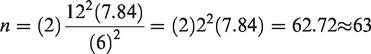Abstract
Objective
This study aimed to investigate the association of TLR3 single-nucleotide polymorphisms 1337 C/T (rs3775290) and 1234 C/T (rs3775291) with the risk of chronic hepatitis B virus infection.
Methods
This case–control study enrolled 136 participants (66 patients with chronic hepatitis B virus infection (cases) and 70 healthy controls). TLR3 polymorphisms were genotyped using polymerase chain reaction–restriction fragment length polymorphism. Demographic/clinical data were collected via standardized questionnaires and analyzed using Statistical Package for Social Sciences (significance level: p < 0.05).
Results
For TLR3 1337 C/T polymorphism, the CT genotype prevalence was significantly higher in cases (47.0%) than in controls (31.4%) (odds ratio = 0.42, 95% confidence interval: 0.20–0.87, p = 0.0187), while the CC genotype demonstrated protective effects (39.4% in cases vs. 62.9% in controls; odds ratio = 0.26, 95% confidence interval: 0.07–0.94, p = 0.0314). The C allele frequency showed significant between-group differences (odds ratio = 0.46, p = 0.0044). No associations were observed for 1234 C/T single-nucleotide polymorphism (p = 1.0). Biochemical analysis revealed significantly elevated alanine aminotransferase (p = 0.0044) and aspartate aminotransferase (p < 0.0001) levels in cases than in controls.
Conclusions
Regarding TLR3 1337 C/T single-nucleotide polymorphism, the CT genotype increases the risk of chronic hepatitis B virus infection (odds ratio = 0.42), while the CC genotype demonstrates protective effects (odds ratio = 0.26) in Sudanese populations, suggesting its utility as a therapeutic target and prognostic marker. No 1234 C/T associations were observed.
Background
Virus-induced hepatitis is a necro-inflammatory liver condition that can vary in severity. Chronic infection with hepatitis B virus (HBV) is often associated with chronic liver disease, which may progress to cirrhosis and hepatocellular carcinoma (HCC).1,2 According to a recent estimate, 296 million people were affected by chronic hepatitis B virus (CHB) infection in 2019, with an annual 1.5 million new infections. 3 This condition is considered a potentially life-threatening liver disease and poses a major global health challenge; among the 2 billion individuals infected by viruses globally, 400 million have CHB infection.2,3 Although most infected individuals can eliminate the virus, approximately 10% may develop CHB infection. 1 Although CHB infection is frequently asymptomatic, it is estimated that 15%–25% of patients may experience severe complications, including HCC and liver cirrhosis (LC). 2 In Sudan, the prevalence of HBV infection exceeds 8%, with hepatitis B surface antigen (HBsAg) positivity rates ranging from 6.8% in the central region to 26% in the southern region. Additionally, among blood donors in northern and central Sudan, HBV infection accounts for approximately 5.1%–5.6% of all infections in the population. 4 A study conducted by Yousif et al. found that of the 404 participants, 6.9% tested positive for reactive HBsAg and 47.5% for hepatitis B core antigen (HBcAg). 5
The Hepadnaviridae family comprises the DNA virus HBV. The HBV genome is characterized by its partially double-stranded DNA structure, with a total length of approximately 3200 base pairs. 6 This genome comprises four overlapping regions that encode the following essential proteins: the viral polymerase (P), surface antigen (HBsAg or S), and core antigen (HBcAg or C). 6 Serological markers such as HBsAg, hepatitis B surface antibody (anti-HBs), hepatitis B e antigen (HBeAg), anti-HBe, hepatitis B core IgM antibody (anti-HBc IgM), and immunoglobulin G serve as indicators of HBV infection.7,8 The identification of these serological markers is critical for diagnosing HBV infections, assessing the progression of CHB infection, determining the clinical stages of the infection, and monitoring the efficacy of antiviral therapies. 9
Toll-like receptors (TLRs) serve as pattern-recognition receptors that initiate the innate immune response by detecting pathogen-associated molecular patterns and damage-associated molecular patterns, thereby activating immune defense mechanisms.6,10 TLR3, also known as CD283, is a key component of the innate immune system responsible for recognizing double-stranded RNA (dsRNA) and initiating an early immune response against external antigens. This receptor is located on chromosome 4q35.6,10 Genetic polymorphisms, particularly single-nucleotide polymorphisms (SNPs) in the TLR3 gene, can influence susceptibility to viral infections, 11 highlighting the importance of TLR3 in the body’s defense against such pathogens. In the context of HBV, TLR3 detects dsRNA, which is formed as an intermediate during viral replication. In the nucleus, HBV DNA undergoes repair to form covalently closed circular DNA, a template for transcription of viral RNA. 12 This RNA, along with the viral polymerase, is packaged into the capsid, where it is reverse-transcribed into negative strand DNA, which is then used to synthesize the positive strand. 13
