Abstract
Objective
To explore the levels of expression and clinical role of peroxiredoxin 6 (PRDX6) in lung adenocarcinoma.
Methods
This retrospective study used a series of bioinformatics methods to detect the levels of expression of and mutations in the PRDX6 gene in a range of cancers and lung adenocarcinoma. Immunohistochemistry was used to verify the levels of expression of PRDX6 protein in samples of lung adenocarcinoma compared with normal adjacent tissue. The effect of PRDX6 gene knockdown on the in vitro proliferation of a lung adenocarcinoma cell line was measured. Bioinformatics methods were used to determine the diagnostic value and impact on survival of the PRDX6 gene in patients with lung adenocarcinoma.
Results
The results showed that the PRDX6 gene was highly expressed in lung adenocarcinoma and there were five mutations at different sites on the gene. PRDX6 promoted the proliferation of the lung adenocarcinoma cell line. The survival duration of lung adenocarcinoma patients with high levels of PRDX6 gene expression was significantly shorter than that of patients with low PRDX6 gene expression.
Conclusion
PRDX6 is highly expressed in lung adenocarcinoma and higher levels of expression of the PRDX6 gene were associated with a poorer prognosis.
Introduction
Lung cancer is a malignant tumour that poses a serious threat to human health and its morbidity and mortality rates are among the highest in all cancers.1,2 The most common histological subtype of lung cancer is non-small cell lung cancer, mainly divided into squamous cell carcinoma and adenocarcinoma.3–5 Despite recent advances in cancer treatment for lung adenocarcinoma, which include surgical resection, immunotherapy, chemotherapy and radiation therapy, the prognosis of lung adenocarcinoma remains disappointing, with a 5-year survival rate <20%.6,7 Therefore, there is an urgent need to identify reliable biomarkers to predict the prognosis of lung adenocarcinoma patients.
Peroxide reductases (PRDXs) are highly conserved members of the peroxidase family and maintain the homeostasis of intracellular reactive oxygen species and family members are expressed in most organisms and participate in various biological processes.8,9 In mammals, six PRDX members have been identified and these are subdivided into three subfamilies: typical 2-Cys (PRDX1, PRDX2, PRDX3 and PRDX4), atypical 2-Cys (PRDX5) and 1-Cys (peroxiredoxin 6; PRDX6).10,11 As one of the most unique members of the PRDX family, PRDX6 is the only member of the 1-Cys subfamily. 10 PRDX6 plays a leading role in maintaining redox homeostasis in mammalian cells by reducing a wide range of peroxides in cells. 12 In recent years, research has shown that PRDX6 is involved in the occurrence and development of various cancers, including liver cancer, 13 colorectal cancer14,15 and cervical cancer. 16 Therefore, further research is needed on the expression levels and clinical role of the PRDX6 gene in lung adenocarcinoma.
This current study investigated the expression levels and clinical role of the PRDX6 gene in lung adenocarcinoma in order to provide information useful to the future clinical treatment of lung adenocarcinoma.
Materials and methods
Bioinformatics analysis in lung adenocarcinoma
Data for 600 files were downloaded from The Cancer Genome Atlas (TCGA) database via the Genomic Data Commons data portal (https://portal.gdc.cancer.gov/), which included 541 lung adenocarcinoma tissues and 59 adjacent tissues. R-4.3.1 software for code visualization processing was used to show the gene expression in lung adenocarcinoma. A volcanic plot was used to identify the top 10 upregulated and downregulated genes in lung adenocarcinoma. The correlation between the PRDX6 gene and these 20 genes was analysed. The expression of the PRDX6 gene in lung adenocarcinoma was demonstrated using a box-plot. The TCGA database was used to assess the diagnostic potential of PRDX6 expression in lung adenocarcinoma. This assessment was followed by plotting the receiver operating characteristic (ROC) curve. The TCGA database was used to analyse the survival duration of patients based on the levels PRDX6 gene expression in lung adenocarcinoma.
cBioPortal database analysis
PRDX6 was entered into the Quick Search module through cBioPortal (https://www.cbioportal.org/). The ‘Cancer styles summary’ module was used to display the absolute counts of mutation types of PRDX6 in Pan-Cancer. The ‘Mutations’ module was used to screen and display the mutation site and mutation type of PRDX6 in lung adenocarcinoma.
UACLAN database analysis
PRDX6 was entered into the TCGA module through UALCAN (https://ualcan.path.uab.edu/index.html). The ‘Pan-cancer view’ module was used to display the expression of PRDX6 in a range of cancers (Pan-Cancer).
Patient information and preparation of tissue specimens
This retrospective study recruited consecutive patients with lung adenocarcinoma from the Fourth Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu Province, China using a random table between January 2022 and January 2023. The experimental group consisted of lung adenocarcinoma tissue from lung adenocarcinoma patients, while the control group consisted of corresponding adjacent normal tissue from the same patients. This process was confirmed by a pathologist. All patients were clinically diagnosed with lung adenocarcinoma. The inclusion criteria were as follows: (i) computed tomography-guided percutaneous lung biopsy for the first confirmed occurrence of lung adenocarcinoma; (ii) an Eastern Cancer Collaborative Group score of 0–2; (iii) an estimated survival time of >3 months; (iv) no serious underlying heart and lung diseases; (v) routine blood analyses and liver and kidney function were normal before treatment. The exclusion criteria were as follows: (i) multiple epidermal growth factor receptor (EGFR) gene mutations; (ii) metastases from the primary lung tumour.
This study was a retrospective study and obtained an exemption from the Ethics Committee of the Fourth Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu Province, China. All patient details were de-identified and patients were required to provide verbal informed consent.
Immunohistochemistry
All paraffin samples were cut into 4-mm thick sections and dewaxed. Antigen retrieval was achieved by four rounds of heat treatment in a microwave oven on a medium setting for 6 min each time. The sections were pretreated with 10% normal goat serum at 37°C for 60 min. The sections were incubated with rabbit antihuman PRDX6 polyclonal antibody (catalogue number PA5-24632; Thermo Fisher Scientific, Rockford, IL, USA; 1:150 dilution) at 37°C for 30 min. Sections were rinsed with 0.01 M phosphate-buffered saline (PBS; pH 7.5) three times at room temperature for 3 min each time. The sections were then incubated with a goat antirabbit immunoglobulin G (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (catalogue number A-11008; Thermo Fisher Scientific; 1:100 dilution) at 37°C for 30 min. Sections were rinsed with 0.01 M PBS (pH 7.5) three times at room temperature for 3 min each time. The sections were stained with 0.01% 3,3′-diaminobenzidine at room temperature for 5 min and then rinsed with 0.01 M PBS (pH 7.5) three times at room temperature for 3 min each time. All samples were dehydrated and imaged through a CX31 optical microscope (Olympus Optical, Tokyo, Japan). Finally, ImageJ software was used to quantitatively evaluate the results.
Cell culture and CCK8 assay
The cell lines used in this study were all from our laboratory. All cell lines were maintained in Roswell Park Memorial Institute 1640 containing 10% heat-inactivated fetal bovine serum, 100 U/ml penicillin and 100 mg/ml streptomycin at 37°C in a 5% CO2 atmosphere. NCI-H1395 cells were seeded into a 96-well plate and cell proliferation was confirmed in each group before transfection. The cells were transfected with 1 µl small interfering RNA (siRNA) (siRNA-PRDX6; experimental group) or 1 µl siRNA-NC (control group). Lipofectamine™ 2000 (catalogue number 11668500; Thermo Fisher Scientific) was used as a transfection reagent in this study. Cell proliferation was measured at 24 and 48 h using a cell counting kit (CCK8 assay kit; C0037, Beyotime, Shanghai, China). An ultraviolet visible spectrophotometer (E0946; Beyotime) was used to measure absorbance in each well at a wavelength of 450 nm to represent cell proliferation.
Statistical analyses
All statistical analyses were performed using IBM SPSS Statistics for Windows, Version 21.0 (IBM Corp., Armonk, NY, USA). Three independent experiments were performed. Data are expressed as mean ± SEM. An unpaired Student’s t-test was used to assess differences between two groups. Survival analysis was determined using the Kaplan–Meier method. A two-sided P-value <0.05 was considered statistically significant. All assays were performed at least three times. The study conforms to STROBE guidelines. 17
Results
This retrospective study included clinical data from patients with lung adenocarcinoma from the TCGA database (Table 1). The study also included 25 patients with lung adenocarcinoma from the Fourth Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu Province, China (Table 2).
Demographic and clinical data for patients with lung adenocarcinoma patients (n = 517) from The Cancer Genome Atlas database.

