, only few studies using animal-derived primary cells or tissues have been conducted in SARS-CoV-2 research, although natural infection has been shown to occur in several animal species. Therefore, the need for in-depth investigations on possible interspecies transmission routes and differences in susceptibility to SARS-CoV-2 is urgent. This review gives an overview of studies employing alternative culture systems like primary cell cultures, tissue explants, or organoids for investigations of the pathophysiology and reverse zoonotic potential of SARS-CoV-2 in animals. In addition, future possibilities of SARS-CoV-2 research in animals, including previously neglected methods like the use of precision-cut lung slices, will be outlined.
Keywords
At the end of December 2019, an unusual human pneumonia occurred in patients in Wuhan, China, which quickly developed into a pandemic causing overwhelming problems in the public health sector and had a significant impact on social and economic life. 170 The causative pathogen was identified by viral genome sequencing and is known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).85,150,152,168 Severe acute respiratory syndrome coronavirus 2 infection in humans can be asymptomatic or may cause mild to severe and fatal Coronavirus Disease 2019 (COVID-19), which can include respiratory symptoms as well as cardiovascular, gastrointestinal, hepatic, renal, neurologic, ocular, and dermatologic manifestations.63,75,147 Many questions regarding reservoir and intermediate hosts, epidemiology, cell tropism, pathogenesis, and potential application of antiviral strategies remain to be answered. The rapid spread of the current pandemic and the emergence of novel variants of concern 1 call for the development of efficient pharmaceutical and nonpharmaceutical intervention strategies, which could also represent a blueprint for other emerging viral infections in the future.
Severe acute respiratory syndrome coronavirus 2 is an enveloped, positive-sense single-stranded RNA betacoronavirus (β-CoV) belonging to the subfamily Orthocoronavirinae, the family Coronaviridae, and the order Nidovirales. 151 It is assumed but still discussed controversially that the virus might have evolved from bat-derived SARS-like coronaviruses because it has greater sequence identity with RaTG13, a betacoronavirus isolated from horseshoe bats (96,2% identity), than with the human SARS-CoV-1 (79% identity) or Middle East respiratory syndrome coronavirus (MERS-CoV) (50% identity).85,168 Although bats are most likely the original source of SARS-CoV-2, several observations indicate that another animal species might have acted as an intermediate host.82,164 Severe acute respiratory syndrome-related betacoronaviruses can infect a broad range of different hosts belonging to 4 mammalian orders, including bats, primates, carnivores, and pangolins.76,161,162,68 The second-closest relatives of SARS-CoV-2 determined so far are 2 sublineages of SARS-related CoVs isolated from Malayan pangolins,76,154 making them a debated potential candidate as an intermediate host.81,82,154 Moreover, natural infection with SARS-CoV-2 has been observed in domestic cats107,160, captive large felids,91,138,139 as well as dogs, 136 ferrets, 51 mink, 111 otters, 141 and gorillas, 140 all of which were most likely infected by close contact with humans. 99 Importantly, the transmission of the virus from mink back to humans highlighted the potential risk of bidirectional interspecies transmission, which could contribute to ongoing virus circulation even after vaccination becomes widely available for the human population, increasing the frequency of virus mutations and the risk of development of virus variants.39,111 The evidence of natural infection in domestic and wildlife animals, as well as the search for suitable animal models for COVID-19, has prompted investigations on the similarity of the receptor for SARS-CoV-2, angiotensin-converting enzyme-2 (ACE-2), between humans and other animal species.24,99,130 Among mammals with a high sequence identity were, for example, primates, baleen and toothed whales, rodents, and deer. Compared to these species, felids and several ruminant species had a lower sequence identity and, notably, all bat as well as pangolin species analyzed showed a low ACE-2 amino acid sequency homology. 24 To further confirm these sequence comparisons and other studies investigating ACE-2 characteristics in several species,84,88,122 numerous in vivo experiments related to SARS-CoV-2 have been conducted in various mammalian and 2 avian species. 68 As a consequence, a high susceptibility has been demonstrated in vivo in rhesus macaques,100,127 Syrian golden hamsters,3,9 domestic cats,43,55 ferrets,129,135 white-tailed deer, 112 American deer mice, 53 and Egyptian fruit bats. 129 The degree of susceptibility varied from moderate to low in raccoon dogs, 40 New Zealand white rabbits, 101 tree shrews, 166 dogs, 135 and cattle 137 while no virus replication was found following inoculation of pigs, chickens, turkeys, and ducks.129,135 Some of the data obtained by these in vivo studies could also have been gathered by in vitro and ex vivo investigations, which are already being used to study host and cell tropism, infection kinetics, and local cellular pathogenesis in humans as well as animals.2,12,15,33,66,93 In vitro and ex vivo culture methods represent highly predictive and versatile approaches in the discovery of newly emerging viral diseases and can be used as the first step in the research of antiviral approaches. Air–liquid interface (ALI) cultures, lung organoids, respiratory tract tissue explants, and precision-cut lung slices (PCLS) are well-established and characterized tools to investigate respiratory tract disease, especially of infectious pathogens. Together, they cover the whole range of upper (nasal cavity) and lower (trachea, bronchi, and alveoli) respiratory tract tissue elements (Fig. 1) and can therefore be applied to identify species and cell types susceptible to emerging respiratory viruses and aid the assessment of the epidemiologic risk potential of animals living in close contact with humans. In addition, they are suitable to characterize certain aspects of the host response in detail (eg, by transcriptomic analysis) and to profile new treatment options, both antiviral and, to a certain extent, anti-inflammatory drugs.