TLR3 activation leads to the stimulation of antiviral signaling pathways, particularly the upregulation of interferon-alpha/beta (IFN-α/β) and the production of antiviral proteins, bolstering the immune response. 14 As an intracellular receptor, TLR3 plays a pivotal role in mounting inflammatory responses to both DNA and RNA viruses, including HBV, through the recognition of viral dsRNA. 15 Consequently, TLR3 is essential in shaping the immune response to viral infections and may contribute to the persistence of these infections, potentially leading to HCC and LC. 16 Furthermore, genetic variations in TLR3 have been associated with increased susceptibility to infections and the progression of their clinical outcomes. 17 Studies have shown that TLR3 expression on peripheral dendritic cells (DCs), Kupffer cells, and hepatocytes is significantly reduced in patients with CHB infection compared with that in healthy individuals, suggesting a strong link between TLR3 and HBV infection. 18 The SNP rs3775290 (1377 C/T), located on exon 4 of TLR3, alters the ectodomain of the receptor, affecting receptor–ligand interactions and impairing its functional activity. 19
This research examined the influence of TLR3 polymorphisms, specifically 1337 C/T and 1234 C/T, on individuals’ susceptibility to CHB infection. These genetic variations were chosen for their potential to disrupt TLR3’s ability to recognize viral dsRNA, which is essential for initiating innate antiviral immune responses. Currently, the relationship between TLR3 rs3775291 polymorphism and HBV remains controversial. Some studies have reported that allele T at the TLR3 rs3775291 locus increases the risk of HBV infection.20,21 However, other studies have suggested that this T allele is not related to the risk of HBV infection.22,23 TLR3 is crucial in coordinating the immune system’s initial defense against viral pathogens, thereby rendering these polymorphisms particularly significant in the context of CHB infection pathogenesis. Through the analysis of a meticulously chosen group of patients with CHB infection alongside healthy controls, this study aimed to identify statistically significant correlations that illuminate the genetic underpinnings of CHB infection susceptibility. This investigation marks an important step forward in comprehending the relationship between host genetic factors and chronic viral infections, especially given the considerable global health challenges posed by this disease. By elucidating the functional consequences of these TLR3 polymorphisms on immune responses, this research lays the groundwork for precision medicine initiatives, facilitating personalized risk evaluations and the formulation of targeted therapeutic interventions. Such findings hold the promise of revolutionizing the clinical management of CHB infection, underscoring the study’s importance and originality.
Methods
Study setting and population
This investigation was conducted as a hospital-based case–control study in Sudan between 2022 and 2024, following the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines for case–control studies.
24
The study comprised 136 participants, including 66 patients diagnosed with CHB infection and 70 healthy controls, selected randomly according to specific inclusion and exclusion criteria. The study population comprised Sudanese individuals of varying ages who were referred to the Tropical Disease Hospital and diagnosed with CHB infection. Diagnosis of CHB infection was established through the presence of HBsAg and the absence of anti-HBs, with some participants exhibiting elevated alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels for more than 6 months. Conversely, individuals presenting with conditions other than CHB infection were designated as controls. Clinical and demographic data were collected from patient records and through a structured questionnaire. Control participants were matched to the patient group based on age, sex, and residential area (urban/rural). The sample size was calculated using the following formula:
For 80% power, Zβ = 0.84
For a 0.05 significance level, Zα = 1.96 r = 1 (equal number of cases and controls)
σ = 12.0
Difference = 6.0
Based on the abovementioned formula, this study included 136 study participants, including 66 cases and 70 controls.
