Abstract
The purpose of the study was to evaluate clonality and presence of numerical chromosomal and centrosomal aberrations in 5 established feline fibrosarcoma cell lines and in a fetal dermal fibroblast cell line as a control. The clonality of all cell lines was examined using limited-dilution cloning. The number of chromosomes was counted in metaphase spreads. The immunocytochemical analysis of centrosome numbers was performed by indirect immunofluorescence using a monoclonal antibody that targets γ-tubulin, a well-characterized component of centrosomes. Monoclonal cell populations could be established from all cell lines. In all feline fibrosarcoma cell lines, the number of chromosomes deviated abnormally from the normal feline chromosome number of 2n = 38, ranging from 19 to 155 chromosomes per cell. Centrosome hyperamplification was observed in all 5 feline fibrosarcoma cell lines with a proportion of cells (5.7 to 15.2%) having more than 2 centrosomes. In the control cell line, only 0.6% of the cells had more than 2 centrosomes. In conclusion, the examinations revealed that centrosome hyperamplification occurs in feline fibrosarcoma cell lines. The feline fibrosarcoma cell lines possessed 10 to 25 times as many cells with centrosome hyperamplification as the control cell line. These observations suggest an association of numerical centrosome aberrations with karyotype instability by increasing the frequency of chromosome missegregation. The results of this study may be helpful for further characterization of feline fibrosarcomas and may contribute to the knowledge of cytogenetic factors that may be important for the pathogenesis of feline fibrosarcomas.
Keywords
Soft tissue sarcomas are common mesenchymal tumors of the cat. 21 Since the early 1990s, an association of solitary subcutaneous sarcomas with vaccinations became obvious.8,22,23,24 These sarcomas have been termed vaccination-associated, vaccine-associated, or injection site–associated sarcomas due to their typical location at common sites of subcutaneous injection of vaccines.
Possible causes proposed and under investigation include chronic inflammation and proto-oncogenic induction at the vaccine site by vaccine adjuvants, especially aluminium. Aluminium-containing vaccines caused greater lesions than vaccines without aluminium. 24 However, the exact role of aluminium in tumorigenesis has not been discovered yet, also because follow-up studies failed to confirm a correlation between aluminium-containing vaccines and tumor formation at injection sites.24,32,51 Vaccination alone will not cause a fibrosarcoma, more like a multistep carcinogenesis model that includes genetic and iatrogenic factors (eg, cytokines) that support a local inflammatory reaction.3,20,24
Additionally, long-term penicillin, methylprednisolone acetate, and lufenuron, injected subcutaneously, have been discussed to induce sarcoma formation in some cats as well.13,16,20,32 These sarcomas are locally invasive and have a high rate of local recurrence.26,32 The estimations of the frequencies of vaccination-associated sarcomas mostly range from 1 per 1000 to 1 per 10 000 vaccinated cats.7,19,35
The pathogenesis of these feline sarcomas is not well understood yet. No sex, breed, or age predisposition has been shown. In addition, a smaller amount of feline fibrosarcomas appear to have no association with vaccinations.3,32
The genetic background of feline fibrosarcomas still remains unclear.3,24 Classic cytogenetic analysis as well as genomic microarray and in situ hybridization studies revealed that feline soft tissue sarcomas may have complex karyotypes with ploidy changes.28,37,38,50
In feline fibrosarcomas, chromosomal instability (aneuploidy) was found in several previous studies. Mayr et al 37 analyzed 4 feline fibrosarcomas cytogenetically. Each cell line showed a moderate number of abnormal metaphases, and on the whole, 20 to 40% of the cells revealed a genetic heterogeneity. Another analysis of metaphase spreads of a feline fibrosarcoma revealed 34 to 115 chromosomes per cell, in which 69.2% of the cells had a karyotype of 51 to 64 chromosomes per cell. 38 Kalat et al 28 found, in addition to structural chromosomal aberrations, a chromosomal hyperdiploidy with 40 to 46 chromosomes per cell in feline fibrosarcoma cell lines.
An extensive range of genomic imbalances was also shown by Thomas et al, 50 who used a genomic microarray to investigate nonrandom chromosome copy number aberrations in 46 feline sarcomas. A further study of Santos et al analyzed the cat satellite DNA family and telomere sequences by in situ hybridization. This repeat sequence was found to be amplified in the cat tumor cells. The amplification was associated with variable marker chromosomes. Some of the most representative tumor clones (more than 50%) showed hypo- and hypereudiploid karyotypes: 2n = 26, 2n = 32, 2n = 49 and 2n = 50. The authors concluded that this amplification might be related to chromosomal instability. The lack of an association between amplification of the cat satellite DNA family and chromosome number alteration suggests that alternative causative mechanisms could induce genetic instability. 45
The instability of chromosome number is considered to be important in multiple-step tumorigenesis. Possible mechanisms that might lead to chromosomal instability include the loss of mitotic checkpoint function and abnormal amplification of centrosomes. 14 The important role of the centrosome as a microtubule-organizing center is the formation of bipolar spindle poles for a symmetric chromosome segregation. The centrosome duplication cycle is closely linked to the cell division cycle. Prior to mitosis, the centrosome duplicates, matures, and separates by moving to the opposite poles of the cell. Since centrosomes duplicate only once during the cell cycle, each daughter cell inherits 1 centrosome. 15 Mitosis requires the presence of a bipolar spindle organized by 2 centrosomes, and in normal cells, the centrosome numbers are regulated by strict control mechanisms.9,27,44 Numerical centrosome aberrations, which lead to the formation of multipolar spindle poles and unequal chromosome segregation, are an important factor for aneuploidy, which is a relevant characteristic of malignant tumors. Failure in these control mechanisms results in abnormal amplification (hyperamplification) of centrosomes with the presence of more than 2 centrosomes per cell. Cells with hyperamplified centrosomes frequently exhibit increased aberrant mitosis and chromosomal missegregation.10,14 These various cytogenetic studies have demonstrated that chromosomal instability may be involved in the pathogenesis of feline fibrosarcomas. A relationship between aneuploidy and centrosome anomalies has also been reported in a number of tumor cell lines and in solid tumors in humans—for example, in cells of mammary carcinomas, carcinomas of the prostate, colon carcinomas, and hepatocellular carcinomas.6,18,34,40,43 Also, precancerous tissue alterations, such as prostate hyperplasia or benign colon adenomas, revealed an association between aneuploidy and centrosome hyperamplification (>2 centrosomes per cell).30,42,43
Chromosome studies provide a relevant approach to the understanding of tumor mechanisms. Based on these findings, it seems to be of considerable interest to examine whether centrosome hyperamplification is coexistent with aneuploidy in feline fibrosarcomas. These phenomena have been demonstrated in feline lymphoma cell lines and in canine tumor cell lines.39,46 In this study, we investigated numerical centrosome aberrations in feline fibrosarcomas as well as a possible relationship between centrosome hyperamplification and chromosomal instability. In addition, we tried to evaluate the origin of karyotype instability in all 5 fibrosarcoma cell lines by the establishment of clonal cell lines. We hypothesized 2 possibilities for the occurrence of aneuploidy in these cell lines. The first possibility was that a fibrosarcoma cell line consists of different cell populations, in which every cell population has a stable karyotype that is given from the mother cell to the daughter cell during every mitosis. The karyotype can deviate from the feline eudiploid karyotype (2n = 38), but it stays stable in each cell clone. Alternatively, we hypothesized that cells of a fibrosarcoma cell line have different karyotypes and that a variable number of chromosomes is delivered to the daughter cells during cell division. Subsequently, the karyotype of the daughter cells can be different from the parental cell.