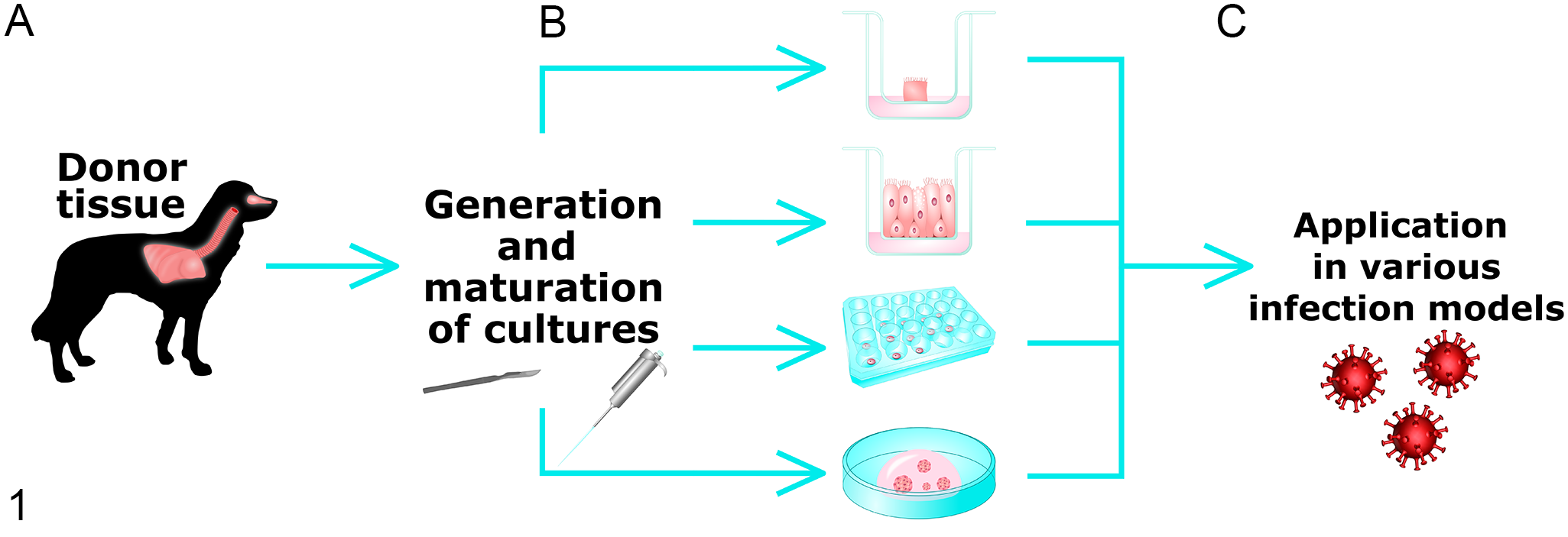
Schematic illustration of process workflow. (A) Native tissue of the respiratory tract such as lung, trachea, and nasal mucosa, originating from various animal species, are processed for different culture models, (B) From top to bottom, nasal mucosa explants (NME), air–liquid interface (ALI) cultures, precision-cut lung slices (PCLS), and organoids generated from donor tissue are maintained under culture-specific conditions, and (C) Fully differentiated cultures can be used in numerous studies concerning respiratory infectious pathogens.
In this review, we first describe selected cell culture models and their application for research of viral and bacterial infectious diseases using companion- and farm animal-derived tissues. Secondly, we outline the role of in vitro and ex vivo models in SARS-CoV-2 research in various species including humans.
Generation and Use of Selected In Vitro Models in Non-SARS-CoV-2 Investigations
ALI Cultures of Tracheal and Bronchial Epithelial Cells
Primary and immortalized bronchial or tracheal cells are usually cultured in submerged monolayers on conventional flasks or Petri dishes. However, these submerged cell culture systems do not entirely reflect the 3-dimensional architecture and multicellular complexity of their in vivo counterparts, as they remain undifferentiated and lack cilia or mucus secretion. Interestingly, there are some known exceptions in certain animal species: in hamsters, epithelial cells can differentiate into ciliated and goblet cells in submerged culture.71,77 In contrast, respiratory tract epithelial cells grown at an ALI efficiently develop into a columnar, pseudostratified epithelium with ciliated cells, mucus-secreting goblet cells and basal cells in most species (Fig. 2).18,21,90,120 These morphological characteristics are essential for a proper function of innate defense mechanisms,8,27,28 and allow the culture system to mimic the in vivo respiratory tract environment more closely. 104 Moreover, numerous tests can be performed simultaneously in many replicates by using ALI cultures derived from a single animal (Table 1).
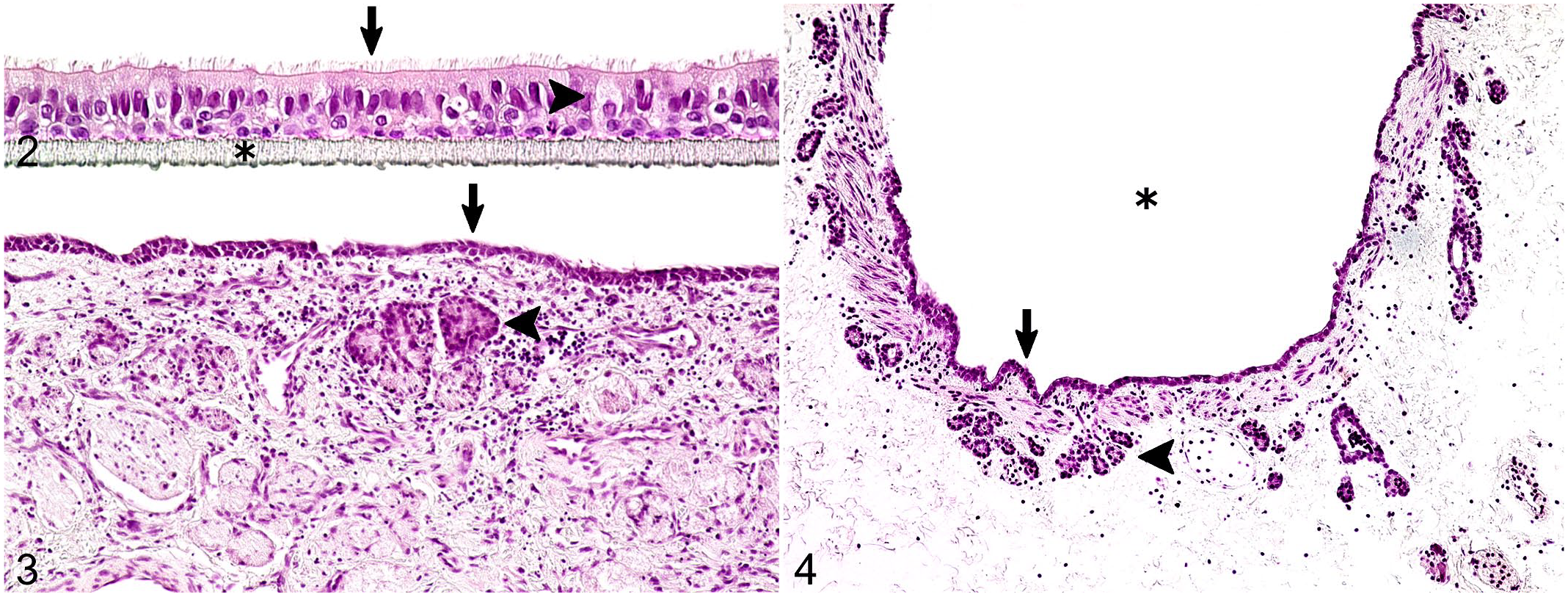
Characteristic morphological features of selected in vitro and ex vivo cell culture methods generated from feline tissues. Hematoxylin and eosin.
