Abstract
Spironucleus spp are parasites of fish and terrestrial vertebrates, including mice and turkeys, that rarely cause extraintestinal disease. Two rhesus macaques (Macaca mulatta) were experimentally inoculated with simian immunodeficiency virus mac251. Both progressed to simian acquired immune deficiency syndrome within 1 year of inoculation and developed systemic protozoal infections in addition to common opportunistic infections, including rhesus cytomegalovirus, rhesus lymphocryptovirus, and rhesus adenovirus. In the first case, the protozoa were associated with colitis, multifocal abdominal abscessation, and lymphadenitis. In the second case, they were one of a number of organisms associated with extensive pyogranulomatous pneumonia and colitis. Ultrastructural, molecular, and phylogenetic analysis revealed the causative organism to be a species of Spironucleus closely related to Spironucleus meleagridis of turkeys. This report is the first of extraintestinal infection with Spironucleus sp in higher mammals and expands the list of opportunistic infections found in immunocompromised rhesus macaques.
Members of the genus Spironucleus are flagellated anaerobic protozoa that parasitize marine and freshwater fish as well as terrestrial vertebrates, including mice and turkeys. 9,19,25,26,31 With other flagellates, including Giardia, Hexamita, Enteromonas, and Trichomonas, they compose a clade known as Fornicata. These organisms are classified as either enteromonads, containing one karyomastigont (nucleus and cytopharynx), or diplomonads, with 2 karyomastigonts. Fornicata are unusual among eukaryotes in that they lack mitochondria, a Golgi apparatus, and cytochromes. Interestingly, some genera are capable of amino acid fermentation for energy production, a trait more commonly found in bacteria than in protozoa. 1,2 These characteristics suggest that Fornicata are among the earliest and most primitive offshoots in the eukaryotic tree. 10
Spironucleus spp are diplomonads, but recently developed molecular phylogenies reveal that reduplication of the karyomastigont has occurred multiple times throughout evolutionary history and that the genus Spironucleus is actually a paraphyletic clade. 15 Based on small subunit ribosomal DNA (SSU rDNA) sequencing, Spironucleus spp cluster into 3 separate groups, the first containing parasites of saltwater fish; the second, parasites of freshwater fish; and the third, parasites of terrestrial animals. 15
Spironucleus and closely related Hexamita spp infect a variety of vertebrate hosts, including birds, fishes, mice, and nonhuman primates. 3,4,8,9,11,17,26,31 Animals become infected after ingesting parasitic cysts, with as few as one cyst being infective in some strains of mice. 12,18 Infected mice and birds develop a clinical syndrome characterized by diarrhea, depression, and weight loss, with mortality ranging from 50% in some colonies of mice to 80% in partridge broods. 4,8 Necropsy of these animals reveals enteritis with fluid- and gas-filled intestinal loops causing gross abdominal dilatation and, occasionally, ascites. 4,8
Rhesus macaques are naturally infected by a variety of intestinal parasites and protozoa, including Trichomonas, Balantidium, Entamoeba, Cryptosporidium, and Giardia. 29 These infections cause few clinical problems in healthy macaques but are more pathogenic in animals that are immunosuppressed (eg, via infection with simian immunodeficiency virus [SIV]). 6,14,20 Enteric infection of nonhuman primates with Spironucleus has been reported in several instances, beginning with a case in 1929 in which the organism was, at the time, referred to as Octomitus pitheci. 5,13,23,29,30 In these reports, infection was confined to the gastrointestinal tract, and animals were asymptomatic. Systemic Hexamita infection in an SIV-infected rhesus macaque has been reported only once and was associated with circulating organisms visible on a blood smear, but there was no evidence of associated pathology. 22
Herein, we describe the pathologic findings of systemic infection of 2 SIV-infected rhesus macaques with a species of diplomonad and the associated pathological findings. Given the ultrastructure and SSU rDNA sequencing, we have identified the organism as a species of Spironucleus.
