Abstract
Siniperca chuatsi rhabdovirus (SCRV) is one of myriad rhabdoviruses recorded in fish. Preliminary data show that inhibition of the SCRV nucleoprotein (N) could significantly reduce the progeny virus titers in infected Epithelioma papulosum cyprinid (EPC) cells. Here, the authors propose that cleavage of the viral 47-kDa N protein is caspase-mediated based on caspase inhibition experiments, transient expression in EPC transfection, and analysis of cleavage sites. Cleavage of the SCRV N protein in culture was prevented by a pan-caspase inhibitor, z-VAD-FMK (z-Val-Ala-DL-Asp-fluoromethyl ketone). Subsequently, N was transiently expressed in EPC cells, the results of which indicated that the specific cleavage of N also occurred in the cells transfected with N-GFP plasmid. Several truncated fragments of the N gene were constructed and transiently transfected into EPC cells. Immunoblotting results indicated that D324 and D374 are the cleavage sites of N by caspases. The authors also found that z-VAD-FMK could inhibit the cytopathic effect in SCRV-infected EPC cells but not affect the production of infectious progeny, suggesting that the caspase-mediated cleavage of N protein is not required for in vitro SCRV replication. To the authors’ knowledge, this is the first report on the cleavage of rhabdovirus proteins.
Siniperca chuatsi rhabdovirus (SCRV) is one of the piscine rhabdoviruses that has been associated with outbreaks of viral disease in aquaculture. 8 Like other rhabdoviruses, the RNA genome of SCRV encodes 5 structural proteins: nucleoprotein N, phosphoprotein P, matrix protein M, glycoprotein G, and RNA-dependent RNA polymerase protein L. 8 SCRV N protein (GenBank No. DQ399789) is 427 amino acids long and is a putative 47.4-kDa protein. 8 Expression of nucleoprotein is essential for the packaging of viral RNA into viruslike particles, and N interacts with the phosphoprotein to fix the nucleoprotein for viral transcription and replication. 5
To survive and replicate, viruses must circumvent host cellular responses that have evolved to limit virus replication; apoptosis is one such cellular response that limits both the time and the cellular machinery available for virus replication. 3 However, as is often the case in biological systems, the relationship between virus replication and apoptosis is quite complex. One mechanism of immune evasion is virus-encoded inhibitors of caspases to escape apoptosis and thereby prolong the viral replication cycle. 2 However, several viruses adopt a seemingly counterintuitive strategy and utilize host-produced caspases to cleave their own proteins. Examples of this pathway include the caspase cleavage of nucleocapsid proteins of influenza A viruses, Aleutian mink disease virus, and feline calicivirus. 1 Caspase-mediated cleavage of viral proteins has been reported for herpesvirus, adenovirus, and orthomyxovirus. This cleavage may be beneficial to virus replication in infected cells or regulate outcome (lytic or persistent infection). 1,6,7 Recently, a mutation within the caspase cleavage sites of the influenza M2 protein could lead to the decline of viral pathogenicity. 9 Therefore, studies on protein cleavage may contribute to clarifying the replication and pathogenetic mechanism of different viruses.
Here we demonstrate that SCRV N serves as a substrate for caspase-mediated cleavage upon viral infection or transient transfection into fish cell lines. z-VAD-FMK (z-Val-Ala-DL-Asp-fluoromethyl ketone), a pan-caspase inhibitor, can apparently inhibit cleavage of N protein. Based on these findings, the cleavage sites in N were investigated. To our knowledge, this is the first report on the specific cleavage of rhabdovirus N protein.