Average number of specimens of air–liquid interface culture inserts and precision-cut lung slices recovered from the trachea and lung of a single animal (own observations).
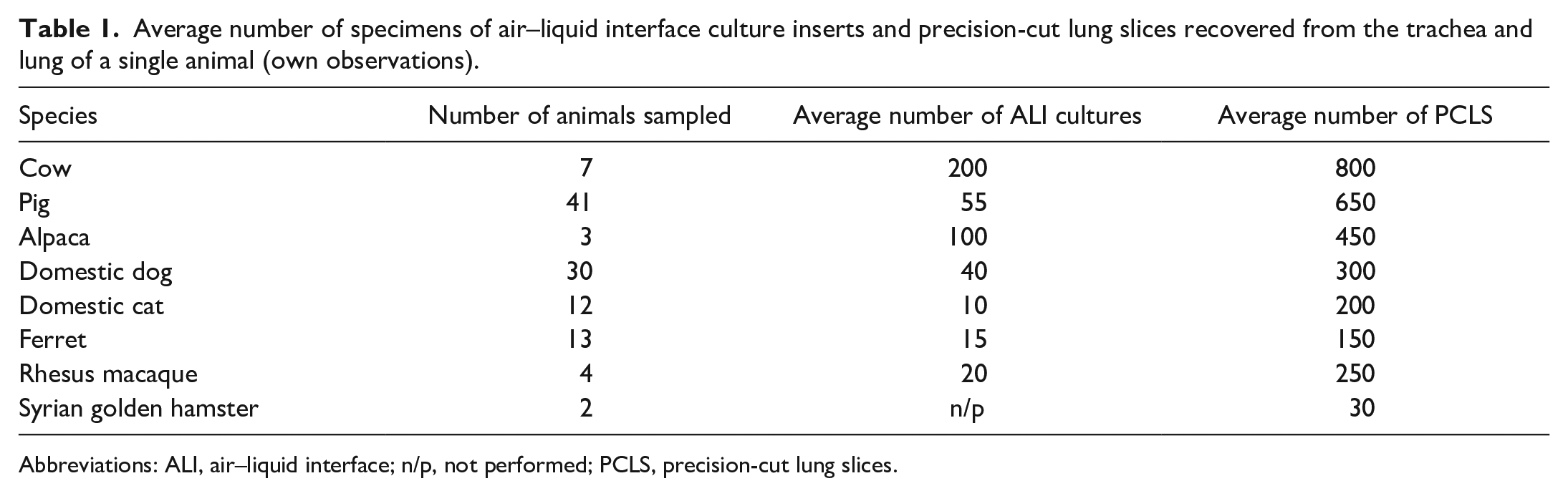
Abbreviations: ALI, air–liquid interface; n/p, not performed; PCLS, precision-cut lung slices.
ALI cultures have been successfully established for a broad variety of different species including humans (Fig. 5).8,37,104,123 For their generation, trachea and/or bronchi are exposed to enzymatic digestion with protease and desoxyribonuclease I, then epithelial cells are obtained by scraping from the tissue, and seeded in type-I collagen-coated flasks until cells reach 70% to 80% confluence. For culture in ALI conditions, primary respiratory cells are seeded on type-IV collagen-coated, semipermeable membranes at a density of 2 to 5 million cells per membrane in a transwell system. Within 3 to 5 days, a confluent monolayer is formed, which is required for establishing ALI conditions. A transepithelial electrical resistance (TEER) voltmeter is used to determine the presence of tight junctions.109,123,128,131
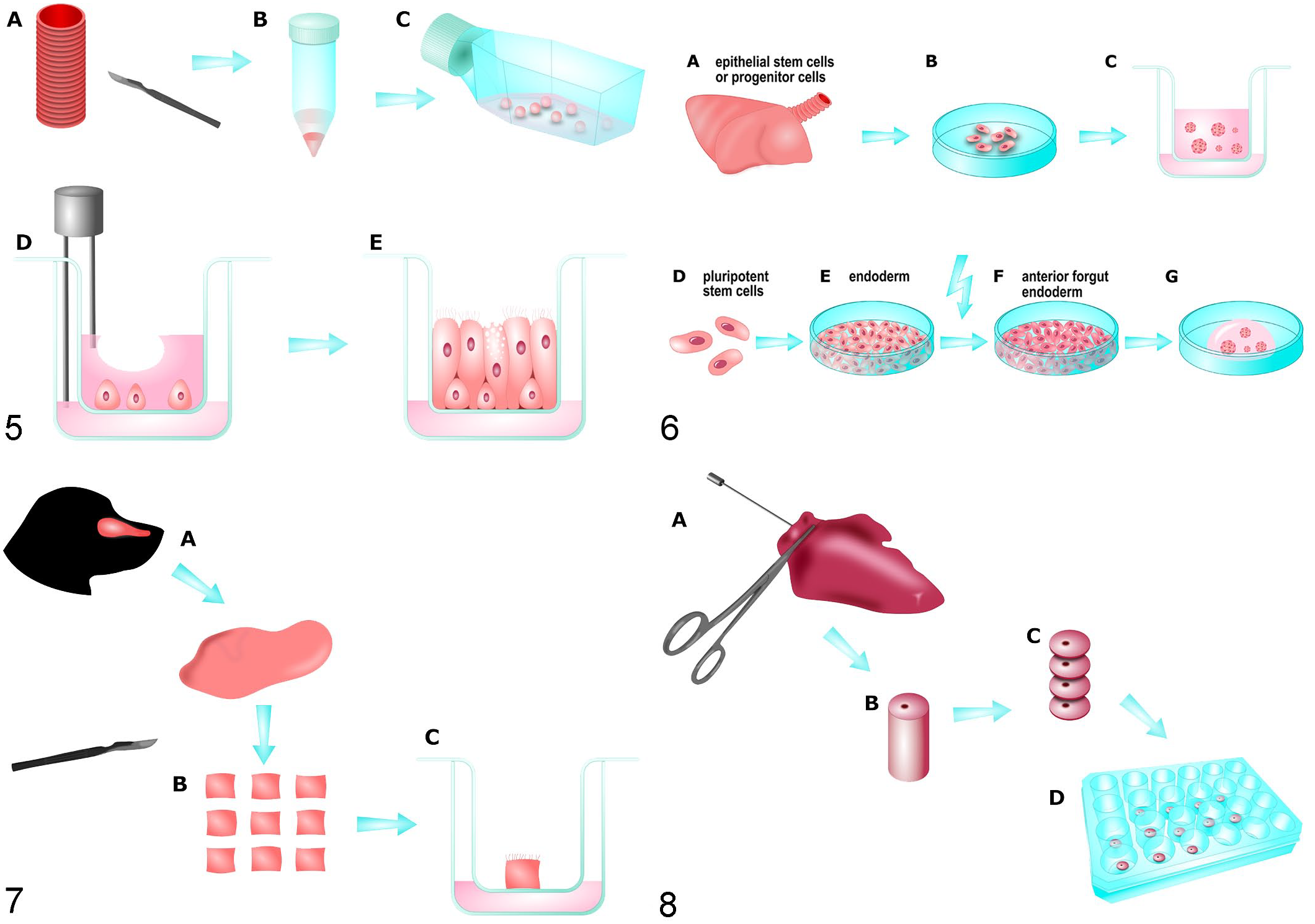
Schematic illustration of the process for producing selected cell culture models.
One typical application of ALI cultures is the investigation of the pathogenesis of airborne viruses and bacteria. An overview of previously performed infection studies is given in Supplemental Table S1. Swine influenza viruses A H1N1 and H1N2 are able to replicate in differentiated porcine ALI cultures with viral particles being released toward the apical surface. 8 In addition, it has been shown that swine influenza A virus infection induces apoptosis in ciliated cells and causes damage to the mucociliary clearance system. 153 Further infection studies have concentrated on bovine ALI cultures with a focus on 3 viruses associated with the bovine respiratory disease complex: bovine parainfluenza virus 3 (BPIV3), bovine respiratory syncytial virus (BRSV), and bovine herpesvirus type 1 (BHV1). These 3 viruses have been compared for their ability to infect bovine bronchial epithelial cells under ALI conditions and for their entry mechanisms to cross the epithelial barrier.50,72 In the family of Herpesviridae, additional studies have been conducted with equine herpesvirus type 1 (EHV1) 142 and feline herpesvirus type 1 (FHV1). 104 These have shown that the integrity of the respiratory epithelium is essential for the innate defense against primary alphaherpesvirus infections. In addition, porcine ALI cultures have been used to study infections with bacterial pathogens like Streptococcus suis, 96 Mycoplasma hyopneumoniae, and Mycoplasma flocculare 158 and bronchial epithelial culture models from cattle and sheep have been applied for the investigation of the host inflammatory response to Mycoplasma sp. infection.90,155