Materials and Methods
Clinical History
Both animals were born and raised at the New England Primate Research Center in a specific pathogen–free breeding colony and screened annually for simian retrovirus D, simian T-lymphotropic virus, SIV, and Macacine herpesvirus 1 (B virus) via serology. Negative tuberculosis status was confirmed via quarterly intradermal testing.
Both animals were inoculated intravenously with SIVmac251 and euthanized after progression to simian acquired immune deficiency syndrome (SAIDS). Subsequent to inoculation, both animals were housed in a biosafety level 2 facility in accordance with the National Research Council's Guide for the Care and Use of Laboratory Animals and the standards of the Harvard Medical School Standing Committee on Animals and the Association for the Assessment and Accreditation of Laboratory Animal Care. Case No. 1, a 5-year-old male, was euthanized 375 days postinoculation. Case No. 2, an 11-year-old female, was euthanized 296 days postinoculation.
Tissue Processing
Necropsies were conducted on each animal within 2 hours of euthanasia. Representative sections of all major organs were collected, fixed in 10% neutral buffered formalin, embedded in paraffin, sectioned at 5 µm, and stained using hematoxylin and eosin (HE). Additional sections containing the unknown organism were stained with periodic acid–Schiff, Ziehl-Neelsen, Gomori methenamine silver, and Gram stain.
Electron Microscopy
Samples from case No. 1 were fixed in 4% paraformaldehyde, 1% glutaraldehyde in 0.1M phosphate buffer. Lung tissue from case No. 2 was fixed in 10% neutral buffered formalin. All samples were postfixed in 1.0% osmium tetroxide, dehydrated through graded alcohol and propylene oxide, and embedded in eponate 12 epoxy resin (Ted Pella Inc, Redding, Calif). Thin sections were prepared on a Sorvall MT-2 ultramicrotome and stained with uranyl acetate and Sato’s lead stain. The grids were examined with a JEOL 1010 Electron Microscope (JEOL Ltd, Tokyo, Japan).
DNA Isolation, Polymerase Chain Reaction, and DNA Sequencing
Isolation of DNA from fresh abscess material from case No. 1 was carried out using the DNeasy Tissue Kit (Qiagen, Valencia, Calif) following the manufacturer’s instructions. Isolation of DNA from stool samples was performed using the QIAamp DNA Stool Mini Kit (Qiagen) following the manufacturer’s instructions.
Broad-based degenerate primers were designed to amplify SSU rDNA sequences from Spironucleus, Hexamita, Trimitus, and Trepomonas species using a prototype version of dprimer (http://sourceforge.net/projects/dprimer/). A multiple-sequence alignment of SSU rDNA was constructed with ClustalX 2.0.5 from Genbank accession Nos. EF551182 (Trimitus sp), EF551176 (Trimitus sp), EF551173 (Trepomonas steinii), EF1174 (Trepomonas sp), EF050054 (Spironucleus meleagridis), EU04320 (Spironucleus muris), EF050053 (Hexamita nelsoni), and Z17224 (Hexamita sp); AY921408 (Enteromonadiae sp) was used as dprimer input. 21 Primers HTTS2-FW (5′-GCC TYA TYG CGT TAA CGA GC-3′) and HTTS2-RV (5′-GAC GGG CGG TGT RTA CAA AR-3′) were chosen from the dprimer output. The product amplified from the DNA of No. 1 was purified from an agarose gel using the QIAEX II Gel Extraction Kit (Qiagen), cloned into pGEM-T Easy vector (Promega, Madison, Wis), and submitted for sequencing (Retrogen Inc, San Diego, Calif).
A second set of nondegenerate primers, Ssp3-FW (5′-AGA CGG CCA GCC CCC CGG GC-3′) and Ssp3-RV (5′-GGT GTG GCC GCG GCC GGA GC-3′), were manually designed against the sequence of the above amplicon. Because of the GC richness of the informative regions of the rDNA, a nested assay consisting of an initial round of amplification using HTTS2 primers and a secondary round with the specific Ssp3 primers was employed to screen fecal samples.
