Abstract
Objective
Colorectal neuroendocrine tumors and polyps share similar endoscopic features, often resulting in misdiagnosis. As neuroendocrine tumors are rare, obtaining a sufficient number of images for deep learning models is challenging.
Methods
This study introduces a few-shot learning model for ternary classification of neuroendocrine tumors, serrated lesions and polyps, and traditional adenomas using endoscopic images. Three groups of images (56 serrated lesions and polyps, 86 adenomas, and 53 neuroendocrine tumors) were collected retrospectively and divided into Support Sets and Query Sets. The proposed few-shot learning model involved transfer learning using ResNet50 V2 pretrained on ImageNet and esophageal endoscopic images, followed by metric learning based on Euclidean distances and K-nearest neighbor classification.
Results
Evaluated across three rounds, the few-shot learning model outperformed conventional deep learning models and both junior and senior endoscopists in several metrics, achieving an average macro–area under the curve of 0.731, macro–F1-score of 0.674, Matthews correlation coefficient of 0.526, and Cohen’s kappa of 0.523. When identifying neuroendocrine tumors specifically, the model achieved the highest accuracy (0.823), sensitivity (0.653), precision (0.673), and F1-score (0.659).
Conclusions
The few-shot learning approach effectively addresses data scarcity issues and improves diagnostic accuracy, offering a promising tool for computer-aided diagnosis of rare gastrointestinal diseases.
Keywords
Introduction
Neuroendocrine tumors (NETs) are well-differentiated neuroendocrine neoplasms originating from neuroendocrine cells and are rarely found in the colorectum. 1 They are generally diagnosed and staged using endoscopic ultrasonography (EUS), imaging techniques such as computed tomography (CT), or biomarkers such as Ki-67 index. 2 White-light imaging (WLI) endoscopy alone cannot determine the nature or origin of lesions; therefore, it cannot be used to diagnose NETs. However, endoscopy serves as the basis for identifying submucosal tumors (SMTs), which allows for an initial assessment. The decision to perform EUS and subsequent imaging examinations often depends on the endoscopist’s preliminary judgment during endoscopy. If NETs are suspected, EUS is typically conducted to confirm the diagnosis.
Colorectal polyps, which protrude into the colorectal lumen, are commonly encountered in clinical practice. According to the 2019 World Health Organization (WHO) classification of digestive system tumors, 3 colorectal polyps can be categorized into serrated lesions and polyps (SLPs) and traditional adenomas. SLPs are characterized by a serrated (sawtooth or stellate) architecture of the crypt epithelium and can be further classified histologically as hyperplastic polyps (HP), sessile serrated lesions (SSL), traditional serrated adenomas (TSA), and unclassified serrated adenomas. Traditional adenomas are benign, premalignant neoplasms composed of dysplastic epithelium and are classified into tubular adenomas, villous adenomas, and tubulovillous adenomas.
Colorectal NETs and polyps exhibit distinct features under endoscopy, enabling endoscopists to conduct a preliminary differential diagnosis. However, NETs <10 mm with atypical endoscopic findings, clear boundaries, and polyp-like protrusions are difficult to differentiate.4,5 Moreover, endoscopy is a labor-intensive procedure, which may lead to misdiagnosis of colorectal NETs as adenomas or atypical HP. 6 A multicenter study reported that fewer than 20% of French endoscopists suspected NETs during endoscopic procedures, resulting in misdiagnosis. 7 Therefore, it is necessary to develop an objective clinical diagnostic technique that can distinguish NETs from polyps under endoscopy to guide subsequent diagnosis and treatment strategies.
Deep learning (DL), which consists of multilayer neural networks, can transform low-level, simple features into high-level, complex features, making it suitable for image processing.8,9 Owing to its excellent performance in image segmentation, detection, and classification, DL has been widely applied in the field of digestive system imaging.10–12 However, conventional supervised DL typically requires large, balanced, and well-annotated image datasets for training. For rare diseases such as colorectal NETs, 13 collecting a sufficient number of endoscopic images is costly and time-consuming, and models trained on small datasets are prone to overfitting. Moreover, subtle interclass and intraclass overlap between NETs and polyps under white-light endoscopy (WLE) limits the benefits of standard augmentation alone.
