Abstract
The authors have determined a consensus sequence for exons 1 and 2 of H-ras from captive lemurs and lorises and evaluated samples of nonneoplastic liver and hepatocellular carcinomas (HCC) from affected animals for mutations in these exons. Frozen liver samples were collected from 20 animals representing 9 different species with a sex distribution of 10 males and 10 females. A total of 26 liver samples, including 11 normal livers, 9 HCC, and 6 samples from nonneoplastic regions of liver from animals with HCC, were evaluated. This is the first report of the consensus sequence for exons 1 and 2 of H-ras in prosimians, and the authors have determined that it is identical to that of human H-ras and differs only slightly from the chimpanzee sequence. Point mutations were identified in 6 of the 9 HCC samples examined with codons 7, 22, 32, 56, 61, 84, and 96 affected. Two carcinomas had double mutations, and one tumor had triple mutations. One HCC had a mutation in codon 61, which is identical to a recognized affected codon for an H-ras “hot spot” in rodent neoplasia that has also been reported in human tumors. Although not statistically different, metastasis occurred in 5 of 6 HCC with H-ras mutation and only 1 of 3 HCC without mutations. There were 4 silent mutations that did not contain changes in the encoded amino acids, 2 of which were found in nonneoplastic regions of tumor-bearing liver.
Introduction
Lemurs and lorises are members of the suborder Prosimii, the most primitive members of the primate family. 20 Lemurs are native only to the island of Madagascar, and lorises live in Southeast Asia and India. 20 Most species are endangered in their natural habitat, and small breeding populations are maintained in various sites around the globe. Given the limited number of these species in captivity, little is known about their propensity to develop neoplastic disease in general and, more specifically, the pathogenesis of hepatic neoplasia.
We previously reported a series of naturally occurring hepatocellular carcinomas (HCC) in a colony of captive lemurs and lorises. 35 The incidence of HCC (14/145, 9.7%) was higher than that reported for other captive primates, 21 and the behavior was relatively aggressive with metastasis evident in approximately 50% (7/14) of cases. 35 The pathogenesis of HCC in lemurs remains unclear. There was no evidence of active hepadnavirus (hepatitis B virus family) infection in lemurs with HCC, nor was there a detectable relationship between hepatic iron levels and the incidence of liver tumors. Both hepadnavirus infection and high liver iron levels are risk factors for HCC in humans. 1
The ras family of proto-oncogenes encodes proteins with guanine triphosphate phosphatase activity. 17 These genes are involved in cellular signal transduction and survival, differentiation, and stimulation of cell proliferation primarily via the extracellular signal-related kinase/mitogen-activated protein kinase (ERK/MAPK) signal transduction pathway in health and in human cancers. The family of ras genes includes H-ras, N-ras, and K-ras. Mutations in ras family genes are recognized in both human and animal HCC. 29 Activated ras family genes with single-point mutations in or near codons 12, 61, and 117 have been demonstrated in chemically induced rodent and human neoplasms. 8,9,22,33,34 Mutated H-ras clearly has a role in murine hepatocarcinogenesis as mice with experimentally induced mutations in both β-catenin and H-ras have a 100% incidence of liver tumor formation compared to β-catenin mutation alone in transgenic mice. 15 Mutations in H-ras are rare in human HCC. 25,30 H-ras expression has been associated with recurrence and extrahepatic metastasis of human HCC. 28 Mutations (A → T transversion) in codon 61 of K-ras and N-ras have been identified in a few human HCC, suggesting that mutation at this site may play a role in tumor formation in humans. 7
Mutations in H-ras are commonly encountered in spontaneous and chemically induced HCC of mice. 16,18,19,23,26 An incidence of 31% of H-ras mutations was described in a group of 40 separate studies of chemical carcinogenesis in mice. 16 The mutation incidence was significantly higher in spontaneous HCC than in chemically induced tumors, and the spectra of mutations differed. 16 Mouse liver epithelial cell lines have been transformed following transfection with an activated c-H-ras with a mutation in codon 12. However, the role of H-ras likely varies with the strain of mouse and the dose of chemical carcinogen used. 11,19 Within the H-ras gene, codon 61 is considered a “hot spot” for mutation in mice due to its frequent involvement in tumorigenesis. 10 H-ras mutations in codon 61 have also been reported in rats but are not common. 2,4,5,12
Because of the frequent involvement of H-ras mutation in spontaneous and chemically induced mouse but not human liver tumors, we elected to evaluate the incidence of mutation of this oncogene in a set of spontaneously developing HCC in prosimians.
