Abstract
Metachromatic leukodystrophy (MLD) is a genetic lysosomal disease. Here, we investigated the role of prosaposin (
Keywords
Introduction
Metachromatic leukodystrophy (MLD) is a type of lysosomal storage disorder that affects sphingolipid metabolism and is classified as both leukodystrophy and sphingolipidosis.
1
MLD is characterized by neurodegeneration with progressive demyelination.
2
It is most commonly associated with a mutation in the
In addition to the
It should be noted that reports of
Materials and methods
Inclusion and exclusion criteria
For this
Family ascertainment
Because of the consanguinity, it was presumed that the inheritance pattern in this family followed an autosomal recessive pattern. The research included four family members: the proband, her parents (who were unaffected), and her brother.
DNA extraction
Peripheral blood samples were collected from the subject, her parents, and her brother after receiving informed consent. Genomic DNA was extracted from peripheral blood leukocytes using the DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s suggested protocol.
Whole exome sequencing (WES)
After fragmenting 1.5 μg of genomic DNA with a Covaris S2 Ultrasonicator (Covaris, Woburn, MA, USA), as recommended by the manufacturer, Agilent SureSelect libraries were generated for the WES analysis of family members II-1 and II-3. The exome capture was performed using the SureSelect Human All Exon V5 kit (Agilent, Santa Clara, CA, USA) for 51 Mb. The paired-end sequencing of barcoded exome libraries was performed on the HiSeq 4000 platform (Illumina, San Diego, CA, USA) using 150-bp + 150-bp reads. Sequences were paired demultiplexed and aligned to the human genome (NCBI build 37/hg19) by Burrows and Wheeler alignment. 11 Further analyses were performed using the Genome Analysis Toolkit (GATK), SAMtools, and Picard, with GATK Unified Genotyper employed for variant calling. Variant annotation was performed with Ensembl release 82 and ANNOVAR Documentation was used to implement filtering using various single nucleotide polymorphism (SNP) databases, such as dbSNP (build 138; http://www.ncbi.nlm.nih.gov/projects/SNP/index.html), Exome Variant Server (release ESP6500SI-V2; http://evs.gs.washington.edu/EVS/), 1000 Genomes Project (released May 2012; http://www.1000genomes.org/), and HapMap CHB (release #28; http://hapmap.ncbi.nlm.nih.gov/). To predict the possible impact of variants, SIFT (http://sift.jcvi.org/), Polyphen-2 (http://genetics.bwh.harvard.edu/pph2/), Mutation-Taster (http://www.mutationtaster.org/), and Human Splicing Finder (http://www.umd.be/HSF3/) were used.
Sanger sequencing
The candidate variants were recognized using Sanger sequencing. Primers were designed using Primer Premier 5.0 to amplify fragments containing the variant with the following sequences: forward, 5′-CCAGAGCTGGACATGACTGA-3′; reverse, 5′-TTCTTAATGGGCTCCACCAG-3′. The PCR products underwent sequencing using the ABI PRISM 3730 DNA Analyzer (Applera Corporation, Norwalk, CT, USA) and the ABI PRISM Big-Dye Terminator Cycle Sequencing v.3.1 Ready Reaction Kit.
Protein structural and functional analysis
To assess the structural consequences of
Cell culture
Cells were maintained in an appropriate culture medium supplemented with necessary growth factors and antibiotics at 37°C in a humidified incubator with 5% CO2.
Plasmid construction and transfection
Immunofluorescence staining
Cultured cells were washed twice with phosphate-buffered saline (PBS), then fixed with 3.7% paraformaldehyde for 10 minutes at room temperature. Subsequently, the cells were permeabilized with 0.1% Triton X-100 for 10 minutes at room temperature. Next, the cells were blocked with 5% bovine serum albumin for 30 minutes at room temperature, then incubated with the primary antibody (anti-Flag, clone M2, Sigma-Aldrich) at a 1:500 dilution overnight at 4°C. An Alexa Fluor 488-conjugated secondary antibody (Invitrogen) was used for detection (1:1000 dilution) and incubated for 1 hour at room temperature in the dark. The sample was examined using a confocal microscope (TCS SP5; Leica, Wetzlar, Germany), which was equipped with a Plan-Apochromat 63×NA 1.4 oil differential interference contrast objective.