To address these issues, few-shot learning (FSL) has been proposed. As the name implies, FSL represents a method that identifies and classifies novel, previously unseen samples by learning rapidly and accurately from a small number of base-labeled examples. 14 It has shown considerable progress in overcoming small-sample and domain-generalization challenges in the medical field. This approach can be applied effectively with few per-class examples and can extend to new classes without full retraining. Therefore, we hypothesized that FSL would be suitable for classifying colorectal NETs and polyps under conditions of limited data availability.
This study aimed to develop an FSL-based model that integrates transfer learning with metric learning for the classification of colorectal SLPs, adenomas, and NETs based on endoscopic images, thereby enabling accurate identification of colorectal SMTs and polyps.
Materials and methods
Study design
The overall design process of this study is summarized as follows: 1. Endoscopic images of SLPs, adenomas, and NETs were collected from multiple centers and preprocessed. 2. A three-way, three-shot dataset was constructed, comprising a Support Set (SS) that included three images each of SLPs, adenomas, and NETs and a Query Set (QS) that included 53 SLPs, 83 adenomas, and 50 NETs. 3. A residual network (ResNet50 V2), initially trained on ImageNet, was selected as the DL model for extracting the eigenvectors. 4. The model was subsequently pretrained using esophageal endoscopic images to capture more detailed endoscopic features, and its performance was visualized through Gradient-weighted Class Activation Mapping (Grad-CAM). 5. The dual-pretrained ResNet50 model was then used to extract eigenvectors from the SS and QS. 6. The Euclidean distance was calculated between each QS image and the nine SS images based on the eigenvectors. 7. K-nearest neighbor (KNN) algorithm was used for classification. 8. The FSL process was conducted in three rounds, and its performance was compared with that of the conventional ResNet50 model as well as junior and senior endoscopists.
Steps 3–5 involved transfer learning, whereas steps 6 and 7 involved metric learning. The FSL model consisted of SS and QS based on the combination of transfer learning and metric learning. The study flowchart is presented in Figure 1. Institutional Review Board approval (approval number: 2022098) was obtained for the retrospective collection and analysis of data. Additional Institutional Review Board approval was not required because the data were de-identified. The study was conducted in accordance with the Checklist for Artificial Intelligence in Medical Imaging (CLAIM).

The flowchart of the study. The three-way three-shot FSL model consists of SS and QS based on transfer learning and metric learning. FSL: few-shot learning; SS: Support Set; QS: Query Set.
Datasets and sets
Datasets
Endoscopic images of SLPs, adenomas, and NETs were obtained from the First Affiliated Hospital of Soochow University, Changshu No.1 People’s Hospital, and Yangchenghu People’s Hospital. All images were captured using Olympus endoscopy systems under white-light and nonmagnifying imaging and confirmed by senior endoscopists and pathologists. Each image corresponded to a single patient. To prevent image-level data leakage, we ensured that the dataset was divided at the patient level. All images were processed as red–green–blue (RGB) three-channel inputs. Each image file was decoded from the JPEG format, resized to 331 × 331 pixels, converted to 32-bit floating-point tensors, and normalized to [0,1] intensity via dividing by 255. No image augmentation was applied. The inclusion criteria are provided in Supplementary Method 1.
In addition, esophageal endoscopic images (normal (n = 429), early cancer (n = 489), and advanced cancer (n = 594)) were obtained under WLI at The First Affiliated Hospital of Soochow University for dual pretraining. These images were augmented and randomly divided into training and validation sets in a ratio of 8:2. The dataset of esophageal endoscopic images used for dual pretraining is shown in Figure 2(a). The definitions of esophageal early cancer and advanced cancer are presented in Supplementary Method 2.