Methods
Case Material
A retrospective review of cases from the Duke University Lemur Center was undertaken to identify cases of HCC for which there were frozen tumor and nonneoplastic liver available. Complete postmortem examinations were performed on animals that were found dead or euthanized due to natural causes. All animals had been maintained in accordance with the existing standards of the Duke University Institutional Animal Care and Use Committee guidelines. For each frozen sample, there was an additional representative sample collected for histopathology. These specimens were fixed in 10% neutral buffered formalin, processed into paraffin, and stained with hematoxylin and eosin to confirm the diagnoses using standards developed for canine and feline liver tumors. 24
Frozen Liver
A total of 26 liver sections were snap frozen within 1 hour of death from 20 animals (10 males and 10 females) that died or were euthanized due to natural causes. Nine animals (8 lemurs and 1 loris) had HCC at the time of death. Normal cohort samples were collected from 11 animals (9 lemurs and 2 lorises) that died of causes unrelated to liver disease. Two liver samples were obtained from the animals with HCC, one from the tumor mass and a second from a nonneoplastic region of the liver. Normal cohort samples were taken from animals that did not have HCC at the time of death (Table 1).
Frozen Lemur Liver Samples Used for Mutation Analysis
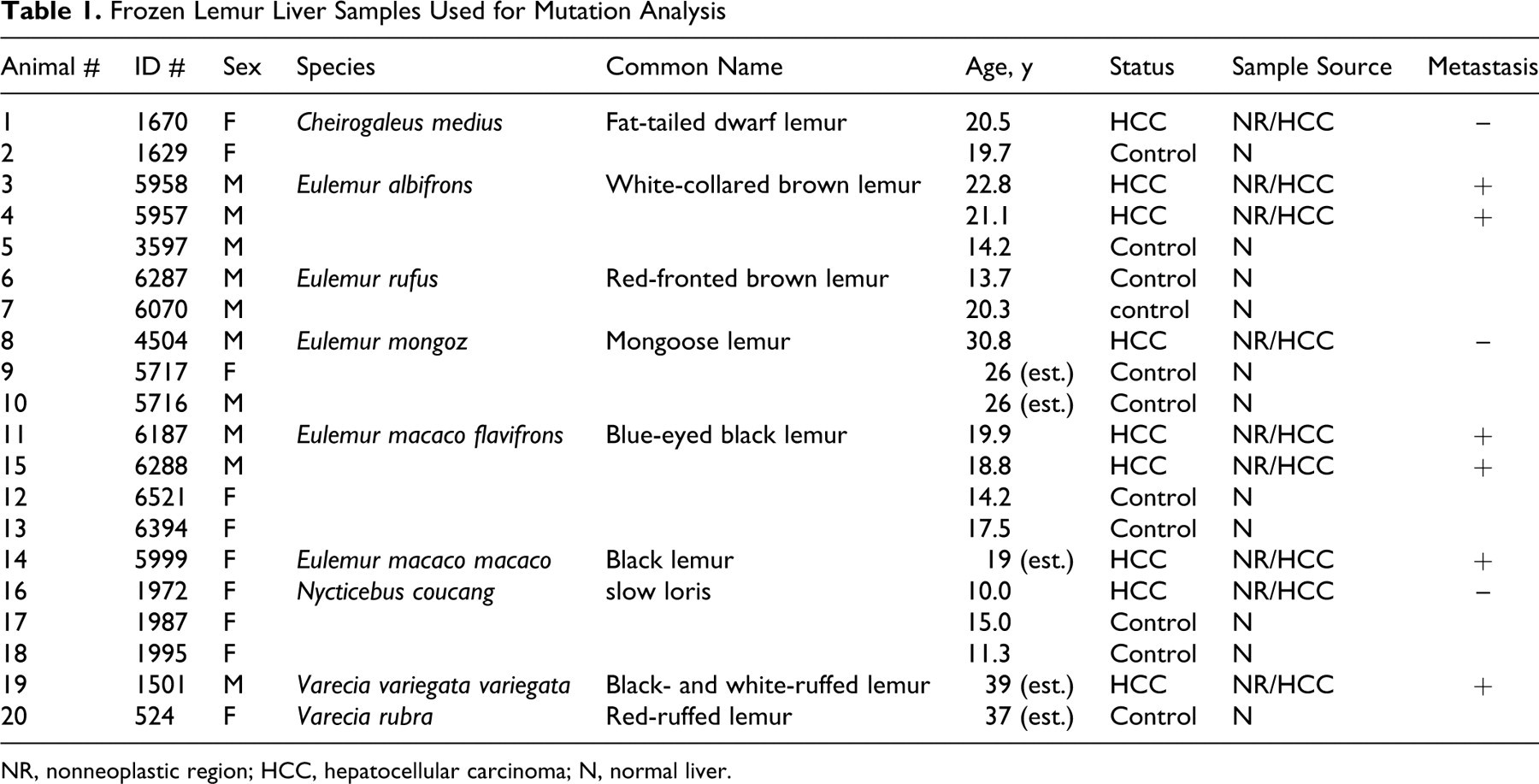
NR, nonneoplastic region; HCC, hepatocellular carcinoma; N, normal liver.