Inclusion and exclusion criteria
Individuals diagnosed with CHB infection lasting for over 6 months and showing the presence of HBsAg alongside the absence of anti-HBs were selected as cases for this investigation. Patients with a viral load greater than 2000 were considered chronically active, while those with a viral load less than 2000 were regarded as chronically inactive. Additionally, control participants were recruited, ensuring that they had no history of HBV infection. They tested negative for HBsAg, anti-HBc, infectious diseases such as human immunodeficiency virus and hepatitis C virus as well as for alcohol dependence, smoking behaviors, cirrhosis, HCC or any other conditions that might affect liver function. Conversely, those with acute HBV infection and individuals who chose not to participate were excluded from the study.
Data collection and analysis
Data collection involved patients’ responses to questionnaires, which could be either self-administered or administered by others. A part of the data was sourced from the patients’ clinical records. The analysis was conducted using SPSS version 20, with the data undergoing two rounds of verification. Descriptive statistics, including means and proportions related to sociodemographic and obstetrical characteristics, were computed. The findings were illustrated through tables and figures. Additionally, the means and standard deviations of frequencies were calculated. The chi-square test was employed to examine the relationships between qualitative variables, and both univariate and multivariate analyses were performed. An independent t-test was also applied to the quantitative data. A 95% confidence interval (CI) was established for the odds ratio (OR), with a significance threshold set at p < 0.05. In this study, statistical tests were deemed significant if the p-value fell below 0.05.
Laboratory methods
Sample collection
Five milliliters of blood were collected from each participant in both the case and control groups. A 2-mL sample was drawn into a lithium heparin tube for serological testing, while a 3-mL venous blood sample was collected in an ethylenediaminetetraacetic acid (EDTA) tube for molecular analysis.
Enzyme-linked immunosorbent assay (ELISA) for HBsAg detection
The blood samples were subjected to centrifugation, leading to the separation of plasma, which was subsequently stored in a sterile container at −20°C within 6 h post-collection. The detection of HBsAg was conducted using ELISA. This assay was executed in accordance with the manufacturer’s guidelines (Fortress Diagnostic Limited), with each microplate analyzed independently for the calculation and interpretation of results. The results were determined by comparing the optical density values of each sample to the established cutoff (C.O.) for the plate (C.O. = NC × 2.1 (where NC refers to negative control optical density)). Samples exhibiting absorbance below the cutoff threshold were classified as negative for HBsAg, indicating its absence.
Polymorphism detection
DNA was extracted from EDTA blood samples obtained from both the case and control groups utilizing the guanidine extraction technique as described in a previous study. 25 The quantity and quality of the extracted DNA were verified using spectrophotometry (NanoDrop™), with samples exhibiting an A260/A280 ratio between 1.7 and 2.0 considered acceptable for downstream analyses. Additionally, gel electrophoresis (1% agarose) was performed to confirm DNA integrity. This process enabled the identification of two polymorphisms within the TLR3 gene through the polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) assay, as described in the previous study. Detailed information regarding the specific primers and PCR products is presented in Table 1.25,26 The amplification was performed using a Px2 Thermo Hybrid thermal cycler (Thermo Electron Co., MA, USA), with each SNP analyzed separately. The thermal cycler was programmed to initiate with a 2-min incubation at 94°C, followed by 35 cycles consisting of denaturation at 94°C for 13 s, annealing at temperatures ranging from 55°C to 60°C for 40 s, extension at 72°C for 1 min, and concluding with a final extension at 72°C for 10 min. After amplification, the PCR products were treated with 10 units of Taq I restriction enzyme (per 1 µg DNA) for the 1337 C/T SNP and 10 units of HincII enzyme (per 1 µg DNA) for the 1234 C/T SNP, followed by incubation at 37°C for 16 h. The resulting digested products were then separated using a 2% agarose Tris–borate–EDTA gel and visualized through ethidium bromide staining.