Furthermore, the present study suggests the presence of a still-unknown mechanism that leads to aneuploidy and centrosome hyperamplification in fibrosarcomas in cats. The cytogenetical characterization of these fibrosarcoma cell lines intends to make a contribution to the diagnosis of cytogenetical factors within the pathogenesis of feline fibrosarcomas.
To the best of our knowledge, this is the first time that centrosome hyperamplification in association with aneuploidy is demonstrated in feline fibrosarcoma cell lines.
Materials and Methods
Cell Lines
Feline fibrosarcomas derived from 5 cats with primary subcutaneous solitary soft tissue sarcomas were collected during surgical treatment and referred to the Department of Veterinary Pathology of the Justus-Liebig-University Giessen, Giessen, Germany, between 2002 and 2003 (Table 1 ). All tumors were presumed to be vaccine associated, based on vaccination history and/or tumor site (lateral thorax, interscapular region). The fibrosarcoma cell lines examined in this study were designated as fibrosarcoma cell line I (FS I) to fibrosarcoma cell line V (FS V).
Origin and Clinical Data of the Feline Fibrosarcoma Cell Lines and the Feline Dermal Fibroblast Cell Line (Control Cell Line) a
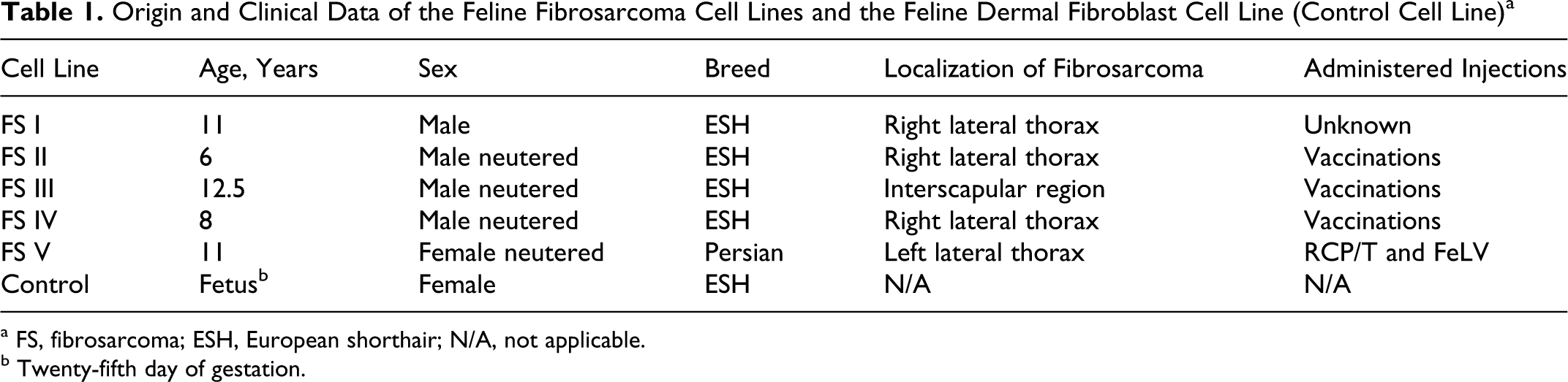
a FS, fibrosarcoma; ESH, European shorthair; N/A, not applicable.
b Twenty-fifth day of gestation.
All examined tumors were located in the subcutis and were poorly circumscribed. Histologically, all tumors were composed of pleomorphic spindle-shaped cells in interlacing bundles and streams. The spindle cells were separated by a moderate amount of collagenous stroma. Oval to fusiform nuclei differed in size, and mitotic figures were common. The tumors from which the cell lines FS I, FS II, and FS III derived revealed the presence of multinucleated giant cells. Additionally, the tumor from which the cell line FS III derived showed a large cavitated center in consequence of extended necrosis. Small vessels in the periphery of the tumors were surrounded by small aggregates of lymphocytes and macrophages. In all tumors, some macrophages enclosed gray-brown intracytoplasmic granular material, presumed to contain vaccination adjuvants (eg, aluminium). These findings support the classification of these tumors as vaccine-associated fibrosarcomas, even though absolute certainty in differentiating from spontaneous fibrosarcomas is difficult to achieve. 36
For tissue culture, pieces of fresh tumor tissue were cut into small pieces and put into tissue culture flasks with complete DMEM, supplemented with 0.1 ml/ml of fetal bovine serum, 0.01 ml/ml of penicillin/streptomycin (10 000 U/10 mg per ml of stock solution), and 1 μl/ml of gentamicin (50 mg per ml of stock solution, sterile filtration). The tissue culture flasks were incubated in a humidified chamber (37°C) with 5% CO2.
When a confluent monolayer had grown in a tissue culture flask, the cells were trypsinized and seeded into another tissue culture flask. As a control, a feline fetal dermal fibroblast (DF) cell line was used, which was established from fetal DFs. The cells were developed under the same conditions as described above. The cell line was brought to passage 1 and then deep-frozen in our cell bank. The control cell line showed a much lower growth rate than the tumor cell lines; they were consequently passaged much less frequently.
All tissue culture reagents were obtained from PPA Laboratories GmbH, Pasching, Austria, unless otherwise stated.
The exclusion of FeLV infection was performed for all feline fibrosarcomas and also for the control cells by immunohistochemistry (envelope glycoprotein 70 and p27) according to the method of Kipar et al 31 and by polymerase chain reaction (PCR). PCR was performed according to the method described by Suntz et al. 49 Since FeLV infection has been shown to induce transformation of mesenchymal cells in feline fibrosarcomas, the absence of FeLV infection in each of the cell lines from the present study was confirmed by immunohistochemistry and PCR. 11
Establishment of Clonal Cell Lines: Limited-Dilution Cloning and Subcloning
Limited-dilution cloning was performed to obtain a monoclonal cell population starting from a polyclonal population. This was achieved by setting up a series of dilutions of the parental cell culture to a final concentration of 100 cells/ml. Afterward, 10 μl of that cell suspension was pipetted into a well of a 96-well microtiter plate and controlled by light microscopy for the presence of 1 cell per well. Wells with 1 cell were marked and controlled daily for the formation of a monoclonal cell colony. Wells with more than 1 cell were destroyed. When a confluent monoclonal monolayer had grown in a well after about 2 to 3 weeks, the cells were trypsinized and seeded into tissue culture flasks (25 cm2).