Lysosome staining
Lysosomes in cells were visualized using LysoSensor™ Green DND-189 (Invitrogen). Cells were seeded on coverslips, allowed to adhere, and then treated or transfected as previously described. Subsequently, the cells were incubated with 1 μM LysoSensor™ Green DND-189 for 30 minutes at 37°C in a humidified incubator with 5% CO2. After PBS washes, the cells were fixed with 4% paraformaldehyde and mounted onto glass slides. DAPI was used for nuclear counterstaining. Images were captured using a confocal microscope, then fluorescence intensity was quantified using ImageJ software (National Institutes of Health, Bethesda, MD, USA). The data are presented as a percentage of the fluorescence intensity relative to the WT group.
Cathepsin D activity assay
A cell lysate-based assay was used to examine cathepsin D activity (ab65302, Abcam, Cambridge, MA, USA) following the manufacturer’s protocol. Fluorescence intensity was measured with an excitation wavelength of 328 nm and an emission wavelength of 460 nm using a fluorometer (Thermo Fisher Scientific). The data are presented as a percentage of fluorescence intensity relative to the WT cohort.
Transmission electron microscopy (TEM)
The cells were fixed with 2.5% glutaraldehyde overnight at 4°C. Following fixation, all specimens were post-treated with 2% OsO4 (Electron Microscopy Sciences, Hatfield, PA, USA). The specimens were dehydrated using a series of fractional acetone, then immersed in a mixture of acetone and resin (1:1). Following this, they were embedded in Ebon (Sigma-Aldrich). A slide containing an ultra-thin section (70 nm) was doubly stained using lead nitrate (supplied by Sigma-Aldrich) and uranyl acetate (provided by Ted Pella, Redding, CA, USA). A Tecnai G2 Spirit transmission electron microscope (FEI Company, Hillsboro, OR, USA) and Eagle 4k HS digital camera (FEI Company) were used to capture the images.
Results
Clinical report
At the time of the report, the female proband was 2 years and 5 months of age. She was born on time after a trouble-free pregnancy and normal delivery. She was diagnosed with motor development regression 7 months prior to the report. She began walking independently at 1 year and 2 months old, began to trot at 1 year and a half, and was able to speak short sentences by 1 year and 9 months of age. However, from about 1 year and 10 months old, her motor development began to regress. She started having difficulty walking, gradually losing her ability to walk, then experienced increased muscle tension in both lower limbs, limb pain, and developed intentional tremors in both upper limbs. Over time, her language capabilities and cognitive development decreased.
Upon hospital admission, a brain magnetic resonance imaging (MRI) scan showed diffuse long T1 and long T2 shadows in the bilateral white matter, indicating potential white matter lesions (Figure 1). Electromyography revealed a slowed conduction velocity in the bilateral common peroneal nerve and a low amplitude of evoked potential. The conduction speed of the motor nerves of the bilateral tibial nerves was also decreased, suggesting neurogenic damage in both lower limbs. Auditory evoked potentials showed that bilateral wave intervals III–IV were greater than those of I–III. A Gesell development assessment indicated a motor score of 32, severe growth retardation, and mild growth retardation in adaptability, speech ability, and social communication.

Brain magnetic resonance imaging (MRI) scan of the proband. The axial image using T2-weighting demonstrates bilaterally high-intensity white matter that is abnormal in the centrum semiovale. This image displays a tigroid pattern with a radial arrangement of hypointense stripes, clearly indicated by the arrows. Importantly, the white matter of the subcortical areas remains unaffected. The FLAIR image in the axial plane displays an extensive region of hyperintense white matter within the centrum semiovale. FLAIR, fluid-attenuated inversion recover.
The patient’s seven-year-old brother exhibited a similar history. He displayed significant spasticity in his limbs, was unable to walk or sit independently, and experienced severe mental and language retardation. As a result, he was unable to care for himself.
Molecular analysis revealed the same homozygous c.643A>G variant in the elder brother, indicating that he also had the same disease.
Molecular analysis
Figure 2(a) displays the family’s pedigree. Targeted sequencing revealed a homozygous c.643A>G mutation within

A novel pathogenic prosaposin (
We submitted these mutation data to the Leiden Open Variation Database (LOVD). The individual ID is #00377058. The p.N215

Crystal structure of the prosaposin (PSAP) protein with mutated amino acids. (a) The multi-species PSAP protein sequence alignment generated using ClustalX2 software. The affected amino acid is marked in red and (b) the crystal structure of PSAP amino acids 198–263 (wild-type PSAP). The impacts of the p.N215D mutation and previously reported p.N215H mutation on PSAP protein structure are predicted. The positions of the mutated amino acids are highlighted by white boxes.