Polymerase chain reaction (PCR) reactions were carried out in 25 µl of 1xGoTaq Green Master Mix (Promega, San Diego, Calif) with 20 pmoles of each primer and 10 to 40 ng of template DNA. The cycling conditions for the HTTS2 primer set consisted of an initial 2-minute denaturation at 95°C; followed by 40 cycles of 20 seconds of denaturation at 95°C, 60 seconds of annealing at 63°C, and 45 seconds of extension at 72°C; followed by a final 10-minute extension step at 72°C. Reactions with the Ssp3 primer set were performed on 1 µl of the HTTS2 product and cycled with a 2-minute denaturation at 93°C; followed by 40 cycles of 20 seconds of denaturation at 93°C, 60 seconds of annealing at 80°C, and 45 seconds of extension at 72°C; followed by a final 10-minute extension step at 72°C.
Phylogenetic Analysis
A phylogenetic tree was produced from the cloned fragment in case No. 1. It was based on the homologous regions of published SSU rDNA sequences from several other fornicates and was created with the ClustalX implementation of the UPGMA (unweighted pair group method with arithmetic mean) clustering algorithm. 21 Finally, the tree was visualized with TreeViewX (http://darwin.zoology.gla.ac.uk/~rpage/treeviewx/).
Results
Gross and Histopathologic Analysis
Case No. 1
Grossly, the caudal abdomen contained multiple large coalescent abscesses arising from the pelvic lymph nodes and invading the caudal abdominal aorta. In addition, the proximal jejunum was corrugated and thickened, and the mesenteric lymph nodes were greatly enlarged. Numerous extracellular and phagocytosed 5- × 4-µm oval protozoa were identified in impression smears of the abscess. Their cytoplasm stained pale blue, and they contained 2 anterior nuclei that sometimes appeared overlapped or fused. Two cytopharynxes extending from the nuclei transversed the cells posteriorly and appeared either separate and divergent or crossed, depending on orientation of the parasite. Occasional organisms contained nonstaining intracytoplasmic vacuoles. Flagella were not visible (Fig. 1). No bacteria or other infectious agents were present. Aerobic and anaerobic cultures of the abscess were negative.
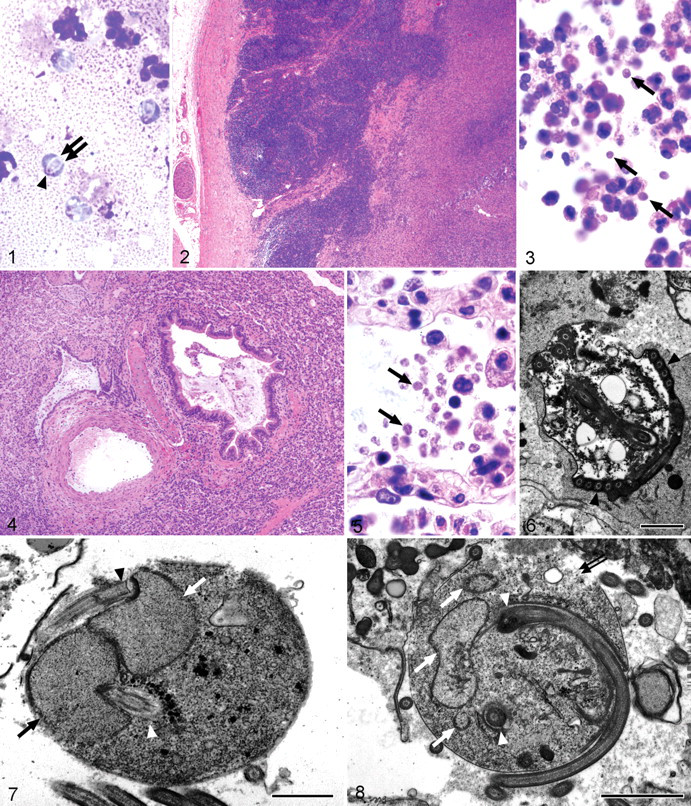
Histologically, the abscesses were composed of large numbers of macrophages that formed well-defined pyogranulomas with intralesional and intrahistiocytic protozoal organisms. With HE staining, they appeared as 5- × 4-µm oval bodies with homogeneous to vacuolar pale eosinophilic cytoplasm and inapparent nuclei. Gomori methenamine silver, Ziehl-Neelsen, periodic acid–Schiff, and Gram stain did not highlight the organism. One section contained scattered remnants of lymph node cortex that had been effaced by sheets of protozoan-laden macrophages (Figs. 2, 3). In other areas, large rafts of histiocytes, many laden with protozoa, invaded through the caudal abdominal aorta where they were intermixed with fibrin and necrotic cellular debris.
