Abstract
In the postgenomic era, the accumulation of massive amounts of molecular and genetic data is quickly transforming the landscape of cancer research. A deeper understanding of cancer biology will allow pathologists studying genetically engineered mouse models and spontaneous tumors to ask novel questions and provide useful insights into disease mechanisms. Although tumor suppressor gene loss is known to be an essential step in the pathogenesis of many tumors, the variety of mechanisms that lead to such loss are poorly understood. This article provides an overview of loss of heterozygosity as an important mechanism of tumor suppressor gene loss; it also emphasizes the importance of loss of heterozygosity detection as a tool for tumor suppressor gene discovery. Understanding the applications of the array of molecular techniques currently available in cancer research will enable pathologists to further contribute to the advancement the field.
Keywords
The genomic revolution has brought a new perspective to research and has dramatically changed the landscape of cancer pathology. Now more than ever before, morphologic findings can be correlated to genetic signatures. The gold standard for cancer diagnosis remains the recognition of morphologic patterns from tissues placed on glass slides. Now more than ever before, distinguishing the molecular characteristics of tumors that present identical morphology is essential to determine prognosis and best treatment. 47 Remaining updated on the fast-changing field of cancer biology is vital for pathologists working with cancer research, whether with animal models or with spontaneous tumors. The factors that lead to cancer are many, and they go beyond loss of tumor suppressor genes and activation of oncogenes. Thus, it is crucial that pathologists understand the molecular makeup of cancer cells to better define the disease state and have useful insights into disease mechanisms.
This review focuses on the genetic mechanisms that lead to loss of tumor suppressor genes other than mutations—namely, the group of events broadly defined as loss of heterozygosity (LOH). Because the observation of LOH in tumor tissues has been pivotal in the discovery of many tumor suppressor genes known to date, I discuss some of the techniques used for LOH detection. In summary, this review offers a historical perspective of LOH and an overview of the molecular techniques commonly used in the study of tumor suppressor genes. The understanding of the biology behind these techniques, as well as their applications, may help the research pathologist ask relevant questions based on morphologic findings and significantly contribute to study design and interpretation.
Coming of Age for LOH: The Concept of Tumor Suppressor Gene Is Established
In the 1970s, researchers studying cancer viruses raised an intriguing question: Genetically speaking, is the neoplastic phenotype dominant or recessive over the normal one? They found that when normal cells and cancer cells were fused, the resulting hybrid tetraploid cell lost its ability to form tumors when injected into proper hosts. 22,23 However, they also found that when the tumor was induced by a viral infection, the opposite happened—that is, the hybrid cells retained their ability to form tumors—thereby indicating that the cancer phenotype was recessive when spontaneously occurring and dominant when induced by viral oncogenes. In the case of virus-induced tumors, neoplastic transformation occurred after “1 hit” but required “2 hits” in spontaneously occurring cancers. These observations led to the hypothesis that in the genome there are genes that carry the ability to suppress tumors and that both copies of such genes had to be lost (2 hits) for spontaneous tumors to arise. These genes were soon called tumor suppressor genes, and with that, a new concept in genetics was born.
However, the tumor suppressor gene theory presumed that cancer resulted only if the 2 copies of the same putative tumor suppressor gene were inactivated by random mutation, which researchers found unlikely. Alternative explanations on how complete tumor suppressor gene inactivation might occur came through the study of the retinoblastoma tumor.