Using Swiss-Model software, we predicted the structural effects of the p.N215D mutation on the PSAP protein. Additionally, we predicted the effects of the p.N215H mutation, another amino acid change in the same codon. This mutation has been previously reported in a Spanish ancestry family by Wood et al. (2000) and is classified as pathogenic in ClinVar. 12 The results showed that p.N215D and p.N215H have a greater impact on the non-helical structure of PSAP. The p.N215D and p.N215H mutants show more structural similarity compared with the WT PSAP protein (Figure 3(b)).
PSAP p.N215D induces lysosomal abnormalities
To explore the effect of the p.N215D mutation on PSAP function, we constructed a mutant expression plasmid and overexpressed it in mammalian cells. The results suggested that the p.N215D mutation had no impact on PSAP distribution, which was correctly localized in the lysosomes (Figure 4).

Lysosomal localization of the p.N215D prosaposin (PSAP) mutation. (a) HeLa cells transfected with either pCMV-PSAP-Flag, a plasmid encoding wild-type (WT) PSAP, or a PSAP mutant (p.N215D) were co-immunostained with p.N215D anti-Flag (green) and anti-LAMP1 (red) antibodies. The scale bar represents 10 μm and (b) a statistical graph of the colocalization of WT PSAP and p.N215D PSAP mutant with the lysosomal marker protein LAMP1.
It should be noted that lysosome staining with pH-sensitive LysoSensor™ dyes showed that the lysosomal pH decreased in cells overexpressing the p.N215D mutant compared with WT PSAP (Figure 5(a,c)). TEM results also showed that the p.N215D mutation caused abnormal lysosome accumulation in the cells (Figure 5(b)). More importantly, the PSAP p.N215D mutant resulted in decreased cathepsin D activity in lysosomes compared with WT PSAP, suggesting impaired lysosomal function (Figure 5(d)).
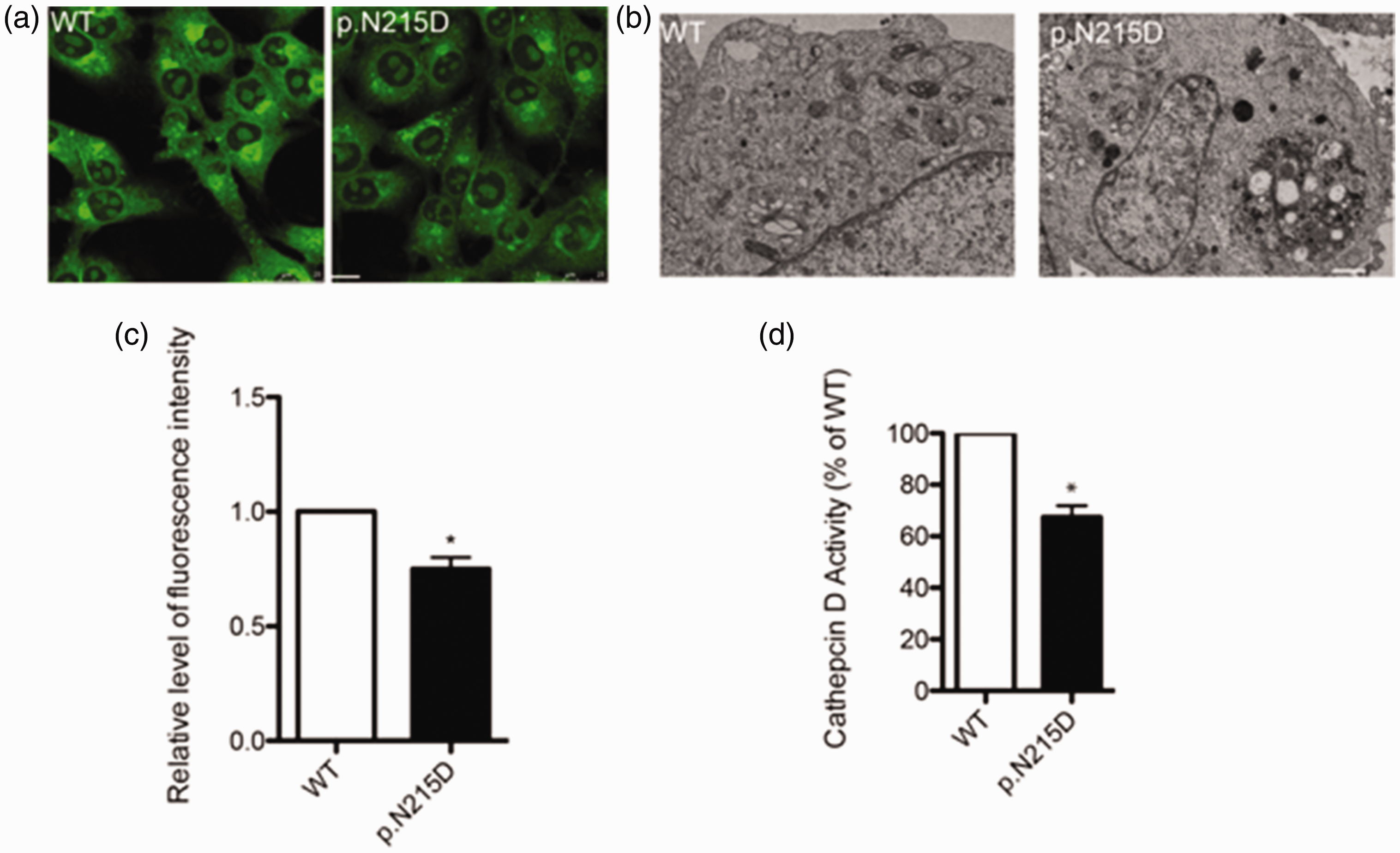
The p.N215D prosaposin (PSAP) mutation induces lysosomal abnormalities. (a) The lysosomes in cells overexpressing wild-type (WT) PSAP or the p.N215D mutant were stained with LysoSensor™ Green DND-189. The scale bar denotes 10 μm. (b) Transmission electron microscopy analysis of the lysosomes in HeLa cells expressing WT PSAP or the p.N215D mutant. Representative images of damaged lysosomes (indicated by white arrows) are displayed. The scale bar signifies 1 μm. (c) The fluorescence intensity statistics from panel a and (d) cathepsin D activity levels after the cells were transfected with either WT PSAP or the p.N215D mutant. *
Discussion
The
This study is the first to report a case in a Chinese family where clinical and neuroradiological features of the proband led to a suspected diagnosis of MLD. Genetic testing was pursued directly without prior measurement of leukocyte arylsulfatase A enzymatic activity. Although it has been documented that arylsulfatase A enzymatic activity may not always correlate with the MLD phenotype,