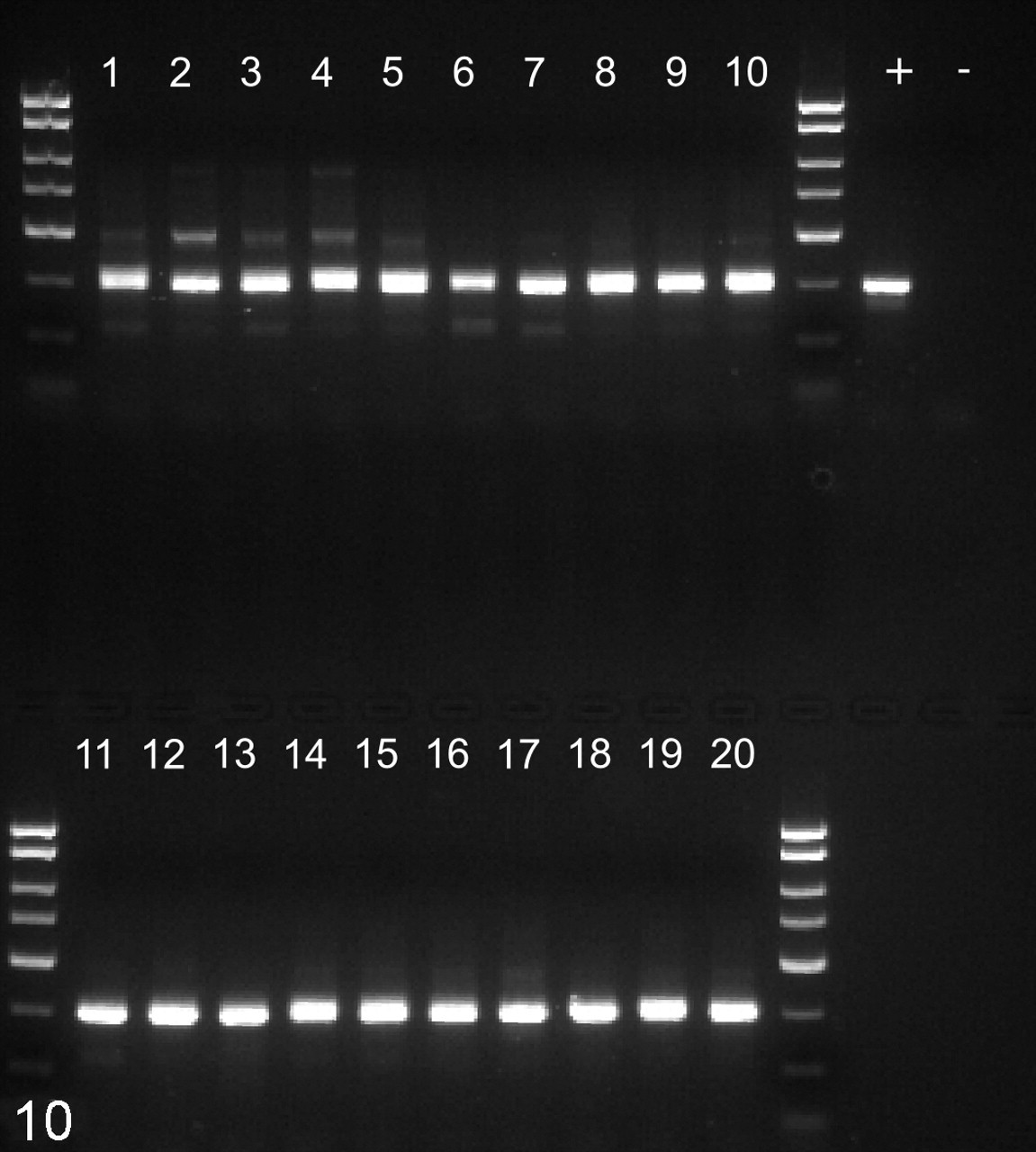
Results of polymerase chain reaction on fecal samples using broad-based HTTS2 primers. Lanes 1-10, uninfected animals. Lanes 11–20, simian immunodeficiency virus–infected animals. +, positive control DNA from case No. 1. –, no template control. Products of the expected 310–base pair size were obtained from all fecal samples.
