Abstract
Epidermal growth factor receptor (EGFR) is overexpressed in many human colorectal cancers and anti-EGFR agents are employed as immunotherapies. However, KRAS, EGFR, and BRAF gene mutations can influence the activity of the anti-EGFR agents. We evaluated EGFR expression at protein and mRNA levels in canine intestinal adenocarcinomas using immunohistochemistry (IHC) and RNA in situ hybridization (RNA-ISH). We also investigated the mutation status of EGFR, KRAS, and BRAF to aid the development of anti-EGFR agents for canine intestinal adenocarcinoma. EGFR expression was highest in adenocarcinoma, followed by intramucosal neoplasia (adenoma and in situ carcinoma), and nonneoplastic canine intestinal tissue, at both protein (P = .000) and mRNA (P = .005) levels. The EGFR, KRAS, and BRAF genes showed wild-type sequences at the mutation hot spots in all 13 specimens. Thus, EGFR might serve as a promising diagnostic marker in canine intestinal adenocarcinoma, and further studies would be needed to develop EGFR-targeted anticancer therapies.
Although the prevalence of intestinal tumors in dogs is only 0.6% to 0.7%, canine intestinal adenocarcinoma accounts for 65% of these tumors. 18 Benign tumors are rare among canine intestinal epithelial tumors. 18 Canine intestinal lesions are diagnosed using endoscopy, analogous to that in humans. 35 Diagnosis are usually based on hematoxylin and eosin (HE) staining, and additional molecular information can provide prognostic or therapeutic information. 34,35
Epidermal growth factor receptor (EGFR) is a transmembrane receptor tyrosine kinase that transduces growth factor–related signaling to downstream molecules like Kirsten rat sarcoma viral oncogene homolog (KRAS) and serine/threonine-protein kinase B-Raf (BRAF), ultimately inducing cell proliferation. 4 In humans, anti-EGFR agents can serve as therapeutics when EGFR is overexpressed in colorectal cancer. 13,26 Despite having different pathological features such as a 50-fold lower incidence than that of colorectal cancer, 19 human small intestinal adenocarcinoma often exhibits EGFR overexpression, 20 indicating that EGFR could also be targeted to treat adenocarcinoma in the human small intestine. 20 Anti-EGFR agents exert their effects via 2 mechanisms. 21 First, monoclonal antibodies specifically bind to EGFR, which prevents growth factor binding and receptor activation, and consequently inhibits tumor growth. 25 For instance, many types of human cancer have been treated with cetuximab, an anti-EGFR monoclonal antibody. 4,25 Second, anti-EGFR tyrosine kinase inhibitors such as gefitinib block the processes downstream of EGFR, resulting in tumor growth inhibition. 26
Human and dog EGFR are significantly conserved at the amino acid level and exhibit 91% homology. 29 Cetuximab targets an EGFR epitope, inhibits the processes downstream of the EGFR pathway, and reduces tumor size in canine mammary carcinoma cell lines in vitro through pharmacological mechanisms that are similar to those in human tumors. 29 In addition, an anti-EGFR canine immunoglobulin for passive immunotherapy is available. 28
However, mutations in KRAS or BRAF in human colorectal cancers result in the EGFR-independent activation of the MAPK pathway, rendering anti-EGFR agents such as anti-EGFR monoclonal antibodies and tyrosine kinase inhibitors ineffective. 12,23 KRAS and BRAF mutations in dogs and humans have been reported in the same mutation hot spots, namely, codons 12, 13, and 61 in KRAS and codon 450 in BRAF, 15,16 and these hot spots share the same amino acid sequences between dogs and humans. 15
EGFR mutations frequently occur between exons 18 and 21 that comprise a kinase domain, in humans. 22 Tumors harboring mutations in kinase domain of EGFR are susceptible to the activity of tyrosine kinase inhibitors, such as gefitinib, as opposed to tumors with KRAS mutations, which are mutually exclusive from EGFR mutation. 3,22
The findings of some studies have enabled the development of canine anti-EGFR agents, 28,29 and EGFR expression and KRAS mutations have been evaluated in canine gastric tumors. 31 However, these have not been evaluated in canine intestinal tumors. Therefore, we evaluated EGFR expression and EGFR, KRAS, and BRAF mutations in canine intestinal neoplasms to determine the feasibility of developing anti-EGFR drugs for canine intestinal adenocarcinoma. In addition, we analyzed EGFR transcripts in formalin-fixed paraffin-embedded (FFPE) tissues of canine intestinal tumors to confirm the role played by EGFR mRNA in the development of canine intestinal tumors as the overexpression of its protein might not be related to the overexpression of mRNA as factors such as posttranscriptional regulation must be accounted for. 2,11,36
Materials and Methods
Ethical Statement
All samples were obtained from local animal hospitals during endoscopy or surgery for enabling the diagnosis at the Department of Veterinary Pathology, Konkuk University. This study did not use any live animals, and thus, approval by the Institutional Animal Care and Use Committee (IACUC) was waived. The owners of the dogs provided informed consent to use the clinical data and biological samples for teaching and research purposes.
Case and Control Tissues
We selected 21 documented canine intestinal tissues including additional samples remaining after the completion of a previous study, 5 from the histopathological database of the Department of Veterinary Pathology, Konkuk University, Seoul, South Korea, between 2005 and 2020. Nonneoplastic intestinal regions of samples 2, 5, 9, and 10 were included as nonneoplastic canine intestinal tissues. Severely inflamed tissues or samples that were <1 cm in diameter were excluded from the analysis. Histopathological parameters in HE-stained slides were evaluated by 2 pathologists in a blinded manner (SH-C, BJ-S), and ambiguous results were resolved by discussion with a third pathologist (JH-S). Clinical information included breed, sex, and age of the dogs; clinical signs; and biopsy site.