To summarize, ALI cultures represent a highly suitable tool to study various respiratory tract diseases and allow detailed investigations of virus infection and spread, cytopathic effects, and certain aspects of the pathogenesis.
Lung Organoids
Organoids are stem cell-derived 3D culture systems35,114 distinguished by the autonomous organization of cells and organotypic architecture with or without supporting mesenchymal cells.19,38 They represent another important culture method to investigate the pathogenesis of infectious diseases, 35 as well as the effects of pharmaceuticals, 148 and environmental influences such as airborne toxic substances 5 on the respiratory tract.
At present, 2 primary ways to generate lung organoids have been described, either from isolated adult stem cells or from pluripotent stem cells including embryonic stem cells and induced pluripotent stem cells (Fig. 6). 6 Lung organoids derived from epithelial stem cell or progenitor cell populations of the adult lung can be developed from basal cells, 126 airway secretory club cells,92,124 or alveolar type-II cells 7 using fluorescence-activated cell sorting or magnetic bead sorting. Primary cells can be directly embedded in Matrigel, in inserts or multiwells, and cultured as self-perpetuating 3D primary cultures in growth factor enriched media. 6
Directed differentiation of pluripotent stem cells into lung organoids has been based on the generation of endoderm and then anterior foregut endoderm, from which the lung is derived, and on the subsequent differentiation of foregut spheroids into lung organoids.23,52 Foregut spheroids are differentiated in serum-free media after 9 to 10 days. If subsequently embedded in a droplet of Matrigel and grown in high levels of fibroblast growth factor 10 and fetal bovine serum, they develop into lung organoids, which form airway-like structures and cell types surrounded by mesenchymal populations. 97 In general, there are numerous differentiation protocols with variations to the components of the growth medium, the extracellular coating, and the stages at which the cells are placed in a 3D environment.6,19,36,65
To date, several organoids derived from companion and farm animals have been successfully generated, including models of the intestine (dog, chicken, pig, and cow),14,29,49,113,116,119 mammary gland (cow),22,86,87 and liver (cat and dog).5,74,102,103 However, no lung organoids obtained from cells derived from companion or farm animals have been generated yet for the investigation of respiratory infectious diseases to our knowledge.