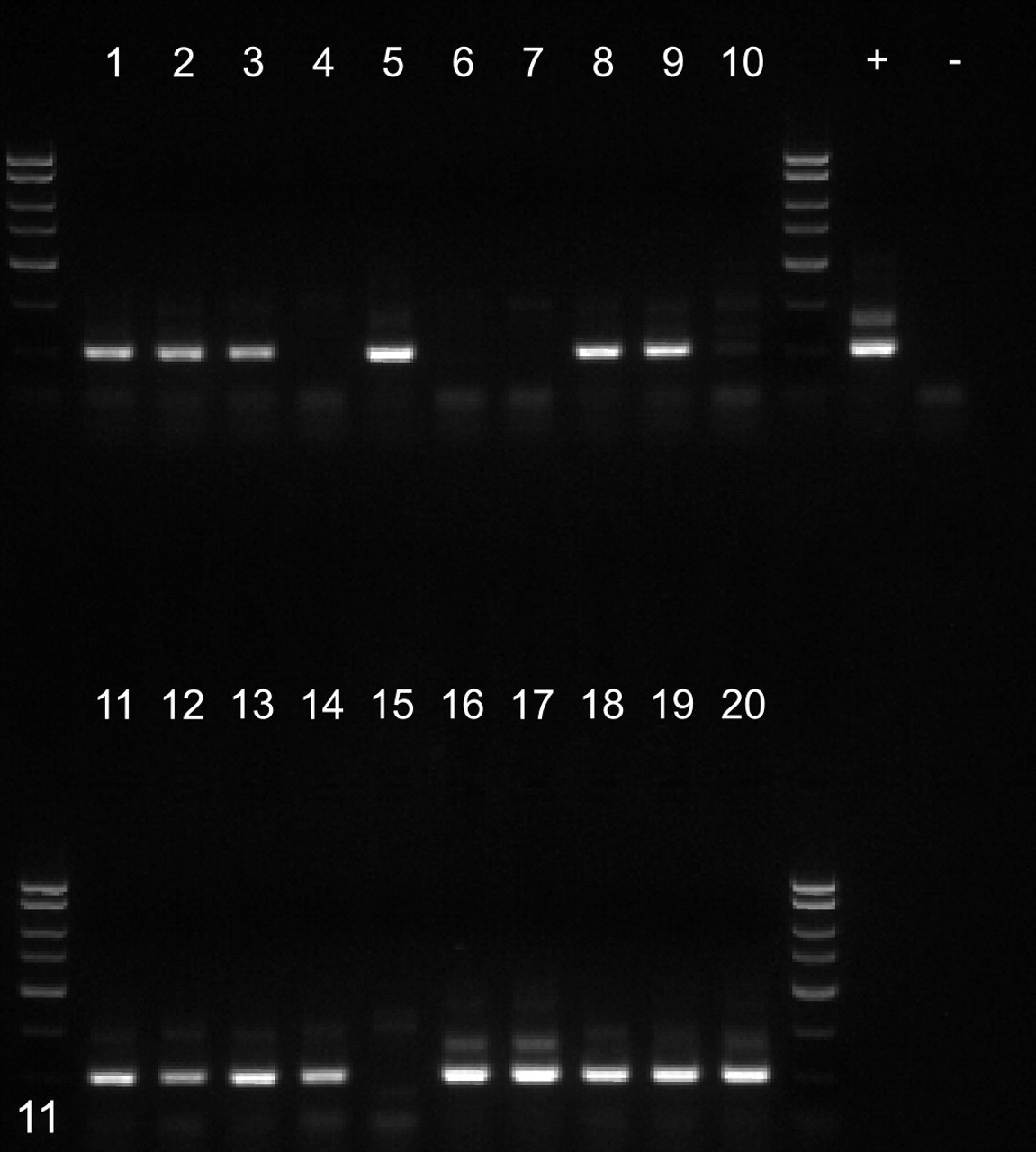
Nested polymerase chain reaction reactions using 1 µl of template from each of the HTTS2 products depicted in Fig. 10. Lanes 1–10, uninfected animals. Lanes 11–20, simian immunodeficiency virus–infected animals. +, positive control DNA from case No. 1. –, no template control. Products of the expected 150–base pair size were obtained from 6 uninfected animals and 9 infected animals.
Crypts throughout the colon contained protozoa similar to those described within the abdominal abscesses. The lymphoid nodules of the rectum contained infiltrates of pyogranulomatous inflammation with scattered intracellular protozoa. Comorbid histologic findings in this animal included adenoviral enteritis, myocarditis and coronary arteriopathy, and lymphosarcoma of the distal jejunum and adrenal medulla.
Case No. 2
Grossly, the lungs contained multiple large red/tan consolidated regions. The left lung was tethered to the chest wall by numerous fibrous adhesions. Impression smears of the affected lung revealed intense pyogranulomatous inflammation with moderate hemodilution. Neutrophils were variably degenerate. Rare clusters of intracellular and extracellular rods and cocci were scattered throughout the smear. Intermixed throughout the affected sections were myriad protozoa identical to those described above.
Histopathology of the lung revealed multifocal to coalescing areas of neutrophilic interstitial pneumonia and alveolitis with intralesional protozoa. The protozoa were identified within alveoli, most noticeable at the margins of the inflammatory lesions, and were occasionally observed within neutrophils and macrophages.
