Abstract
Animal models have historically informed veterinary and human pathophysiology. Next-generation genomic sequencing and molecular analyses using analytes derived from tissue require integrative approaches to determine macroanalyte integrity as well as morphology for imaging algorithms that can extend translational applications. The field of biospecimen science and biobanking will play critical roles in tissue sample collection and processing to ensure the integrity of macromolecules, aid experimental design, and provide more accurate and reproducible downstream genomic data. Herein, we employ animal experiments to combine protein expression analysis by microscopy with RNA integrity number and quantitative measures of morphologic changes of autolysis. These analyses can be used to predict the effect of preanalytic variables and provide the basis for standardized methods in tissue sample collection and processing. We also discuss the application of digital imaging with quantitative RNA and tissue-based protein measurements to show that genomic methods augment traditional in vivo imaging to support biospecimen science. To make these observations, we have established a time course experiment of murine kidney tissues that predicts conventional measures of RNA integrity by RIN analysis and provides reliable and accurate measures of biospecimen integrity and fitness, in particular for time points less than 3 hours post–tissue resection.
Keywords
A biorepository can be defined as the storage of any biological material in an organized manner that contains value for past, present, or future use in a research or clinical application. The College of American Pathologists (CAP) defines a biorepository as an entity that receives, stores, processes, and/or disseminates biospecimens, their derivatives, and relevant data, as needed. It encompasses the physical location as well as the full range of activities associated with its operation. Biorepositories are the functional units involved in the collection process and act as custodians of the biospecimens by implementing procedures and biorepository management practices. Over the past decade, research investigators and funding agencies such as the National Institute of Health have realized a significant lack of high-quality biospecimens being used, especially in National Institute of Health–funded clinical research, largely the result of nonstandardized standard operating procedures (SOPs) and the lack of knowing the effect of processing on the quality of the biospecimens during the collection and management processes. Pathologists and biobanking experts understood these issues and, as a result, formed the International Society of Biological and Environmental Repositories (ISBER) in 1999. After a decade of activity to improve the management of biospecimens, the CAP initiated its Biorepository Accreditation Program (BAP) in 2012. The CAP checklist for accreditation covers a range of activities and broadly references checklists from the ISBER best practices 3,4 and the National Cancer Institute best practices for biospecimen resources. 61,93
High-quality diseased pathogenic tissue samples derived from patients are required sources of material for molecular analysis of disease in the research setting. However, the diagnostic standard in anatomic pathology remains the formalin-fixed, paraffin-embedded (FFPE) tissue sample, which is less than ideal for molecular analyses. 17,32,42,78 Quality assurance (QA) practices for tissue collection to molecular phenotyping are critical to implement for the postgenomic era of basic and clinical research. 35,36,64 These QA practices must include participation of trained pathologists as an essential link in the chain of events to obtain reliable and reproducible research findings and, ultimately, appropriate clinical care. There is an ongoing need for evidence-based data that define the minimal requirements and best practices of tissue sampling for molecular diagnosis. Best practices can then be incorporated into certified/accredited quality control (QC) programs of sampling procedures. 3,39,62,63 The sequence of tissue collection, transport, processing, and distribution requires critical monitoring by detailed protocols and QA/QC practices associated with various biospecimen collections. The QA procedures include not only biospecimen storage but macroanalyte extraction, quantitation, and quality assessment. 17 Attention to QA procedures is critical because of the labile nature of protein 25,86 and RNA analytes. 45,77 To develop and validate the best practices for the management of tissues, we believe that evidence-based and validated data from controlled experimental model systems should be established. Controlled model systems could use cell lines and mouse models with morphologic and molecular phenotypic correlates that can then be extended to human clinical sample collections.