Tissue samples were fixed in 10% neutral-buffered formalin, processed routinely, and embedded in paraffin wax. Sections (4 µm thick) were stained with HE. The samples were assigned to “nonneoplastic canine intestinal tissue,” “intramucosal neoplasia,” or “adenocarcinoma” groups. Tissue samples from dogs that were assessed by endoscopy during regular health checks or in response to the manifestation of clinical signs such as vomiting—but no epithelial proliferation—were included as nonneoplastic canine intestinal tissues. Adenomas and in situ carcinomas were grouped together as “intramucosal neoplasia,” because they are difficult to differentiate from each other in HE-stained sections and in situ carcinomas are not considered clinically malignant due to the lack of extension past the basement membrane. 9,18 When investigated using the 2003 classification of the World Health Organization, the following histological subtypes were identified in the adenocarcinoma group: acinar, papillary, and mucinous adenocarcinoma, signet ring cell carcinoma, and adenosquamous carcinoma. 10
Immunohistochemistry
All samples were immunolabeled for EGFR using an antibody with validated immunohistochemistry (IHC) reactivity in canine tissues. 24,27 The homology of the EGFR antibody-binding site between dogs and humans is 94.71%, a relatively high conservation rate as confirmed by amino acid sequence alignment using BLAST (Basic Local Alignment Tool; http://blast.ncbi.nlm.nih.gov/Blast.cgi). 1,8 Sections immobilized on slides were deparaffinized in xylene and rehydrated in graded ethanol before staining. Endogenous peroxidase activity was blocked by immersing the sections in 3% H2O2 prepared in phosphate-buffered saline (PBS; 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4; pH 7.4) for 20 minutes at room temperature. Antigen retrieval was performed by incubating the sections with proteinase K (Dako) for 15 minutes at room temperature. Nonspecific binding to sections was blocked using 5% normal goat serum prepared in PBS, and then the sections were incubated overnight (15 hours) with an EGFR-specific primary antibody (clone 111.6, Thermo Fisher Scientific) at 4 °C. The sections were washed with PBS, incubated with secondary antibodies (Dako REAL EnVision kit, Dako) for 40 minutes at room temperature, stained with 3,3′-diaminobenzidine (Dako REAL Envision kit, Dako), and counterstained with Gill hematoxylin. A mouse IgG1 kappa isotype-matched antibody (eBioscience) was used as a negative control for IHC.
Images of tumor cells were acquired at 400× magnification using an ocular FN 22 mm objective. Necrotic and highly inflamed areas were excluded from interpretation. Digital images were acquired using an Olympus BX51 microscope (Olympus) and image transfer software (Olympus). The expression of EGFR in tumor cells was semiquantified as described for human colorectal cancer. 6 Membrane staining intensity was scored in the following manner: 0, no stained tumor cells; 1, <10% of the tumor cells stained at any intensity or <30% of the tumor cells stained at weak intensity; 2, 10% to 30% of the cells stained at moderate to strong intensity or 30% to 50% of the tumor cells stained at weak to moderate intensity; 3, >30% of the tumor cells intensely stained or >50% of the tumor cells stained at any intensity. A pathologist (SH-C) scored the staining, and ambiguous scores were resolved by discussion with a second pathologist (JH-S).
RNA In Situ Hybridization (RNA-ISH)
The expression of EGFR mRNA was analyzed by in RNA-ISH using a probe for EGFR and a positive control probe against the reference gene (POLR2A). Samples that did not express POLR2A were excluded from RNA-ISH analysis due to low mRNA retention. Finally, 15 samples (nonneoplastic canine intestinal tissue, n = 3; intramucosal neoplasia, n = 3; adenocarcinoma, n = 9) were evaluated. We used RNAscope 2.5 High Definition (HD)—BROWN Assays (Advanced Cell Diagnostics [ACD]) for performing RNA-ISH with some modifications. 33
Sections (4 µm thick) were cut from FFPE and placed on slides that were dried at room temperature overnight. The slides were then heated at 60 °C for 120 minutes, deparaffinized in xylene twice for 5 minutes each, then dehydrated twice in 100% EtOH for 1 minute each. The sections were dried for 5 minutes and incubated with hydrogen peroxide (ACD) for 10 minutes at room temperature. The sections were then heated at 60 °C for 60 minutes, immersed in Target Retrieval Reagent (ACD) at 95 °C for 15 minutes, then washed twice with distilled water and once with 100% EtOH. The sections were air-dried, then a barrier was drawn twice (ImmEdge, Vector Laboratories) around the tissue samples, which were then completely dried. The slides were placed in a slide rack, and an appropriate number of drops of Protease Plus (ACD) was added to each section based on the size of the tissues. The slide rack was incubated at 40 °C for 30 minutes and then rinsed in distilled water twice for 2 minutes each and incubated with the probes for 2 hours at 40 °C. A target probe against canine EGFR (Probe-Cl-EGFR, ACD), positive control probe (positive control probe-Cl-POLR2A, ACD), and negative control probe (negative control probe-DapB, ACD) were used. GenBank accession numbers and probe regions were as follows: EGFR (GenBank, XM_533073.4; target region, 725-1660), POLR2A (GenBank, XM_852751.3; target region, 1846-2924), and DapB (GenBank, EF191515; target region, 414-862). After incubation, the slides were rinsed twice with Wash Buffer Reagent (ACD; 2 minutes each).
The signal was amplified and detected with 2.5 HD Detection Reagent–BROWN (ACD). The tissue sections were then incubated with Amp 1 (preamplifier) for 30 minutes at 40 °C, followed by incubation with Amp 2 (background reducer), Amp 3 (amplifier), and Amp 4 (label probe) at 40 °C for 15 minutes each, then the sections were serially incubated with Amp 5 (30 minutes) and Amp 6 (15 minutes) at room temperature. After each step, the sections were rinsed twice with wash buffer (2 minutes each). Signals were detected using a DAB mixture (according to the kit), and sections were counterstained with 50% Gill’s hematoxylin. After rinsing with distilled water, the sections were washed with 0.02% ammonia water for 20 seconds, and then the slides were dehydrated and mounted with coverslips.
Interpretation of RNA-ISH
The EGFR/POLR2A ratio was measured to compare the level of EGFR expression (Fig. 1), and the same areas that had been evaluated using IHC were examined. Because the retention of RNA by FFPE tissues varies after fixation and storage, 17 we quantified target and reference genes in serial sections and determined the target gene (EGFR)/reference gene (POLR2A) ratio as follows. Each section was photographed for 5 areas with the same field of view for EGFR and POLR2A. The RNA-ISH images were acquired using the Olympus BX51 microscope and analyzed using the FIJI analysis software ImageJ win64 (National Institutes of Health, Bethesda, MD). We used the Image, Color, and Color Deconvolution modules, selected colour_3 from “H&E DAB” as the stain, then separated RNA-ISH-positive signals (dots) from the background. The dots were counted using ICY software. 7 Tumor sites (intestinal epithelial areas) were selected using the Region of Interest and Polygon tools. The dots were then counted using Detection & Tracking and Spot Detector. We set the dot metric by setting SizeFiltering to 10-300. EGFR and POLR2A signals were counted in 5 fields of view per section, then the EGFR/POLR2A ratio was calculated.