26
examining this factor remains valuable when studying MLD and
PSAP protein is primarily localized to the lysosomes, but evidence has suggested that it is present in other cellular organelles and locations. For example, PSAP has been identified in the Golgi body and endoplasmic reticulum, 3 which are involved in protein synthesis, modification, and transportation, which is consistent with its role in the cell. PSAP can also be secreted outside of cells, where it can act as a neurotrophic factor influencing cell survival, differentiation, and myelination. 3 PSAP can bind to the sortilin receptor, which is also a binding partner of progranulin (PGRN). When PSAP is bound to sortilin, it prevents PGRN from binding to the same receptor and being internalized and degraded in the lysosome. This leads to increased PGRN levels outside of the cell. 27 PGRN has also been linked to certain neurodegenerative conditions, including Alzheimer’s disease and frontotemporal dementia.28,29 In the current study, the p.N215D mutation did not alter the lysosomal localization of PSAP. However, whether it affects the PSAP distribution in other organelles or extracellular secretion remains to be further studied. Moreover, the mechanisms and physiological significance of this extracellular role are not as well understood as its intracellular functions. Given the abnormal lysosome accumulation observed via TEM and decreased cathepsin D activity, our data suggest that the p.N215D mutation can impact PSAP functionality.
Footnotes
Acknowledgements
We thank the patient and her family for participating in the present study and all those who helped us complete the research.
Author contributions
SY and GH conceived and designed the study. XL and ZL curated the data. XK and GH performed the formal data analysis. SY and XK developed the methodology. XL and SY performed the investigations and wrote, reviewed, and edited the manuscript.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Ethics statement
The Institutional Ethics Committee of Changsha Hospital for Maternal & Child Health Care of Hunan Normal University approved this study for publication (approval no. 2020002). Written informed consent was obtained from all participants prior to enrollment in this study, with all individuals consenting to publish their images. The reporting of this study conforms to CARE guidelines. 30
Funding
This work was supported by the Health Commission of the Hunan Province (B2019138) and Major Scientific and Technological Projects for collaborative prevention and control of birth defects in Hunan Province (2019SK1010).