SCRV was isolated from diseased mandarin fish (S. chuatsi Basilewsky) by several cycles of cell plaque identification in grass carp fin cell line. 8 The infection spectrum analysis demonstrated that SCRV could also infect the Epithelioma papulosum cyprinid (EPC) and Chinook salmon embryo cell lines in vitro. Prior to being challenged with SCRV (300 TCID50), EPC and grass carp fin cells were grown at 25°C in TC199 medium supplemented with 10% fetal bovine serum (Gibco, Invitrogen), while the Chinook salmon embryo cell line was incubated at 15°C with the same medium. After 70% of the virus-infected cells showed cytopathic effect, the cultures were collected and lysed for Western blotting analysis according to previous methods. 8 The primary and secondary antibodies are the anti-N protein antibody (dilution of 1-500; mouse antiserum against prokaryotic recombinant N protein) 8 and the alkaline phosphatase-conjugated goat anti-rabbit IgG antibody (dilution of 1-1000, Vector Laboratories, Burlingame, CA). The results showed the anticipated full-length 47-kDa N protein band. However, 2 slightly faster-migrating bands (Fig. 1a) were also detected in the identical PVDF membrane in the different infected fish cells. The 2 protein bands were approximately 41 and 35 kDa, demonstrating that the cleavage phenomenon of N existed, though the extent of cleavage was not identical in different cell lines.
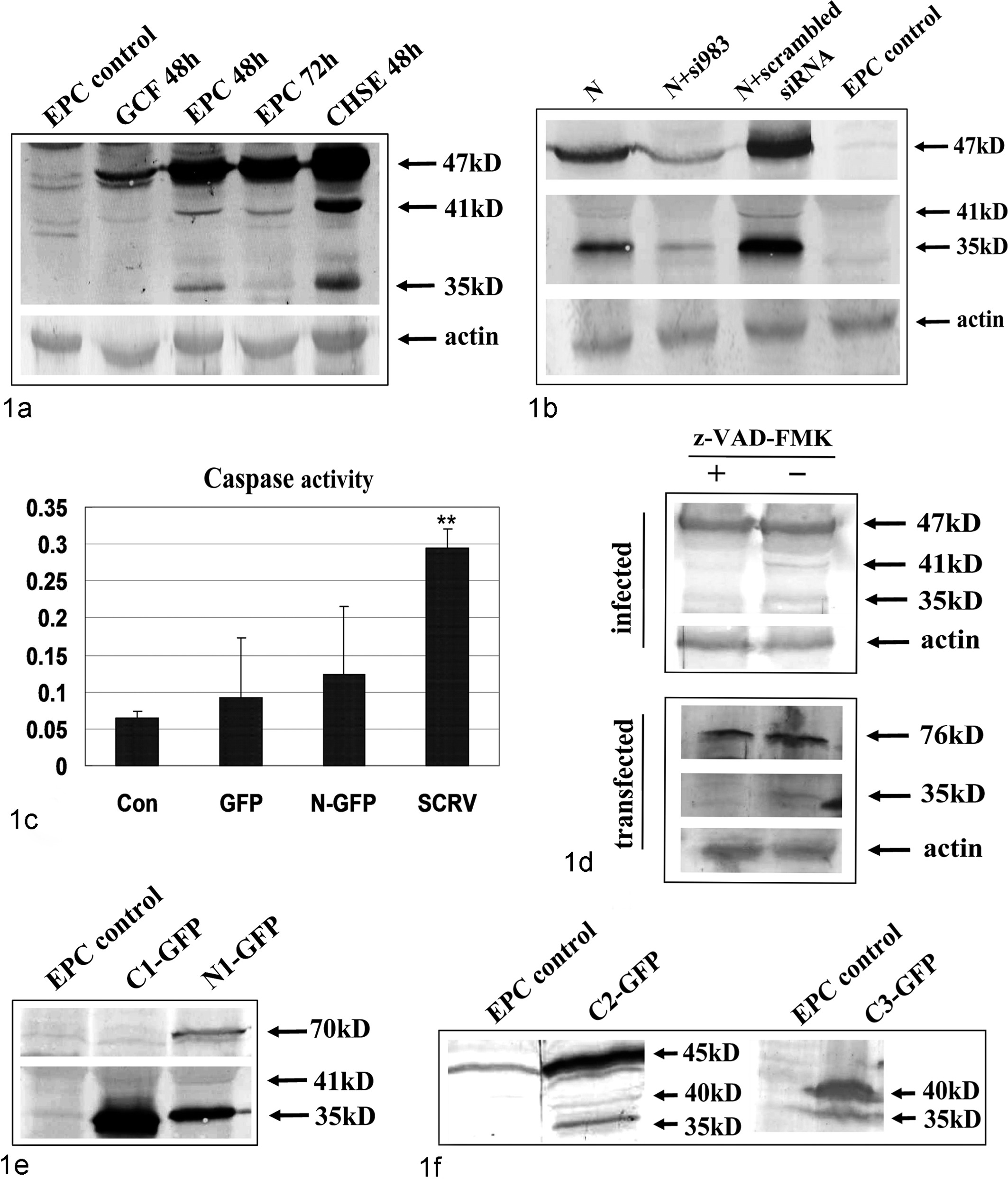
Identification of caspase cleavage of N protein. Lysates of Siniperca chuatsi rhabdovirus (SCRV)–infected cells (Epithelioma papulosum cyprinid [EPC], Chinook salmon embryo [CHSE], and grass carp fin [GCF] cell lines) were subjected to immunoblotting analysis using the anti-N antibody (a). SCRV N-GFP (b) and other truncated N mutant expression vector (e, f) were