Biobank QA/QC Practices and Biospecimen Science
Tissue-based molecular research applications in veterinary and clinical arenas have historically relied on archival FFPE tissues, as they are associated with attached clinical outcomes data and represent cost-effective long-term storage of clinical and veterinary tissues. 29,30,58,96 However, the long-term preservation function is due to extensive cross-linking by formalin of histone and nonhistone proteins and makes characterization of effects at the macromolecular level challenging and at times nearly impossible. 17,34,83,89,91 The deleterious effects of formalin fixation are most pronounced in the most labile analytes of RNA and protein and have often limited whole genome and proteome surveys in FFPE tissues. Additional preanalytic variables inherent to retrospective studies, along with the lack of protocols in biospecimen collection, transport, and processing, historically have led to widely variant nucleic acid analyses and disparate molecular signatures, which have become a critical element in reviewing genomic studies. The historical lack of real standards in tissue fixation and poor documentation in the FFPE repositories in veterinary and human clinical samples further challenge the utilization of this resource and the reporting of molecular findings. 20,84 Thus, leading organizations at the national level 49,61,63 and the international level 5,47,62 address such preanalytic, analytic, and postanalytic variables in systematic formulations of best practices to address the complete spectrum of biospecimen collection, processing, and distribution, including data elements generated from that biospecimen. The fields of biospecimen science and biobanking practices provide protocols and standard practices to maximize the validity of studies whether applied to veterinary or human disease processes. Standard QA/QC practices 9,98 are of great importance to ensure the highest quality of research and, eventually, clinical care. While QA/QC practices are more strictly utilized in human pathology, we argue that to provide excellent research data, the QA/QC practices in human and animal tissue management are critical, and often the methodologies of best practices can or should be tested in an animal model and utilized in veterinary-based sample collection. 5,95 At the international level, the ISBER provides best practices and harmonization that connect repositories and varied biological collections. The National Cancer Institute (http://biospecimens.cancer.gov/practices/) has also developed a set of best practices, and, as noted, the CAP has embraced the importance of QA needs in biospecimen management through the implementation of the BAP, which is patterned from the accreditation of clinical laboratories and requires organized and validated procedures and QA/QC protocols to document the steps associated with maintaining tissue sample integrity and characterization of prospective tissue collections. 35,68 The need of a BAP was emphasized by the CAP’s experience that a “specimen” can mean different things to different disciplines and that those objectively measureable standards of FFPE samples can vary from institution to institution and even within institutions. 26 The concept of a quality specimen is now essential, and the certification of biorepositories requires that organizations define the adequacy of the specimen used in research and clinical practice. In our laboratory, we have implemented detailed SOPs for every aspect of tissue collection, processing, analyte extraction, and analysis (Supplementary Table 1, available at http://vet.sagepub.com/supplemental). These SOPs help to ensure specimen quality and the reliability of the data obtained from the specimens.
Anatomic Pathology: Morphology to Molecular Data
Historically, the anatomic pathology report was a qualitative description of characteristic histology, and it provided the forum for the anatomic pathologist to render a diagnosis based on representative sampling. The anatomic diagnosis, by nature, was limited by sampling and varied degrees of subjective determination of tissue histology. Morphologic correlates of tissue sample integrity were not required, as morphology was the endpoint. Moreover, the lack of objective criteria to assess molecular integrity skewed the molecular characterization of the majority of both fresh-frozen and FFPE tissues. Currently, advances in radiographic imaging and resultant smaller and smaller biopsy samples push the consideration for molecular testing, which makes processing and documentation of tissue sampling increasingly critical. The evaluation of tissue specimens to be used for clinical or research assessment needs to ensure that morphologic and molecular characterizations match. If the selection process for tissue is not managed well, inadequate amounts of sample to select a representative region for morphologic and molecular analysis may result in inaccurate information. Currently, it is common for each discipline to obtain its own sample without determining whether the molecular specimen is an appropriate representation of the histologic component on which the diagnosis and treatment are based. Notwithstanding, most genetic/genomic observers assume that probe and associated data points are correct (ie, reflect cellular biology and the tissue level) and that they do not need to examine the sample for viability (eg, amount of necrosis); percentage of normal and tumor histology; tumor heterogeneity, such as benign versus atypical versus neoplastic tumor tissue; or epithelial tissue and stromal ratios. 1,2,28 Histology also serves as a guide to genomics in the integration and correlation to the diagnostic imaging component. 9,11,19,55 If one compares a non–small cell lung carcinoma expression assay, the optimal control would be the presumed normal tissue from which it originates, preferably bronchus lined by ciliated bronchial epithelium. However, even if the pathology sample is labeled correctly in the surgical pathology lab as an adjacent control, it might include a variety of heterogeneous tissue components including, but not limited to, pulmonary artery, potential non–small cell lung carcinoma contaminant, normal alveoli, lobular septae, or lymph node. The histologic admixture of these various cell types may obscure differences in disease-related mRNA expression-profiling experiments. The inherent contribution of molecular signals from normal tissue collected and assessed differently in various laboratories may reflect the variation of differences in collection procedures that causes discordance in analytic measurements. Although the genomic research data may be considered reportable, the fact remains that if histologic verification of the original tissue material was not performed, association back to the tissue region of interest cannot be verified. Therein lies the value of a trained anatomic and clinical pathologist and biobanking technologist to help critically manage the practice and logistics of tissue samples to ensure that the genomic analysis is appropriate, correct, and helpful to research and, ultimately, clinical care. 2,36,44 Moreover, best practices in the research setting include a thorough documentation of the observations of the trained pathologist using defined data fields for the observations (eg, percentage tumor, percentage necrosis). These observations can now objectively be determined with digital- or spectral-based imaging and inexpensive software to replace or augment the less accurate qualitative immunohistochemistry (IHC) scoring.