Retinoblastoma is a tumor that occurs in children and originates from photoreceptor precursors in the retina. It can be sporadic or familial. In the sporadic form, tumors are unilateral, and children do not develop other retinoblastomas after treatment and are not at a higher risk to develop other tumors later in life. Children with the familial form, however, develop multiple bilateral tumors and have a 500-fold higher risk of developing osteosarcomas during adolescence as well as other tumors later in life, when compared to the normal population. 11,16,19
The incidence rate of retinoblastoma in children with spontaneous and familial forms indicates that a single event is responsible for the familial form whereas 2 random events (2 hits) are necessary for the sporadic form to develop. 29,49 This incidence pattern agrees with the existence of a putative tumor suppressor gene, subsequently named RB, which needed to be completely inactivated for tumors to arise. In the sporadic form, 2 somatic mutations, each one affecting one allele of RB, were required to completely inactivate the gene. Conversely, children with the familial form were thought to carry a germ-line mutation of one of the alleles of the RB gene and so required a single somatic mutation affecting the remaining normal allele to develop retinoblastoma. These speculations are in line with results obtained from the cell fusion experiments mentioned above but were confirmed only after the chromosomal localization of the RB gene was established and followed by its cloning. 15,32,33,55
Definition and Mechanisms of LOH
Geneticists' struggle with the extremely low probability of 2 random mutations happening within the same cell population (calculated as 10–12) led to alternative explanations for the occurrence of 2 nonfunctional RB alleles in the same cell in the case of the sporadic form of retinoblastoma. One explanation predicted that if the first allele of a tumor suppressor gene such as RB was lost by a random mutation, then the loss of the second allele would occur through another mechanism, such as homologous recombination. Because recombination between homologous chromosomes was known to occur almost exclusively in meiosis, the term mitotic recombination was proposed for recombination that occurs during mitosis (Fig. 1 ). Because the first random mutation resulted in a heterozygous cell for the RB gene, the second event, the mitotic recombination, would result in the LOH of RB—hence, the term LOH. 41
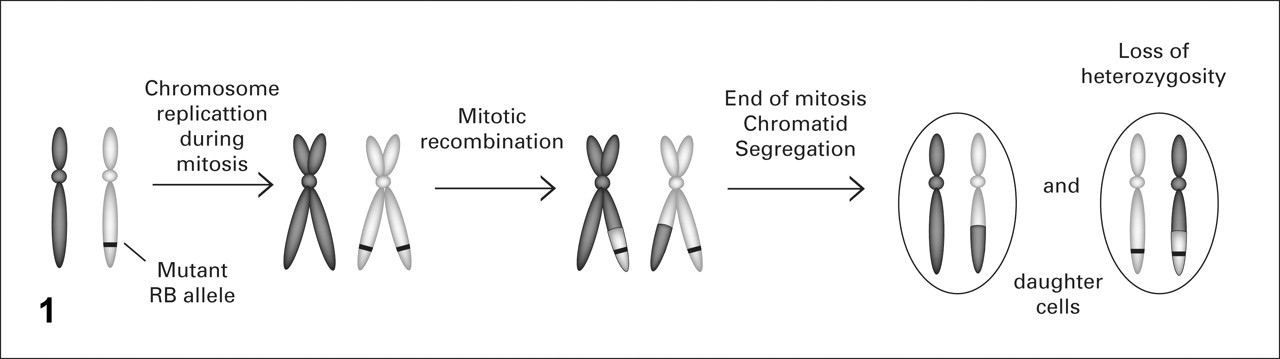
Mitotic recombination. During the S phase of mitosis and after chromosome duplication, stretches of DNA can be exchanged between the paired chromatids from both parental chromosomes. In this manner, a mutant gene can become part of both parental chromosomes and end up in the same daughter cell, resulting in loss of heterozygosity.
Later, LOH was found to occur through other mechanisms as well, such as gene conversions and interstitial deletions. 37 Gene conversion can lead to the loss of chromosome segments during DNA replication when the DNA strand being elongated forms a hybrid with its complementary strand present on the homologous chromosome. As a consequence, the DNA polymerase will use the second, homologous strand as a temporary template until the pairing is undone (Fig. 2 ). This jump from one strand to the other results in the replacement of whole stretches of DNA in a chromosome with DNA containing information from its homologous chromosome, which can lead to LOH of tumor suppressor genes and to loss of large amounts of flanking DNA. Consider the example of a cell carrying the RB tumor suppressor gene with one mutant allele. If chromosomal recombination occurs during mitosis and the DNA polymerase uses the mutant RB allele as a template during replication, then the daughter cells will have suffered LOH of RB. In interstitial deletions, whole segments of chromosomes are deleted, and flanking chromosomal regions are rejoined. This type of genetic error usually involves double-strand DNA breakage and the nonhomologous, end-joining apparatus 21 and can account for the loss of a chromosomal region carrying the remaining normal allele of a tumor suppressor gene leading to LOH.
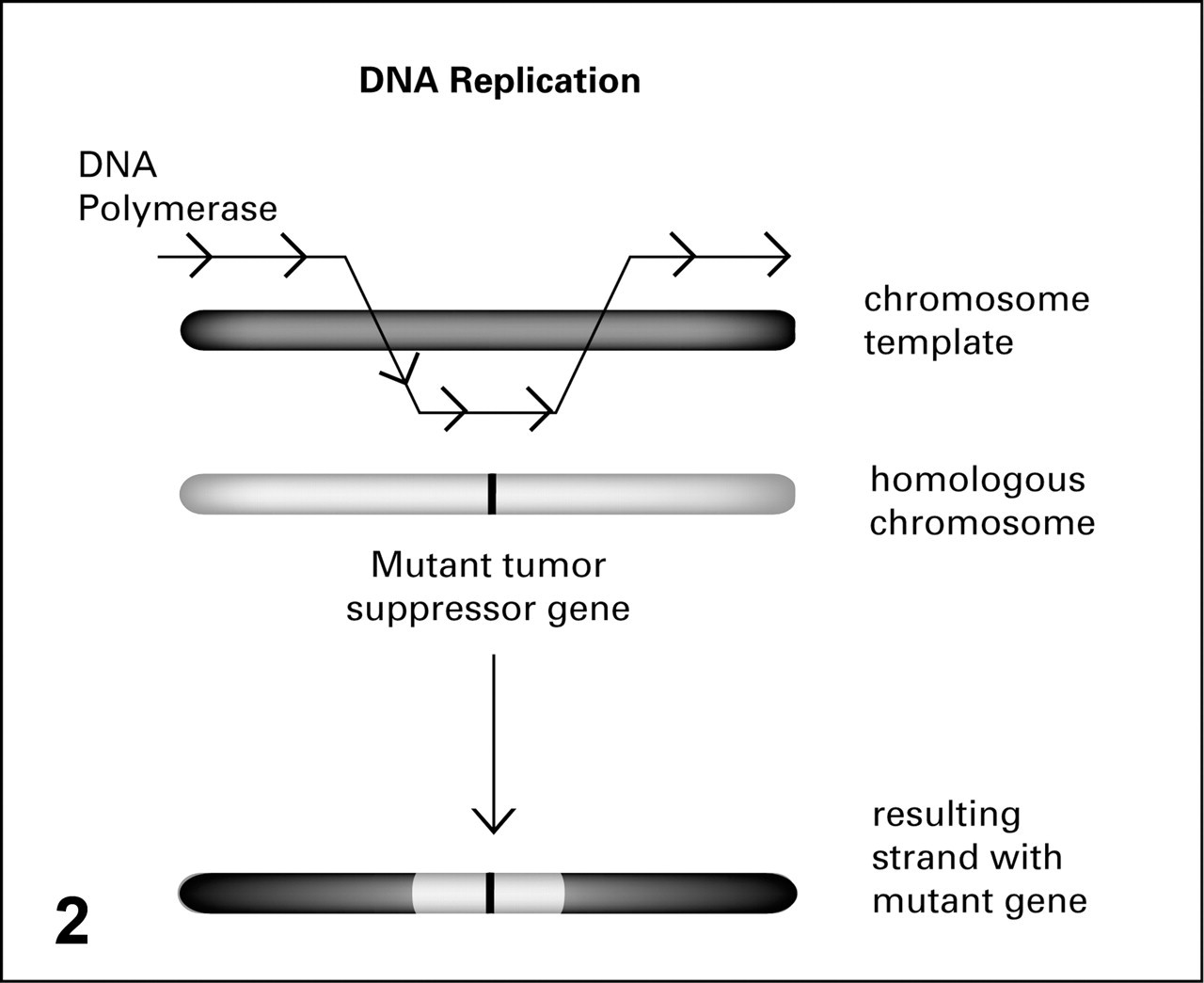
Gene conversion. When DNA polymerase jumps from the chromosome template being duplicated to its homologous chromosome, a large portion of the homologous gene can serve as an alternate template, resulting in a hybrid strand potentially carrying a mutant gene and resulting in loss of heterozygosity for the daughter cell.