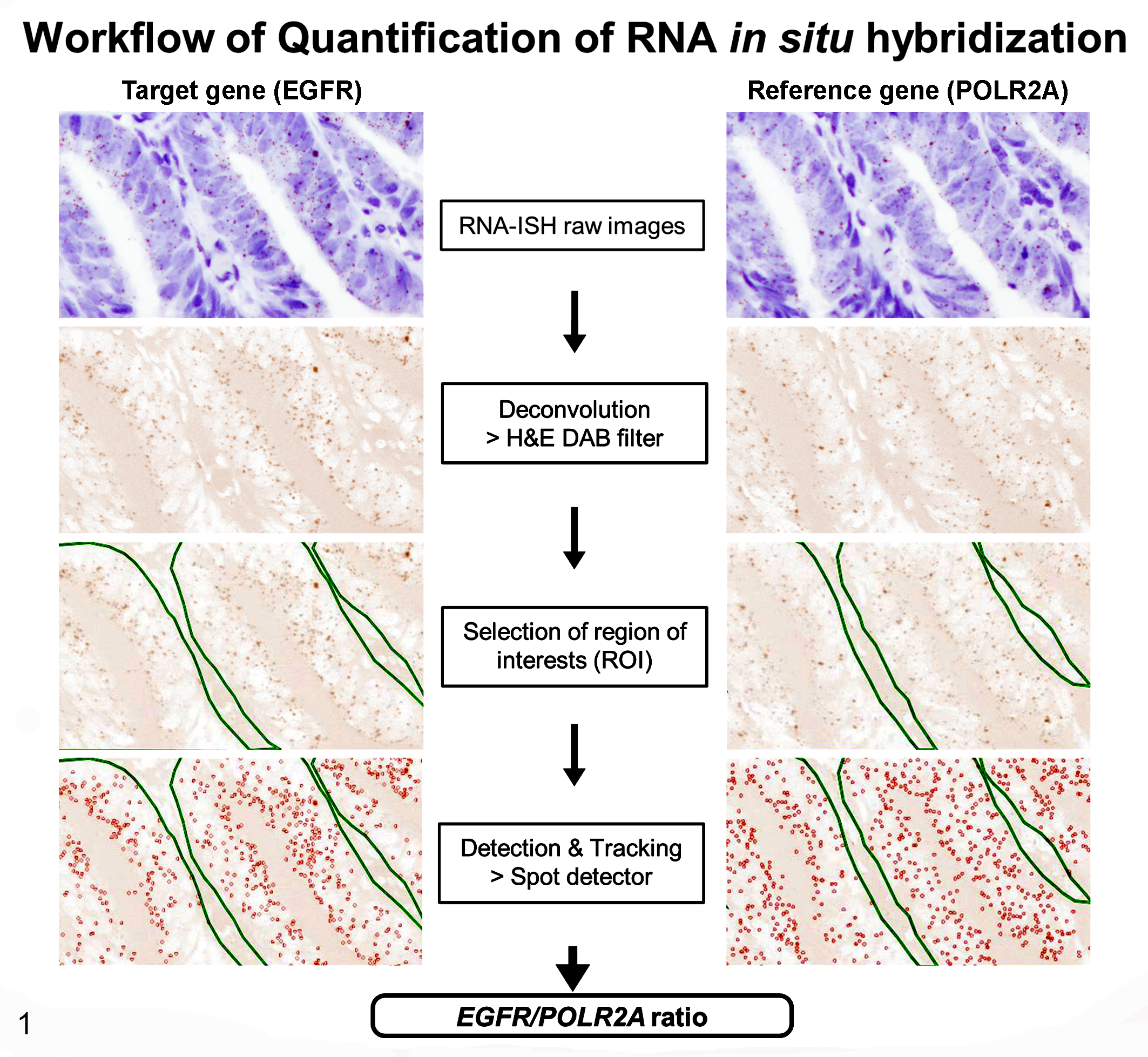
Flow chart depicting the workflow used for performing quantitative RNA in situ hybridization (RNA-ISH). Epidermal growth factor receptor (EGFR) mRNA signals appear as brown dots in the raw RNA-ISH images. These were converted by ImageJ using the “H&E DAB” filter. Regions of interest were selected and quantified using ICY software. Signals corresponding to EGFR mRNA were counted and circled in red, then EGFR/POLR2A ratios were calculated.
Statistical Analysis of IHC and RNA-ISH Data
Comparison of the results of IHC and RNA-ISH among the 3 groups (nonneoplastic canine intestinal tissue group, intramucosal neoplasia group, and adenocarcinoma group) were analyzed using Kruskal-Wallis tests followed by post hoc analysis with Dunn’s multiple comparison test. Association between IHC result and clinicopathological parameters (sex, age, location, subtype, invasion) in the adenocarcinoma group was assessed using Mann-Whitney U test and Kruskal-Wallis tests followed by post hoc analysis with Dunn’s multiple comparison test. Only surgical biopsy samples were used for the comparison between IHC result and tumor invasion. Comparisons between IHC and RNA-ISH results were assessed using Spearman correlation analysis. All data were statistically analyzed using Prism 7 software (GraphPad). Values with P < .05 were considered significant.
PCR and Sequencing for EGFR, KRAS, and BRAF