In one section of colon, the crypts were packed with protozoa of similar size and nondescript staining characteristics. The underlying submucosa was infiltrated by macrophages and neutrophils, some of which contained engulfed protozoa. Colonic lymph nodes also contained foci of pyogranulomatous inflammation, although no organisms were definitively present.
Comorbid histologic findings of animal No. 2 included lymphocryptoviral esophagitis, cryptosporidial cholecystitis, choledochitis, pancreaticodochitis, and tracheitis, cytomegaloviral enteritis, and adenoviral enterocolitis.
Electron Microscopy
Ultrastructural analysis of the parasite revealed the presence of numerous flagella, often arranged in parallel rows lined up along the lateral margins of the cell membrane, suggesting a coiled arrangement (Fig. 6). The 2 nuclei were closely apposed to the cell membrane and to one another. They were arranged in an S-shaped configuration (Fig. 7 ). A recurrent flagellum arose from an indentation of the nuclear membrane near the apex of each nucleus. The cytoplasm was coarse, grainy, and moderately electron dense, containing occasional membrane-bound vacuoles and cisternae but lacking mitochondria. Deposits of electron-dense material interpreted as glycogen were found at the origin of the recurrent flagella and sometimes scattered throughout the cytoplasm (Fig. 8 ).
Sequencing and Phylogenetic Analysis
BLAST searches with the partial SSU rDNA amplicon obtained from case No. 1 as input showed the highest-scoring alignment to the SSU rDNA sequence of S meleagridis, Genbank ID No. EF050054 (90% identity, 4 gaps, 320 bits). A phylogenetic tree constructed with SSU rDNA sequences of several other fornicates clustered the unknown sequence in a clade comprising S muris and S meleagridis, with S meleagridis as its closest relative (Fig. 9).