transiently transfected into EPC cells. Lysates were analyzed by immunoblot as described above. Caspase 3 activity was also detected in N-GFP-transfected and SCRV-infected EPC cells (c). SCRV-infected (d, above) or N-GFP-transfected (d, below) EPC cells were assessed by immunoblot after treating with z-VAD-FMK (z-Val-Ala-DL-Asp-fluoromethyl ketone; 100µM).
To analyze whether this cleavage of N in fish cell cultures occurred only during viral infection, the expression pattern of N protein was assessed in EPC cells transfected with N-GFP plasmid. First, several fusion plasmids—N-GFP (amino acids [aa] 1-429), N1-GFP (aa 1-376), C1-GFP (aa 377-429), N2-GFP (aa 1-281), C2-GFP (aa 282-429), and C3-GFP (aa 325-429)—were generated by fusing different polymerase chain reaction fragments of SCRV N gene with in-frame restriction sites into pEGFP-N3 vector (Clontech) using standard cloning procedures with XhoI and KpnI. The relative sense and antisense primers are shown in Supplemental Table 1. The EGFP gene was located downstream of the target genes. Figure 2 shows the schematic diagram. The SCRV N-GFP plasmid was used for transfection analysis using Lipofectamine 2000 (Invitrogen) according to the previous instructions. 10 Transfected cells were examined under bright-field illumination and fluorescence microscopy (Leica Microsystems, Wetzlar, Germany) after 24 hours. Immunoblotting analysis showed a similar cell-specific processing of N protein (Fig. 1b), and the additional 2 bands of N were still detectable. Furthermore, 1 siRNA molecule (si983) 10 specifically targeted to N gene apparently inhibited N protein expression in cotransfected EPC cells, which then led to a decrease of 2 small bands’ amounts of expression (Fig. 1b). Controls were performed using scrambled siRNA and nontransfected cells.