Species tested include fat-tailed dwarf lemurs (Cheirogaleus medius; n = 2), white-collared brown lemurs (Eulemur albifrons; n = 3), red-fronted brown lemurs (Eulemur rufus; n = 2), mongoose lemurs (Eulemur mongoz; n = 3), black lemurs (Eulemur macaco macaco; n = 1), blue-eyed black lemurs (Eulemur macaco flavifrons; n = 4), black- and white-ruffed lemurs (Varecia variegata variegata; n = 1), red-ruffed lemurs (Varecia rubra; n = 1), and slow lorises (Nycticebus coucang; n = 3).
DNA Isolation and PCR Amplification
DNA was isolated and extracted from normal and tumor-bearing liver with the DNeasy Tissue Kit (Qiagen, Valencia, CA). Amplification reactions were carried out by touch-down polymerase chain reaction (PCR) using the primer sets for H-ras at exons 1-2 using chimpanzee sequences from GenBank (NCBI; Table 2). Briefly, a 50-μl reaction mixture containing 100 ng of DNA template in 10× PCR buffer (200 mM Tris-HCl [pH 8.4], 500 mM KCl), 2.5 mM MgCl2, 0.6 mM dNTP, 5 μl dimethyl sulfoxide (DMSO), with each primer at 1 μM, and 1 unit of Platinum Taq DNA Polymerase (Invitrogen, Carlsbad, CA) was subjected to 30 cycles of amplification in a 9700 Thermal Cycler (PE Applied Biosystems, Foster City, CA) as follows: soak at 95°C for 7 minutes, 20 cycles at 94°C for 30 seconds each, 68°C (–0.5°C/cycle) for 30 seconds, 72°C for 1 minute, then 10 cycles at 58°C, annealing for 30 seconds, and at 72°C for 7 minutes. Human genomic DNA and controls lacking DNA were run with all sets of reactions. PCR products were purified using a QIAquick Gel Extraction Kit (Qiagen, Valencia, CA).