Other processes can lead to LOH, such as loss of whole chromosomes owing to mitotic nondisjunction (when sister chromatids or chromosome pairs fail to separate during mitosis) or defective chromosomal segregation (when daughter cells fail to receive an equal number of chromosomes during mitosis). In both cases, the daughter cells are aneuploid instead of diploid because they carry only one chromosome of a certain pair. If the cell survives in this hemizygous state 1,30,38 and happens to carry the mutant allele of a tumor suppressor gene in the remaining unpaired chromosome, then LOH has taken place. The unpaired chromosome can be duplicated in the cell, leading to 2 identical chromosomes, both lacking a functional tumor suppressor gene. Translocation—the recombination between fragments of nonhomologous chromosomes—and other karyotypic changes can result in LOH. 51 In summary, independent of the mechanism involved, LOH happens whenever the remaining wild-type allele of a tumor suppressor gene is lost or inactivated.
Find LOH in Tumor Cells and You May Find a Tumor Suppressor Gene
LOH seemed to be a plausible mechanism to explain the loss of the remaining tumor suppressor gene allele in multistep carcinogenesis; thus, detection of LOH became the chosen strategy to discover new tumor suppressor genes. The strategy initially concentrated on polymorphic genetic markers.
Certain polymorphic stretches of DNA can be used as genetic markers because they are inheritable, they occur frequently throughout the genome, and they are highly variable among individuals. This means that an individual will have 2 versions of a certain polymorphic DNA sequence, one inherited from each parent. The chances of being homozygous for such variable fragments are very low. 46
The ability of restriction enzymes to cut DNA at specific sites and produce fragments of variable lengths, depending on the DNA sequence, allowed the detection of polymorphic fragments so named restriction fragment length polymorphisms (RFLP; aka, “rif-lip”). For example, researchers could apply a restriction enzyme such as EcoR1, which can cleave DNA at only the GAATTC sequence, to DNA extracted from a tumor and from normal tissues of the same patient. Assume that this patient has a polymorphic DNA sequence with a GAATTC sequence (EcoRI cleavage site) in one strand and a GGATTC sequence in its homologous strand. Once treated with EcoRI, normal DNA from this patient gives rise to 2 sizes of fragments because the enzyme can only cleave one of the strands. If the DNA in the tumor is mostly intact, then the fragments obtained from the cleavage are of identical length in both tumor and normal tissues. When DNA fragments obtained from cleaved tumor DNA are of different lengths than those obtained from cleaving normal DNA with the same enzyme, what researchers suspect is mutation or deletion of a tumor suppressor gene containing that polymorphism (Fig. 3 ). The survey of RFLPs by cleaving DNA from tumor and normal tissues with an assortment of restriction enzymes and by comparing the sizes of the fragments on a Southern blot represented the first step into a systematic search for tumor suppressor genes because LOH-rich chromosome regions in tumors were considered likely to harbor putative tumor suppressor genes. 3,24,53
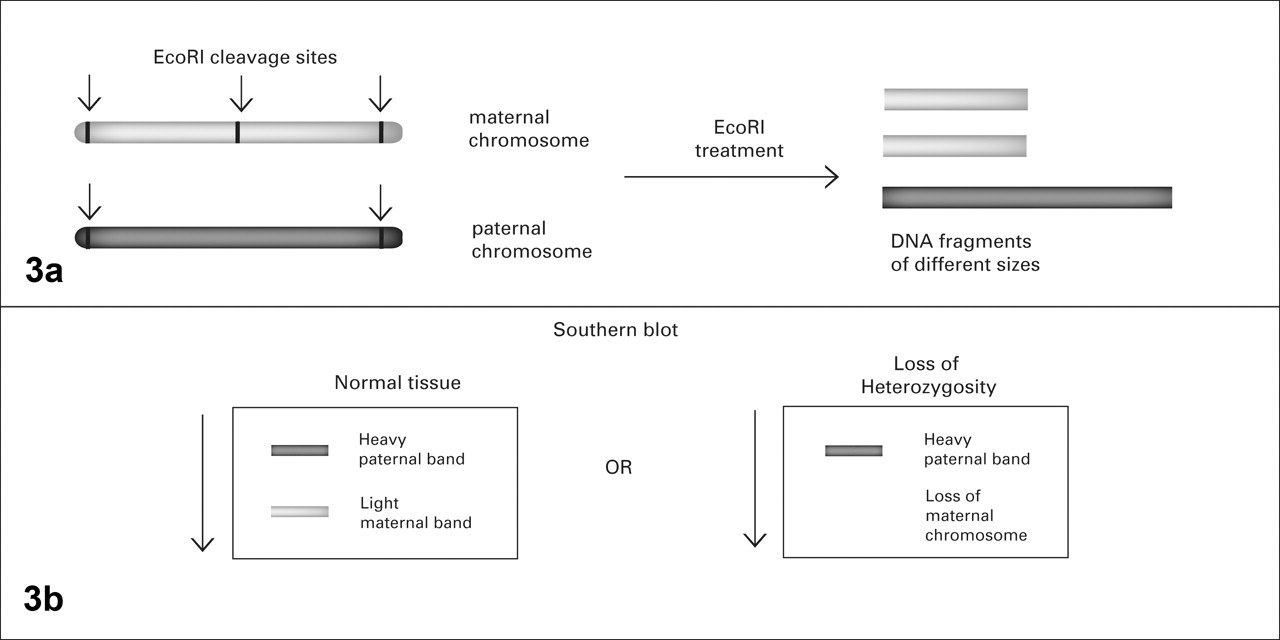
Restriction fragment length polymorphism. A, restriction enzymes, such as EcoRI, recognize specific DNA motifs that may not be present in regions of paternal and maternal chromosomes owing to nucleotide polymorphisms. The resulting cleaved fragments will then vary in length. B, cleaved DNA from heterozygous cells will resolve as different bands on a cellulose membrane. Loss of heterozygosity will be noted by the absence of a band.
With the establishment of the polymerase chain reaction (PCR) in the late 1980s, the RFLP technique was used with PCR or sometimes replaced by it. 31,36,43 –45 The ability to amplify specific segments of DNA via PCR made the survey of polymorphic regions much easier. For example, instead of treating normal and tumor DNA from the patient mentioned above with EcoR1, researchers could survey the same polymorphism by designing PCR probes to amplify sequences containing both GAATTC and GGATTC. Normal DNA would produce the 2 bands because the polymorphic region is intact, but tumor DNA would produce only one band on an agarose gel.