Primers and enzymes for detecting TLR3 SNPs.

SNPs: single-nucleotide polymorphisms.
Quality control
Appropriate protocols were implemented throughout the processes of specimen collection, transportation, storage, and processing to ensure optimal outcomes and uphold stringent conditions. Over 15% of the DNA samples that underwent PCR amplification were analyzed through DNA sequencing to confirm the genotyping results. The comparison of outcomes from PCR and DNA sequencing demonstrated complete agreement, achieving a concordance rate of 100%.
Ethics approval and consent to participate
This study was conducted in full accordance with the ethical principles of the Declaration of Helsinki (1975, as revised in 2013). Ethical approval was obtained from Omdurman Islamic University (Approval No. OIU/FMLS/DO/C17). All participants provided written informed consent after receiving detailed explanations of the study’s purpose, risks, benefits, and voluntary participation. All patient data were rigorously de-identified to ensure confidentiality, with all personal identifiers removed from the datasets, analysis records, and publications to prevent any possibility of participant identification, in compliance with both international standards and institutional requirements.
Results
Table 2 displays the mean ages and liver enzyme levels of patients with CHB infection and controls. There was no statistically significant difference in age between cases and controls (33.73 ± 1.346 vs. 33.09 ± 1.060 years; p = 0.8937), indicating proper age matching between the two groups. The significantly elevated liver enzyme levels in cases (ALT: 33.43 ± 3.585 vs. 22.37 ± 0.7794 U/L, p = 0.0044; AST: 34.89 ± 4.053 vs. 17.64 ± 0.6216 U/L, p < 0.0001) were consistent with the expected hepatic inflammation in CHB infection. Table 3 illustrates the demographic features of the study population, with males being substantially more infected than females (66.7% cases vs. 85.7% controls; χ2 = 6.078, p = 0.014, OR = 0.33 (95% CI: 0.14–0.78)), a pattern commonly observed in HBV