Primer Sets for PCR Amplification of Lemur H-ras Exons 1 and 2
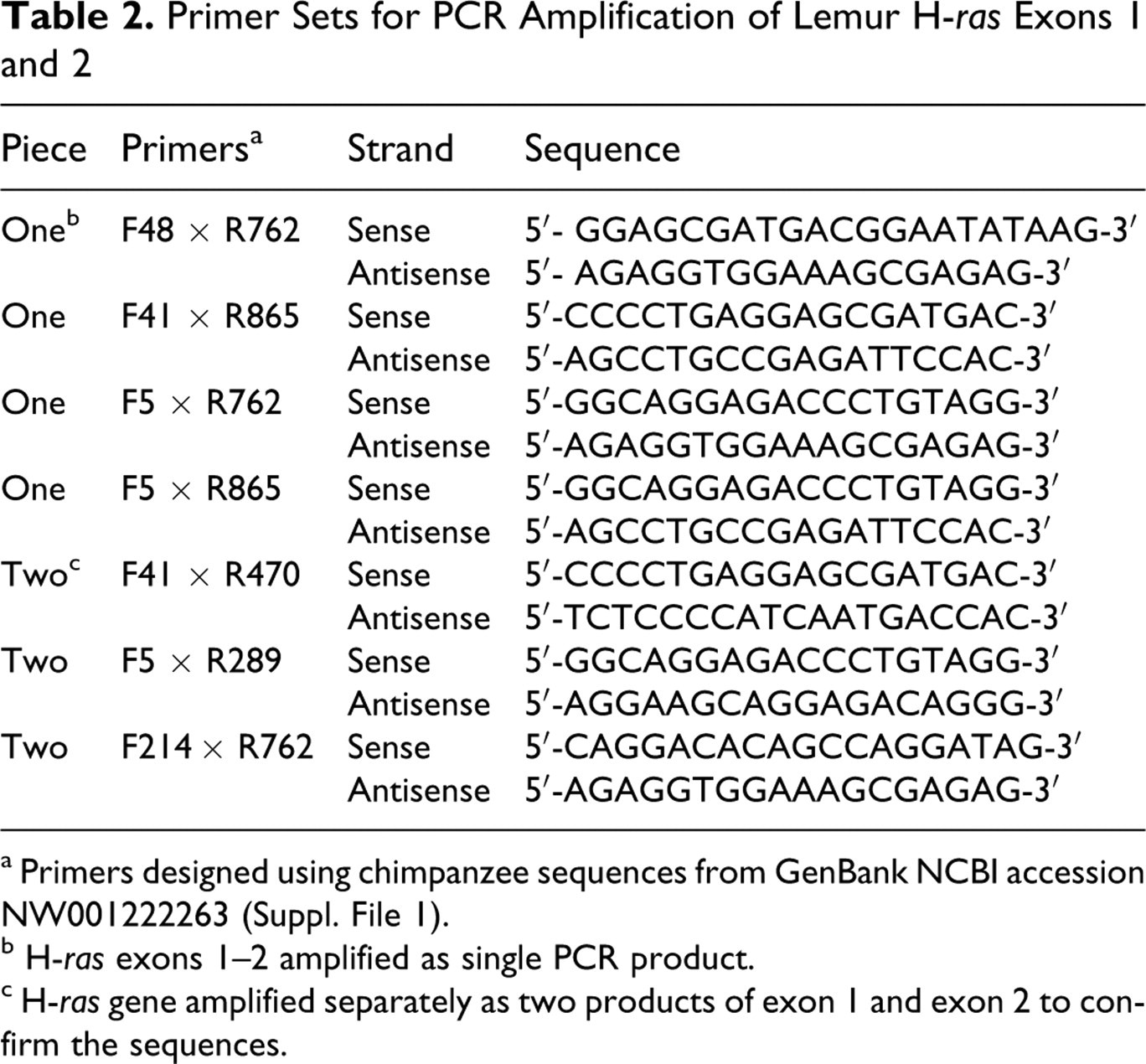
a Primers designed using chimpanzee sequences from GenBank NCBI accession NW001222263 (Suppl. File 1).
b H-ras exons 1–2 amplified as single PCR product.
c H-ras gene amplified separately as two products of exon 1 and exon 2 to confirm the sequences.
Cycle Sequencing and Autosequencing
The purified PCR products were cycled with Terminal Ready Reaction Mix-Big Dye (PerkinElmer, Foster City, CA), and then extension products were purified with DyeEx 2.0 Spin Kit (Qiagen, Valencia, CA). The lyophilized PCR products were sequenced with an automatic sequencer (PerkinElmer ABI Model 3100).
Statistics
Fisher’s exact test was calculated to determine the potential association of the presence or absence of the ras mutation and metastasis. 6 A P value of <.05 was considered significant.
Results
A consensus sequence of H-ras at codons 1 and 2 was developed using liver samples from 17 lemurs and 3 lorises representing 9 different species and 4 families of prosimians. The consensus sequence for these 2 exons was identical to the human sequences and very similar to the chimpanzee sequence, which had 6 silent mutations compared to the human and lemur sequences (Table 3). There were 4 base insertions at introns 2 and 3 in the chimpanzee sequence compared to the human and lemur sequences (Table 4).