Generation and Use of Selected Ex Vivo Models in Non-SARS-CoV-2 Investigations
Nasal Mucosa Explants
The nasal mucosa constitutes a dynamic interface constantly exposed to the external environment with myriads of airborne pathogens. It simultaneously acts as a physical and immunological barrier to protect the host against undesirable invaders or toxins. 11 Moreover, the mucosal epithelial cells are a common site of virus invasion and replication after infection. As an example, MERS-CoV infection of dromedaries leads to a high virus load in nasal turbinates. 58
Nasal mucosa explants (NMEs) can be obtained from nasal turbinates or the nasal septum (Fig. 3).48,145 The respiratory mucosa is stripped from the underlying cartilage using surgical blades and washed several times in phosphate-buffered saline (PBS). After washing steps are completed, the respiratory mucosa is cut into 16 to 25mm2 pieces and placed on meshed gauze with the epithelial surface facing upward (Fig. 7). Explants are cultured at an ALI with serum-free medium (50% Dulbecco’s Modified Eagle’s Medium [DMEM] and 50% Roswell Park Memorial Institute [RPMI] Medium) and, variably, additional antibiotics (gentamycin, streptomycin, and penicillin).
So far, only a few infection studies have been conducted in NME (Supplemental Table S2). In farm animals, most studies using NME have investigated viral infection of porcine tissue with suid herpesvirus 1 (SHV1) to study host–virus interaction 48 and to gain a better understanding of viral pathogenesis 45 as well as to clarify molecular mechanisms of viral penetration. 46 Similar complementary approaches including the 2 ex vivo system models NME and PCLS have been used in another study to compare swine, human, and avian influenza isolates and confirm susceptibility of porcine respiratory tissues to infection with avian influenza virus. 145 The infectivity pattern in this study was consistent with results of in vivo experiments,31,79 indicating that appropriate ex vivo models offer useful alternatives to the use of live animals. Nasal mucosa explants have also been used to investigate viral entry of Japanese encephalitis virus in pigs and thus provided insights into nonvector-borne direct transmission of this virus. 42 EHV1 has been used to infect equine NME and to study viral replication kinetics as well as disruption of intercellular junctions by the virus, which is used to overcome respiratory epithelium integrity.47,142 –144
Precision-Cut Lung Slices
Precision-cut lung slices (PCLS) consist of various pulmonary cell types arranged in the organotypic 3D multilineage architecture of the lung (Fig. 4) and can model certain vital functions of the respiratory tract, such as a local immune response, 80 mucus production, 26 ciliary activity, and bronchoconstriction.4,78 Precision-cut lung slices can be maintained for at least 1 week in culture conditions.80,105 Furthermore, many lung-slice cultures can be generated from one organ (Table 1), reducing the number of animals required to obtain meaningful data while allowing a sufficient number of replicates for statistical analysis. In addition, cross-species comparisons 132 allow the assessment of translation aspects of preclinical animal data to the human situation. 105
The generation of agarose-inflated lung slices of a variety of species has been extensively investigated and described with protocols available in which the slices remain viable for at least 1 week (Fig. 8).32,72,94 Briefly, removed lung lobes are stabilized by filling the airways with low-melting-temperature agarose followed by solidification on ice. After trimming the samples with a Krumdiek tissue slicer to thin, equal slices of approximately 250 µm thickness, the PCLS are transferred and maintained in culture medium (50% DMEM and 50% Ham’s F12 media) containing essential maintaining factors and antibiotics (streptomycin and penicillin). The intactness of PCLS can be monitored by observing the ciliary activity of bronchial epithelial cells via light microscopy. 149
Precision-cut lung slices have been used to investigate viral tropism and pathomechanisms of various infectious diseases in domestic and farm animals (Supplemental Table S3). In pigs, most groups have utilized different subtypes of influenza A virus primarily targeting respiratory epithelial cells to investigate the replication efficiency and ciliostatic effects of virus infections.41,73,94,95 Due to homologies of sialic acid receptor distribution in the respiratory tract of humans and pigs, porcine PCLS have been shown to represent a promising culture system for the modeling of susceptibility, virulence, and replication of influenza A viruses. 121 In addition, studies have found that pigs are susceptible to nonspecies-specific subtypes of influenza A viruses like avian or even human subtypes under experimental and natural circumstances.115,121,145 An influence of primary viral infection on the susceptibility for bacterial co-infection has been determined by a study using influenza A virus followed by infection with Streptococcus suis. 95 In another study investigating bacterial pathogens in pigs, Bordetella bronchiseptica infection in porcine PCLS promoted adherence and colonization of Streptococcus suis. Notably, the prior infection with Bordetella bronchiseptica supported cytotoxic effects of Streptococcus suis. 146 Other infection studies on PCLS derived from farm and domestic animals have so far been restricted to nonbovine ruminants, to investigate the Jaagsiekte sheep retrovirus, 20 and respiratory disease complex in goats. 72 In addition, investigations on the pathogenesis of paramyxoviruses have been performed using PCLS of ferrets and dogs infected with canine distemper virus, macaques with measles virus, as well as cotton rats with metapneumovirus and respiratory syncytial virus. 108 Consistent with the in vivo situation, all viruses showed different replication efficiency in ex vivo lung slices.
To summarize, PCLS provide an important bridge between in vivo and in vitro experiments by enabling detailed investigations in an organotypic cellular composition setting without the need for live animals.