Polymorphic DNA regions may contain a variable number of tandemly repeated nucleotide sequences, of which the most common types are the minisatellites (9 or more base pairs [bp]) and shorter microsatellites (fewer than 9 bp, usually around 5). The term short tandem repeats is often used interchangeably with microsatellites (2–6 bp). Another polymorphism studied in many fields of genetics is the single-nucleotide polymorphism (SNP), which represents a single–base pair variation. SNPs are frequent throughout the genome and form patterns that can be associated to predisposition to certain diseases, cancer, and drug resistance, to mention a few. PCR-based microsatellite analysis is currently one of the methods of choice to detect LOH because it can be used to survey small and therefore specific chromosomal regions (< 100 bp) with scant amounts of DNA. When microsatellites are used as genetic markers, LOH is defined by a significantly weaker or absent band for one version of the microsatellite amplified from tumor DNA versus the 2 bands amplified from normal cells of the same patient. That means that one of the alleles of the gene in which the microsatellite is located is missing from the tumor. For example, allelic loss of chromosome 10q is common in patients with glioblastoma. The tumor suppressor gene PTEN is one of the genes located in chromosome 10q. To determine if PTEN has suffered LOH in a glioblastoma sample, researchers can design PCR primers to amplify microsatellites D10S185 and D10S2491, which are located within 10q23–24, near PTEN. 2 These microsatellites can then be analyzed in the tumor sample and in normal tissue. If the bands corresponding to the microsatellite amplified from tumor DNA are absent or much weaker in an agarose gel when compared to the bands obtained from normal DNA, then PTEN has probably suffered LOH in that specific tumor. Another way of determining LOH in a tumor is by analyzing at least 3 microsatellite markers—one located within, another centromeric, and the last telomeric to the gene of interest. If a tumor sample is hemizygous for these 3 markers it is considered to have suffered LOH, since the probability of simultaneous hemizygosity for the 3 markers in normal tissue is less than 0.01%. 9,48,50 The advantage of this technique is that it does not require normal DNA for comparison. LOH results based on microsatellite markers have to be interpreted carefully because of the possibility of contamination with benign DNA from stroma or neighboring normal tissue. 9,54
Laser Capture and LOH Analysis
Contamination of tumor tissues with normal adjacent tissue can pose a challenge in interpreting LOH results. Fortunately, the advent of laser capture microdissection in 1996 has solved this problem. 8,12 With this system, a small group of cells can be isolated and removed from a thick section of tissue placed on a glass slide so that DNA can be extracted from tumor cells only. This technique decreases contamination of the sample with normal tissue and allows researchers to obtain DNA and RNA from different tissue compartments, such as stroma and epithelium, and elucidate genetic heterogeneity in different lesions present on the same slide. 13,14,18,28,39,40 For example, a section of human prostatic adenocarcinoma can be used to dissect epithelial cells from invasive tumors and from preneoplastic lesions, which are commonly found adjacent to adenocarcinomas in the prostate. DNA extracted from preneoplastic lesions can then be compared to DNA from the tumor cells so that the function of genes involved in the malignant transformation can be investigated. Laser capture microdissection also enables scientists to study LOH and other genetic lesions in tumor tissues from paraffin-embedded archival materials, which is extremely useful in retrospective studies. It is important to mention that successful genetic analysis of archival materials depends on the length of fixation and type of fixative used, as well as other factors. LOH analysis based on genomic DNA can be frequently achieved, whereas survey of transcription levels of tumor suppressor genes is usually difficult because RNA promptly degrades upon formalin fixation.