Comparison of H-ras Sequences at Exons 1 and 2
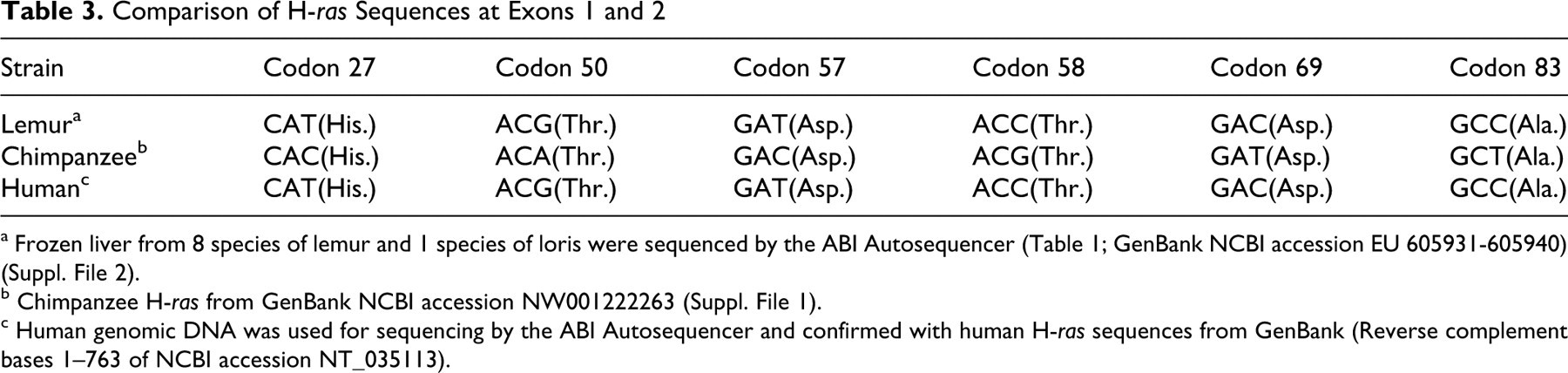

a Frozen liver from 8 species of lemur and 1 species of loris were sequenced by the ABI Autosequencer (Table 1; GenBank NCBI accession EU 605931-605940) (Suppl. File 2).
b Chimpanzee H-ras from GenBank NCBI accession NW001222263 (Suppl. File 1).
c Human genomic DNA was used for sequencing by the ABI Autosequencer and confirmed with human H-ras sequences from GenBank (Reverse complement bases 1–763 of NCBI accession NT_035113).
Comparison of H-ras Sequences at Introns 2 and 3
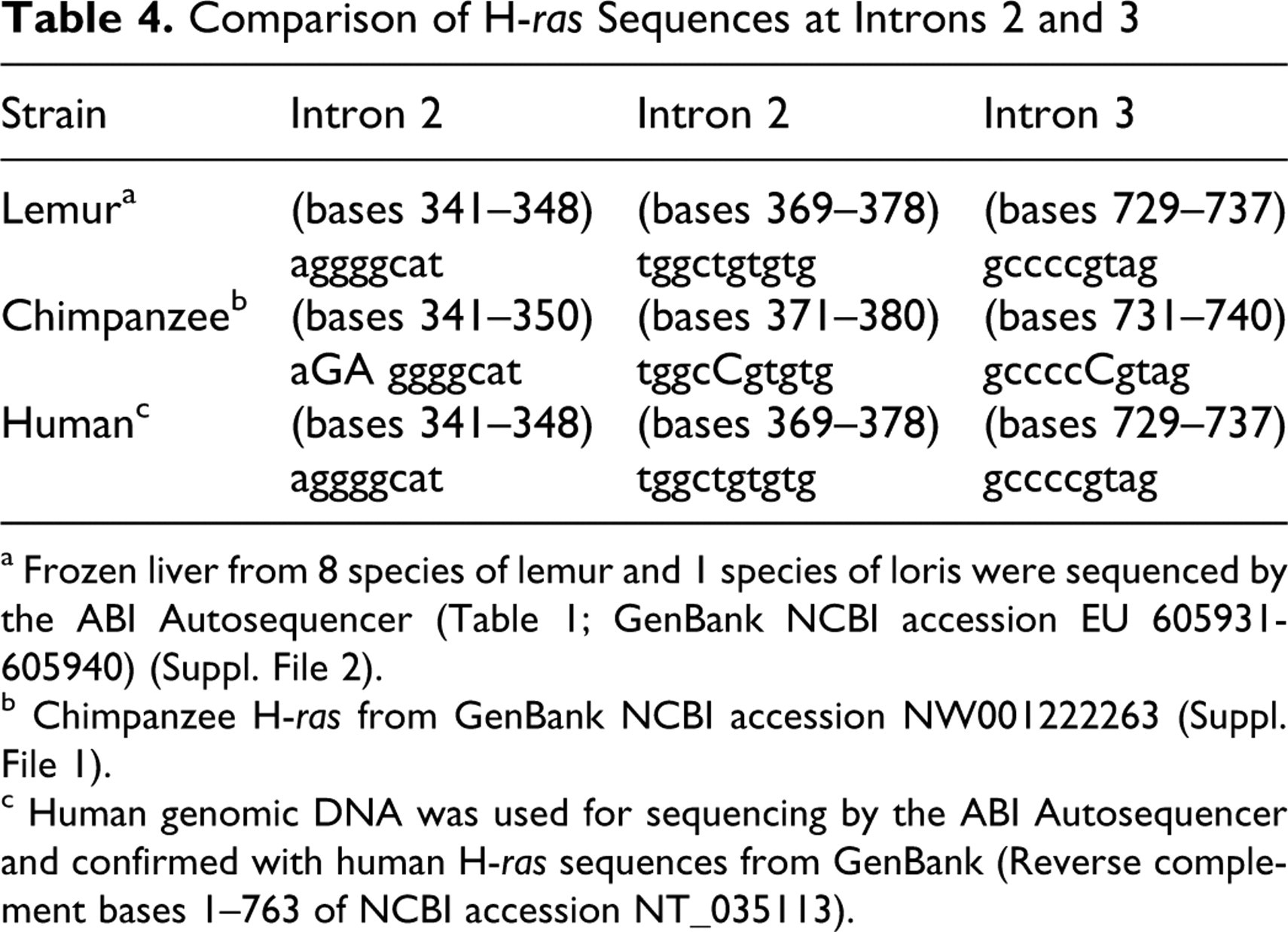

a Frozen liver from 8 species of lemur and 1 species of loris were sequenced by the ABI Autosequencer (Table 1; GenBank NCBI accession EU 605931-605940) (Suppl. File 2).