Datasets of endoscopic images. (a) Esophageal endoscopic images were randomly allocated into training set and validation set for subsequently pretraining the DL model and (b) the SS consisted of three base classes of SLPs, adenomas, and NETs, with each class containing three images. The QS consisted of 186 images to be classified. DL: deep learning; SLPs: serrated lesions and polyps; NETs: neuroendocrine tumors; SS: Support Set; QS: Query Set.
SS and QS
FSL is a form of supervised machine learning (ML) composed of an SS used for training and a QS used for testing. The SS contains a K-way N-shot setting, where K represents the number of classes and N represents the number of images in each class. The QS contains images from the remaining samples.
In this study, the SS consisted of three classes, with each class containing three images (SLPs, n = 3; adenomas, n = 3; and NETs, n = 3), forming a three-way, three-shot setting. The remaining images were assigned to the QS (SLPs, n = 53; adenomas, n = 83; and NETs, n = 50). The SS and QS settings used in the FSL model are shown in Figure 2(b). Additional shot settings were explored, including 3-way 5-shot, 3-way 7-shot, and 3-way 10-shot settings.
Transfer learning
DL models trained on large datasets such as ImageNet (https://image-net.org/) are generally highly transferable to image classification and recognition tasks. In this study, because the images in the source and target datasets differed in type, transfer learning was performed to achieve a domain shift and enhance the feature extraction capability for endoscopic images. The transfer learning process, as illustrated in Figure 3, involved the original ResNet50 V2 model, dual pretraining, and feature extraction.

The structure of transfer learning. (a) Structure of the dual-pretrained ResNet50 V2 model: The model employed residual blocks as the fundamental module with an output of a 1 × 64 eigenvector. In each residual block, the BN-ReLu-Conv structure is used. (b) Feature extraction: For nine images in SS (S1–9), their corresponding eigenvectors FS1–9, each containing 64 features, were extracted through the dual-pretrained ResNet50 V2. For any image Qn (n = 1–186) in QS, its corresponding eigenvector FQn was extracted. SS: Support Set; QS: Query Set; BN: batch normalization; ReLU: rectified linear unit; Conv: convolutional layer.
ResNet50 V2, a DL model, was used to extract eigenvectors from endoscopic images. 15 It was first pretrained on ImageNet and subsequently pretrained on a ternary classification task using esophageal endoscopic images to better identify endoscopic features. The last fully connected layer (FCL) and classification layer (1000 categories) of the single pretrained ResNet50 V2 model were truncated and replaced with a new FCL and classifier layer (three categories of real-world endoscopic images). For each image, a 1 × 64 eigenvector was obtained using the dual-pretrained ResNet50 V2 model. In the SS, nine images were labeled S1–S9, and their corresponding eigenvectors were labeled FS1–FS9. In the QS, 183 images were labeled Q1–Q183, with corresponding eigenvectors labeled FQ1–FQ183. The performance of the dual-pretrained model was further validated using Grad-CAM to visualize the extracted features. 16
Metric learning
In image classification, selecting appropriate metrics to evaluate the distance or similarity between images is both essential and challenging. This has led to the development of metric learning. 17 The objective of metric learning is to learn a similarity or distance metric that maximizes interclass variation and minimizes intraclass variation, allowing images to be classified based on their distance values or similarities. A classical prototypical network for FSL based on metric learning has been proposed, 18 wherein the mean eigenvector of each SS image is calculated as its corresponding prototype representation. The similarity between each QS image and prototype is then used for classification using a nearest neighbor classifier.
In this study, the similarity between QS and SS images was calculated using Euclidean distance. For each QS image, nine Euclidean distance values (d1, d2…d8, d9) were computed based on the eigenvectors between the QS image and nine SS images, which were then used for further analysis. A classifier based on the KNN algorithm19,20 was subsequently applied for classification according to similarity. For each QS image, its category was determined by the k-nearest SS images. Figure 4 illustrates the overall process of metric learning. A detailed description of the methodology and model parameters is provided in Supplementary Method 3.