To increase the cloning efficiency, the cell medium was supplemented with 20% fetal calf serum. The procedure of cloning was repeated once for all established clones from the fibrosarcoma cell lines and the control cell line. The resulting subclones were subjected to cytogenetic analyses. The subclones in this study are designated FS-SC-1 to FS-SC-5, corresponding to the cell line of origin. The subclone of the control cell line was termed DF-SC.
Fibrosarcoma cell line FS-SC-1 derived from FS I, passage 90; FS-SC-2 derived from FS II, passage 81; FS-SC-3 derived from FS III, passage 55; FS-SC-4 derived from FS IV, passage 53; FS-SC-5 derived from FS V, passage 43; and DF-SC derived from the control cell line (DF), passage 28.
Cytogenetic Analysis
Chromosome preparation
Metaphase spreads were prepared according to the method of Barch 2 with the modification described by Henegariu et al. 25 To enrich mitotic cells, the cells were incubated for 2 hours prior to harvesting by addition of 50 μl of colchicin (Colcemid, Biochrom AG seromed, Berlin, Germany; 10 μg/ml in Hanks balanced salt solution) in 5 ml of culture fluid. Then, the cells were transferred into 15-ml conical tubes for centrifugation. The cells were pelleted by centrifugation, and the supernatant was removed. Subsequently, the cell pellet was gently resuspended in a hypotonic solution (0.075M potassium chloride) and allowed to stand for 10 minutes at 37°C. Afterward, it was pelleted again (225 × g, 4°C, 10 minutes). After removal of the hypotonic solution, a methanol–acetic acid (3:1) fixative with a temperature of –20°C was added, and the cells were allowed to stand for 10 minutes at room temperature. Centrifugation, removal of the supernatant, and resuspension in a new charge of the fixative was repeated 2 times. After the last centrifugation step, the pellet was resuspended in 0.5 ml of fixative, and 2 drops were put onto a frozen (–20°C) glass slide. Air-dried samples were Giemsa stained. Subsequently, the metaphase spreads were analyzed under a light microscope.
Chromosome preparation of the feline fibrosarcoma cell lines before cloning
With regard to the feline fibrosarcoma cell lines, the following metaphases were Giemsa stained and microscopically analyzed: in cell line FS I, 15 metaphases from passage 82; in cell line FS II, 25 metaphases from passage 74; in cell line FS III, 25 metaphases from passage 48; in cell line FS IV, 25 metaphases from passage 47; in cell line FS V, 25 metaphases from passage 36; in the control cell line (DF), 25 metaphases from passage 23.
Chromosome preparation of the subclones
With regard to the subclones, the following metaphases were Giemsa stained and microscopically analyzed: in cell line FS-SC-1, 71 metaphases from passage 118; in cell line FS-SC-2, 60 metaphases from the passages 28, 37, and 65; in cell line FS-SC-3, 112 metaphases from passage 53; in cell line FS-SC-4, 63 metaphases from the passages 21 and 66; in cell line FS-SC-5, 120 metaphases from the passages 10, 20, 27, and 44; in the control cell line DF-SC, 39 metaphases from passage 13.
Indirect immunofluorescence
With regard to indirect immunofluorescence, the following cells were examined by fluorescence microscopy: in cell line FS-SC-1, 151 cells from passage 134; in cell line FS-SC-2, 151 cells from passages 67; in cell line FS-SC-3, 151 cells from passage 66; in cell line FS-SC-4, 151 cells from passages 69 and 84; in cell line FS-SC-5, 151 cells from passage 54; and in the control cell line DF-SC, 151 cells from passage 13.
To determine centrosome numbers, cells were immunostained for γ-tubulin, which is a well-characterized component of centrosomes. Immunostaining of γ-tubulin detects centrosomes in all phases of the cell cycle.4,48,53 To perform immunocytochemical examination, the cells were seeded on chamber slides (Lab-Tek II, Nalge Nunc International Corp., Naperville, IL). Subconfluent monolayers were fixed at –20°C in methanol (99.8%) for 10 minutes. The fixative was removed, and the cells were additionally fixed at –20°C in acetone (100%) for 1 minute. After removal of the second fixative, the slides were air-dried at room temperature. The slides were washed with Tris-buffered saline (TBS; pH 7.6, Sigma-Aldrich Chemie GmbH, Taufkirchen, Germany) and permeabilized with 1% Nonidet P-40 (Sigma-Aldrich Chemie GmbH) in TBS (pH 7.6) for 12 minutes at 37°C. Then, the slides were rinsed in TBS. To avoid nonspecific background staining, 40% swine serum in TBS was applied to the slides for 1 hour at room temperature. Subsequently, the cells were incubated with mouse anti-γ-tubulin monoclonal antibody (Clone GTU-88, Sigma-Aldrich, Steinheim, Germany) diluted 1:800 in TBS overnight at 4°C. The samples were rinsed 3 times in TBS. Incubation with the Cy3-conjugated secondary antibody (AffiniPure Goat Anti-mouse IgG (H+L), Dianova GmbH, Hamburg, Germany), diluted 1:800 in TBS, followed for 1.5 hours at room temperature. The samples were rinsed in TBS and counterstained with Sybr Green (Invitrogen GmbH, Karlsruhe, Germany).
Statistical Analysis
Statistical analysis was performed using the nonparametric Kruskal–Wallis rank sum test (H test). In case of significant differences, this test was followed by the pairwise Nemenyi test to compare the mean chromosome numbers between 2 cell lines. The Levene test was used to examine the equality of variances of the chromosome numbers of the different cell lines. The overall level of significance used was P ≤ .05. A possible correlation between the distribution of chromosome number per cell and percentages of centrosome hyperamplification was examined by Spearman rank correlation. The computations were performed using the program BMDP/Dynamic 7.0 (BMDP Statistics Software, Inc., Los Angeles, CA).
Results
Chromosome Analysis of the Feline Fibrosarcoma Cell Lines Before Cloning
The number of chromosomes per cell showed in all 5 feline fibrosarcoma cell lines, before cloning, an abnormal deviation from the eudiploid number of 2n = 38. The chromosomal analysis revealed 23 to 156 chromosomes per cell (Fig. 1 ). The control cell line (DF) revealed 23 metaphases with 2n = 38 and 1 metaphase with 37 chromosomes per cell and 1 metaphase with 36 chromosomes.