b Chimpanzee H-ras from GenBank NCBI accession NW001222263 (Suppl. File 1).
c Human genomic DNA was used for sequencing by the ABI Autosequencer and confirmed with human H-ras sequences from GenBank (Reverse complement bases 1–763 of NCBI accession NT_035113).
Point mutations in H-ras were identified in 6 of the 8 lemur HCC samples, and there were no mutations in the one loris HCC examined. The 6 animals with H-ras mutations in their HCC (animals 1, 3, 4, 11, 14, 15) had mutations within codons 7, 22, 32, 56, 61, 84, and 96 (Figs. 1–2, Table 5). Two lemurs (animals 3, 4) had 2 point mutations, and one animal (animal 11) had 3 mutations. One of the lemurs (animal 3) had a point mutation at codon 61. The point mutation was a transversion from CAG to CTG, which changes the encoded amino acid from glycine to leucine (Gln → Leu). This mutation location is identical to the “hot spot” H-ras frequently found in liver tumors of mice and reported in human HCC. This animal was one of those with pulmonary metastasis. There were 4 silent mutations (did not alter the encoded amino acids). Two of these were found in nonneoplastic regions of tumor-bearing lemurs (animals 8 and 19). The third silent mutation was the only mutation found in the one HCC that did not metastasize (animal 1), and the fourth silent mutation occurred in a metastatic HCC with 2 additional mutations that did alter the encoded amino acids (animal 11; Table 5). No mutation was detected in the normal liver.
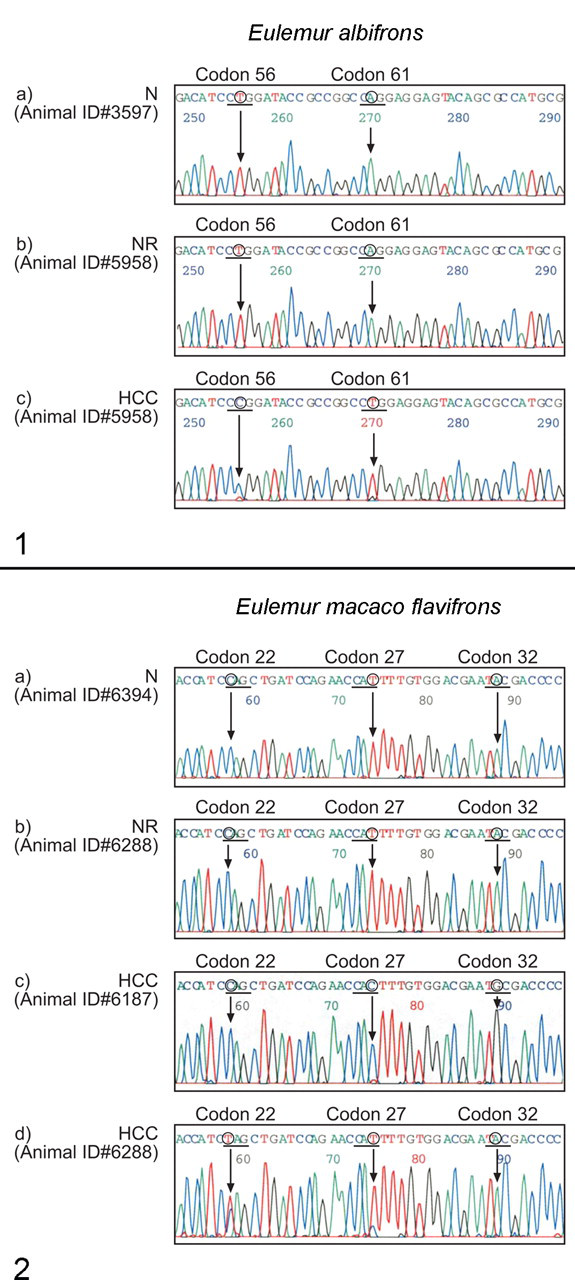
Examples of H-ras mutations at exon 2 in hepatocellular carcinoma (HCC) from animal #5958, a white-collared brown lemur (Eulemur albifrons). Sequencing panel (a) is a normal H-ras sequence from a normal (N) lemur of this species, panel (b) sequence is from a nonneoplastic region (NR) from animal #5958 with HCC, and panel (c) demonstrates mutated sequences at indicated codons 56 and 61 in an HCC from animal #5958. The mutation in codon 61 CAG → CTG (Gln → Leu) is identical to a recognized mutational hot spot for H-ras in rodents and humans.