Other Techniques to Detect and Confirm LOH
Fluorescent in situ hybridization (FISH) can be used to identify or confirm LOH in tumor samples. The technique relies on the ability of DNA and RNA to bind to their complementary sequences. Specific nucleotide sequences can be labeled and then used as selective probes. The application of these probes to glass slides allows chromosomal regions of 50 to 300 bps to be surveyed within different tissues and cell types. 7,25,26 For instance, say that the 5′-TCC… TATG-3′ sequence (between 50 and 300 nucleotides long, to ensure specificity) is exclusively found within the PTEN gene. A complementary sequence (3′-AGG… ATAC-5′) labeled with a fluorophore can be used as a gene-specific FISH probe to locate both alleles of PTEN in tumor and normal tissues.
In dual-staining FISH, a centromeric probe and a probe specific to the gene of interest are labeled with 2 fluorophores and applied to the glass slide containing the tumor sample. The nucleus of a normal diploid cell containing 2 alleles of a tumor suppressor gene—say, TP53—would be labeled with 2 centromeric probes and 2 TP53-specific probes. LOH would be defined by a decreased number of TP53 signals in comparison to the centromeric signals (Fig. 4 ). 5,56
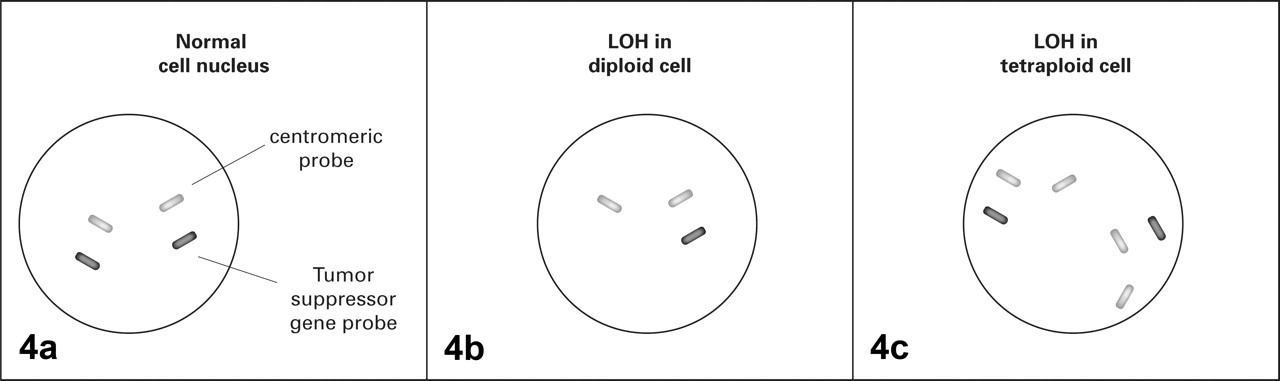
Dual-staining fluorescent in situ hybridization. Cells are treated with probes for a specific tumor suppressor gene and probes for centromeric DNA from the same chromosome and examined under the fluorescence microscope. Centromeric probes are used to evaluate ploidy. A, normal diploid cells will show not only centromeric probes but also probes for the gene of interest from both the maternal and the paternal chromosomes. Loss of the signal for the gene of interest indicates loss of heterozygosity (LOH) in diploid cells (B) and tetraploid cells (C).
FISH is the ideal method for LOH detection because it allows the researcher to confirm the absence of a gene or locus in a specific cell population. Consider the example of prostate adenocarcinoma mentioned above. If researchers found a decreased number of TP53 FISH signals only in invasive tumors but not in adjacent preneoplastic lesions, they might conclude that this tumor suppressor gene plays a role in disease progression rather than initiation. They would also know that ultimate loss of TP53 protein in this specific tumor was due to allelic loss rather than to mutation or posttranscriptional mechanisms. If, however, DNA extracted from homogenates of this same tumor had been used for PCR-based microsatellite analysis, the results would be different. Microsatellites near or within the TP53 gene would amplify from cells in the stroma and in the adjacent preneoplastic lesions, and researchers might have concluded that TP53 had not suffered LOH in this tumor. The principal disadvantage of the FISH technique is that nuclear truncation owing to plane of section can lead to false positive results.