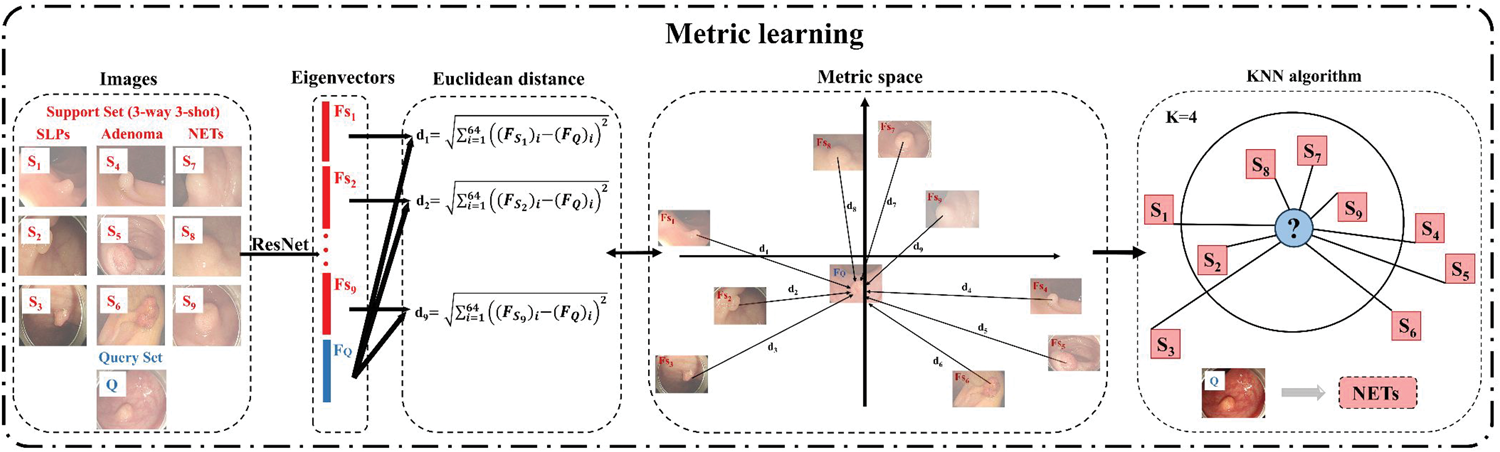
The process of metric learning. (a) Euclidean distance: For nine images S1–9 in SS and one image Q in QS, the corresponding eigenvectors FS1–9 and FQ were obtained. The Euclidean distance d1 =
Statistical analysis
Python software (version 3.9) and TensorFlow (version 2.8.0) were used to fit the FSL model. Statistical analysis was performed using R (version 4.2.2).
A confusion matrix was used to visualize the classification performance of DL models and endoscopists. Accuracy, recall, specificity, precision, and F1-score were used to evaluate the classification performance for each category (SLPs, adenomas, and NETs). The Matthews correlation coefficient (MCC), macro–area under the curve (macro-AUC), macro–F1-score, and Cohen’s kappa 21 were used to comprehensively evaluate and compare the performance between the FSL model and other models. Details of each indicator are provided in Supplementary Method 4.
Two endoscopists (one junior endoscopist with <5 years of experience and one senior endoscopist with ≥15 years of experience) were blinded to the diagnoses and independently classified the QS images into SLPs, adenomas, and NETs. McNemar’s test was used to compare the FSL model with both the conventional ResNet50 V2 model and endoscopists.
Results
Baseline characteristics of the collected NETs and polyps
A total of 195 eligible images (SLPs, n = 56; adenomas, n = 86; and NETs, n = 53) were included in this study. Among the SLPs, HP (n = 36), SSL (n = 18), and TSA (n = 2) accounted for 64.3%, 32.1%, and 3.6%, respectively. Among the adenomas, tubular (n = 35), villous (n = 21), and tubulovillous (n = 30) adenomas accounted for 40.7%, 24.4%, and 34.9%, respectively. Among the NETs, 71.7% (n = 38) were well-differentiated, 24.5% (n = 13) were moderately differentiated, 3.8% (n = 2) were poorly differentiated. Detailed characteristics of the included images are provided in Supplementary Table 1.