To investigate the mutation hot spots of EGFR, KRAS, and BRAF known in human (exons 18 to 21 in EGFR; codons 12, 13, and 61 in KRAS; codon 600 in BRAF), we compared the amino acid residues in corresponding loci between dogs and humans. EGFR, KRAS, and BRAF exhibited a homology of 99.47%, 100%, and 100%, respectively, between dogs and humans, indicating a high conservation rate. The mutation hot spots in human genes were identified in the following loci in dogs: exons 17 to 20 of EGFR, codons 12, 13, and 61 of KRAS, and codon 450 of BRAF. In particular, the nucleotide sequence of the mutation hot spot region was 100% identical between dogs and humans (Fig. 2). Polymerase chain reaction (PCR) primer pairs were designed for each mutation hot spot region in dogs using Primer3plus (Table 1). 32
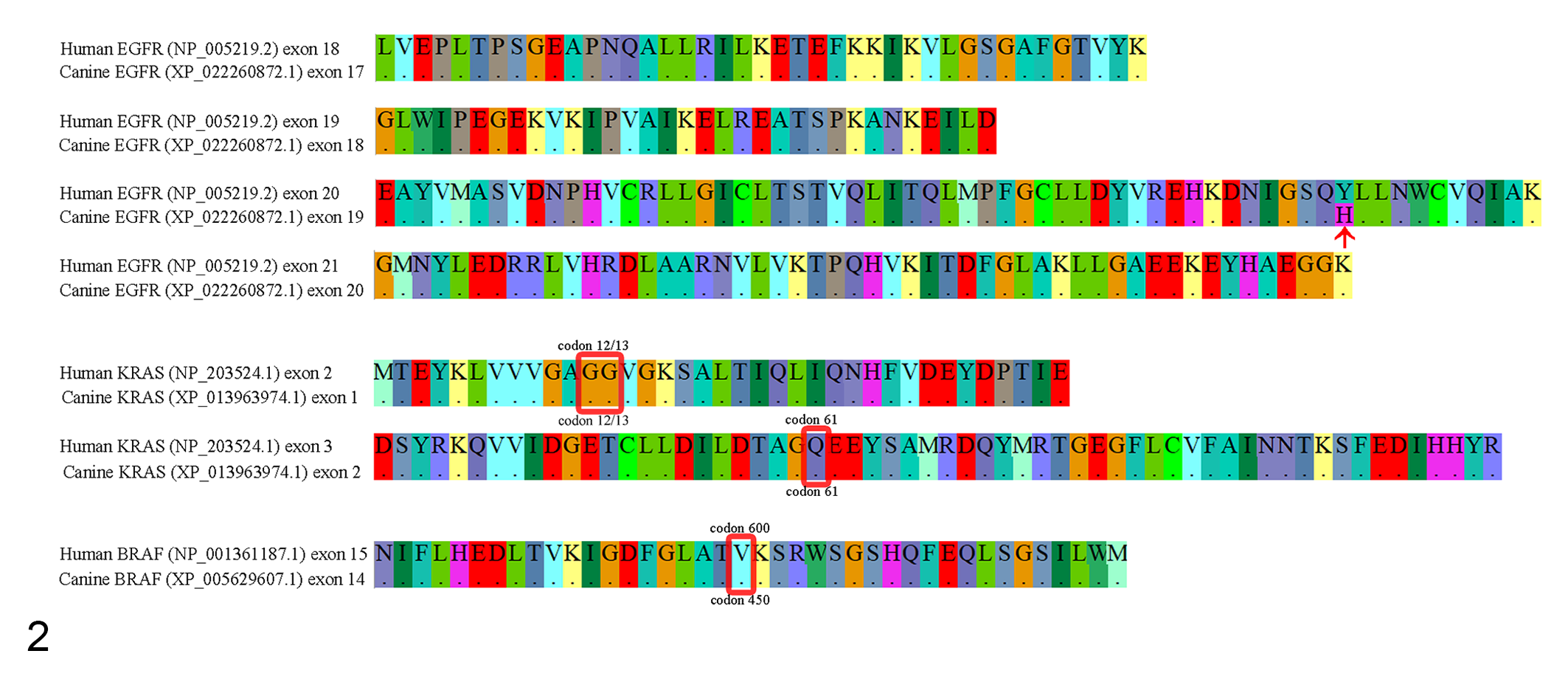
Multiple sequence alignments of human and canine EGFR, KRAS, and BRAF proteins. Sequences were highly conserved across the hot spots between human and dog homologs of these proteins, except for one amino acid in exon 19 of canine EGFR (red arrow). Mutation hot spots in human and dog KRAS and BRAF (codons 12, 13, and 61 in KRAS, codon 450 in BRAF; red squares) exhibit 100% homology.
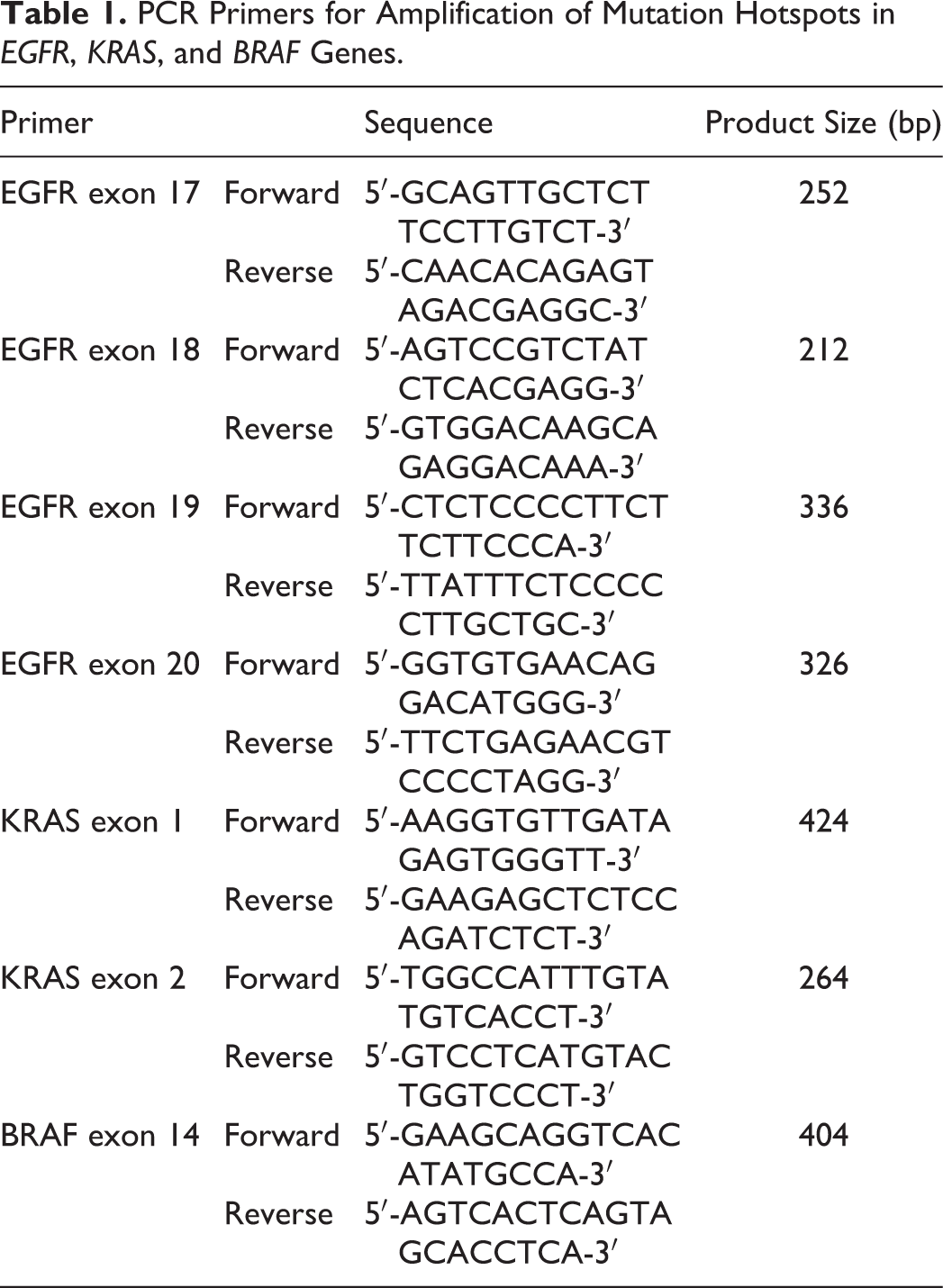
PCR Primers for Amplification of Mutation Hotspots in EGFR, KRAS, and BRAF Genes.

Since sequencing cannot be performed in FFPEs that had been stored for a long period, 19 we extracted DNA from 13 tissues in adenocarcinoma group that had been stored for up to 5 years. DNA was extracted, only from tumor areas detected via HE staining using QIAamp DNA FFPE tissue kit (Qiagen).