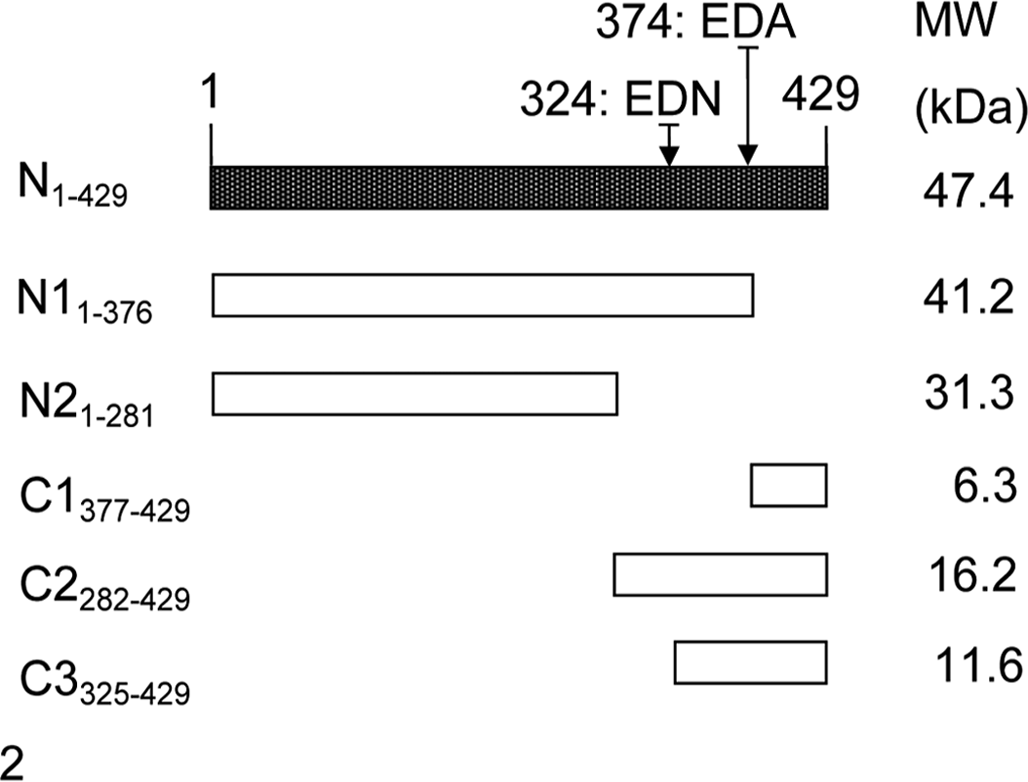
Schematic diagram showing the constructed Siniperca chuatsi rhabdovirus (SCRV) N-GFP expression vector and its deletion mutants and their predicted molecular masses (MW). The 2 caspase cleavage sites (EDA, EDN) of nucleoprotein are also shown.
Full-length N protein and 2 additional bands were detected in virus-infected and N-transfected EPC cells. We wanted to characterize the caspase-mediated cleavage of SCRV N. Fish rhabdovirus induces cellular apoptosis. 2 In this case, the activation of caspase 3, a key mediator of apoptosis, was first assayed in SCRV-infected or N-transfected EPC cells. Mock-infected or SCRV-infected (300 TCID50) EPC cells (48 hours postinfection) were harvested by scraping, washed once with phosphate buffered saline, frozen for 30 minutes, and centrifuged (16 000 g for 15 minutes). The supernatants of these cell lysates were assayed for caspase 3 activity in the presence of 20µM Ac-DEVD-pNA (Beyotime, China) using a plate reader (Model 550 Microplate Reader; Bio-Rad, Hercules, CA) at 405 nm. Caspase 3 activity was also measured in N-GFP-transfected EPC cells (48 hours posttransfection). These experiments were conducted 3 times each. The results showed that the values of OD405 with SCRV-infected or N-transfected cell supernatants increased between 5 and 2 times the OD405 value of control cell supernatant, respectively (Fig. 1c). This revealed that caspase pathways could be activated in virus-infected or N-transfected cells and that caspases may be involved in the process of N cleavage.
Alternatively, for further investigation of the role of caspases in N protein cleavage, the infected or N-transfected EPC cells were treated with pan-caspase inhibitor z-VAD-FMK (Beyotime; 100µM), which prevents activation of caspase 1- and caspase 3–related caspases. Infections (300 TCID50) were carried out in the absence or presence of 100µM z-VAD-FMK. Lysates were subjected to SDS-PAGE and immunoblotting for SCRV N protein as described above. Similarly, the transfected cell lysates with N-GFP expression vector were also evaluated in the same manner. The results showed that only full-length N was detected in the transfected cells upon pan-caspase inhibitor treatment, while the 2 additional bands were much fainter when compared to the untreated control in the infected cells (Fig. 1d). This indicated that the lower bands may be the result of caspase-mediated cleavage.