epidemiology. The presence or absence of HBV infection was unaffected by marital status (p = 0.883). Geographic variations were evident, with Khartoum residents showing lower infection rates (84.8% cases vs. 94.3% controls; p = 0.0421). Occupation had a significant impact on HBV susceptibility (p = 0.006), particularly for farmers (9.1% vs. 0%), homemakers (19.7% vs. 2.9%), and students (15.2% vs. 10.0%), likely reflecting differential exposure risks. The 20–30 years age group showed the highest infection rate (47.0% cases vs. 32.9% controls; p = 0.071), possibly corresponding to periods of increased exposure.
Comparative variables of age and liver enzymes (mean ± standard deviation) in the study population.

AST: aspartate aminotransferase; ALT: alanine aminotransferase.
Demographic characteristics of the study population.
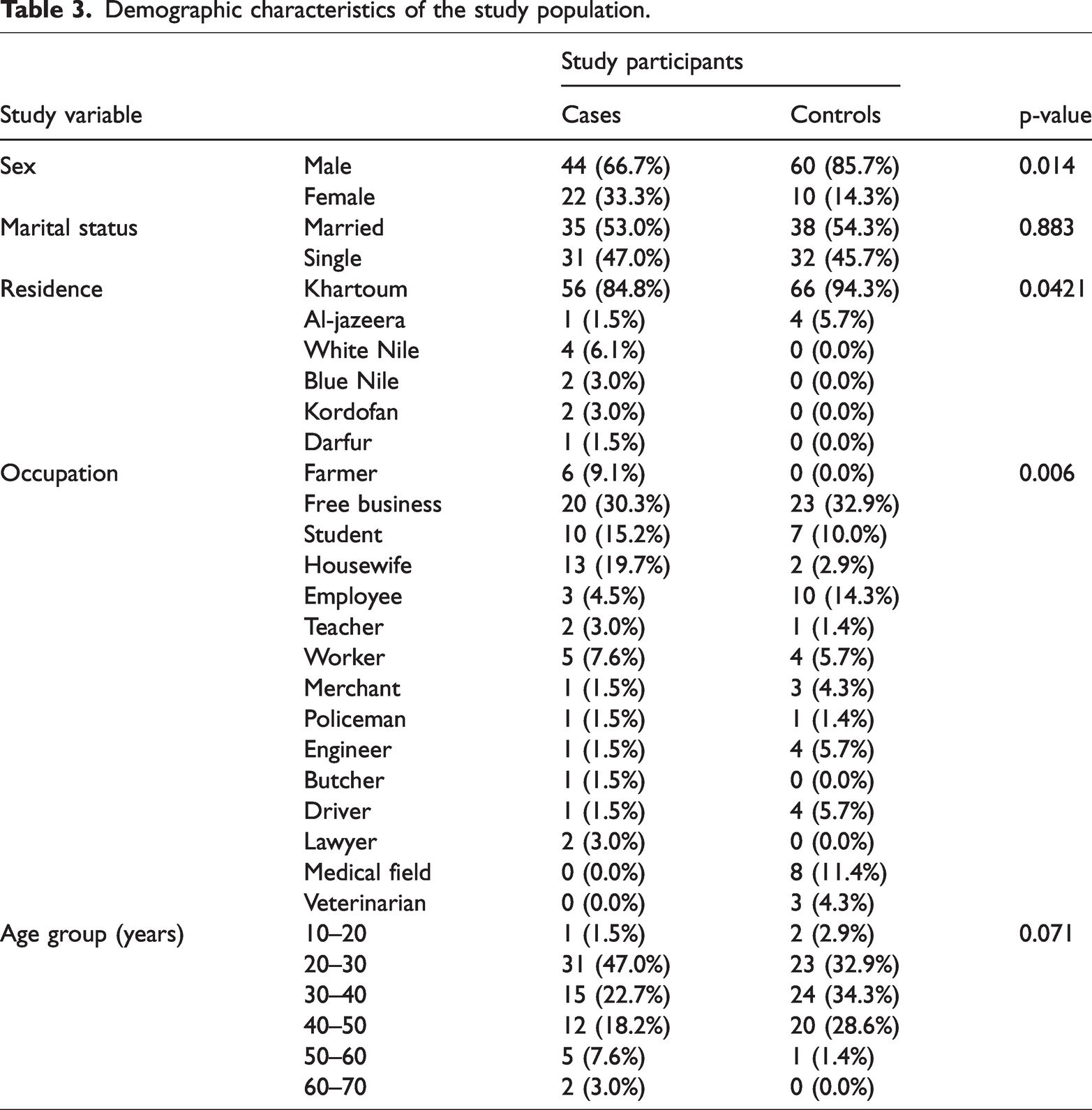
Figure 1 shows the PCR–RFLP results for TLR3 SNPs. For the 1337 C/T SNP (Table 4), the CT genotype was significantly more frequent in cases (47.0%) than in controls (31.4%; χ2 = 5.528, p = 0.0187, OR = 0.42 (95% CI: 0.20–0.87)), suggesting a heterozygote disadvantage. The CC genotype showed a protective effect, being less frequent in cases (39.4% vs. 62.9%; χ2 = 4.629, p = 0.0314, OR = 0.26 (95% CI: 0.07–0.94)). The C allele predominated in both groups (χ2 = 8.119, p = 0.0044, OR = 0.46 (95% CI: 0.27–0.79)), indicating selection pressure. All participants exhibited the TT genotype for 1234 C/T (p = 1.0), suggesting that this SNP is evolutionarily fixed in our population.

Agarose gel electrophoresis (2%) image of TLR3 1234 C/T and 1337 C/T SNP PCR–RFLP products. DNA ladder (L) is included for size estimation. (a) PCR product showing only the TT genotype (337 bp) for SNP 1234 C/T (b) PCR products displaying TT (337 bp), CT (337/275/62 bp), and CC (275/62 bp) genotypes for SNP 1337 C/T. DNA markers ranged from 50 bp to 1000 bp in panel A and up to 500 bp in panel B. PCR–RFLP: polymerase chain reaction–restriction fragment length polymorphism; SNP: single-nucleotide polymorphism.