Implementing biobanking protocols and active QA/QC programs offers clear benefits for uniform biospecimen collections and subsequent macroanalytes that reflect biological alterations. One critical application of cellular and genetic profiling in personalized medicine practice remains the critical QA measure of positive and negative baseline controls or degradation profile that could be documented and reported as specimen integrity status. 14,24,30,36 The documentation and reporting of cell populations of interest is becoming more routine in molecular assays in cytologic and tissue-based procurement, especially in heterogeneous tumor tissue samples. The percentage tumor content provided by the pathologist is a critical assessment in cancer research based on genomic and proteomic assays. Additional methods of cellular enrichment include laser capture microdissection and flow cytometry.
We propose that anatomic pathology verification of tissue samples utilized in human and veterinary research adopt a standard for QC and QA of specimen aliquots destined for high-throughput molecular analysis. Our laboratory, having received CAP BAP accreditation, is developing a clearer standard practice for QA of tissue-based high-throughput assays. These practices are particularly important where “diseased” and “normal” tissues are being compared, as well as in cases where the disease tissue has a high probability of being heterogeneous. The procedures that we use promote assay validity and improve the confidence and accuracy in genomic analysis by measuring biospecimen appropriateness of fit for use. The protocols include the following.
The histologic characteristics of the sample submitted for molecular assay must be verified by a trained pathologist by either of the 2 protocols described below. In either case, the protocol for the molecular assay must include parameters describing to what extent the sample analyzed must be histologically homogeneous and what is to be done if those parameters are exceeded. These parameters must be experimentally defined. The responsibility of the pathologist is to determine whether the parameters are met. The 2 acceptable methods for verifying the histologic characteristics of material sent for molecular assay are as follows.
Serial Cryostat Sectioning
A section from the frozen specimen is cut and prepared for histologic examination. This control section should come from tissue directly adjacent to the tissue to be used in the molecular analysis. An additional 5 to 10 sections of tissue (10 μm each) are then submitted for molecular analyses. The process of alternating sections for histologic examination and molecular assay can be repeated as necessary to obtain the required mass of tissue for assay. After the last set of tissue sections are obtained for molecular assays, a final section should be obtained for histologic examination to document the consistency of the tissue’s morphologic content.
Gross Sectioning
If a frozen tissue section is not being sectioned for immediate confirmation of tissue content at the time of acquisition, a thin gross specimen can be obtained and placed, freshly cut surface down, in a histology cassette and submitted for fixation, processing, and embedding (FFPE). The mirror-image frozen tissue section (or sections) is then submitted for molecular analysis. The FFPE specimen block is sectioned for hematoxylin and eosin staining, and the morphologic examination can then be used to confirm histologic equivalence for the tissue used for molecular analysis.
Suitable positive and negative control samples must be used in molecular assay protocols. The SOP should ensure that the analytic results obtained from subsequent test samples fall within the defined ranges identified by the control samples or within the dynamic range of the procedures.
Anatomic Pathology Tools and Microscopy to Evaluate Tissues for Use in Molecular Analysis
Histologic QC in Gene Expression Studies
Diagnostic pathologists are well aware of the need for rigorous application of standards and QC to ensure that accurate diagnosis is obtained from clinical samples and communicated to caregivers and investigators. However, in clinical research, systematic QCs have been the exception rather than the rule. Thus, in the assessment of high-throughput expression profiling (RNA) and gene dosage (DNA) assays in clinical samples, the lack of QCs (eg, the lack of determination in the tumor sample content) can lead to inaccurate analyses. Our recent analysis of the literature 45 revealed that among 100 of the most recent papers reporting results from microarray analysis on human malignancies, only 13% described serial sectioning of samples destined for molecular profiling to verify the histology of those samples and 40% did not document QC measures to ensure accurate pathologic review of the tissue aliquot destined for molecular analyses. To highlight the importance of the role of the pathologist in molecular diagnostics, we randomly selected 18 paraffin blocks of tumor and adjacent normal specimens that were designated by the surgeon or pathology assistant for a recent cancer study at our institution. Upon microscopic examination, several tumor samples proved in fact to be admixtures of normal and tumor tissue, and one normal tissue was clearly mislabeled and was in fact tumor. The failure for uniform histologic evaluation of tissue samples utilized for a given study results in (1) the lack of identification of significant differences between tumor and normal expression of receptors, (2) nonviable or inappropriate tissue in sample for quantification (eg, adipose or muscle tissue), and (3) contaminating benign cells and would ultimately lead to adoption of IHC analysis for simultaneous detection of receptor status and visualization of the histology. 27 For microarray analysis to be a reliable tool in either research or clinical decision making, it is imperative that the classification of the source sample be confirmed by histologic analysis. Histologic verification of the sample, as demonstrated in the “Histologic Review” protocol in Supplementary Table 2 (available at http://vet.sagepub.com/supplemental), will promote accurate and reproducible results of such analyses.