Having demonstrated that the N protein cleavage was caspase mediated, the next step was to identify the specific cleavage sites in N proteins. It is well known that caspases are aspartate-specific cysteine proteases, with the cleavage occurring after aspartate residues. The N1-GFP (1-376 aa, 41.2 kDa) and C1-GFP expression plasmids were constructed in accordance to the putative amino acid sequence of N protein (1-429 aa) and the position of the Asp (D) residues (Fig. 2). 8 The 2 constructs were each transfected transiently into EPC cells, and the cell extracts were used for immunoblotting analysis. The additional 41- and 35-kDa fragments were both detected in the N1-GFP transfected cells, while one 35-kDa protein band was found in the C1-GFP transfected cells (Fig. 1e). In view of Asp374 in N protein sequence, 1 of the N cleaved sites should be located at D374 from the N-terminus of N protein according to the above results. To identify the next cleavage site, N2-GFP, C2-GFP and C3-GFP (325-429 aa, 11.6 kDa) plasmids were constructed and used in experiments similar to those described. However, no protein band was detected in N2-GFP transfected cells, which may account for an absence of antigenic epitopes recognized by the anti-N protein antiserum employed. Two additional bands (40 and 35 kDa) to the full-length 45-kDa signal of C2-GFP (Fig. 1f) were detected in transfected