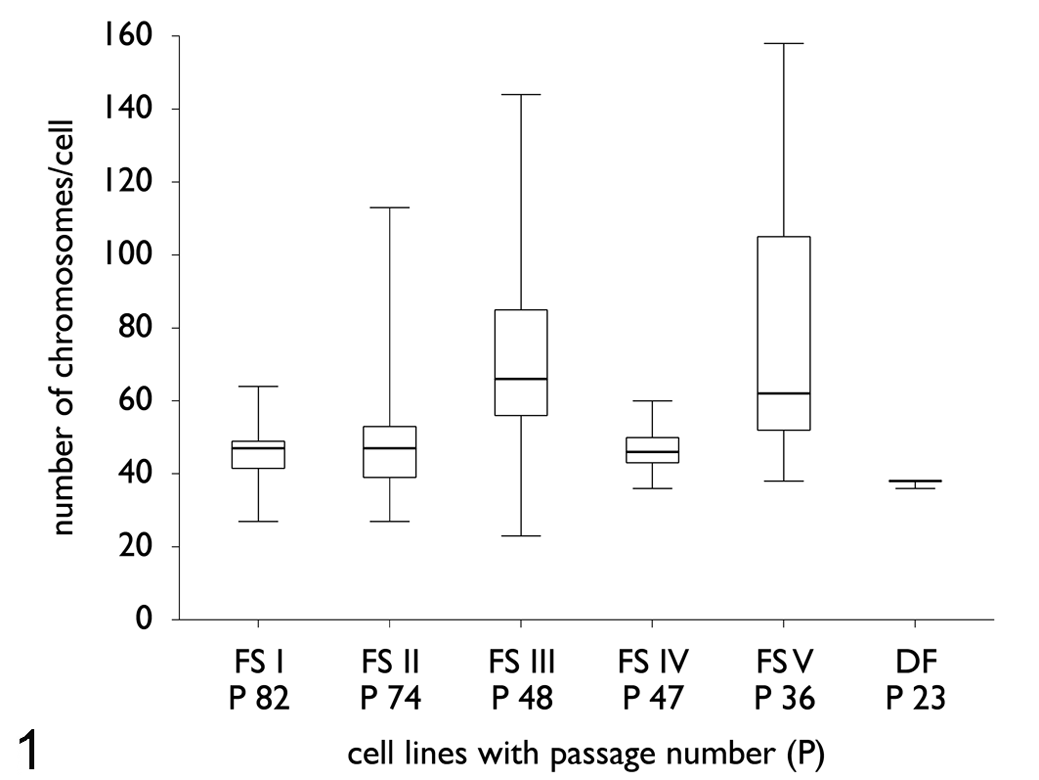
Box and whisker plots of the chromosome numbers per cell in all feline fibrosarcoma cell lines before cloning. The analysis of metaphase spreads displayed 23 to 156 chromosomes per cell, an extreme deviation from the normal number of chromosomes in cats (2n = 38). The number of chromosomes per cell revealed a widespread variability in chromosome number (aneuploidy) within all feline fibrosarcoma cell lines. In contrast, the control cell line revealed predominantly a eudiploid karyotype (2n = 38). FS, fibrosarcoma; DF, dermal fibroblast (control).
Establishment of Clonal Cell Lines
From each fibrosarcoma cell line, clonal growth could be achieved. A clonal cell line was also established from the control cell line. The cloning efficiency of all fibrosarcoma cell lines differed considerably from the control cell line (Fig. 2 ). The fibrosarcoma cell lines showed an average percentage of clones of 3.4%. In comparison, the control cell line had a threefold-higher cloning efficiency, with an average percentage of 11%. The cloning efficiency of all subclones was similar to that of the respective original clones.
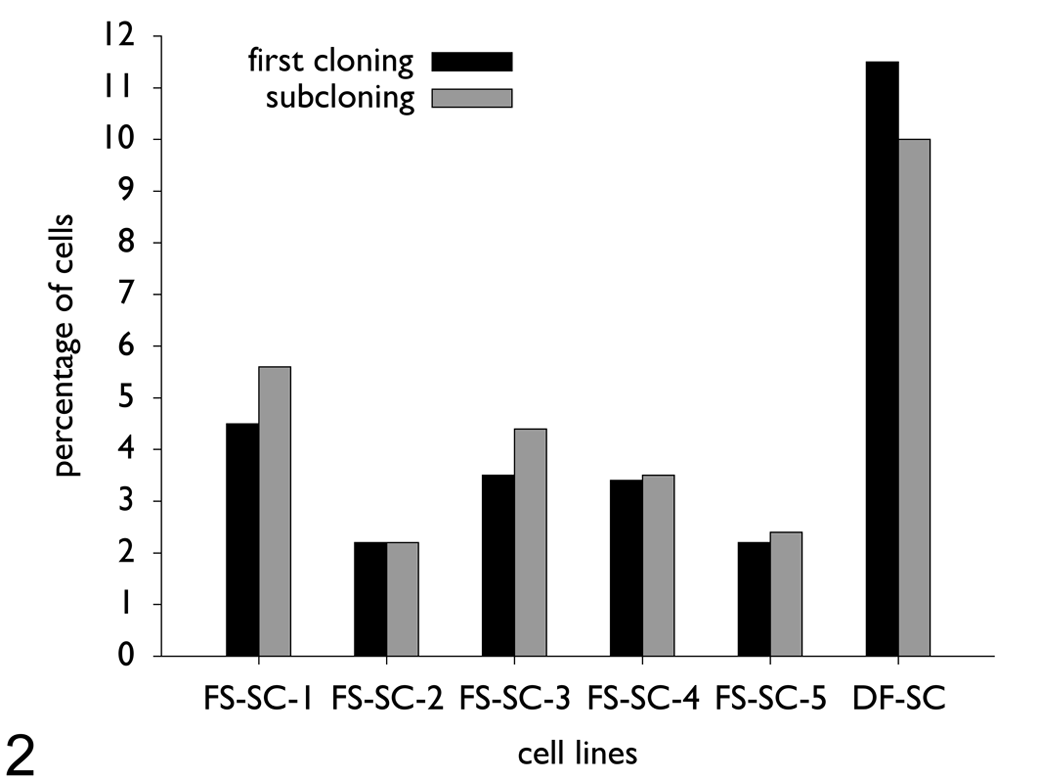
Cloning and subcloning efficiency of fibrosarcoma cell lines and control cell line. The cloning efficiency of the tumor cell lines varied considerably from the control cell line. The monoclonal fibrosarcoma cell lines (FS-SC-1 to FS-SC-5) showed an average percentage of clones of 3.4%, whereas a threefold-higher efficiency (11%) was present in the control cell line (DF-SC).
Cytogenetic Analysis
Chromosome analysis of the 5 subcloned feline fibrosarcoma cell lines
The feature of chromosomal instability (aneuploidy) was examined separately from numerical centrosome aberrations. The comparison of the karyotypes of all 5 feline subcloned fibrosarcoma cell lines, in part from different passages (FS-SC-2, FS-SC-4, and FS-SC-5), revealed an extensive karyotype instability (Table 2 ). The chromosome number per cell (19 to 155) deviated substantially from the normal species-specific number (2n = 38). Numbers of chromosomes per cell of the monoclonal feline fibrosarcoma cell lines and the control cell line are shown in Table 2.