Generation of Tissue Microarrays
Histopathology-based techniques are the gold standard for diagnosis in anatomic pathology. However, the century-old protocols lack preservation in tissue quality often needed for many molecular analyses. Furthermore, for research, the highest-quality tissues are tethered to the clinical data. 33,43,44 Although standardized collection practices of FFPE samples are not able to meet the high-quality standard for tissues in molecular and genomic testing, FFPE tissues still have a use. 45,48 Despite limitations of the FFPE sample, these retrospectively collected tissue samples can be utilized in research design and biomarker validation. To maintain low costs for screening and to increase the number of cases reviewed in studies, high-density tissue microarray (TMA) 43 is a portable tool that has proven highly effective for evaluation of tissue biomarkers. 8,10 TMAs are able to maintain the targeting representative tissue, 74,76 conform to standardization of in situ assays, 7,10 fit the evaluation of molecular targets in context of tissue morphology, and provide molecular stratification of human and veterinary samples. 75,87,99 The value of the FFPE tissue sample and associated macroanalytes has risen significantly with the widespread implementation of TMAs over the past 15 years. 37,90,98 The TMA platform has demonstrated the need for control of preanalytic variables, even in FFPE collected tissues. This standardization includes the collection of research tissues in 10% neutral buffered formalin, documented fixation time of 12 to 72 hours, and transfer of fixed tissue to 70% ETOH solution up to 1 week before processing. It is also recommended that, in the assessment of the effects of preanalytic variables (eg, fixation and collection times), a control TMA be used with cores of tissues that have been optimally collected, to control for optimal IHC labeling and localization. This technique has proven extremely valuable in novel antibody optimization and biomarker validation. The TMA format offers critical advantages in the standardization for assay performance in FFPE and has brought increased attention and utilization of the clinical paraffin block archives. To better determine deleterious fixation effects in a given case, a normal adjacent core is punched and placed alongside tumor cores. This direct comparison of patient-matched tumor–normal pair provides clues in expression level between diseased and nondiseased tissue, especially in the contextual interpretation of protein expression, either gain or loss, in a tumor-specific TMA. IHC is widely used in both the research setting and the hospital-based anatomic pathology department. Validation of novel and known biomarkers by IHC labeling offers valuable cross-institutional and translational resource in the evaluation of novel diagnostic and prognostic biomarkers. The primary value and contribution of TMA methods have been in gauging relative abundance of protein expression in tumor versus nontumor tissues by quantitative measures of protein abundance in a continuous manner. Automated algorithms for quantitative analysis of IHC or immunofluorescence-labeled tissues are largely demonstrated in the research setting, where there is much better-controlled collection and processing practices. Conversely, in diagnostic pathology, predominantly qualitative measures in FFPE archival samples remain the common clinical practice. In addition, the TMA format offers an optimal platform to evaluate novel fixatives, provided that the tissue is processed in the paraffin block format.