Performance of the dual-pretrained ResNet50 V2 model
As shown in Figure 5, Grad-CAM heatmaps highlighted the abnormal regions of the endoscopic images based on the dual-pretrained ResNet50 V2 model, demonstrating its high reliability.

Grad-CAM of dual-pretrained ResNet50 V2. Heatmaps of six examples generated using the Grad-CAM algorithm. The left panels show the original endoscopic images, the middle panels show heatmaps predicted by the dual-pretrained ResNet50 V2 model, and the right panels show combined original and predicted heatmap images. Grad-CAM: Gradient-weighted Class Activation Mapping.
Performance of the three-way three-shot FSL model
Three rounds were conducted to evaluate the performance of the proposed three-way three-shot FSL model. Figure 6(a) to (c) shows the confusion matrices of the FSL model in each round, which correctly identified an average of 29 SLPs, 67 adenomas, and 33 NETs.

Confusion matrices of the DL models and endoscopists. (a) FSL model, round #1. (b) FSL model, round #2. (c) FSL model, round #3. (d) Traditional ResNet50 V2 model. (e) Senior endoscopist (experience ≥15 years) and (f) junior endoscopist (experience <5 years). FSL: few-shot learning; DL: deep learning.
When distinguishing among SLPs, adenomas, and NETs, the FSL model achieved average accuracies and F1-scores of 0.783 and 0.593, 0.785 and 0.772, and 0.823 and 0.659, respectively, as shown in Table 1.
Performance of the models and endoscopists in distinguishing between SLPs, adenomas, and NETs.

FSL: few-shot learning; SLPs: serrated lesions and polyps; NETs: neuroendocrine tumors; DL: deep learning.
The overall process of the three-way three-shot FSL model was conducted in three rounds, where it was compared with traditional ResNet50 V2, senior endoscopist, and junior endoscopist, respectively. The performance of the FSL model, traditional model, and endoscopists in identifying each class was presented.
Figure 7 illustrates the comprehensive classification performance. The FSL model achieved an average macro-AUC of 0.731, macro–F1-score of 0.674, MCC of 0.526, and Cohen’s kappa of 0.523, as shown in Table 2. Performance under different shot settings is presented in Supplementary Table 2 and Supplementary Figure 1, which showed no consistent improvement compared with the three-way three-shot setting. In addition, correctly and incorrectly classified cases are presented in Supplementary Figure 2, along with the corresponding Grad-CAM heatmaps and annotations of potentially misclassified cases.

Visual performance of the ternary classification. The FSL model achieved superior classification performance compared with the traditional ResNet50 V2, senior endoscopist, and junior endoscopist. FSL: few-shot learning.
Performance of the models and endoscopists during the ternary classification task.

AUC: area under the curve; MCC: Matthews correlation coefficient; FSL: few-shot learning; DL: deep learning.
The three-way three-shot FSL model was conducted in three rounds, where it was compared with traditional ResNet50 V2, senior endoscopist, and junior endoscopist, respectively. The comprehensive performance of the FSL model, traditional model, and endoscopists during the ternary classification task was evaluated using four indicators: macro-AUC, macro–F1-score, MCC, and Cohen’s kappa.
Comparison of the FSL model with the conventional ResNet50 V2 model and endoscopists
Figure 6(d) to (f) displays the confusion matrices of the conventional model and endoscopists. Table 1 presents their performance in distinguishing SLPs, adenomas, and NETs, while Figure 7 and Table 2 depict the comprehensive classification performance.
As shown in Table 1, when identifying SLPs, the best performance was achieved by the FSL model (accuracy: 0.783, recall: 0.554, precision: 0.643, and F1-score: 0.593), followed by senior endoscopist (accuracy: 0.774, recall: 0.491, precision: 0.634, and F1-score: 0.553). The FSL model also achieved the highest accuracy (0.785), specificity (0.763), precision (0.736), and F1-score (0.772) for adenoma identification. When identifying NETs, the FSL model achieved the highest accuracy (0.823), recall (0.653), precision (0.673), and F1-score (0.659).