Use of In Vitro Models in SARS-CoV-2 Research
Cell Lines Derived From Various Tissues
Human-derived, immortalized cell lines such as Calu-3, Huh-7, Caco-2, and HEK293, generated from various organs, have been used for isolation of SARS-CoV-257,70,168 and have also played a vital role in the establishment of ACE-2, transmembrane protease serine subtype 2 (TMPRSS2), and furin as important factors for SARS-CoV-2 entry.62,67,89,133
There are also numerous cell lines derived from species, such as nonhuman primates (most importantly Vero E6 cells110,168), bats, hamsters, and cats, that have been inoculated with SARS-CoV-2 to describe virus-host cell interactions and to determine possible susceptibility of several species: in 6 out of 16 tested animal-derived cell lines from one study (originating from bats, non-human primates, dogs, cats, pigs, rabbits, chicken, mice, and hamsters), virus replication was found to be most effective in nonhuman primate and pig cells. 17 Another study also reported successful infection of porcine cell lines, which stands in contrast to the fact that experimental infection of domestic pigs in the same study as well as in others has shown that pigs are not susceptible to SARS-CoV-2 infection.93,129,135 This strongly implies that there is a discrepancy between immortalized cell lines and in vivo models, exemplified by phenotypic alterations59,69 and a variable expression of proteins in cell lines compared to primary cells, 44 making the search for more suitable in vitro or ex vivo models a necessity. Surprisingly, in primary kidney cells derived from Chinese rufous horseshoe bats (Rhinolophus sinicus), SARS-CoV-1 but not SARS-CoV-2 was able to replicate, even though Rhinolophus sinicus bats have been identified as a highly probable reservoir of SARS-CoV-2.17,168 It was hypothesized by the authors that the inability of the virus to infect bat cells was due to adaptation of the virus to humans; 17 however, a disparity between the in vivo and in vitro situation could also have been responsible, because in another study, fruit bats were susceptible to infection using a SARS-CoV-2 isolate. 129
Primary Cell Culture of Respiratory Cells
To imitate in vivo conditions more precisely, human primary airway cells derived from tracheal and/or bronchial mucosa and cultured at an ALI have been used to study viral replication and changes in gene expression following infection with SARS-CoV-2.56,118,134 As a result, detailed analyses of tracheobronchial epithelial cells cultured at ALI conditions have revealed that ciliated cells constitute a primary target for infection.56,169 In a comparable study using nasal primary human airway epithelial cells, similar results were observed, with a notable lack of viral replication in goblet cells. 117
In animals, primary lung cells derived from white-tailed deer, one of the species that shares high uniformity with the human ACE-2, have been inoculated with a SARS-CoV-2 isolate, resulting in productive viral replication. 112 Showing that experiments with primary cell cultures can be used as a first-line screening method, susceptibility of deer fawns was confirmed in vivo by intranasal infection that resulted in efficient virus replication and shedding. Notably, infected white-tailed deer were able to transmit the virus to naïve deer by indirect contact, 112 making this species a potential source of virus spread.
The use of primary tracheobronchial ALI cultures for SARS-CoV-2 research has only recently been described in animals: of 12 animal species including primates, ferrets, cats, and bats, efficient replication of SARS-CoV-2 was only seen in primary cells derived from rhesus macaques and cats. 54 Cells obtained from other species that were reported to be susceptible to infection in vivo, such as dogs, ferrets, and rabbits,51,101,136 did not show efficient viral replication in vitro. A proposed explanation was that due to differences in viral cellular tropism, SARS-CoV-2 infection in these animal species mainly occurs in other parts of the respiratory tract (such as the nasal tissue) that show a different cellular composition than the investigated cell culture. 54 This should be confirmed with further studies using different organotypic culture systems, as it would expand the knowledge about the behavior of SARS-CoV-2 in different host species and more generally contribute to a better understanding of host factors determining susceptibility to viral infection. However, since SARS-CoV-2 often affects cells located in the distal lung tissue, especially alveolar epithelial type-II cells, 64 which are not easily maintained as primary cultures, 157 other culture methods also have to be considered.
Organoids Generated From Various Cell Types
To investigate the viral tropism and cytopathogenic effects of SARS-CoV-2 on alveolar epithelial type-II cells, induced pluripotent stem cells can be used to generate ALI lung cultures or epithelial lung organoids. Both types of culture methods have been successfully infected in vitro.64,114,125,157 By analyzing human-induced pluripotent stem cell-derived alveolar epithelial type-II cells, certain cell-type-specific transcriptomic reactions to infection were detected, such as a reduced expression of surfactant genes. These were not previously anticipated using primary airway cells or cell lines. Importantly, it was noted that interferon responses to SARS-CoV-2 infection were delayed with only moderate induction of interferon signaling, which, if confirmed in vivo, could present possible treatment targets. 64 In addition, the use of human pluripotent stem cells in SARS-CoV-2 research is not only confined to the respiratory tract but can also be extended to other organ systems, which has revealed that intestinal, vascular, renal, brain, and liver tissues are also susceptible to infection.98,156,159,163,167
SARS-CoV-2 research in organoids has not only been accomplished using human-derived tissue but also by using cells from several animal species. For example, with the establishment of an adult stem cell-based organoid culture of bat small intestinal epithelium, virus isolation was shown to be more efficient than with the conventionally applied Vero E6 cell line. Furthermore, analysis of distribution of ACE-2 and TMPRSS2 by immunofluorescence was conducted in this model. 167 In addition, mouse lung organoids have been cultured to test several pharmacological agents for their effect on growth and differentiation of progenitor cell types responsible for epithelial repair in the lung in an effort to contribute to the development of effective therapeutics against SARS-CoV-2. 148
Use of Ex Vivo Models in SARS-CoV-2 Research
Tissue Explants of the Respiratory Tract