cells, which suggested that the other cleavage site should be located on D324. Furthermore, one 35-kDa band was detected in C3-GFP-transfected cells in addition to the expected 40-kDa protein (Fig. 1f). The results further confirmed the D374 cleavage site of N protein and another, D324.
Finally, when SCRV-infected cells were treated with the broad-spectrum caspase inhibitor (z-VAD-FMK) for 48 hours, the level of cytopathic effect was markedly reduced in comparison with untreated cells (Fig. 3). The output of infectious SCRV progeny was also determined. However, treatment of cells with z-VAD-FMK did not alter the amplification of SCRV compared to control cultures, with titers between 106.4 (with inhibitor) and 107.4 TCID50/ml (without inhibitor) for cell supernatant and 107.5 (with inhibitor) and 107.8 TCID50/ml (without inhibitor) for cell lysates, respectively (Fig. 4), which indicated that caspase activation is not required for SCRV replication in EPC cells.
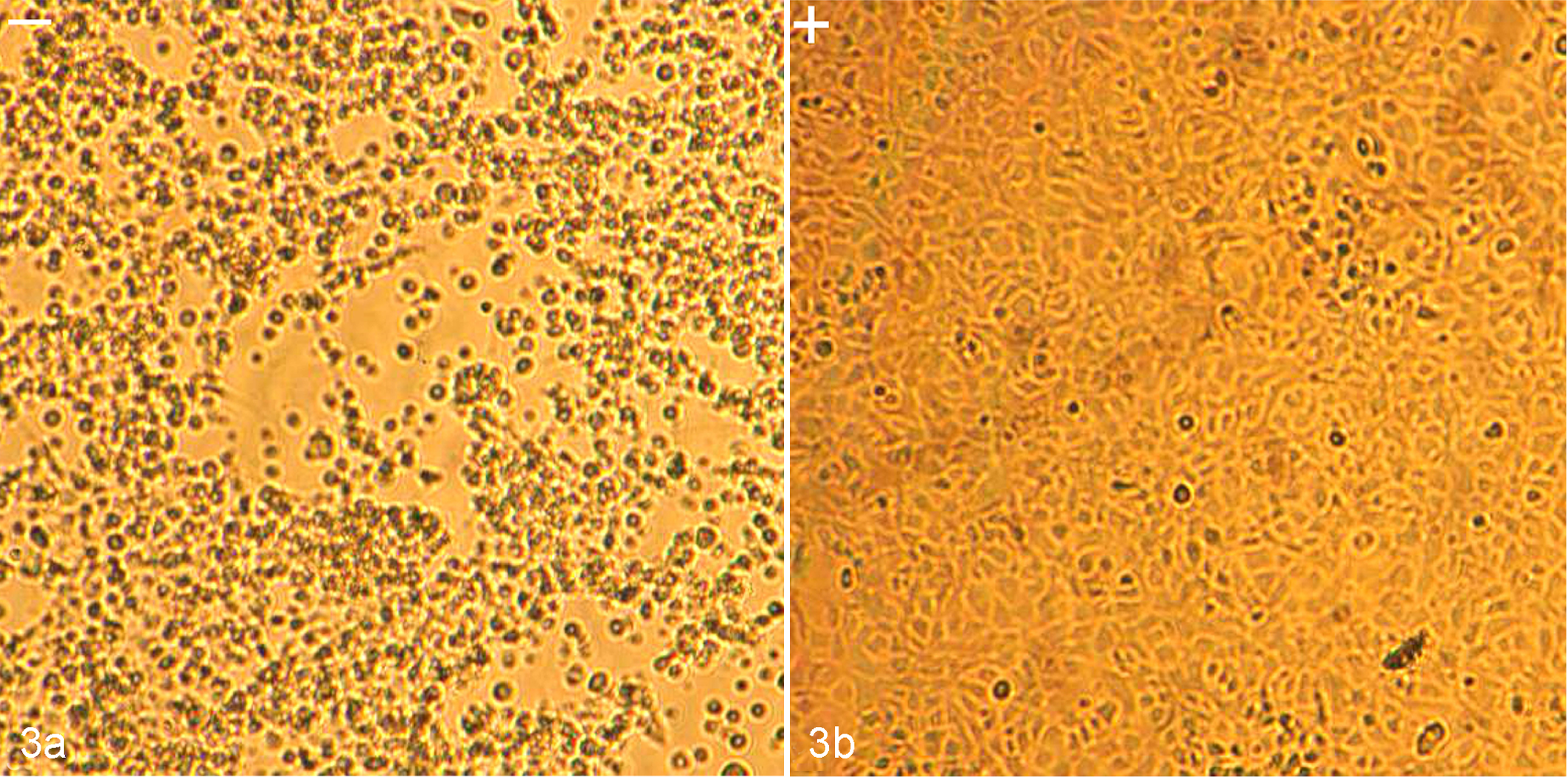
Characterization of alterations in cell culture morphology of Siniperca chuatsi rhabdovirus (SCRV)–infected Epithelioma papulosum cyprinid (EPC) cells treated with z-VAD-FMK (z-Val-Ala-DL-Asp-fluoromethyl ketone). (a) At 48 hours postinfection, the SCRV-infected cells were shrunken and detached (left, –); (b) SCRV-infected EPC cells treated with z-VAD-FMK showed little change (right, +).