Spectral Imaging
Spectral imaging provides another method of analyzing tissue components with the natural emissions of light expressed through the tissue without having to specifically label the tissue or to better discern the emission of light from a fluorophore from other background fluorescence. An example of a spectral imaging system is the Nuance Cri (Perkin Elmer, Waltham, Massachusetts), which provides moderate content image files with spectral separation of fluorescent and nonfluorescent images. A broad use is the spectral separation of various chromogenic signals, such as separation of hematoxylin-and-eosin spectra, or the dual-stained IHC slide to calculate colocalized signals and quantitative signal intensity. Spectral imaging provides a tool to simplify the extraction of morphologic and molecular information by use of modified fluorescent tools that mount on standard microscopes and can generate spectral image data sets with excellent spatial and spectral resolution. The Nuance platform segments image data in 3 dimensions as a set of pixels into regions of cellular significance, such as nucleus versus cytoplasm. After segmentation and cataloguing of data coordinates, evaluation of the spectral and spatial properties is applied to identify differences/similarities of specimens, which can be used to analyze biomarkers that may be useful in potential prognosis or predictive drug treatment with a given pharmacologic agent. The example of anti-VEGF or similar antiangiogenic marker is illustrated in Figure 1 with CD31 staining of the highly vascular renal cell carcinoma to calculate vascular density and integrity. 71,79,100 Spectral imaging has been widely applied in cytometry for cell morphometric measures, such as cell size/shape, nuclear composition, and dimensions. 22,53
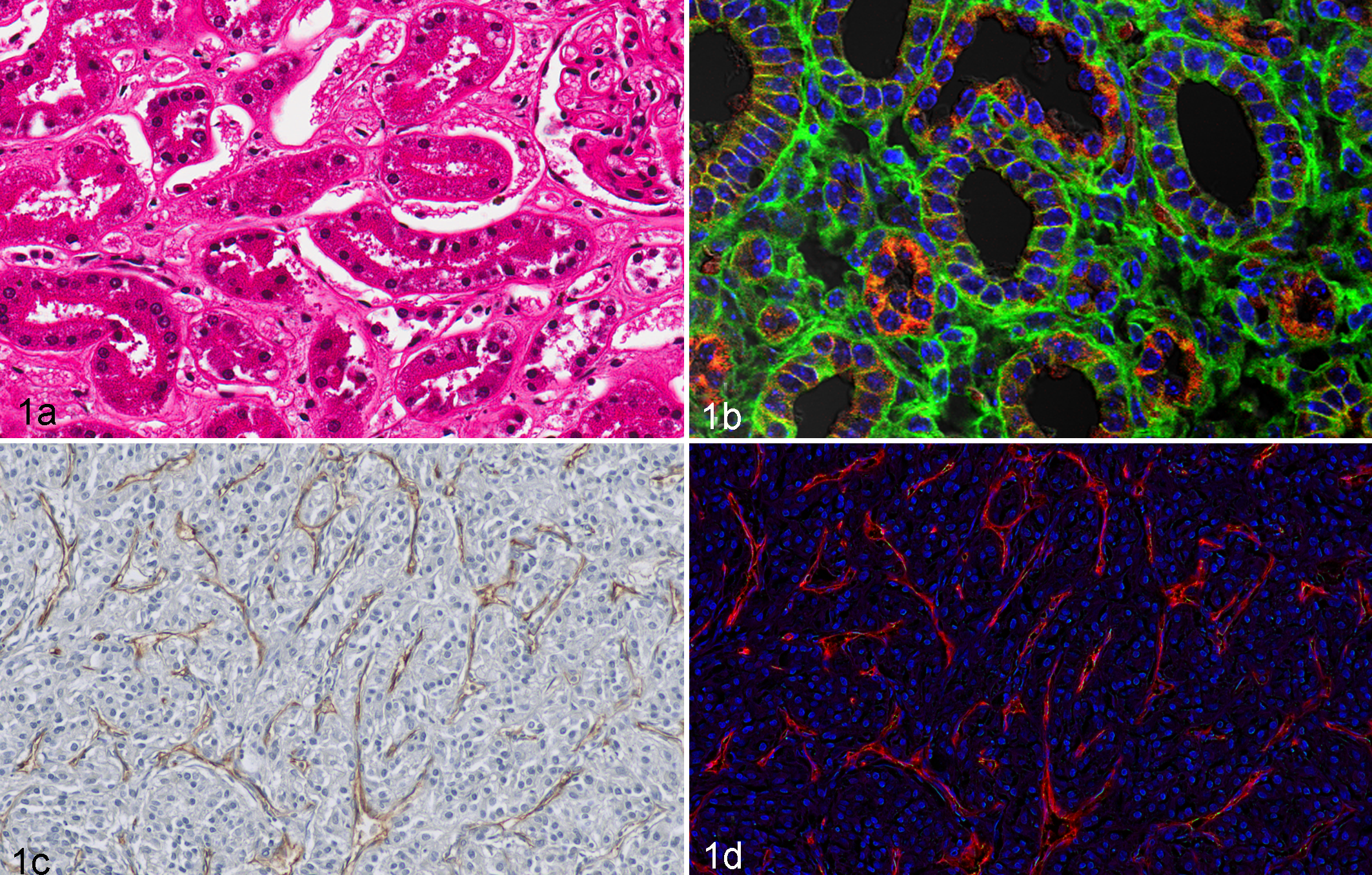
Normal murine samples: (a) Hematoxylin-and-eosin staining; (b) fluorescent-labeled alpha catenin (TRITC, red) and beta catenin (FITC, green). (a) Proximal tubules susceptible to hypoxia show disrupted cell borders and (b) are rich in TRITC-labeled mitochondria (red). Distal tubules with absorptive and secretory functions show contiguous FITC-labeled cell membranes (green). Renal cell carcinoma human samples with CD31 staining (c, d) illustrate the use of spectral imaging to quantitate stromal elements. (c) Spectral imaging allows use of standard immunochemistry-stained tissue sections for quantitative measures of blood vessels and pericytes with DAB chromagen–labeled CD34 marker (brown) and hematoxylin (blue) counterstained. (d) Blood vessel number and halo positivity were calculated by Nuance camera system, in which emissions were collected between 420 and 720 nm in 20-nm increments and unmixed images were quantified using specialized analysis software.