Number of Chromosomes per Cell of the Monoclonal Feline Fibrosarcoma Cell Lines and the Control Cell Line a
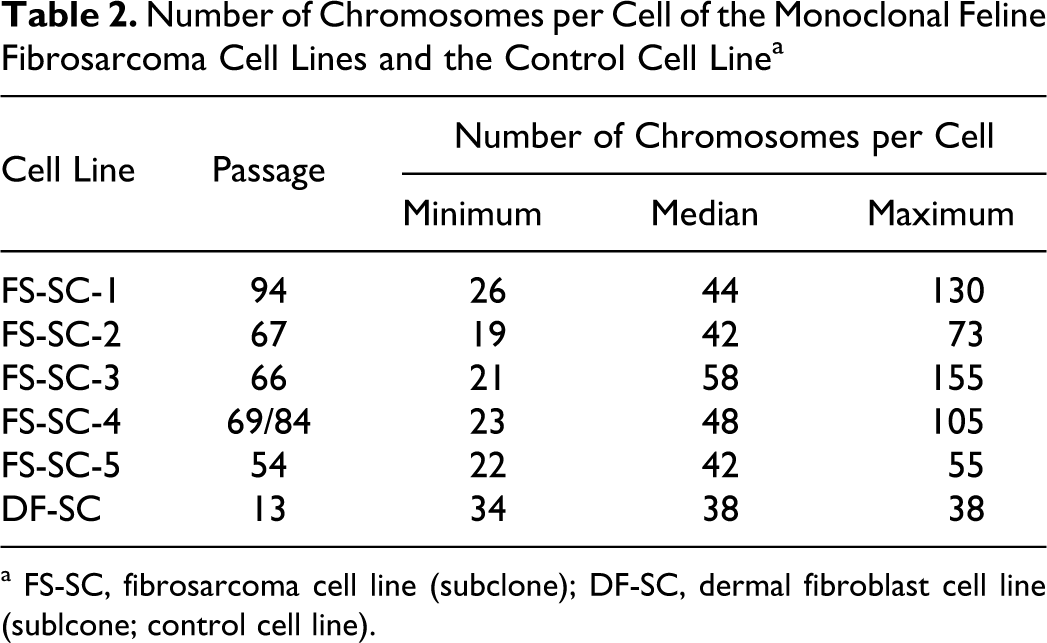
a FS-SC, fibrosarcoma cell line (subclone); DF-SC, dermal fibroblast cell line (sublcone; control cell line).
In Figures 3–6, examples of Giemsa-stained metaphase spreads of fibrosarcoma cell line FS-SC-1, passage 118, and the control cell line (DF-SC, passage 13) are shown. In addition to the metaphase spread of feline fibrosarcoma cell line FS-SC-1, a normaleudiploid metaphase spread (2n = 38) of the control cell line is shown.
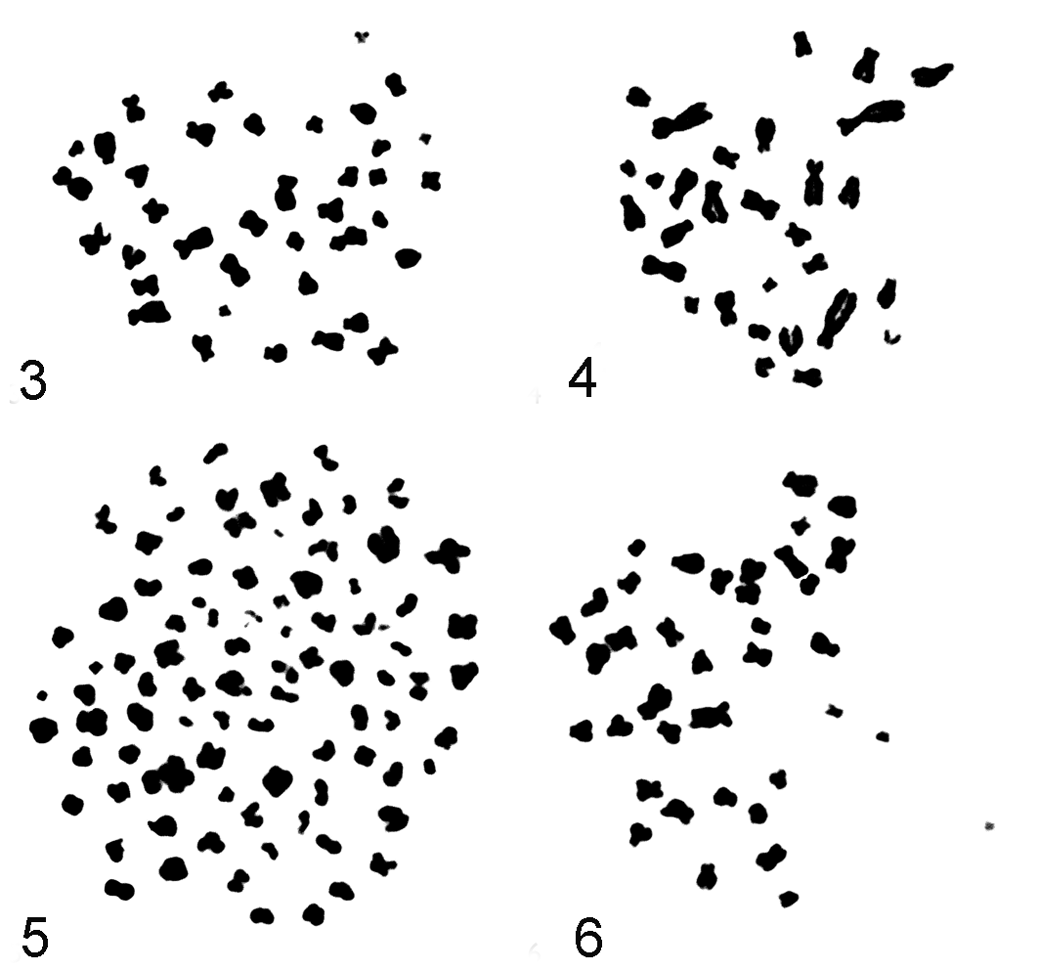

Examples of Giemsa-stained metaphase spreads of monoclonal fibrosarcoma cell line FS-SC-1, passage 118, and the control cell line (DF-SC, passage 13). FS, fibrosarcoma; SC, subclone; DF, dermal fibroblast (control).
Chromosomal analysis of cell line FS-SC-3 revealed an extensive chromosomal instability. On the whole, 112 metaphases from passage 53 have been evaluated by light microscopy. In Figure 7, the distribution of the chromosome number per cell is shown for FS-SC-3. In conclusion, the chromosome number per cell was not stable. None of the clonal cell lines had a constant number of chromosomes per cell; on the average, a clearly hyperdiploid status existed. In contrast, karyotyping of the control cell line at passage 13 showed predominantly a diploid karyotype of 2n = 38. There were significant differences in the median chromosome number per cell between all cell lines (P < .0001, Kruskal–Wallis test; Fig. 8). The Levene test, which examined the equality of variances of the chromosome numbers of the different cell lines, revealed significant differences (P < .0001).