H-ras Mutations in Lemur Liver
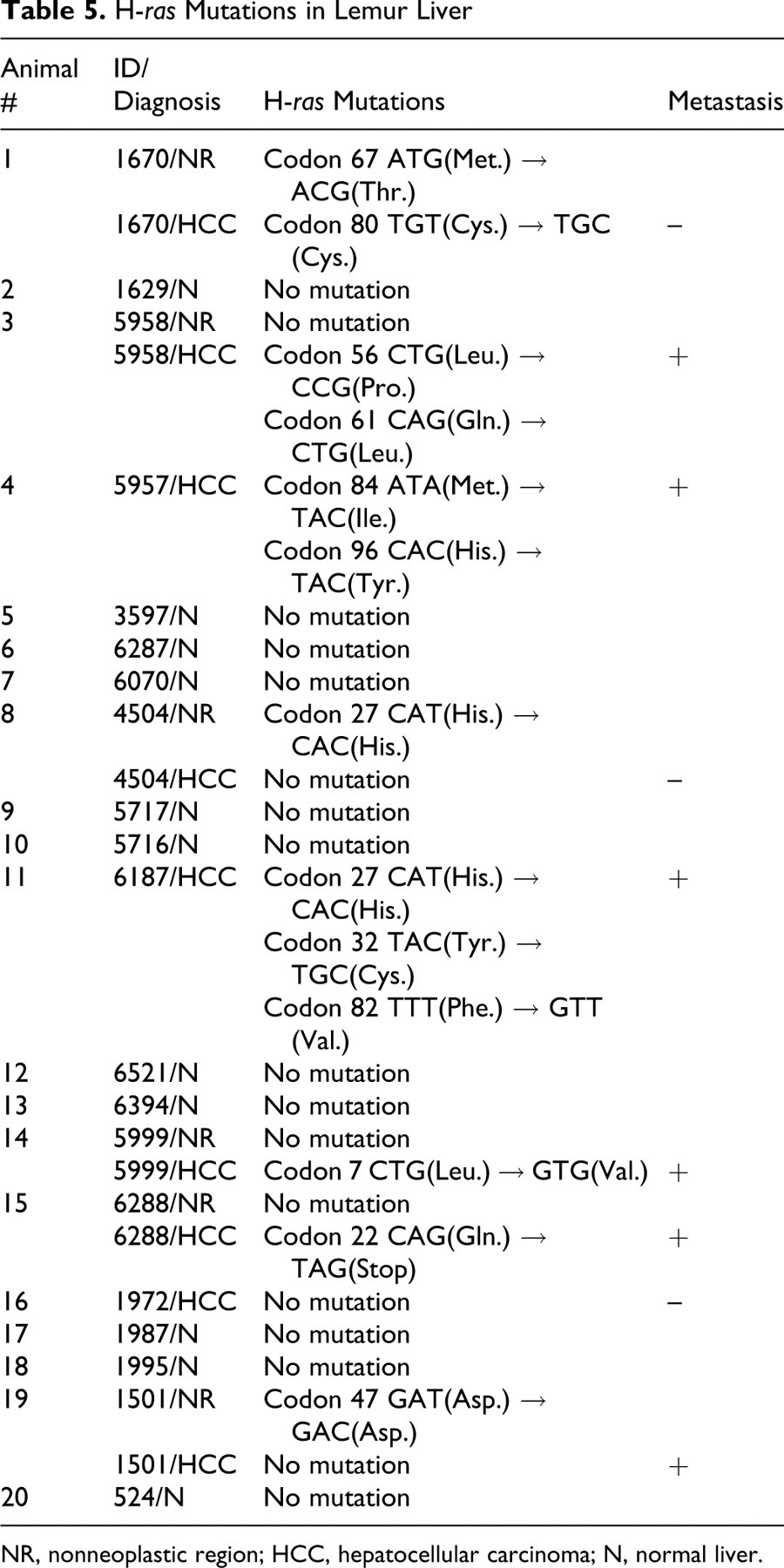

NR, nonneoplastic region; HCC, hepatocellular carcinoma; N, normal liver.