Comparative genomic hybridization (CGH), array CGH, and other microarrays are valuable techniques to assess chromosomal losses and gains. For CGH, DNA from tumor cells and normal cells from the same individual are labeled with different fluorophores and then hybridized to a normal chromosomal metaphase preparation. 6 Regional gains and losses can be detected on the basis of fluorescence intensity within different chromosomal segments. Note that CGH does not detect chromosomal abnormalities that do not involve changes in copy number. Furthermore, the region of loss has to be relatively large (5 to 10 Mb) to be detected by traditional CGH analysis. 27,52 More recently, array CGHs have been developed in which labeled DNA from tumor cells and normal cells can be hybridized to a large number of probes printed on a glass or silicone surface (chip). Array CGHs are also called dual-channel DNA microarray (or two-channel microarray) and so provide a comparative analysis of copy number but with much higher resolution than that of traditional CGH analysis given that probes may be as small as 5 to 10 Kb. 10,35 The term DNA microarray usually refers to one-channel detection in which tumor DNA is hybridized to specific probes on the chip without having the normal tissue for comparison. SNP microarray is a type of DNA microarray made with SNP probes and used to search for genomewide LOH and other disease-associated genetic signatures. 20
While microsatellite-based LOH analysis can be used to survey a specific gene or a region of interest, traditional CGH, array CGH, and DNA microarrays are high-throughput techniques, more suitable to large-scale surveys of chromosomal imbalances and LOH. As such, they require complex image analysis and statistical software for interpretation of data. The large volume of data obtained in arrays is usually clustered hierarchically and displayed in heat maps. Hierarchical clustering means that genes with similar copy number or expression levels, depending on the type of array, are placed close to one another and apart from genes with distinct expression levels (most common application) or copy number. Heat maps are then used to color-code gene expression with red and blue, illustrating overexpression or loss, respectively. For instance, researchers analyzed cDNA from normal mammary gland, lactating mammary gland, preneoplastic mammary lesions, and mammary adenocarcinomas to determine expression of a variety of target genes in a mouse model. All genes were then grouped according to expression levels and represented in shades of red or blue in the heat map (see Fig. 5 ). Note that the heat map highlights gene expression patterns unique to tumors. Genes depicted in blue have low expression in tumor tissues and may function as tumor suppressors in this model.
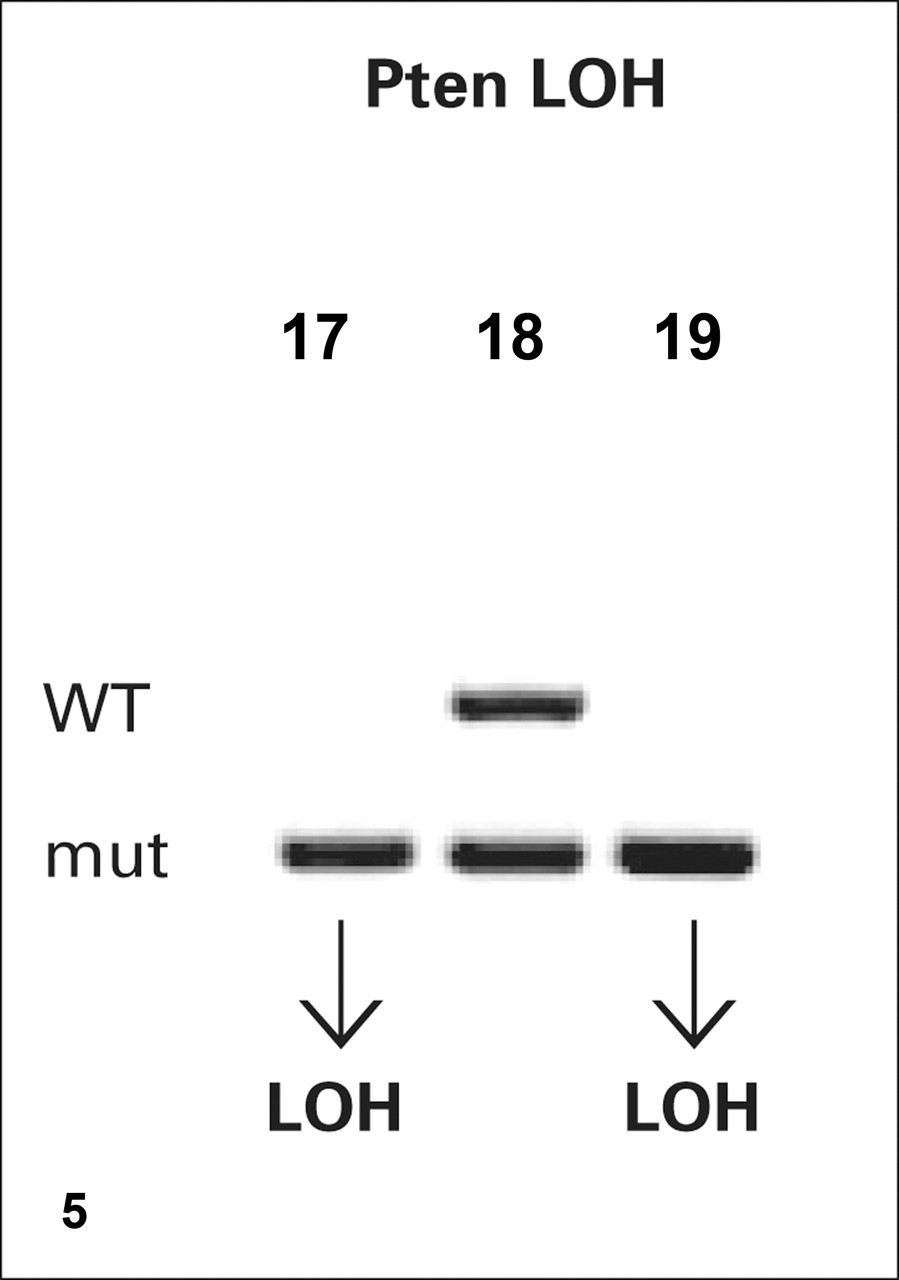
Loss of heterozygosity (LOH) analysis based on genotyping probes. DNA extracted from 3 prostatic lesions derived from Pten heterozygous mice is examined with genotyping primers for the wild-type (WT) and mutant (mut) Pten exon used to construct the mouse. Heterozygous DNA produces two bands on agarose gel (lane 18), whereas LOH is represented by the loss of the wild-type band (lanes 17 and 19).