The genotype and allele frequencies of TLR3 1337 C/T SNPs between patients with CHB infection and controls (chi-square test).

CHB: chronic hepatitis B virus; N.A.: Not applicable.
We have used N.A., as a zero value in the row prevents calculation.
Table 5 demonstrates that occupation significantly influenced CC genotype distribution (p = 0.025), with complete penetrance in farmers (100% cases) and high frequency in homemakers (80% cases), highlighting potential gene–environment interactions. Other demographic factors showed no significant associations (sex: p = 0.227–1.000; marital status: p = 0.622–1.000; residence: p = 0.159–0.497). Table 6 presents important clinical correlations, with cases showing abnormal ALT/AST levels exclusively possessing CC or CT genotypes (ALT-CC: p = 0.007; ALT-CT: p = 0.028; AST-CC: p = 0.002; AST-CT: p = 0.015), strongly suggesting that these variants influence disease severity. The TT genotype distribution remained stable regardless of liver function (p ≥ 0.05), possibly representing a neutral variant in disease progression. These findings collectively indicate that the TLR3 1337 C/T polymorphism significantly modulates both HBV susceptibility and clinical presentation in our population, with the CT genotype marking a particularly high-risk subgroup.
Study variables vs. TLR31337 (C/T) polymorphisms’ genotype distribution.

Study participants’ clinical data vs. TLR3 1337 (C/T) polymorphism distribution.

AST: aspartate aminotransferase; ALT: alanine aminotransferase.
Discussion
TLR3 plays a pivotal role in detecting dsRNA either within the viral genome or as a byproduct of replication, making it a key component of the host’s antiviral defense. 20 Recent studies have demonstrated that activating TLR3 with a specific ligand can suppress HBV replication in the liver of HBV transgenic mice. 27 In patients experiencing acute-on-chronic hepatitis B liver failure, the expression levels of TLR3 and IFN-β in DCs are significantly reduced compared with those in healthy individuals. 15 Similarly, individuals with CHB infection exhibit markedly lower TLR3 protein levels in peripheral blood mononuclear cells. 28 This case–control study examined 1337 C/T SNPs and found that the TT genotype was relatively rare in both CHB infection cases and controls. Notably, the CT genotype was significantly more prevalent in patients with CHB infection than in controls, while the CC genotype was considerably less frequent in patients with CHB infection than in healthy individuals. For the first time in Sudan, we identified a significant association between the CT genotype and susceptibility to CHB infection. Conversely, individuals with the CC genotype exhibited a reduced likelihood of developing CHB infection. In contrast, the TT genotype showed no correlation with HBV infection progression. Our findings indicated that the CT genotype of a functional TLR3 SNP is linked to an increased risk of CHB infection, reinforcing the potential role of TLR3 in HBV infection prevention and management. Moreover, genetic polymorphisms within the TLR3 gene may influence disease progression in patients with infectious diseases. 29 Although numerous SNPs have been identified in the human TLR3 gene, only a select few are predicted to induce structural modifications in the TLR3 protein. 30 Among these, the 1337 C/T polymorphism has been shown to impact TLR3 function, with the nucleotide substitution leading to an amino acid change from leucine (Leu) to phenylalanine (Phe) at position 412. 20 These in vitro findings suggest a clinical relevance for this SNP. Our study further revealed that although TT and CC genotypes exhibited no significant correlation and lower susceptibility to CHB infection, respectively, the CT genotype was associated with an elevated risk of CHB infection. However, given the low prevalence of the TT genotype, further research with a larger population sample is essential to confirm these observations.