There were no differences in the histologic appearance of the HCC with point mutations in exons 1 and 2 of H-ras and those that did not have mutations. The incidence of metastasis was higher in animals with H-ras mutations, as 5 of 6 HCC with H-ras mutations metastasized to the lungs, whereas only 1 of 3 HCC without mutations metastasized (Tables 1 and 5). More than 83% (5/6) of the animals with an H-ras mutation developed metastasis. However, there was no statistically significant difference (P > .23) observed between the presence and absence of H-ras mutation and metastasis.
Discussion
To the best of our knowledge, this is the first study to determine that the consensus sequence for the prosimian H-ras gene is identical to the human sequence and was surprisingly a better match for the human sequence than published sequences for the chimpanzee. We also evaluated the incidence of mutations in H-ras in the HCC in a previously unstudied group of primates using a collection of tissue that includes primary HCC, adjacent nonneoplastic liver, and normal liver from 9 different prosimian species.
H-ras mutations were relatively common in the HCC of the primates in this study, as they are in mice. The cause of the mutations is not clear. Many of the lemurs and lorises in this study had histologic evidence of excess iron storage in the liver (hemosiderosis) in addition to high hepatic iron measurements. 35 This raises the possibility of iron-induced oxidative injury producing genetic injury, as has been suggested by several groups. 3,13,27 Although iron-induced mutations cannot be excluded, our previous analysis did not show any relationship between hepatic iron levels and the incidence of HCC, 35 nor was there a relationship between hepatic injury and iron levels in lemurs in another study. 32
There was no clear pattern of mutation sites in the samples examined, although the number of HCC available for analysis was relatively small, making patterns of mutations difficult to discern. Interestingly, one of the mutations occurred in codon 61, a recognized hot spot for rodents and a site reported as one of the few H-ras mutations in human HCC, although K-ras and N-ras, rather than H-ras, are typically affected in human HCC. 7,31 Other similarities with mice included the presence of a mutation in the second base of codon 61 (A → T) as seen in 3 murine cell lines. 14 Although more than 83% of the animals with mutation in the H-ras and 1 of 3 without the mutation developed metastasis, the difference in metastatic rate was not statistically significant. The number of animals available for study, however, was small, and a larger, more representative population may have yielded different results. The trend suggests that additional study of the relationship between the ras gene and metastasis is warranted.
Although there was no consistent pattern of mutation and the study size was relatively small, a relationship between RAS mutation and metastasis has been reported in human HCC. 28 Since many of the mutations were found in animals with HCC at end-stage disease, it is not clear whether some of the mutations are causal or due to a late tumor stage loss of genetic integrity. The functional consequences of most of the mutations identified are not clear as they occur in sites that are not well characterized in other studies. However, the determination of an identical consensus sequence for exons 1 and 2 of H-ras for prosimians and humans may provide insight into the role of H-ras mutations and their relevance to human cancer. Moreover, there was no difference in the histologic appearance of the HCC with point mutations in exons 1 and 2 of H-ras and those without mutations, which underscores the importance of examining liver tumors at the molecular level to better understand the pathogenesis of cancer. Further evaluation of these mutations and mutations in other ras family members in neoplasms of liver and other organs would be interesting given the recognized role of ras family mutations and activation in human HCC. 29 In addition, evaluation of the biology of H-ras mutations and mutations in other ras family members in prosimians and humans would be interesting given the important role of the ras family and the related ERK/MAPK signal transduction pathway in human cancers.
Footnotes
Acknowledgements
We acknowledge Dr. Jay Levine and Carie Kimbrough for advice with statistics.
The authors declared that they had no conflicts of interests with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