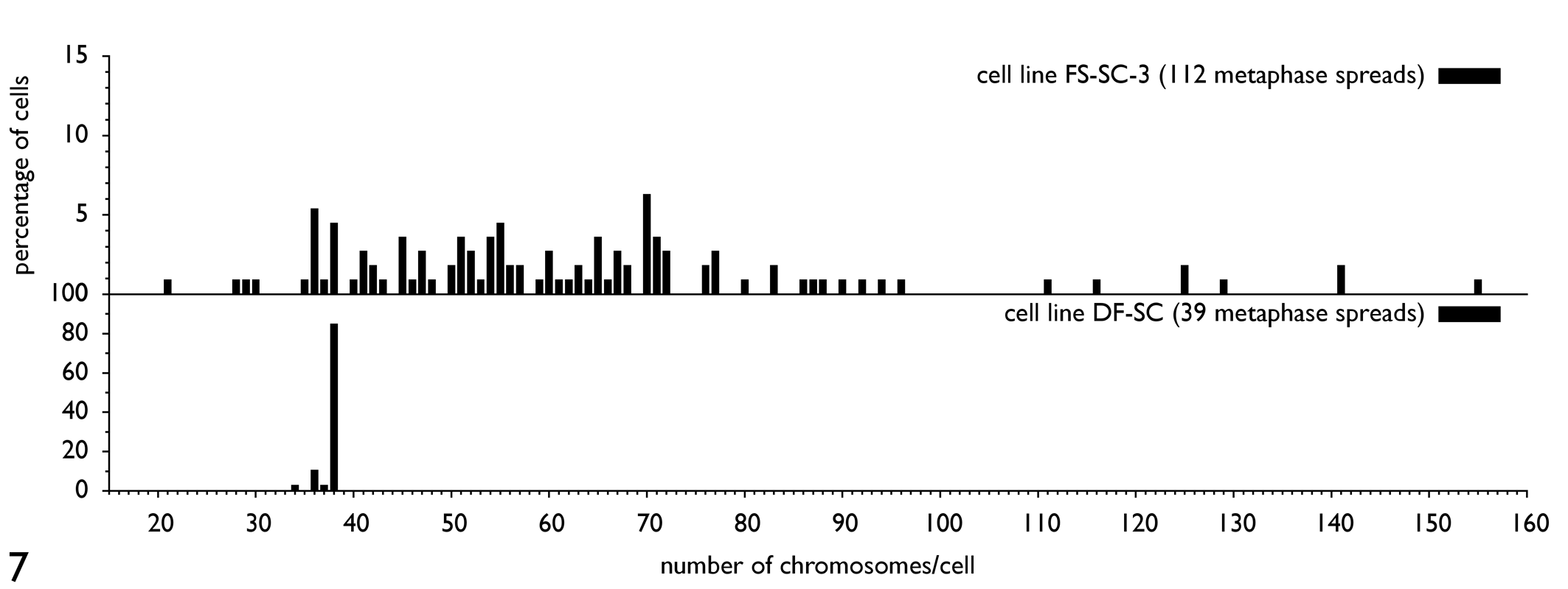
Distribution of the chromosome number per cell in the fibrosarcoma cell line FS-SC-3, passage 53, and the control cell line (DF-SC, passage 13). The control cell line showed a normal feline eudiploid spread of chromosomes per cell. Cell line FS-SC-3 revealed an extensive chromosome number alteration, with 21 to 155 chromosomes per cell. In sum, 112 Giemsa-stained metaphase spreads of fibrosarcoma cell line FS-SC-3 and 39 metaphase spreads of the control cell line (DF-SC) have been examined. FS, fibrosarcoma; SC, subclone; DF, dermal fibroblast (control).
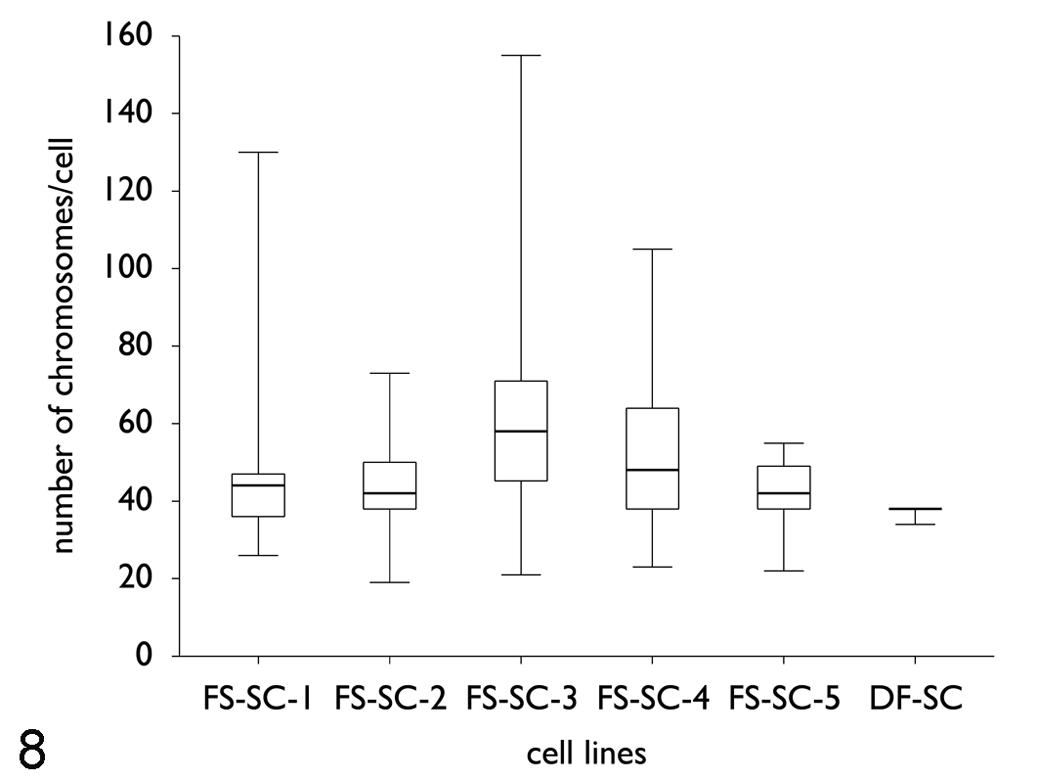
Box and whisker plots of the chromosome numbers per cell in monoclonal fibrosarcoma cell lines (FS-SC-1, P118; FS-SC-2, P28/37/65; FS-SC-3, P53; FS-SC-4, P21/66; FS-SC-5, P10/20/27/44) and the control cell line (DF-SC, P13) after cloning. In all feline fibrosarcoma cell lines, the number of chromosomes deviated abnormally from the normal feline chromosome number of 2n = 38, ranging from 19 to 155 chromosomes per cell. In contrast to that, the control cell line was eudiploid (2n = 38). Between all fibrosarcoma cell lines, significant differences of the mean number of chromosomes were observed (P < .0001, Kruskal–Wallis test). FS, fibrosarcoma; SC, subclone; DF, dermal fibroblast (control); P, passage.