The association of CHB infection with risk polymorphisms in TLR3 1337 C/T SNPs challenges previously reported findings.21,31,32 Sá et al. 22 reported no significant differences in the SNP between a cohort of 35 patients suffering from chronic infections and 299 healthy controls in Brazil. Variations in the demographic characteristics of the study population, sample size, geographic context, and the prevalence of HBV infection may have influenced these results. Nevertheless, the findings of the current study, along with several prior investigations, suggest a relationship between TLR3 1337 C/T SNPs and susceptibility to CHB infection.21,26 Notably, Goktas et al. 26 identified that the TLR3 SNP C1337 (rs3775290) with the CC genotype appears to be a risk factor among Turkish patients with active CHB infection, positing that the wild type may enhance the immune response against HBV. Conversely, in our study, the prevalence of the CC genotype was significantly lower among individuals with normal ALT and AST levels than in the control group, while the CT genotype was notably more prevalent in these cases. In contrast, patients exhibiting abnormal ALT and AST levels demonstrated a higher frequency of both CC and CT genotypes than controls. This observation suggests that individuals with the CT genotype experience diminished TLR3 functional activity, which could result in impaired HBV detection and an increased rate of HBV replication within the liver.
In this study, the prevalence of the C allele was significantly greater in both the case and control groups compared with the T allele frequencies associated with the 1337 C/T SNP (p-value = 0.005). This observation implies that these specific alleles may not play a role in the progression of CHB infection. Notably, these results were contradictory to those of a Tunisian investigation, which suggested a slight association between the T allele and CHB infection as well as an elevated risk of HBV-related HCC. 32 This discrepancy may be due to ethnic differences in the polymorphism, underscoring the importance of conducting validation studies across various population groups. In this investigation, both cases and controls exclusively presented the TT genotype for the 1234 C/T SNP, suggesting a lack of association between this SNP and CHB infection. These results are consistent with those of a Brazilian cohort study, which determined that the rs3775291 polymorphism did not significantly influence susceptibility to or chronicity of HBV infection. 22 In contrast, another research indicated that individuals possessing the CT and TT genotypes exhibited a 1.4- and 2.3-fold increased risk of developing CHB infection, respectively, compared with those with the CC genotype. 20 Li and Zheng 25 identified a link between the 1234 C/T SNP and an increased risk of HBV-related HCC; however, another study from China proposed that this polymorphism might serve as a protective factor against HBV-related HCC. 23 Given that the current study did not establish a correlation between rs3775291 and CHB infection, it is noteworthy that previous research has indicated a strong association between this SNP and HCC.
The rs3775291 variant is a missense mutation that results in the substitution of Phe with Leu at position 412 in the TLR3 protein. 33 In vitro studies have indicated that this SNP substantially impairs TLR3 functionality. 34 Moreover, TLR3 expression within HCC tissues appears to influence apoptosis, thereby restricting both HCC cell proliferation and angiogenesis. 35 Consequently, it is hypothesized that TLR3 dysfunction may play a role in vascular remodeling and liver tumor progression. Additionally, the minor A allele of rs3775291 has been linked to a 50% decrease in NF-κB activation compared with the G allele, along with a suppression of type I interferon signaling.36–38 These observations indicate that diminished TLR3 activity and related genetic alterations may contribute to the reduction of cytokine and chemokine production.
It is important to highlight that seven SNPs within the TLR3 gene have been linked to HBV infection. A study conducted among a Saudi Arabian cohort identified a significant correlation between the T allele of the rs1879026 polymorphism and an increased susceptibility to HBV. 38 However, research involving a Chinese cohort found no such association between this polymorphism and HBV-related liver disease. 36 These findings suggest that rs1879026 may have ethnic variability or may primarily influence HBV susceptibility rather than disease progression. Haplotype analysis further revealed that in the Saudi Arabian population, the GCGA haplotype (rs1879026-rs5743313-rs5743314-rs5743315) was significantly associated with HBV infection. 38 Meanwhile, individuals carrying the GT haplotype (rs1879026-rs3775290) exhibited a lower risk of developing CHB infection, LC, and HCC, whereas the GC haplotype was strongly linked to an increased likelihood of LC. 31
Our research did not identify any significant correlations between the TLR3 1337 C/T polymorphism genotypes and demographic variables such as sex, marital status, age categories, or residential location. However, it is essential to validate these results in larger cohorts to ensure their reliability. Additionally, although we explored the occupational links to TLR3 polymorphisms, we did not evaluate other vital socioeconomic factors, including income, education, access to healthcare, and living conditions. These elements are known to have a considerable impact on both susceptibility to HBV and the effectiveness of treatment. 39 To address these limitations, future studies should incorporate larger and more diverse cohorts to improve statistical power and reduce potential confounding effects. Additionally, longitudinal study designs could help assess disease progression over time and account for dynamic socioeconomic influences. Future studies should integrate these factors and increase sample sizes, as this approach would help validate the current genetic findings, analyze the interplay between socioeconomic conditions and genetic factors, and develop more effective, multifaceted interventions that address both biological and social determinants of HBV outcomes. Such thorough investigations could ultimately connect precision medicine with public health initiatives aimed at controlling HBV. Furthermore, although our study utilized rigorous data collection methods, any reliance on self-reported demographic or clinical data may introduce recall bias or inaccuracies. Future research should prioritize objective measurements and standardized data collection protocols to minimize such biases.