Evidence-Based Best Practices for Collection of Tissues for Molecular Analysis
Tissue-processing data that have not been routinely captured in anatomic pathology of clinical tissues include the time of surgical excision to fixation. These data may not affect the morphologic analyses of tissues in traditional pathology, but it will presumably have a significant effect on analysis of RNA and labile proteins that are needed for the interrogation of complete molecular signatures. Evaluation of RNA templates is increasingly important, as many tissue types, including FFPE materials, have been adapted to next-generation sequencing platforms and technologies. Suboptimal nucleic acid sources, including FFPE tissues and partly degraded templates, are increasingly utilized in next-generation sequencing applications due to improved technology, special reagents, alternate library preparation schemes, and powerful alignment informatics. 50,69 From earlier experiences in gene expression profiling, the technological advances are not necessarily matched by sample QA and documented RNA quantity or quality. In addition, the higher sensitivity of recent platform measures may accentuate the disparate reporting of findings and so limits even a moderate degree of analytic confidence.
Tissue Degradation Time Course
To model the effects on tissue quality or integrity that occur during the course of routine tissue procurement, we have performed a time course study to mimic time-to-fixation or cryostabilization of tissue RNAs and proteins that may be affected by enzymatic degradation or other signals to activate biological pathways such as cell death. Athymic nude mice were obtained under Van Andel Research Institute Vivarium Core under an Institutional Animal Care and Use Committee–approved study (XPA-12-07-001). Mice were euthanized by standard procedures, and kidney organs were removed and dissected into 4-mm cube aliquots, each to include cortex and medulla. Tissue aliquots were incubated at 37°C in a humid environment and then flash-frozen starting at time intervals of 0, 3, and 6 hours postresection. All tissues were stored in a mechanical –80°C upright freezer. Similar aliquots for each time point were fixed in 10% neutral buffered formalin for 12 to 24 hours, followed by automated tissue processing and paraffin embedding. RNA was extracted by a manual method (RNeasy Mini Kit, Qiagen, Inc); 260/280 ratios were measured with the Nanodrop; and 28S/18S ratios were calculated with the Agilent Bioanalyzer 2100 and are summarized in Figure 2.