Conventional PCR was performed in a total volume of 40 μl; 20 µl of 2× Taq PCR master mix (Qiagen), 11 µl of RNase-free distilled water, 2 µl of each primer (10 pmol, Table 1), and 5 µl of genomic DNA. The following cycling conditions were used: initial denaturation at 94 °C for 3 minutes, 35 cycles of 94 °C for 1 minute, at 58 °C for 30 seconds, 72 °C for 1 minute, and final elongation at 72 °C for 7 minutes.
Electrophoresis was performed in a 1.5% agarose gel at 60 V for 70 minutes using a PowerPac Basic Power Supply (Bio-Rad). DNA was extracted from the gel using QIAquick Gel Extraction Kits (Qiagen). Then, Sanger sequencing was performed by Macrogen, Inc. This sequence was then compared with the reference sequence using BioEdit 7.2 software (Ibis Biosciences). The following reference sequences were used: EGFR (GenBank, NC_006600.3; location, 5983669-6044092), KRAS (GenBank, NC_006609.3; location, 22261753-22296704), and BRAF (GenBank, NC_006598.3; location, 8222909-8318179).
Results
Characteristics of Intestinal Tissues
We analyzed 47 canine intestinal biopsies, including endoscopic and surgical biopsy samples. There were 27 (54%) cases of adenocarcinoma (papillary adenocarcinoma, n = 15; mucinous adenocarcinoma, n = 6; acinar adenocarcinoma, n = 3; signet ring cell carcinoma, n = 2; adenosquamous carcinoma, n = 1); 6 (13%) of intramucosal neoplasia (adenoma and in situ carcinoma); and 14 (30%) of nonneoplastic canine intestinal tissues. Supplemental Table S1 summarizes the details and tumor characteristics of the dogs.
Immunohistochemistry
Membrane expression of EGFR was confirmed by IHC, and EGFR expression was highest in adenocarcinomas, followed by intramucosal neoplasia (adenoma and in situ carcinoma), and nonneoplastic canine intestinal tissue (P = .000, Kruskal-Wallis tests; Fig. 3, Table 2). A significant difference in EGFR expression was not observed between intramucosal neoplasia and nonneoplastic canine intestinal tissue (P = .776) or between adenocarcinoma and intramucosal neoplasia (P = .228; Dunn’s multiple comparison test). In the adenocarcinoma group, there was no statistical correlation between EGFR expression (IHC) and the sex or age of the dog, or the location, subtype, or invasiveness of the adenocarcinoma (P > .05, Table 3; Supplemental Table S1).
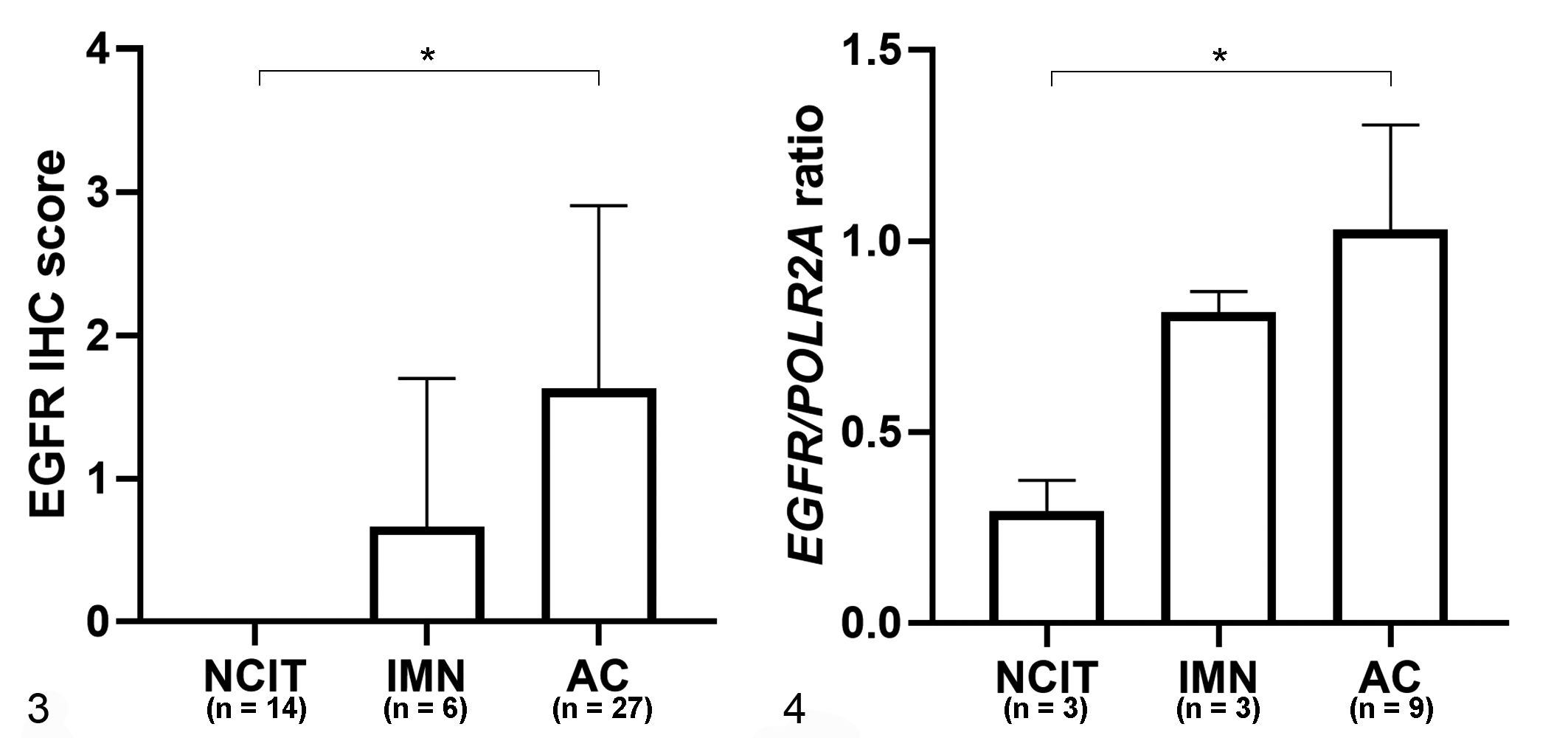
Immunohistochemistry (IHC, n = 47; Fig.3) and RNA in situ hybridization (RNA-ISH, n = 15; Fig. 4) scores for epidermal growth factor receptor (EGFR) in nonneoplastic canine intestinal tissue (NCIT), intramucosal neoplasia (IMN adenoma and carcinoma in situ), and adenocarcinoma (AC). The EGFR IHC scores (Fig. 3) and the EGFR/POLR2A ratios based on RNA-ISH (Fig. 4) were significantly higher in AC compared to NCIT (P < .05, Kruskal-Wallis test).