Between and within every fibrosarcoma cell line, an extreme heterogeneity of karyotypes was observed. Between all fibrosarcoma cell lines, significant differences of the mean number of chromosomes were observed (P < .0001, Kruskal–Wallis test). The pairwise Nemenyi test, to compare the mean chromosome number of each fibrosarcoma cell line with every other fibrosarcoma cell line and with the control cell line, showed significant differences (P < .0001) between all cell lines.
Centrosome Analysis
Centrosome hyperamplification was found to be present in all feline cell lines derived from feline fibrosarcoma. Centrosomes were examined by immunostaining of γ-tubulin (mouse monoclonal antibody, CLONE GTU-88) and visualized by Cy3-conjugated goat anti-mouse IgG. As determined by immunohistochemistry, all fibrosarcoma cell lines displayed hyperamplification of centrosomes, with 5.7 to 15.2% of the cells having more than 2 centrosomes per cell. Those centrosome numbers ranged from 3 to 8 centrosomes per cell, whereas only 0.6% of the control cells revealed 3, but never more, centrosomes per cell (Figs. 9–13 ).
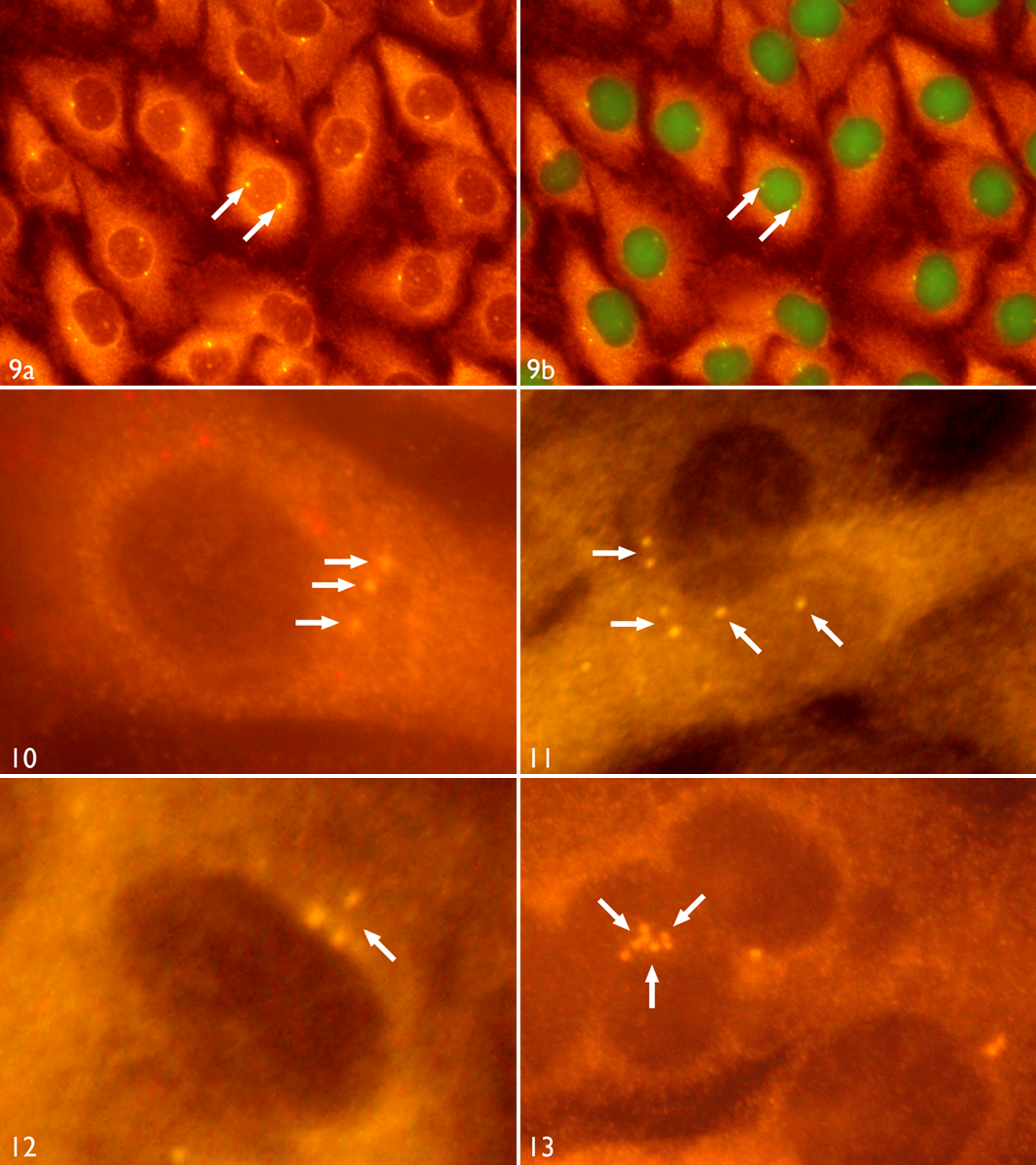
Demonstration of centrosomes in monoclonal feline fibrosarcoma cell lines. FS, fibrosarcoma; SC, subclone; P, passage.
On the whole, 151 cells of each fibrosarcoma cell line and also the control cell line were examined by fluorescence microscopy. In comparison, all cell lines revealed significant differences (Fig. 14 ) in their centrosome number per cell (P < .0001, Kruskal–Wallis test).