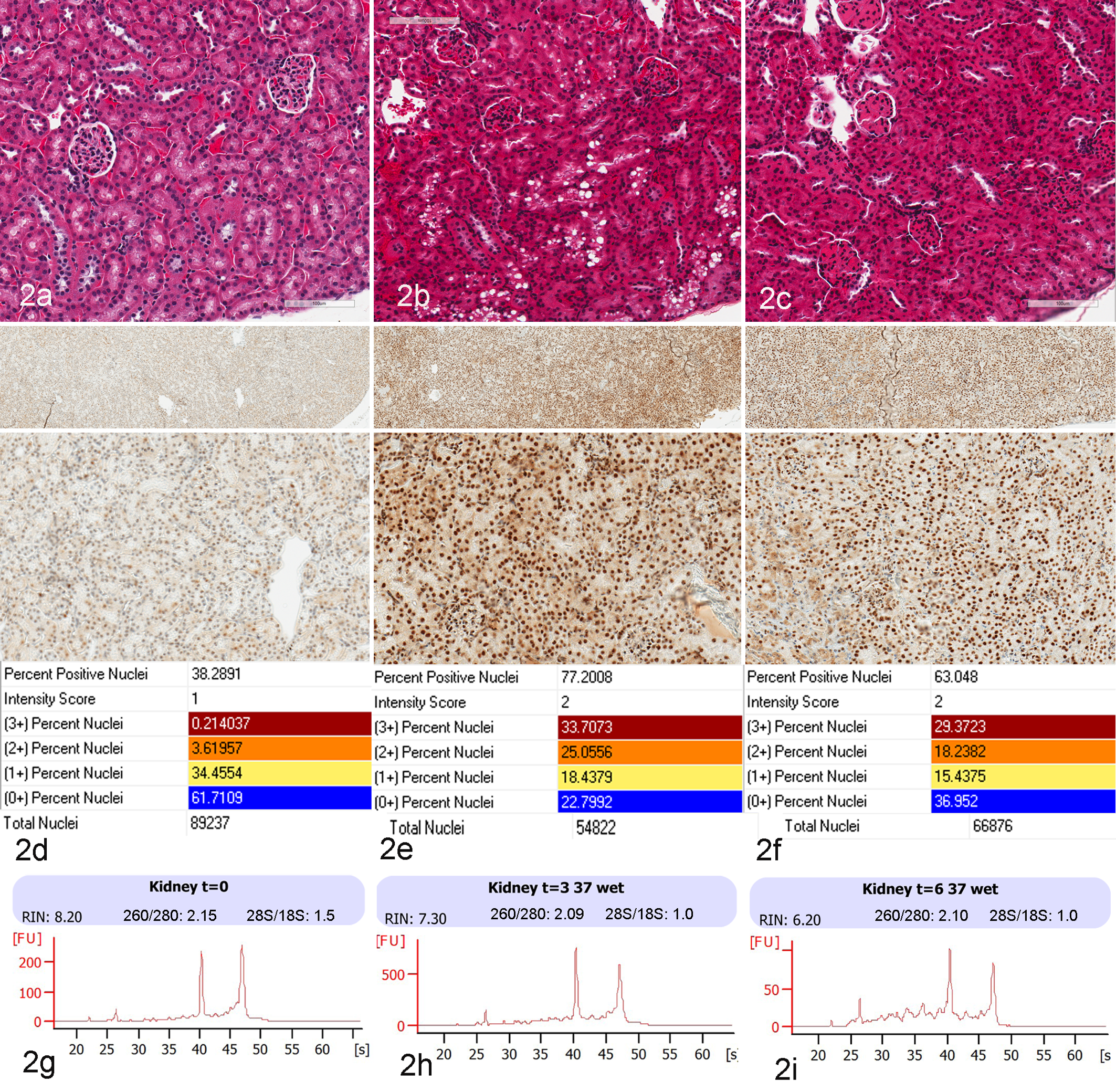
Murine kidney time-course tissue degradation study with compiled images and data. (a–c) Hematoxylin-and-eosin stain of kidney cortex at 0, 3, and 6 hours post–animal sacrifice with autolytic change of cytoplasmic vacuolization (b) and smudged nuclei (b, c). (d–f) Immunohistochemistry staining for XBP-1, a critical marker of endoplasmic reticulum stress, is shown at 20× magnification in the upper panel and 200× magnification in the lower panel, with minimal staining at time 0 (d) and marked upregulation with diffuse nuclear labeling at 3 and 6 hours (e, f), and is confirmed by quantitative image analysis. Correlative morphologic and RNA/protein degradation measures are shown for 0 (a, d, g), 3 (b, e, h), and 6 hours (c, f, i). RNA integrity number profiles by the Agilent 2100 Bioanalyzer (g–i) indicated that RNA degradation was limited to the 6-hour time point. Standard nucleic acid measures of 260/280 and 28S/18S at each time point are included.
To correlate morphologic changes and loss of RNA integrity, IHC was performed against XBP-1, a specific marker for endoplasmic reticulum stress; slides were digitally scanned with an Aperio ScanScope (Leica, Vista, California) and evaluated with automated image analysis software (Aperio, Leica). Over the past decade, high-resolution digital image scanning has led to markedly improved cost-effective storage, retrieval, and transmission of large image files for use in research and, as a result, continue to show promise as a tool for tissue-based diagnosis and consultations. 31,97 A digitized high-resolution electronic image, analyzed with advanced image analysis algorithms, provides accurate and reproducible measurements of pathologic microscopic changes at the tissue and cellular levels. Whole-slide imaging is steadily gaining in popularity as daily practice in research pathology; however, the adoption of digital pathology has lagged in clinical pathology departments. 41 Digital images of human and murine kidney samples that depict known functional features of normal kidney (murine) and renal cell carcinoma (human) are presented in Figure 1. These include the histologic images and a markup image following image analysis with a nuclear algorithm to quantitate nuclear IHC immunolabeling in murine kidney cortex for XBP-1. The IHC staining as shown in Figure 2 is presented with correlative RNA measures, including 260/280 (by Nanodrop) and 28S/18S ratio and RNA integrity number (RIN; by Agilent Bioanalyzer 2100).