Expression of EGFR in Samples of Intestinal Adenocarcinoma, Intramucosal Neoplasia (Adenoma and Carcinoma In Situ), and Nonneoplastic Intestinal Tissue from Dogs. EGFR protein and gene expression were determined by scoring histologic sections labelled by immunohistochemistry (EGFR IHC scores) and in situ hybridization (EGFR/POLR2A ratios), respectively.
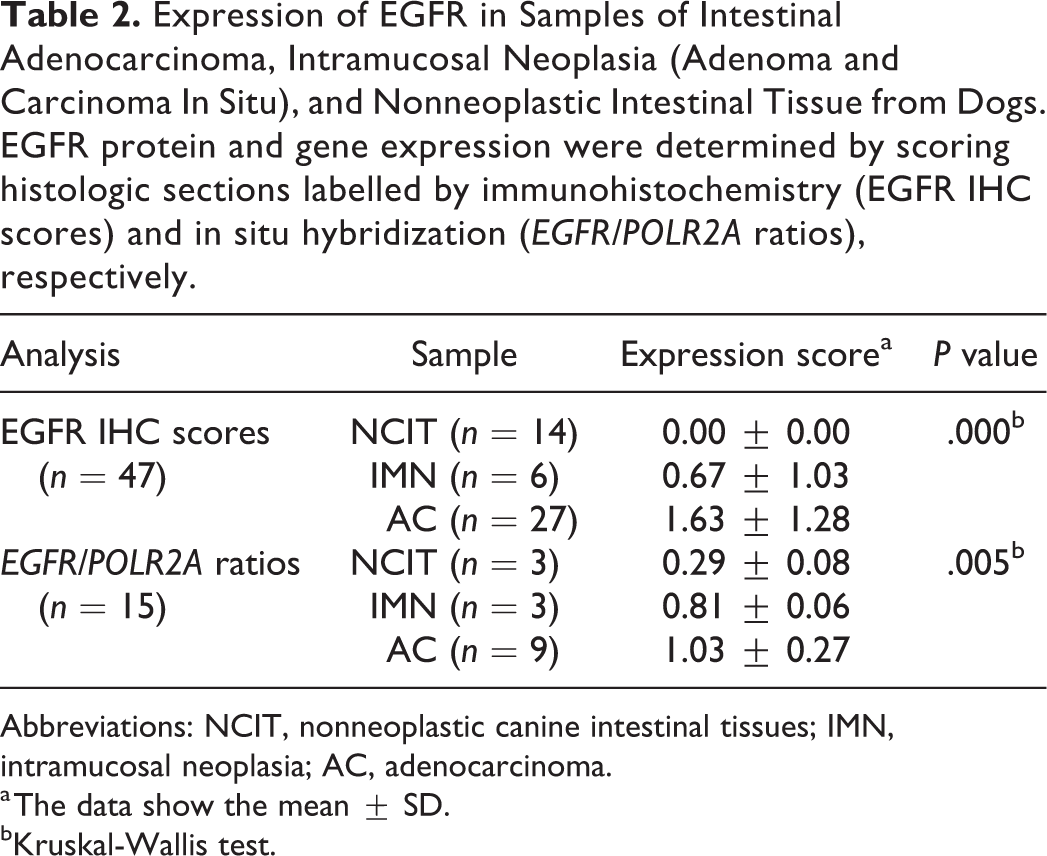
Abbreviations: NCIT, nonneoplastic canine intestinal tissues; IMN, intramucosal neoplasia; AC, adenocarcinoma.
a The data show the mean ± SD.
bKruskal-Wallis test.
Association Between EGFR Protein Expression Based on Immunohistochemistry (EGFR IHC Score) and Clinicopathological Parameters in Canine Intestinal Adenocarcinoma.
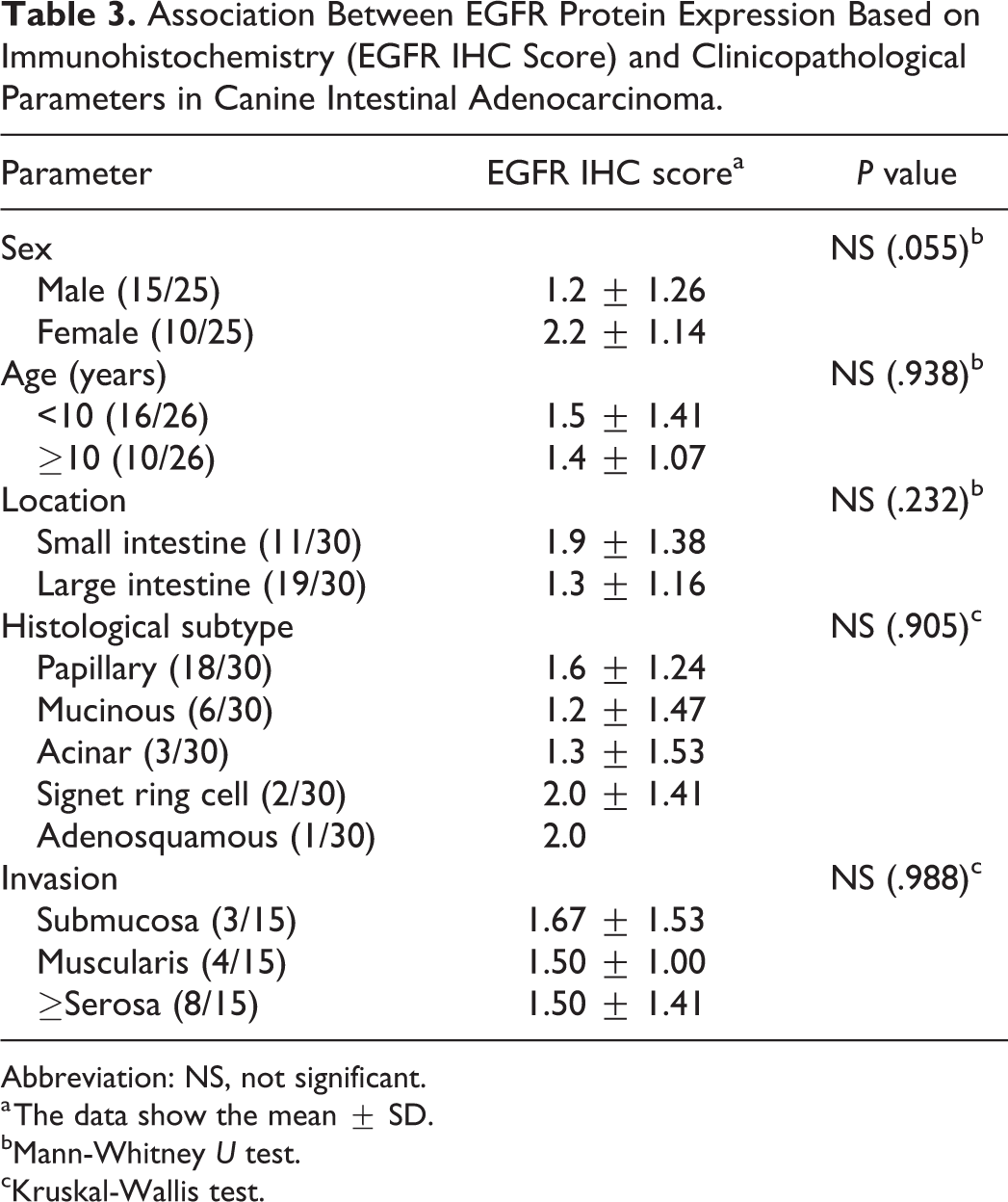
Abbreviation: NS, not significant.