As shown in Figure 7, the FSL model demonstrated better overall performance in terms of macro-AUC, macro–F1-score, MCC, and Cohen’s kappa than the conventional DL model and endoscopists. For the ternary classification task, the FSL model, conventional DL model, senior endoscopist, and junior endoscopist achieved macro-AUC values of 0.731, 0.549, 0.631, and 0.526; macro–F1-scores of 0.674, 0.311, 0.581, and 0.462; MCCs of 0.526, 0.108, 0.363, and 0.175; and Cohen’s kappa values of 0.523, 0.085, 0.350, and 0.173, respectively. McNemar’s test was used to compare the FSL model with both conventional DL model and endoscopists (Supplementary Table 3). Significant differences in classification performance were observed between the FSL model and conventional DL model (χ2 = 7.21, p < 0.001) and between the FSL model and junior endoscopist (χ2 = 4.21, p = 0.005). However, no significant difference was observed between the FSL model and senior endoscopist (χ2 = 1.95, p = 0.051).
Discussion
To the best of our knowledge, this is the first study to use a three-way three-shot FSL strategy based on transfer learning and metric learning to distinguish between colorectal NETs and polyps. The ResNet50 V2 extractor was applied to obtain eigenvectors of endoscopic images, which were then converted into Euclidean distance vectors. A KNN classifier was used to categorize SLPs, adenomas, and NETs. Moreover, a dual pretraining method was innovatively adopted in this study to enhance the model’s ability to recognize and classify endoscopic images. Overall, the proposed method outperformed both the conventional DL model and endoscopists, demonstrating its feasibility for computer-aided diagnosis of NETs.
Differentiating colorectal NETs from polyps during endoscopy has long been a challenge in clinical practice, with important implications for subsequent diagnosis and treatment. The proposed FSL model provides a novel approach to assist endoscopists in the endoscopic diagnosis of colorectal NETs and polyps.
Endoscopic resection and biopsy are the preferred methods for diagnosing and treating polyps. Cold snare polypectomy (CSP) is an effective and safe resection option for small SLPs (<10 mm). For SLPs >10 mm, preoperative biopsy is necessary to determine the pathological type and assess malignancy before performing endoscopic mucosal resection (EMR). Endoscopic submucosal dissection (ESD) is recommended for larger SLPs (>20 mm), including those associated with dysplasia or invasive carcinoma.22–24 For traditional adenomas—the most important precancerous lesions of colorectal cancer—endoscopic resection and biopsy are recommended regardless of size.
In contrast to the treatment strategy for polyps, the management of NETs depends on accurate preoperative diagnosis and malignancy assessment. The invasive nature of preoperative submucosal biopsy may cause mucosal damage or adhesions, which increases the risk of perforation, bleeding, or tumor spread. Hence, it is not recommended for definitive diagnosis. Therefore, preoperative evaluation is needed for submucosal lesions. 25 Endoscopic resection is contraindicated if NETs invade the muscularis propria. Magnetic resonance imaging (MRI) is necessary for NETs ≥2 cm, particularly when muscularis propria invasion or metastasis is suspected. 26 There are no clear guidelines or randomized controlled trials that have formalized the therapeutic interventions for colorectal NETs. EMR is effective for low-risk NETs <0.5 cm. For NETs measuring 0.5–2 cm, ESD offers the advantage of wider and deeper resection margins for adequate removal. For NETs ≥2 cm or those with intermediate or high risk, surgical resection is generally performed. Formal partial colectomy is usually performed for colonic NETs, whereas right hemicolectomy is indicated for NETs near the ileocecal valve. Local transanal resection is recommended for lower rectal NETs (<5 cm from the anal verge), while transanal endoscopic microsurgery is used for middle and upper rectal lesions.27–29 Pathological evaluation is recommended for both surgical and endoscopic resections to assess vascular invasion, muscularis propria invasion, and margin status.26,30,31 Because NETs misdiagnosed as polyps may delay EUS evaluation and increase the risk of complications or metastasis, recall is a key performance metric. The proposed FSL model achieved higher recall than the conventional DL model and both senior and junior endoscopists, with only a small reduction in specificity, highlighting its potential clinical applicability.