In our time course study of mouse kidney and collection variables upon RNA integrity, we determined historical RNA measures to include 28S/18S and 260/280 ratios and RIN scores. As shown in Figure 2, at time intervals of 0, 3, and 6 hours, the 28S/18S ratio was 1.5, 1.0, and 1.0, respectively, and the 260/280 ratio remained quite high at 2.15, 2.09, and 2.10, respectively. The RIN score, however, showed linear downward measures of 8.20, 7.30, and 6.20, respectively, consistent with progressive degradation. Degradation at the RNA level was validated by IHC for a marker of cellular stress, XBP-1, critical to the autophagy pathway. The changes in protein expression were quantitatively assessed by digital imaging algorithms and appeared to match or even precede the reported degradation by RIN score. The analysis confirms the upregulation and production of the XBP-1 protein as a measure of the endoplasmic reticulum stress mechanism as the related unfolded protein response in autophagy. 46,51,52 The experimental model organ of mouse kidney contains heterogeneous cell populations in glomeruli, proximal, and distal tubules that have diverse functions and energy requirements. Here, we utilize an experimental model system to assess macromolecular degradation (RNA) within the context of known cellular biology response (hypoxia) with correlative tissue and cellular product (in situ protein expression) that can support biospecimen science initiatives to influence tissue sample collection and qualification for downstream biomarker and genomic assays. To date, our mouse model studies suggest that additional measures of RNA could better inform sample integrity at the transcription level of highly sensitive assays, such as RNA sequencing. 16,18,101 From these data, we have shown the early developments of a murine tissue model where we can determine the level of degradation by RIN and IHC, which can now be extended and eventually help to evaluate tissue management procedures and establish best practices based on experimental evidence.
Gene Expression Profiling and Veterinary Disease
Recent reviews of expression profiling in veterinary research include canine cardiomyopathy, 64,65,67 degenerative mitral valve disease, 66 atopic dermatitis, 57 pancreatic acinar atrophy, 12 and malignancies of the breast 73 and central nervous system. 88 Disease in other species include bovine mastitis with detailed interactions between host and infectious organisms 85 and osteoarthritis in horses. 82 The goals of these studies are similar to those in human cancer and disease investigations—namely, identification of diagnostic markers, therapeutic targets, or factors leading to disease resistance in diseases such as mycobacteriosis, salmonellosis, trypanosomiasis, 13,80 avian infections, 40,59 infectious anemia virus in salmon, 37 and nematode infections in sheep. 38 Genomics have improved our understanding of the biology of these diseases and have helped determine which novel diagnostic or therapeutic strategy could improve the quality of life of companion animals and/or improve the efficiency of commercial production of animal products. There have been proposals for incorporating gene expression profiling into certain aspects of the clinical care of animals, 21 and this is already taking place to a limited extent in the clinical care of humans—most notably with respect to breast cancer prognosis. 89,91 Molecular medicine holds the promise to augment anatomic pathology by increasing its objectivity—an important goal in light of documented intersite variation in subjective interpretation of pathology samples and inherent discrepancy in semiquantitative methods. 6,15,23,81 The combination of histology and gene expression analysis has the power to improve the accuracy of clinical diagnostics and prognostics by precise segregation of individuals into meaningful groups, allowing for better-informed therapeutic decision making. 70,72 However, molecular techniques themselves have been plagued with reproducibility challenges, 90 including the fact that parallel studies frequently identify different gene sets as a result of variations in sample processing, array platform, endpoints of interest, or computational analysis method 54,92,94 and often failed Federal Drug Administration current review paradigms. These documented sources of variability highlight the need for more research to derive better practices of biospecimen preservation, followed by improved standardized procedures to ensure that appropriate samples are being analyzed when molecular techniques are employed. The need for standardization of reporting and analysis has been recognized in related contexts, including clinical trials and tumor marker reporting. 56,60 Details regarding tissue identity, viability, and homogeneity must be verified when molecular profiling data are reported to ensure valid interpretation of results. We can assume that previous collection and preservation practices for tissues have only added to the variability in early transcriptome analysis because the degree of preanalytic variables has most probably hindered uniform and high-quality measures of RNA integrity. The revitalization of biospecimen science emphasizes that collection and storage protocols should include better assessment of histologic equivalence of the molecular aliquot and more thorough documentation of times to preservation and fixation.
Conclusions
High-throughput molecular assays are playing an ever-growing role in research and into understanding the pathobiology of human and animal diseases. There is an expectation that developing protocols to ensure the collection of quality tissues will define new methods for good clinical practice. The implementation of QA practices for the collection of tissues must be completed by a trained pathologist as an essential link in the chain of events leading ultimately to reliable and reproducible research findings and, eventually, appropriate clinical care. The implementation of these procedures will also need to expedite the collection and fixation of tissues for use in molecular analysis. Furthermore, the integration of pathology-guided selection of targeted sampling and biomarker analysis in heterogeneous tissue samples highlights the need for traditional morphologic evaluation to provide for more informative genomic or molecular analyses. The use of high-resolution digital imaging systems should be encouraged in providing quick and continuous links of tissue morphology to guide tissue sampling for downstream molecular and genomic assays. Subsequently, the refinements in tools and methods will continue to advance experimental design and success in both the human and veterinary research fields.
Footnotes
Acknowledgements
We thank Sabrina Noyes for her editorial and administrative assistance in the preparation of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared receipt of the following financial support for the research, authorship, and/or publication of this article: The authors would like to thank the Van Andel Research Institute for financial support for this research.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