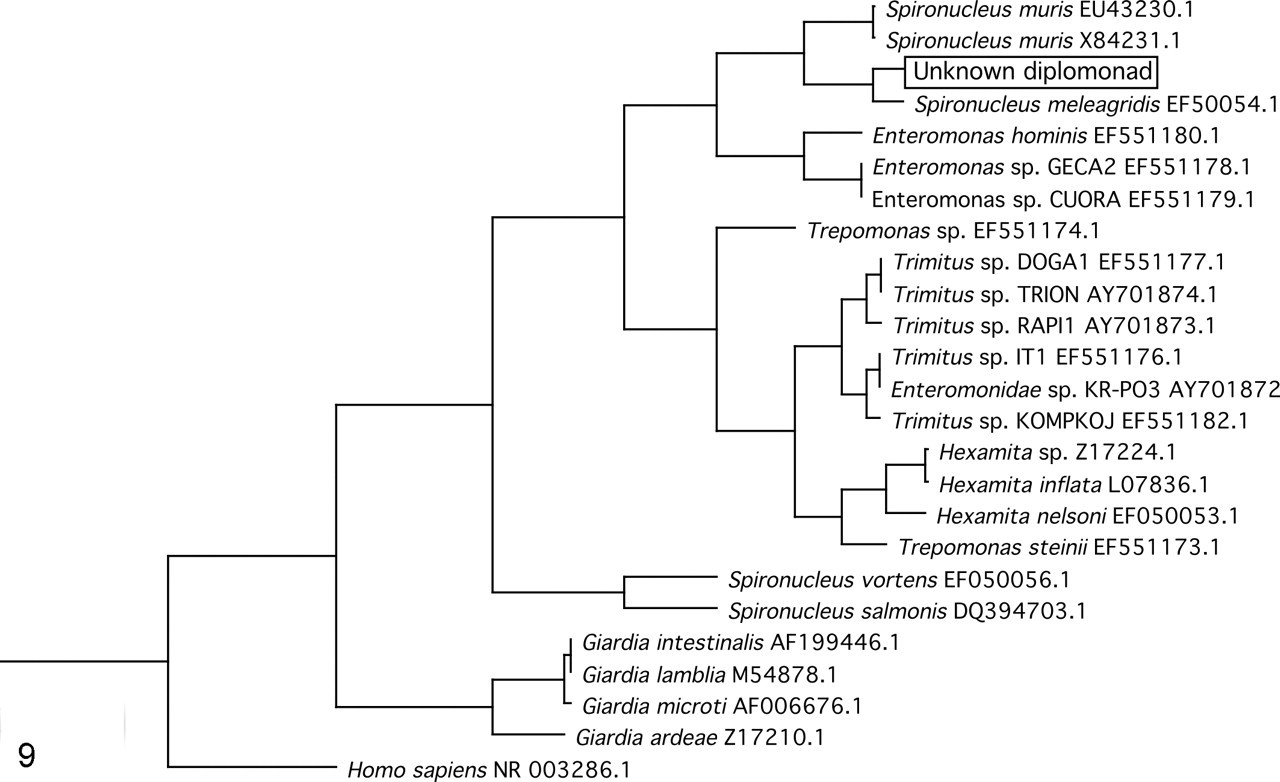
A phylogenetic tree clusters the unknown with Spironucleus meleagridis.
Colony Screening
DNA isolated from fresh fecal samples from ten SIV-infected and ten normal rhesus macaques was screened for the presence of Spironucleus DNA using nested PCR. Positive results were obtained from every animal using the outer set of broad-based HTTS2 primers (Fig. 10 ). Positive reactions were obtained from 9 SIV-infected animals but only 6 uninfected controls, with the inner specific Ssp3 primer set (Fig. 11 ). Although the prevalence among SIV-infected macaques was higher, the difference was not statistically significant (P = .15, one-tailed Fisher exact test).