Prior reviews have indicated that the application of DL to gastroenteropancreatic NETs remains limited.32,33 For instance, Udristoiu et al. 34 used a convolutional neural network and long short-term memory (CNN-LSTM) model to distinguish pancreatic NETs from ductal adenocarcinoma and chronic pseudotumoral pancreatitis, achieving an AUC >90%. For colorectal NETs, most studies have relied on structured clinical or biomarker data. Zhao et al. 35 developed a Surveillance, Epidemiology, and End Results (SEER)–based nomogram to predict 1-, 3-, and 5-year overall survival in patients with NETs. To the best of our knowledge, no prior study has applied DL to classify or detect colonoscopic images of colorectal NETs, likely due to the lack of large, curated image datasets for this rare disease. Furthermore, most studies applying DL to NETs have focused on CT or MRI using small, pancreas-predominant cohorts, leaving endoscopic imaging of colorectal NETs largely underexplored. 33 Our contribution is therefore application-oriented: we proposed an FSL framework based on transfer learning and metric learning to differentiate colorectal NETs and polyps on endoscopic images under conditions of real-world data scarcity, rather than introducing a new algorithm.
Currently, FSL strategies are mainly categorized into three types: transfer learning, 36 metric learning, 18 and meta-learning. 37 Transfer learning utilizes models previously trained on a sourced domain and adapts them to a target domain to improve performance. Metric learning evaluates similarities between samples by calculating distances or optimizing a metric space, allowing each unlabeled image to be classified according to intrinsic differences or similarities. Meta-learning, which consists of meta-training and meta-testing phases, is a task-level optimization approach that transfers knowledge from base-learning tasks to target tasks through a two-loop optimization process.
Based on these strategies, FSL is well-suited for model training with limited available data, particularly for the identification, segmentation, and classification of rare diseases across medical datasets. Another study 38 proposed an FSL technique to classify the severity of coronavirus disease 2019 (COVID-19), achieving an accuracy of 0.954. Lin et al. 39 developed a two-way three-shot framework to differentiate Crohn’s disease from intestinal tuberculosis, which outperformed both junior and senior endoscopists. Yin et al. 40 applied a metric learning–based FSL model to classify endoscopic images of gastric signet ring cell carcinoma, gastric adenocarcinoma, and gastric ulcers, achieving a classification accuracy of 0.794. In the field of polyp classification, FSL has also gained increasing attention. Krenzer et al. 41 proposed a deep metric learning–based FSL pipeline for polyp Narrow-band Imaging International Colorectal Endoscopic classification (NICE) I versus II classification using narrow-band imaging (NBI), reporting an accuracy of 81% after triplet-loss pretraining and fine-tuning, which demonstrated the feasibility of the FSL paradigm for polyp classification in data-scarce environments. For polyp segmentation, Wang et al. 42 addressed few-shot domain adaptation; on a cross-center external test set (CVC-300), it achieved a Dice coefficient of 93.6% with 20-shot supervision, outperforming traditional segmentation models. Although differences in label space, modality, and protocol preclude a direct numerical comparison, these studies collectively demonstrate the potential and feasibility of FSL for endoscopic image analysis under real-world data scarcity, laying the foundation for our research.
The proposed FSL classifier can serve as a second-reader aid during colonoscopy by flagging frames that exhibit NET-like mucosal patterns and providing explanatory heatmaps to assist endoscopists in their review. Beyond lesion identification, these outputs can support clinical decision-making by triaging cases for targeted EUS assessment and by preselecting lesions suitable for EMR or other polypectomy procedures, subject to clinician judgment.
This study has several advantages. First, it is the first study to employ an FSL model to distinguish between colorectal NETs, SLPs, and adenomas and utilize unstructured data from colorectal NETs for DL model development. Second, we conducted a rigorous multicenter epidemiological analysis based on endoscopic images and pathological diagnoses. Third, we used a dual-pretrained method to enhance generalization and verify cross-domain transferability from ImageNet to endoscopic images. Finally, we used metric learning to reduce training costs and minimize the risk of overfitting.