a The data show the mean ± SD.
bMann-Whitney U test.
cKruskal-Wallis test.
RNA In Situ Hybridization
The EGFR/POLR2A ratio as determined by RNA-ISH (Supplemental Table S2) was highest in adenocarcinoma group, followed by intramucosal neoplasia group, and nonneoplastic canine intestinal tissue group (P = .005, Fig. 4, Table 2). The expression of mRNA (based on RNA-ISH) tended to be high in regions with high EGFR expression based on IHC (Figs. 5 –8). A linear relationship between CDX-2 protein and mRNA levels was observed (P = .000, Spearman correlation analysis; Fig. 9).
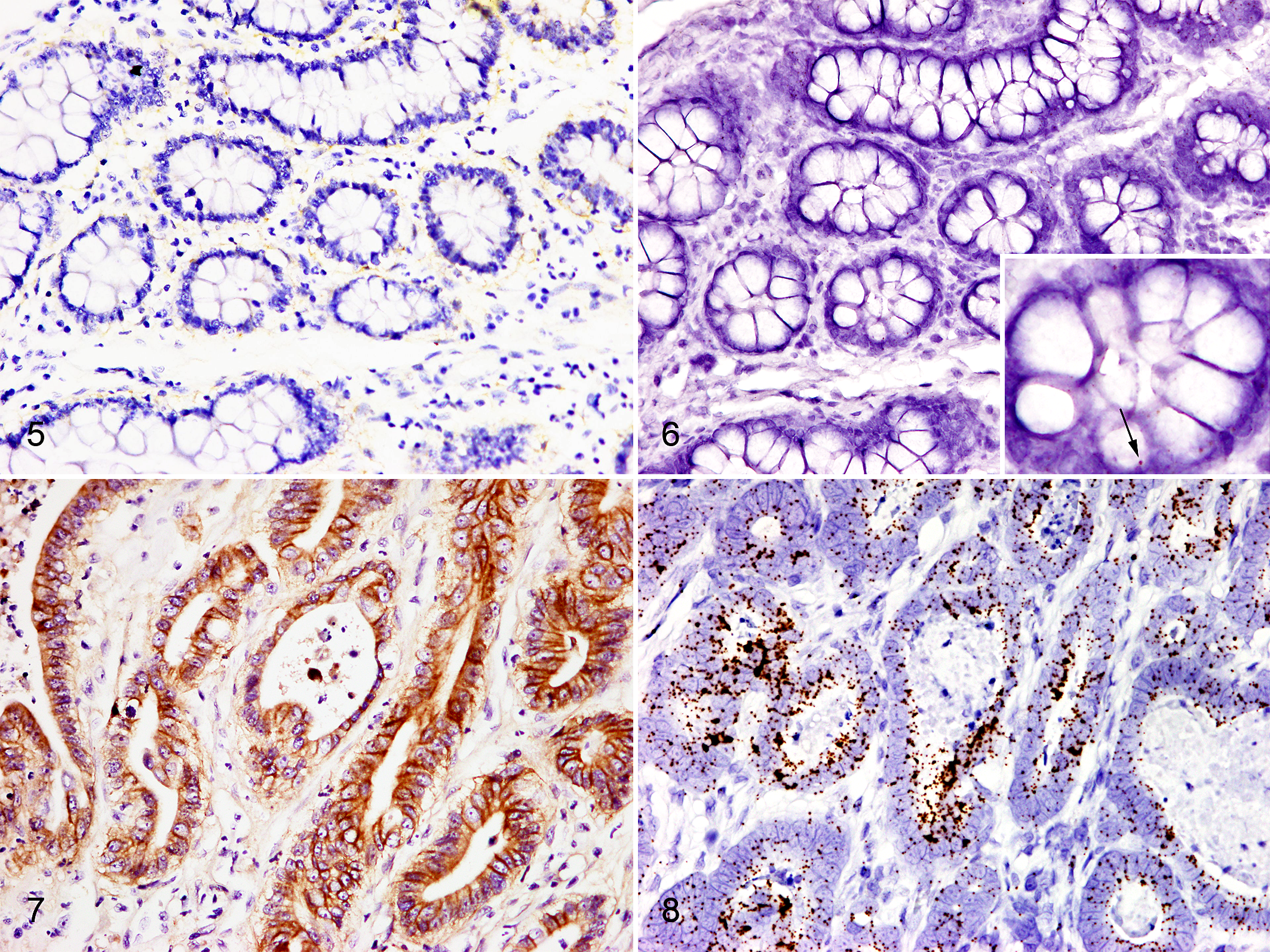
Intestine, dog.