Using ex vivo tissue explants such as human lung tissue in the research of SARS-CoV-2 has presented multiple important and unique insights: by infecting lung tissue samples from the same donor with both SARS-CoV-2 and SARS-CoV-1, a direct comparison of the viral kinetics, cell tropisms and host immune response of the 2 viruses was enabled. 15 As a result, it was clearly shown that SARS-CoV-2 was able to produce a more robust infection and replication in infected lung explants, even though expression of interferons and proinflammatory mediators was more pronounced in SARS-CoV-1 infection. 15 This suppressed activation of the innate immune response has been given as a possible explanation for the mild symptoms observed in most patients with COVID-19 compared to patients with SARS 15 and has also been noted in in vitro experiments utilizing organoids. 64 Furthermore, using infected lung tissue explants, it has been discovered that both SARS-CoV-1 and SARS-CoV-2 display similar cell tropisms in the lung, with type-I and -II pneumocytes and alveolar macrophages being affected. 15 Other tissue explants, such as the nasal mucosa derived from nasal turbinates 2 and conjunctiva, 66 have also been generated for subsequent SARS-CoV-2 infection. Interestingly, upon infection of NME with SARS-CoV-2, which primarily targeted respiratory epithelial cells, a strong innate immune response was elicited, implying that there are tissue-specific response patterns to SARS-CoV-2 infection that could be further investigated using ex vivo and in vitro culture methods. 2 This was further legitimized by another study focused on human intestinal tissue explants, where SARS-CoV-2 induced a stronger innate immune response than SARS-CoV-1 and showed less replication and cytopathic effects in the intestinal epithelium. 16
Similar to the studies conducted using human tissues, tissue explants derived from several species have been applied in the research of SARS-CoV-2 for evaluation of a possible zoonotic potential of these animals: In line with previous in vivo studies of SARS-CoV-2 in dogs,10,135 low susceptibility of canine tissue explants derived from the nasal cavity, soft palate, trachea, and lung was demonstrated, with only nasal cavity explants showing limited replication. 12 This stands in contrast to the substantial expression of ACE-2 in the nasal and palatal epithelium exhibited by immunohistochemical labeling in the same study. It was proposed by the authors that additional experiments using tissue explants and focusing on different isoforms of ACE-2 as well as other host activating proteases like furin and TMPRSS2 could be conducted to better understand this discrepancy. 12 In another investigation, productive replication of SARS-CoV-2 was seen in bovine and ovine tracheal and lung tissue explants, with higher kinetic titers in bovine lung explants compared to the tracheal explants. 33 For the ovine model, no differences were found between viral replication in tracheal and lung tissue explants. Porcine explants derived from these localizations were determined not susceptible to infection with SARS-CoV-2, correlating with a missing ACE-2 immunoreaction, in contrast to bovine and ovine tissue, where viral antigen was associated with ACE-2 positive cells in double-labeling immunofluorescence. 33
Precision-cut lung slices (PCLS)
As another, more complex ex vivo culture method, human PCLS have been successfully infected with SARS-CoV-2 and have consequently been used for pharmacological studies such as treatment with agents like fluoxetine, which caused a significant reduction of viral load. 171 In a further investigation, PCLS were incubated with the protease inhibitors camostat mesylate, its metabolite FOY-251, and nafamostat mesylate, 61 to confirm previous results showing that camostat mesylate treatment can efficiently inhibit SARS-CoV-2 infection in lung cell culture. 62 Indeed, in this study, it was shown that all 3 substances can inhibit viral infection, indicating that there is potential for additional treatment options which should further be examined in in vivo studies. 61
Conclusions and Outlook
Various well-characterized and established in vitro and ex vivo systems of the upper and lower respiratory tract have played an important role in the research regarding SARS-CoV-2 in humans as well as animals, complementing or even substituting in vivo experiments. Questions about the host and cell tropism, virus-triggered cellular pathology, associated virulence, and efficacy of potential treatments and vaccines have been investigated by using cell lines, primary bronchoepithelial cells cultured at ALI conditions, organoids, tissue explants, and PCLS. However, more research concerning SARS-CoV-2 needs to be carried out, specifically regarding the still-unknown intermediate host, patterns of infection with respect to the upper and lower respiratory tract between different animal species, and varying isoforms of ACE-2 potentially limiting SARS-CoV-2 entry in certain cell types.
When considering these investigations, multiple factors that limit the applicability of culture models have to be taken into account: immortalized cell lines are broadly accessible and easy to maintain in culture but they often do not reflect the physiology of the cells they originated from, for example in regards to gene expression patterns. In addition, some cell lines are derived from neoplastic cells and therefore have to be considered genomically abnormal. This may result in passage-dependent morphological and phenotypical changes which can limit their comparability to in vivo circumstances.13,59 In that regard, primary cell culture models are a more accurate representation.
However, a limiting factor when using these culture systems for statistical analysis is the number of samples available. For ex vivo systems like PCLS, this is not a major concern as large amounts of samples can be obtained even from limited tissue supply. In contrast, generating in vitro primary cell cultures derived from smaller animal species like cats, ferrets, or hamsters is often challenging due to the low cell count available (Table 1). A major disadvantage to tissue explants is the fast tissue deterioration rate, making long-term studies challenging. For example in PCLS, the composition of present cells and their functionality changes over time in culture, leading to a reduced production and secretion of cytokines, which as a result might alter the effects of an infectious agent on the cells.105,106 For all the above mentioned culture techniques, the lack of vasculature and inability to recruit nonresident immune cells as well as the missing interactions between different cell types are factors that currently limit their use as replacements for in vivo experiments performed on animal models (Supplemental Table S4).2,30,83 In the future, some of these challenges will likely be overcome.25,34,60,165.
The potential of additional aids when working with cell cultures, such as the performance of (semi-) automated analyses of PCLS, should be considered. The mucociliary transport system, which includes efficient mucus production and secretion as well as coordinated ciliary beating (Supplemental File 1), plays a major role with respect to the prevention of virus-triggered lesions and virus clearance. 82 It has been discovered that ciliary loss and consequent disturbances of mucociliary clearance are vital parts in SARS-CoV-2 pathogenesis.150,198 Therefore, by enabling precise measurements of oscillation frequencies and patterns to evaluate ciliary beating using video processing techniques (Figs. 9–10), as has already been described for other purposes,39,84,164 the value of PCLS in the discovery of viral effects on cilia would be significantly increased.