However, this study has several limitations. First, colorectal NETs are rare in routine endoscopy, which limited our case accrual and statistical power. In addition, pathological confirmation relied on endoscopically obtained specimens (biopsy/polypectomy), leading to the exclusion of cases diagnosed only after surgical resection. Second, confidence intervals for class-wise and composite metrics were not reported due to design and data constraints. Third, because the final classifier is a nondifferentiable KNN operating on ResNet50V2 embeddings, the Grad-CAM visualizations were computed on the backbone only, reflecting feature saliency rather than the exact decision basis of the KNN head. Finally, this was a retrospective, in silico analysis of still endoscopic images; thus, the findings are hypothesis-generating and require prospective, multicenter, video-based validation for clinical application. In the future, we aim to enhance endoscopy image quality, perform analyses of additional subtypes, incorporate binomial and bootstrap confidence intervals for improved uncertainty quantification, and apply the FSL framework for real-time, video-based recognition during endoscopy. To strengthen generalizability, we plan to conduct a multicenter, multivendor prospective study with external validation.
Conclusions
In this study, an FSL model integrating transfer learning and metric learning was developed to classify colorectal NETs, SLPs, and adenomas on white-light colonoscopy under real-world data scarcity. The proposed model outperformed a conventional DL model and achieved performance comparable to that of endoscopists. It may serve as a second-reader aid during colonoscopy by highlighting NET-like mucosal patterns and supporting downstream clinical decision-making. Overall, these findings demonstrate the feasibility of applying FSL to computer-aided diagnosis of rare or low-incidence diseases; however, clinical effectiveness remains to be established. Future work will focus on assessing the model’s generalizability and effectiveness for real-time, video-based recognition in clinical practice.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605251395564 - Supplemental material for Few-shot learning for the classification of colorectal neuroendocrine tumors and polyps on endoscopic images
Supplemental material, sj-pdf-1-imr-10.1177_03000605251395564 for Few-shot learning for the classification of colorectal neuroendocrine tumors and polyps on endoscopic images by Shiqi Zhu, Hailong Ge, Yu Wang, Jiaxi Lin, Rufa Zhang, Minyue Yin, Jinzhou Zhu and Chen Chao in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605251395564 - Supplemental material for Few-shot learning for the classification of colorectal neuroendocrine tumors and polyps on endoscopic images
Supplemental material, sj-pdf-2-imr-10.1177_03000605251395564 for Few-shot learning for the classification of colorectal neuroendocrine tumors and polyps on endoscopic images by Shiqi Zhu, Hailong Ge, Yu Wang, Jiaxi Lin, Rufa Zhang, Minyue Yin, Jinzhou Zhu and Chen Chao in Journal of International Medical Research
Footnotes
Acknowledgments
The authors are grateful to Elsevier Language Editing Services for their support.
Author contributions
Shiqi Zhu: Writing–original draft, Formal analysis, Data curation. Jiaxi Lin: Writing–original draft, Formal analysis. Rufa Zhang: Data curation. Minyue Yin: Formal analysis. Hailong Ge: Data curation. Yu Wang: Writing–review and editing. Jinzhou Zhu and Chen Chao: Funding acquisition, Conceptualization.
Data availability statement
Declaration of conflicting interests
The authors declare that they have no competing interests.
EQUATOR network guidelines
This study was performed in accordance with the Checklist for Artificial Intelligence in Medical Imaging (CLAIM).
Funding
This study was supported by Science and Technology Projects of Jintan District Health Bureau (JTYXH-2025-1-02), Medical Education Collaborative Innovation Fund of Jiangsu University (No. JDYY2023042), Frontier Technologies of Science and Technology Projects of Changzhou Municipal Health Commission (No. QY202309), Suzhou Clinical Center of Digestive Diseases (Szlcyxzx202101), and Youth Program of Suzhou Health Committee (KJXW2019001).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