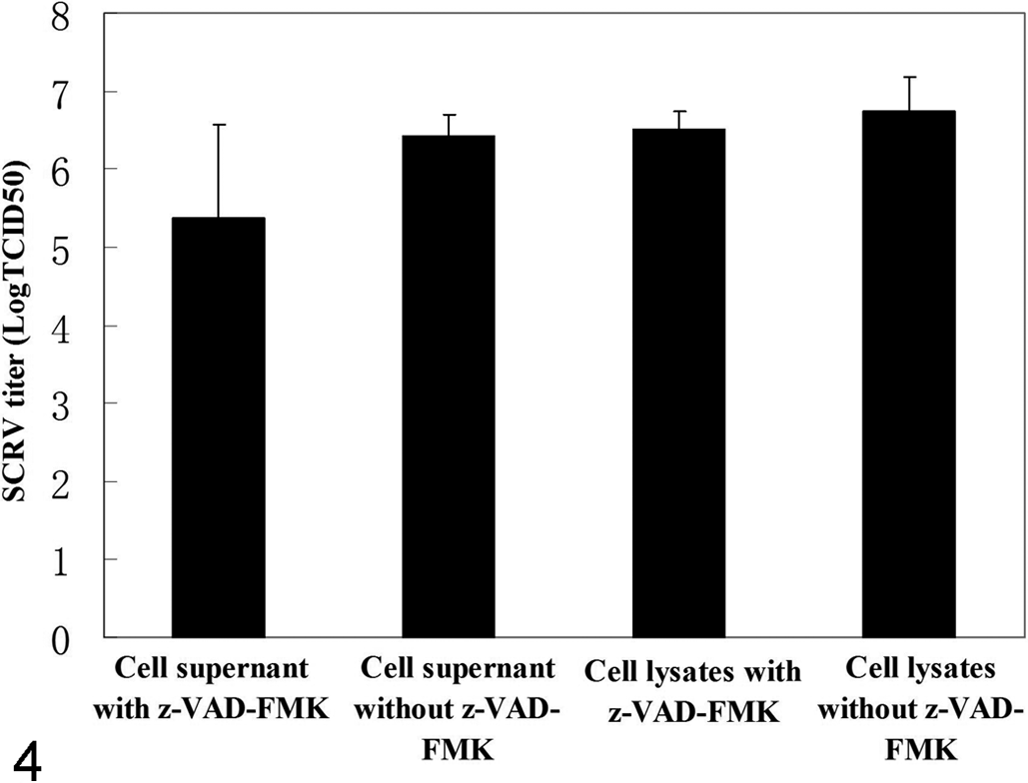
Identification of virus titers of Siniperca chuatsi rhabdovirus (SCRV)–infected Epithelioma papulosum cyprinid cells treated with z-VAD-FMK (z-Val-Ala-DL-Asp-fluoromethyl ketone). z-VAD-FMK does not influence the virus titers in SCRV-infected E. papulosum cyprinid cells after 48 hours postinfection. The differences are not statistically significant (P > .05).
The rhabdovirus N protein forms a viral nucleocapsid by interacting with its viral RNA. The resultant nucleoprotein complex associates with the RNA-dependent RNA polymerase subunits to initiate intracellular virus replication. 5 As such, the N protein of rhabdoviruses likely plays essential roles in viral RNA protection and virus replication. Previous research had suggested that N proteins of several strains of viruses were cleaved by host caspases during viral infection. 1 In this study, the specific caspase-mediated cleavage of SCRV N protein in cell cultures was observed and described. It is suggested that caspase 3 plays a key role in the induction of apoptosis in mammalian rhabdovirus VSV-infected cells, while it does not appear to play any significant role in in vitro viral replication but is nonetheless only 1 reaction of many in the apoptotic cascade. 4 When the EPC cell cultures were infected with SCRV or transfected with N-GFP expression plasmid, caspase 3 was activated. Although z-VAD-FMK could abrogate caspase-mediated N cleavage in SCRV-infected EPC cells as well as cell shrinkage and detachment, it had no significant effect on progeny SCRV virus titers. The results presented in this study are consistent with this observation.
Although we have carried out some identification of cleavage sites on SCRV N protein, more work still needs to be done. For example, which apoptotic pathway was activated—the extrinsic cascade or intrinsic pathway? Which specific caspase may be involved in viral N cleavage? What is the effect on SCRV replication in the fish cell line for nucleoprotein cleavage? More information on the role of caspases in viral replication may be helpful for favorably understanding the pathogenesis of SCRV.
Footnotes
Acknowledgement
We thank Dr Eric Lombardini (AFRIMS) for careful reading of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by the National Natural Science Foundation of China (31101931), Plan for Scientific Innovation Talent of Henan University of Technology (11CXRC13), Training Program for Youth Backbone Teacher of Henan University of Technology (to Guangzhou Zhou), and Earmarked Fund for Conventional Freshwater Fish Industries of Henan Province (S2014-10-G01).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