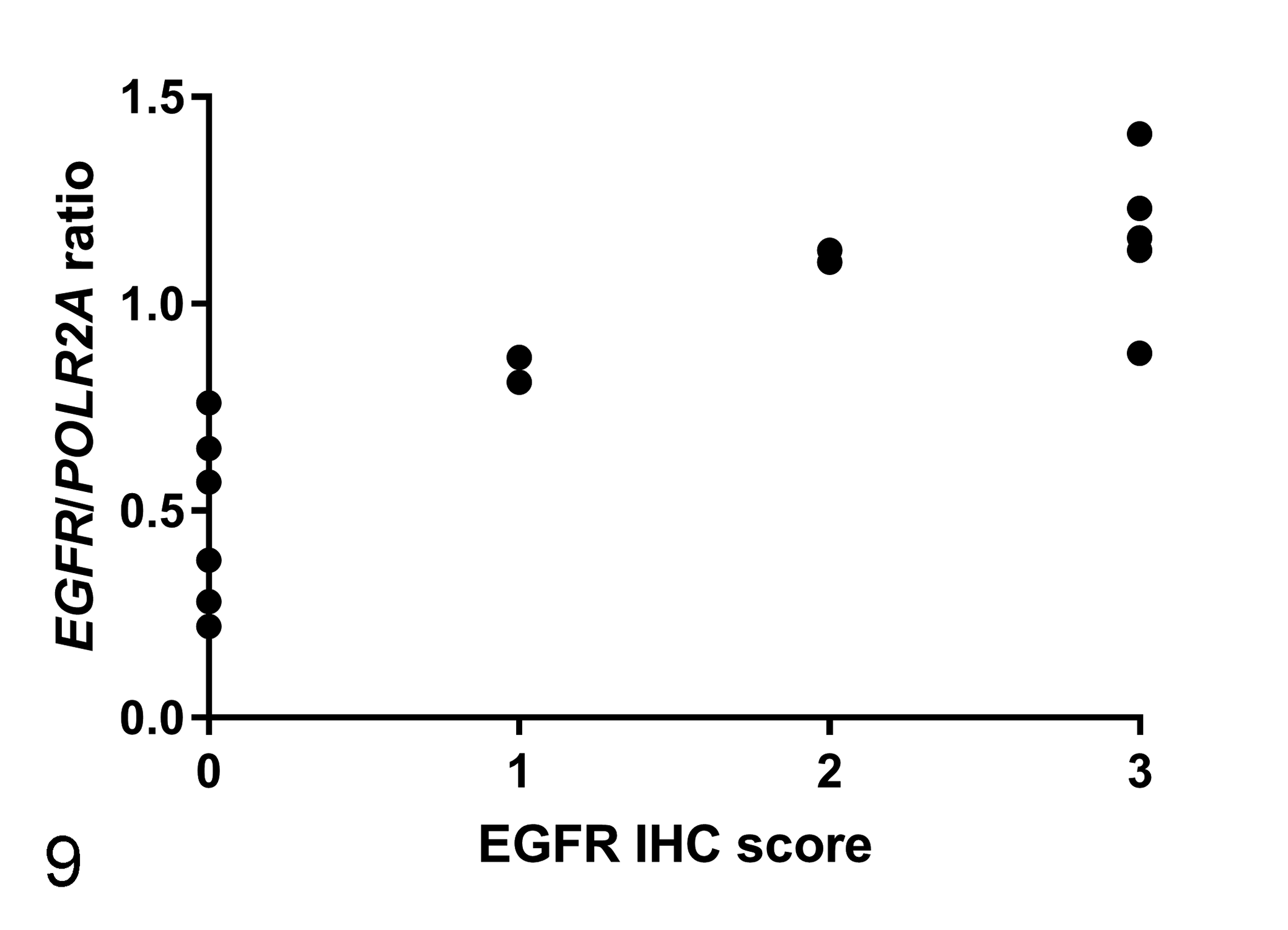
Correlation between Epidermal growth factor receptor (EGFR)/POLR2A ratios obtained using RNA in situ hybridization (RNA-ISH) and EGFR scores obtained from immunohistochemistry in canine intestinal tissues. Spearman correlation analysis identified a correlation between EGFR/POLR2A ratio and the levels of EGFR protein. P < .05.
DNA Sequencing of EGFR, KRAS, and BRAF
No functional mutations that lead to polymorphism of the amino acid sequence were identified in any of the mutation hot spots. However, single nucleotide polymorphisms (SNPs) were identified at 4 sites (Supplemental Figures S1–S4). 30 A synonymous SNP at Chr18.6015600 (A/T, A669A) and an intron SNPs at Chr18.6016238 (C/T) and Chr18.6022295 (G/A) were identified in EGFR from cases 3 and 8. An intron variant at Chr27.22277809 (KRAS; G/A) was identified in cases 2 and 6 (Supplemental Table S3).
Discussion
Many human cancers, including colorectal carcinoma, have been treated using anti-EGFR agents. 13,26 Here, we examined the feasibility of developing EGFR inhibitors for canine intestinal adenocarcinoma. Canine intestinal adenocarcinomas overexpressed EGFR in both small and large intestine, as in human, and no significant difference was observed between EGFR expression of adenocarcinomas in the 2 locations. This result is consistent with EGFR overexpression in cancers of both small and large intestine in humans. 20,26
We investigated the correlation between EGFR mRNA and protein levels in FFPE tissues. When the number of EGFR and POLR2A mRNA signals was counted in the region identified by IHC as expressing high levels of EGFR, a high EGFR/POLR2A ratio was confirmed. This suggests that there is a linear correlation between EGFR protein and EGFR mRNA levels.
The canine intestinal adenocarcinoma samples analyzed herein did not harbor any EGFR, KRAS, or BRAF mutations that would be expected to interfere with the effects of anti-EGFR agents. These findings are in line with the results of a study on cetuximab therapy in humans. 13 The present results support the strategy that anti-EGFR agents could be developed for treating canine intestinal adenocarcinoma, like strategies that have been studied in other canine tumors. 24,27 –29
We found the following SNPs: A/T at Chr18.6015600, C/T at Chr18.6016238, G/A at Chr18.6022295, and G/A at Chr27.22277809, which reportedly occur in dogs in ratios of 0.27, 0.25, 0.04, and 0.17, respectively. 30 Specifically, the 3 spontaneous SNPs identified in EGFR were concurrent in each of the 2 dogs.
The present study has some limitations. Although there was an overlap between KRAS and BRAF mutations in canine and human tumors—in terms of sites of mutation 15 —we cannot rule out the possibility that KRAS or BRAF mutations might occur outside the mutation hot spots or elsewhere, such as codon 146 in KRAS. 14 Furthermore, the number of samples used in present study was too small to derive meaningful conclusions. In addition, we analyzed some endoscopic samples, and some of them which had a limitation that the evaluation of invasion may be unclear because of the small size of the tissue sample. Therefore, further studies that include large surgical biopsy samples are needed.
In conclusion, our results indicate expression of EGFR in canine intestinal adenocarcimoma. This finding supports the development of anti-EGFR agents as a promising therapeutic strategy to control progression of intestinal adenocarcinoma in dogs.
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211009778 - EGFR Overexpression and Sequence Analysis of KRAS, BRAF, and EGFR Mutation Hot Spots in Canine Intestinal Adenocarcinoma
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211009778 for EGFR Overexpression and Sequence Analysis of KRAS, BRAF, and EGFR Mutation Hot Spots in Canine Intestinal Adenocarcinoma by Seung-Hee Cho, Byung-Joon Seung, Soo-Hyeon Kim, Min-Kyung Bae, Ha-Young Lim and Jung-Hyang Sur in Veterinary Pathology
Footnotes
Acknowledgements
We thank Ms E.-M. Yu for excellent technical assistance. This report represents a part of a PhD thesis of Seung-Hee Cho.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Bio & Medical Technology Development Program of NRF funded by the Korean government (2016M3A9B6903437) and by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2017R1D1A1B03032057).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