Hierarchical clustering may also reveal previously unknown gene associations or networks and may help guide further research to elucidate certain pathways. For example, examine the heat map for the mammary tumors mentioned above. Suppose that Socs3, which is overexpressed in tumors, is a known oncogene but that Col18a1, also overexpressed in tumors, is a newly mapped gene of unknown function. By examining the hierarchical data, researchers can then hypothesize that Col18a1 may also act as an oncogene. Experiments can then be designed to test this hypothesis.
LOH Analysis in Mouse Models of Human Cancer
With the complete sequencing of the mouse genome, strain-specific data are readily available. It is now possible to evaluate mouse tissues for LOH with virtually the same techniques as described for humans. Because genetically engineered mouse models are generated to study genetic traits of specific human cancers, LOH analysis should be directed to tumor suppressor genes suspected to be involved in lesion initiation or progression based on data from the human tumors they emulate. An ever-growing list of catalogued mouse microsatellites, SNPs, and other sequences containing polymorphic tandem repeats is currently available, and mouse-specific primers for use in PCR-based LOH analysis can be easily designed. 42 In addition, specific murine microarray chips can be purchased or custom-made to fit the needs of each study.
LOH in tissues from mice heterozygous for certain tumors suppressor genes can also be investigated with genotyping probes. For example, if a researcher suspects that LOH of the second allele of Pten is necessary for tumor progression in Pten +/– mice, then the DNA of the tumor can be evaluated by PCR with genotyping primers. If the tumor remains heterozygous, two bands of different sizes are present in the gel, one for the mutant allele and one for the wild-type allele. 4,17,34 If LOH took place, then only the mutant band is seen in the gel (Fig. 6 ).
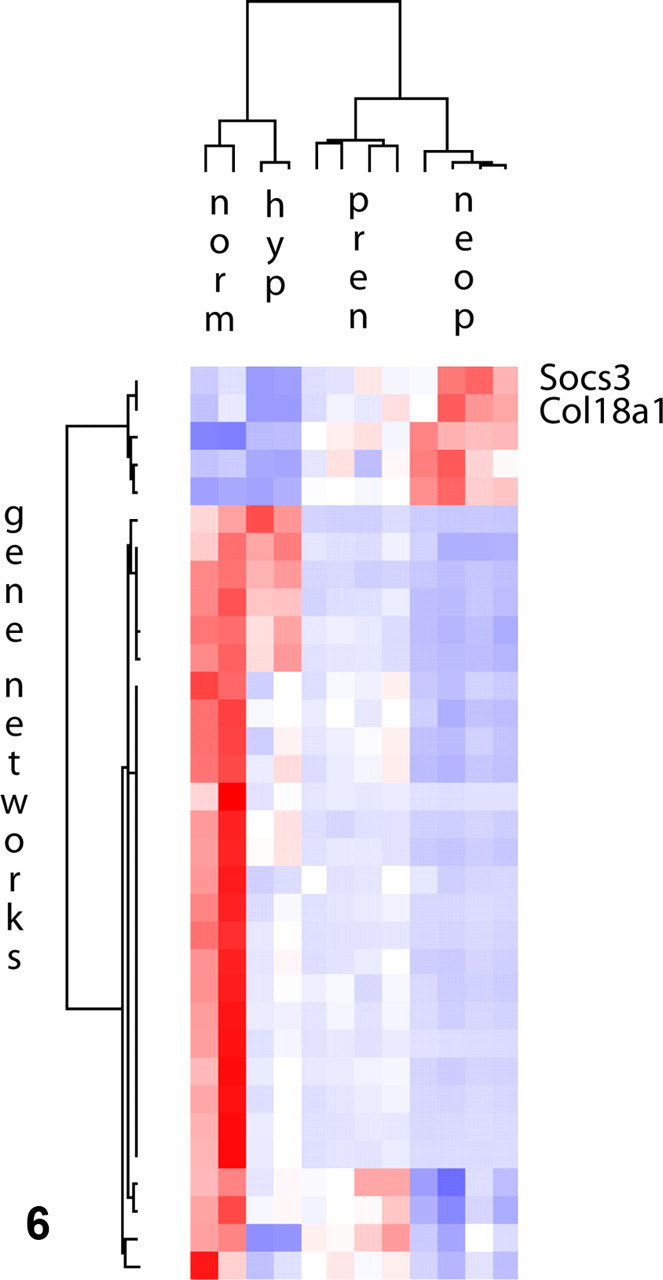
Heat map. Hierarchical clustering of a panel of genes expressed in normal mammary gland (norm), lactating hyperplastic mammary gland (hyp), preneoplastic mammary lesions (pren), and mammary tumors (neop). Overexpressed genes are represented in red, and downregulated genes are represented in blue. Socs3 and Col18a1, highlighted in this example, are both overexpressed in mammary tumors when compared to the normal gland. The expression of a limited number of genes changes in the transition from preneoplastic to neoplastic lesions. Adapted from Namba R, Maglione JE, Young LJ, Borowsky AD, Cardiff RD, MacLeod CL, Gregg JP. Molecular characterization of the transition to malignancy in a genetically engineered mouse-based model of ductal carcinoma in situ. Mol Cancer Res
It is important to consider that only a specific area of the gene—usually spanning one single exon or coding region—can be surveyed with genotyping primers. Thus, LOH may be missed if there is loss of another functionally important exon that is not specifically amplified with genotyping primers.
As with any other technique used to determine LOH in tumor tissues, contamination of tumor samples with normal cells can lead to false-negative results. Thus, sample collection needs to be carefully done to avoid contamination. Also, PCR-based LOH analyses should always start with the same concentration of DNA from tumor and normal tissue controls for the intensity of the bands to be informative.