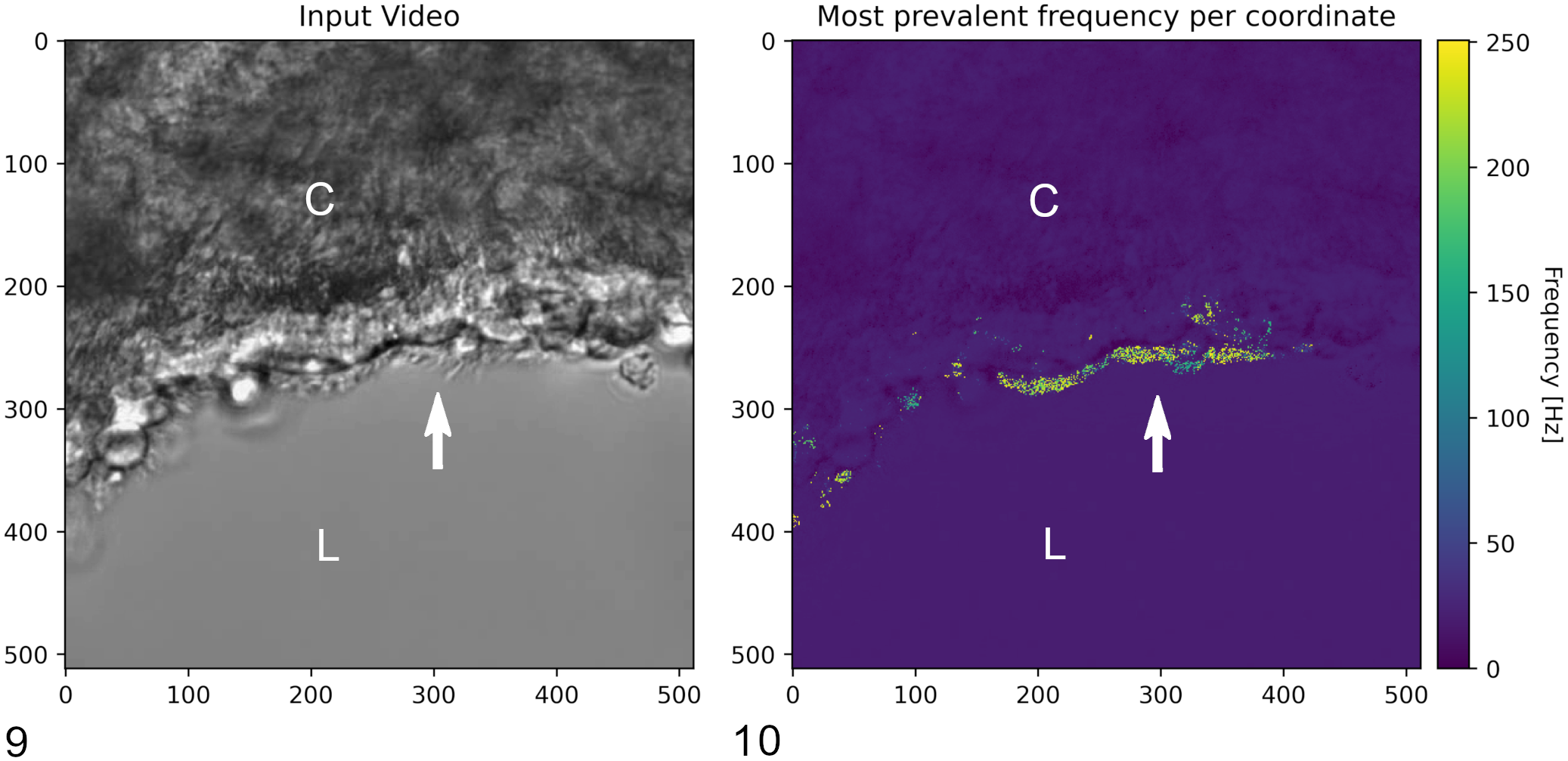
Video-assisted analysis of precision-cut lung slices (PCLS).
To summarize, in vitro and ex vivo models derived from humans as well as animals build an important foundation for in vivo experiments and clinical application. They can be considered essential for preclinical pathogenetic studies of infectious diseases with and without zoonotic potential, whilst championing the 3R principles.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858211073678 – Supplemental material for Alternatives to animal models and their application in the discovery of species susceptibility to SARS-CoV-2 and other respiratory infectious pathogens: A review
Supplemental material, sj-pdf-1-vet-10.1177_03009858211073678 for Alternatives to animal models and their application in the discovery of species susceptibility to SARS-CoV-2 and other respiratory infectious pathogens: A review by Sandra Runft, Iris Färber, Johannes Krüger, Nadine Krüger, Federico Armando, Cheila Rocha, Stefan Pöhlmann, Laura Burigk, Eva Leitzen, Malgorzata Ciurkiewicz, Armin Braun, Daniel Schneider, Lars Baumgärtner, Bernd Freisleben and Wolfgang Baumgärtner in Veterinary Pathology
Footnotes
Acknowledgements
The authors thank Kerstin Schöne, Danuta Waschke, Kerstin Rohn, Julia Baskas, Petra Grünig, Caroline Schütz, Christiane Namneck, Jana-Svea Harre, Siegfried Jelitto, Dunja Hoffmann, and Silvana Böhmecke for their exceptional technical assistance.
Supplemental material for this article is available online.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was in part supported by the R2N Project funded by the Federal State of Lower Saxony, the ANI-CoV Project funded by the German Ministry for Education and Research (BMBF: ZA_FKZ: 01KI2074A-B), a BMBF (Federal Ministry of Education and Research) project entitled RAPID (Risk assessment in re-pandemic respiratory infectious diseases), 01KI1723G and by the Ministry of Science and Culture of Lower Saxony in Germany (14 - 76103-184 CORONA-15/20).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