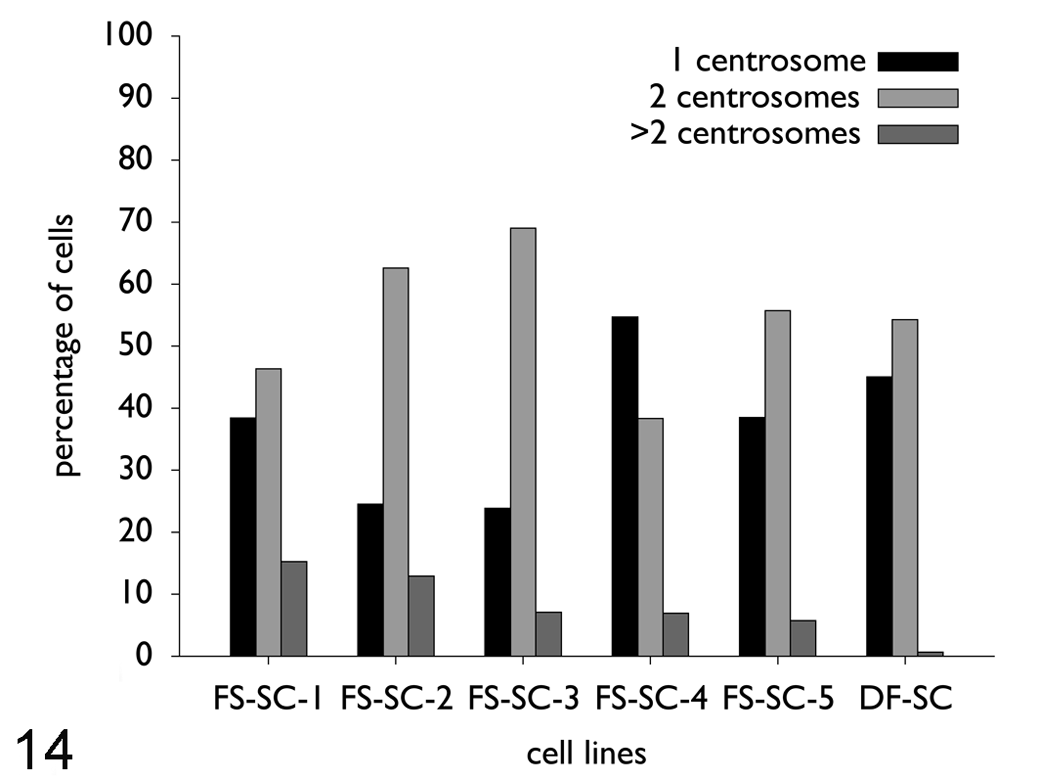
Number of centrosomes per cell in the monoclonal fibrosarcoma cell lines (FS-SC-1, P134; FS-SC-2, P67; FS-SC-3, P66; FS-SC-4, P69/84; FS-SC-5, P54) and the control cell line (DF-SC, P13). On the whole, 151 cells of each cell line were evaluated. About 5.7 to 15.2% of the cells had more than 2 centrosomes per cell. Abnormal centrosome numbers ranged from 3 to 8 centrosomes per cell. Only 0.6% of the control cells revealed 3, but never more, centrosomes per cell. FS, fibrosarcoma; SC, subclone; DF, dermal fibroblast (control); P, passage.
All cell lines revealed significant differences in their centrosome number per cell (P < .0001, Kruskal–Wallis test). At least 151 cells were examined for each cell line.
Correlation Between Aneuploidy and Centrosome Hyperamplification
No significant statistical correlation could be demonstrated between karyotype instability (aneuploidy) of the feline fibrosarcoma cell lines and centrosome hyperamplification (rs = .154, P = .80).
Discussion
In this study, a “clonal cell line” is defined as a cell line that originates from a single cell. The chromosome analysis of all 5 fibrosarcoma cell lines revealed before, as well as after, cloning a widespread variability in chromosome number (aneuploidy). This observation supports our hypothesis that every cell of a fibrosarcoma cell line has an instable karyotype and, consequently, a permanent missegregation of chromosomes occurs. In contrast, the control cell line revealed in 85% of the examined cells a eudiploid karyotype (2n = 38). Only 15% of control cells displayed a hypodiploid karyotype with 34 to 37 chromosomes per cell. This apparent hypodiploidy can be explained by a possible preparation artifact in the last step of the chromosome preparation.2,25
In the last step of the chromosome preparation, the fixed chromosomes are dropped on glass slides. During this procedure, several chromosomes can move away from the metaphase, and a pseudohypodiploid metaphase spread is to be seen. However, on the basis of the control cell results, the preparation artifact is supposed to be, at most, 15% because 85% of examined control cells were eudiploid. Therefore, the extreme karyotype instability of all fibrosarcoma cell lines, typically exceeding the normal chromosome number in most cells, cannot be explained by preparation artifacts. A hyperdiploid chromosome number was never observed in the control cell line. All 5 fibrosarcoma cell lines showed a remarkably lower cloning efficiency (3.4%) than the control cell line (11%). It is likely that the high karyotype instability of each fibrosarcoma cell is responsible for this phenomenon.
Chromosomal heterogeneity was observed in all 5 feline fibrosarcoma cell lines associated with centrosome hyperamplification (>2 centrosomes per cell). Karyotype instability is thought to arise as a result of aberrations in mitotic checkpoints. A possible reason that all feline fibrosarcoma cell lines had heterogenous chromosome profiles may be attributed to centrosome aberrations as an important cause of chromosome missegregation and aneuploidy.5,12,14,29,33,52 All 5 fibrosarcoma cell lines showed in 5.7 to 15.2% of the cells an extensive centrosome hyperamplification with 3 to 8 centrosomes per cell, while only 0.6% of the fibroblast control cells revealed a hyperamplification of centrosomes with not more than 3 centrosomes per cell.
Concluding, the feline fibrosarcoma cell lines revealed 10 to 25 times as many cells with centrosome hyperamplification as the control cells. These findings correspond well with data from tumor cell lines and solid tumors in humans—for example, in mammary gland carcinomas, carcinomas of the prostate, colon carcinomas, and hepatocellular carcinomas.1,5,12,30,41,42
Centrosome hyperamplification was also described for several canine tumors (eg, myxosarcoma, osteosarcoma, chondrosarcoma). Setoguchi et al 46 found in 9.5 to 48.1% of the examined tumor cells centrosome hyperamplification (> 2 centrosomes/cell). Similar characteristics have been found in feline lymphoma cell lines with 10 to 30% of the lymphoma cells possessing 3 or more centrosomes per cell. 40
It has not been entirely evaluated if the role of numerical centrosome aberrations generates chromosomal instability. Several studies, conflicting with our data, showed a direct role of centrosome hyperamplification and chromosomal instability by statistical correlation.17,34 It has been demonstrated that centrosome hyperamplification occurs in malignant mesenchymal tumors as osteosarcomas. 1 To our knowledge, this is the first study examining and characterizing feline fibrosarcoma cell lines by chromosomal instability and numerical centrosome aberrations. However, similar to our findings with feline fibrosarcomas, other malignant epithelial and mesenchymal tumors demonstrated no correlation of aneuploidy and amplification of centrosomes.18,47 We concluded that numerical centrosome aberrations alone may not be responsible for the formation of aneuploidy in feline fibrosarcoma cell lines. In summary, we demonstrated the occurrence of aneuploidy associated with aberrations in centrosome number in 5 feline fibrosarcoma cell lines.
On the whole, we strongly hypothesize that the observed chromosomal instability of all feline fibrosarcoma cell lines are related to numeric centrosome alterations. However, several spontaneous mutations (eg, spontaneous chromosome instability) during in vitro evolution can never be excluded completely in permanent growing cell lines.
Although a significant correlation between karyotype instability and centrosome hyperamplification could not be demonstrated statistically, these findings contribute to a further cytogenetic characterization of feline fibrosarcomas to identify factors regulating centrosome function and centrosome duplication cycle. The causes and consequences of numeric centrosome abnormalities in feline fibrosarcomas require further investigation of factors regulating centrosome function and duplication cycle.
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Corina Löhberg-Gruene received a postgraduate grant from Justus-Liebig-University Giessen, Giessen, Germany.