Discussion
This report is the first description of disseminated spironucleosis in immunocompromised rhesus macaques, and it expands the list of potential opportunistic infections in this species. Both animals had progressed to SAIDS by the time of euthanasia, as evidenced by multiple opportunistic infections. In the first case, no bacteria were observed, and aerobic cultures of the abscess were negative. In the second case, the organisms were associated with severe multifocal pyogranulomatous pneumonia but were accompanied by a mixed population of bacteria, thus rendering their role as the primary etiology of the pneumonia unclear.
Phylogenetic analysis of the organism in this report placed it in close relation to Spironucleus melagridis of turkeys and firmly in the clade occupied by similar parasites of terrestrial animals, including S muris. This finding is in concordance with evolutionary data indicating that organisms infecting terrestrial animals evolved distinctly from those infecting salt and freshwater fishes.
Screening of fecal material from colony animals revealed that the organism is common in rhesus macaques but occurs in a higher proportion of SIV-infected animals, as compared to controls, although none of these animals had clinical signs associated with infection. Given the prevalence of the organism in the colony, extraintestinal infection is likely a rare event. Pathogenic infection with gastrointestinal commensals is relatively common in SIV-infected macaques, secondary to progressive CD4 cell depletion. Indeed, Trichomonas, Entamoeba, and Balantidium coli are commonly observed in normal rhesus macaques but may cause severe illness with progression to SAIDS in the face of SIV infection. 13,16
Reports of disseminated Spironucleus in other species are rare. There is a single report of disseminated hexamitiasis in Siamese fighting fish, which proposed that the organism gained entry through damaged gastric mucosa, causing fatal peritonitis, hepatitis, nephritis, and splenitis. 7 Others have postulated that the organism spreads via the bloodstream, and they have documented widespread infection by Hexamita salmonis in the blood and multiple organs of farmed Atlantic salmon. 24,27 This finding is supported by findings in farmed Arctic char in which the organism was widespread throughout the bloodstream but caused little systemic disease. 28 In mice, the disease generally causes enteritis with poor weight gain in pups, but invasion into the lamina propria has been reported, providing further support for an enteric portal of entry. 8 In athymic mice, S muris has been documented to contribute to a reduced life span. The mortality was reduced or delayed with antiprotozoal treatment and thymic transplant. 3
Given the current findings and previous reports, we suggest that the organism is a common gastrointestinal commensal of rhesus macaques, presumably acquired early in life. The organism is unlikely to be associated with clinical disease except in immunocompromised individuals where systemic involvement is a potential consequence. Based on the presence of the organism in the rectal lymphoid tissue with abscessation of pelvic lymph nodes in case No. 1, in addition to the focal colitis with invasive protozoa in case No. 2, the organism likely gains systemic entry through invasion of the colonic mucosa. That lesions were present in the draining lymph nodes rather than the liver suggests that dissemination occurs via lymphatics rather than the portal circulation; however, vascular dissemination cannot be definitely ruled out. Pulmonary involvement may result from lymphatic drainage into right-sided circulation via the thoracic duct. This mechanism would be unique among enteric protozoal infections of macaques, which generally cause local disease but even with severe immunosuppression tend not to disseminate.
Spironucleus should be included as a differential diagnosis for diarrhea, systemic illness, and abdominal lymph node abscesses in SIV-infected rhesus macaques. Future work is ongoing to determine the overall incidence and potential correlates of infection in SIV-infected rhesus macaques. To our knowledge, Spironucleus infection of the human has not been reported; however, care should be taken to prevent possible exposure, especially in immunocompromised laboratory workers.
Footnotes
Acknowledgements
We would like to thank Elizabeth Curran for her assistance with necropsies and Kristen Toohey for photographic assistance.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
This research was funded, in part, by National Institutes of Health grant Nos. RR00168, RR07000, and U42 RR016020.