The recognition of the TLR3 C1337T CT genotype as a marker of susceptibility for CHB infection among Sudanese individuals underscores its potential applications in clinical and public health settings. Genotype-based stratification may facilitate earlier detection via enhanced monitoring (e.g. routine HBV screening) and timely antiviral treatment, which could help reduce the risk of progression to cirrhosis or HCC. 40 The influence of the SNP on TLR3-mediated immunity might also inform personalized treatment approaches, such as the selection of immune-modulating therapies for CT carriers. On a population scale, incorporating this biomarker into public health initiatives—through focused vaccination efforts, genetic counseling in areas with high prevalence, and improved resource distribution—could strengthen HBV prevention and management. 39 Additional validation across varied cohorts is essential to convert these insights into practical strategies for precision management of HBV susceptibility.
The current findings provide important groundwork for understanding genetic susceptibility to HBV infection. Although this study identifies associations between specific SNPs and TLR3 expression, the lack of functional validation remains a key limitation that should be addressed in future research. Further studies are needed to elucidate how these SNPs influence TLR3 expression and function, particularly in the context of HBV infection. These genetic variants may affect regulatory regions of the TLR3 gene or alter protein structure, potentially modulating receptor activation and downstream signaling. In silico analyses suggest that certain variants have functional consequences, such as changes in protein stability or binding affinity.29,41 However, experimental validation—including gene expression profiling and functional assays—is essential to confirm their precise role in immune responses to HBV. Additionally, expanding the analysis to include other TLR3 polymorphisms as well as SNPs in related immune genes would enhance the study’s scope. The current focus on only two TLR3 polymorphisms may have limited the generalizability of our findings regarding genetic susceptibility to CHB infection. A more comprehensive investigation, incorporating a wider range of genetic variations, could provide deeper insights into the genetic factors influencing CHB infection susceptibility and progression. Such efforts may uncover novel genetic markers associated with chronic infection and clinical outcomes.
Conclusion
This is the first study to establish a significant association between the TLR3 1337 C/T SNP and increased susceptibility to CHB infection in the Sudanese population. These findings suggest that this genetic variant can serve as both a prognostic marker and therapeutic target for CHB infection. However, the study’s limitations include analysis of only two TLR3 polymorphisms and a modest sample size. Further research with larger cohorts is needed to validate these findings and explore additional CHB-associated variants in the TLR3 gene.
Footnotes
Acknowledgments
We would like to thank the participants of our study.
Author contributions
K.E., N.A.M.O., and B.M.T.G. contributed to study design, data collection, and manuscript writing. A.M.A.N. and W.I.Y.A. performed laboratory analyses and data interpretation. N.A. supervised the research, conducted statistical analysis, and finalized manuscript editing. All authors have reviewed and approved the final manuscript.
Availability of data and materials
All data and materials related to the study are included in the current manuscript.
Declarations
This case–control study was conducted in accordance with the STROBE guidelines. All patient details have been de-identified. The patients provided written informed consent for sample collection and publication of the report.
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Ethics approval
The study was conducted according to the Declaration of Helsinki. Ethical approval for this study was obtained from Omdurman Islamic University (Approval number OIU/FMLS/DO/C17). All participants provided written informed consent before participation in the study.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.