Each above-mentioned method available for LOH analysis has its advantages and weaknesses. For instance, PCR-based LOH analysis is neither sensitive nor specific, because contamination with normal tissue is a potential problem. On the other hand, material collected from laser capture microdissection may yield very little DNA. Therefore, it would be ideal to repeat LOH analyses by at least two techniques, such as PCR analysis of microsatellite primers and FISH, to confirm results.
Final Remarks
A better understanding of the mechanisms that lead to LOH and the subsequent design of techniques to detect it have come hand-in-hand with the discovery of many important tumor suppressor genes. By understanding the application, the value, and the shortcomings of the molecular techniques available, the pathologist can provide significant input into the experimental design and interpretation of genetic data in light of specific morphologic features of tumors. For instance, if a Trp53 heterozygous mouse develops a tumor (with Trp53 being the mouse version of the human TP53 gene), it is reasonable to inquire whether the remaining normal allele of this important tumor suppressor gene is lost in focal areas of prominent atypia or in more anaplastic or invasive fronts of the tumor. To answer this question, one could combine laser capture microdissection of the areas of interest and perform an LOH analysis with genotyping probes if quality genomic DNA from the samples is obtained. Evaluating LOH in different areas of the tumor could also be achieved via FISH. As another example, the pathologist could explore the role of a tumor suppressor gene in tumor progression by correlating LOH of the gene of interest with the transition from in situ to invasive cancers. Notice that in most examples mentioned in this review, the motivation to apply a certain molecular technique came from initial morphologic observations and was driven by the insights that a pathologist may be able to provide.
Finally, looking at the big picture and thinking beyond DNA is crucial to elucidate the role of certain genes in cancer. Genes may be physically intact but silenced through epigenetic events such as promoter methylation. Thus, investigating protein and RNA levels of candidate tumor suppressor genes is essential whenever possible. Additionally, a variety of micro RNAs have been recently implicated in the regulation of tumor suppressor genes and oncogenes and so may play an important role in cancer pathogenesis. Unfortunately, epigenetic mechanisms of gene silencing and regulation of micro RNA transcription are beyond the scope of this review. In summary, by acquiring a deeper knowledge of cancer biology and understanding commonly used molecular techniques, the pathologist can have an evermore significant impact in cancer research in the postgenomic era.
Footnotes
Acknowledegments
I would like to thank Robert Cardiff (University of California, Davis), Eva Hernando-Monge (New York University), Lloyd Freeman (Cold Spring Harbour) for valuable discussions of the loss of heterozygosity detection methods currently available; Ron Firestein (Genentech, Inc) for verifying the accuracy of the concepts presented in the article; Carlos Cordon-Cardo, Tulio Matos, and Maika Mitchell (Columbia University) for invaluable help on the bench; Sunny Yeung (Memorial Sloan-Kettering Cancer Center) for skillful assistance with figures; Marcia Triunfol (Publicase) for suggestions and review; and Carol Ross (Hyperlife Editing) for proofreading. Figures 1, 2, and ![]() were inspired by the book The Biology of Cancer, by Robert Weinberg.
57
were inspired by the book The Biology of Cancer, by Robert Weinberg.
57
The author declared no conflicts of interest with respect to the authorship and/or publication of this article.
The author received no financial support for the research and/or authorship of this article.