Clinical data for 541 lung adenocarcinoma samples were downloaded, but 24 had incomplete or missing clinical data so they were excluded from these analyses.
Demographic and clinical data for patients with lung adenocarcinoma patients (n = 25) who were enrolled at the Fourth Affiliated Hospital of Nanjing Medical University, Nanjing, Jiangsu Province, China.

The levels of PRDX6 gene expression in lung adenocarcinoma was moderate in a range of cancers (Figure 1a). Moreover, the mutation frequency of the PRDX6 gene in lung adenocarcinoma ranked fourth among a range of cancers (Figure 1b).

Levels of expression of the peroxiredoxin 6 (PRDX6) gene in a range of cancers based on samples in The Cancer Genome Atlas (TCGA) database (a). Mutation of the PRDX6 gene in a range of cancers (b). The colour version of this figure is available at: http://imr.sagepub.com. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma; CHOL, cholangiocarcinoma; COAD, colorectal adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, oesophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and Neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LGG, brain lower grade glioma; OV, ovarian serous cystadenocarcinoma; MESO, mesothelioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; PAAD, pancreatic adenocarcinoma; PRAD, prostate adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; READ, Rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; LAML, acute myeloid leukaemia; TGCT, testis germ cell tumours; THCA, thyroid carcinoma; THYM, thymoma; STAD, stomach adenocarcinoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
With regard to gene expression levels in lung adenocarcinoma, there were genes with high levels of expression and those with low levels of expression (Figure 2). The correlation between the PRDX6 gene and 10 genes with high expression and 10 genes with low expression in lung adenocarcinoma was analysed (Table 3). The results showed that the PRDX6 gene was positively correlated with nine out of 10 genes with high expression, of which seven were significant (P < 0.05 for all comparisons). The PRDX6 gene was inversely correlated with nine out of 10 genes with low expression, of which eight were significant (P < 0.05 for all comparisons).
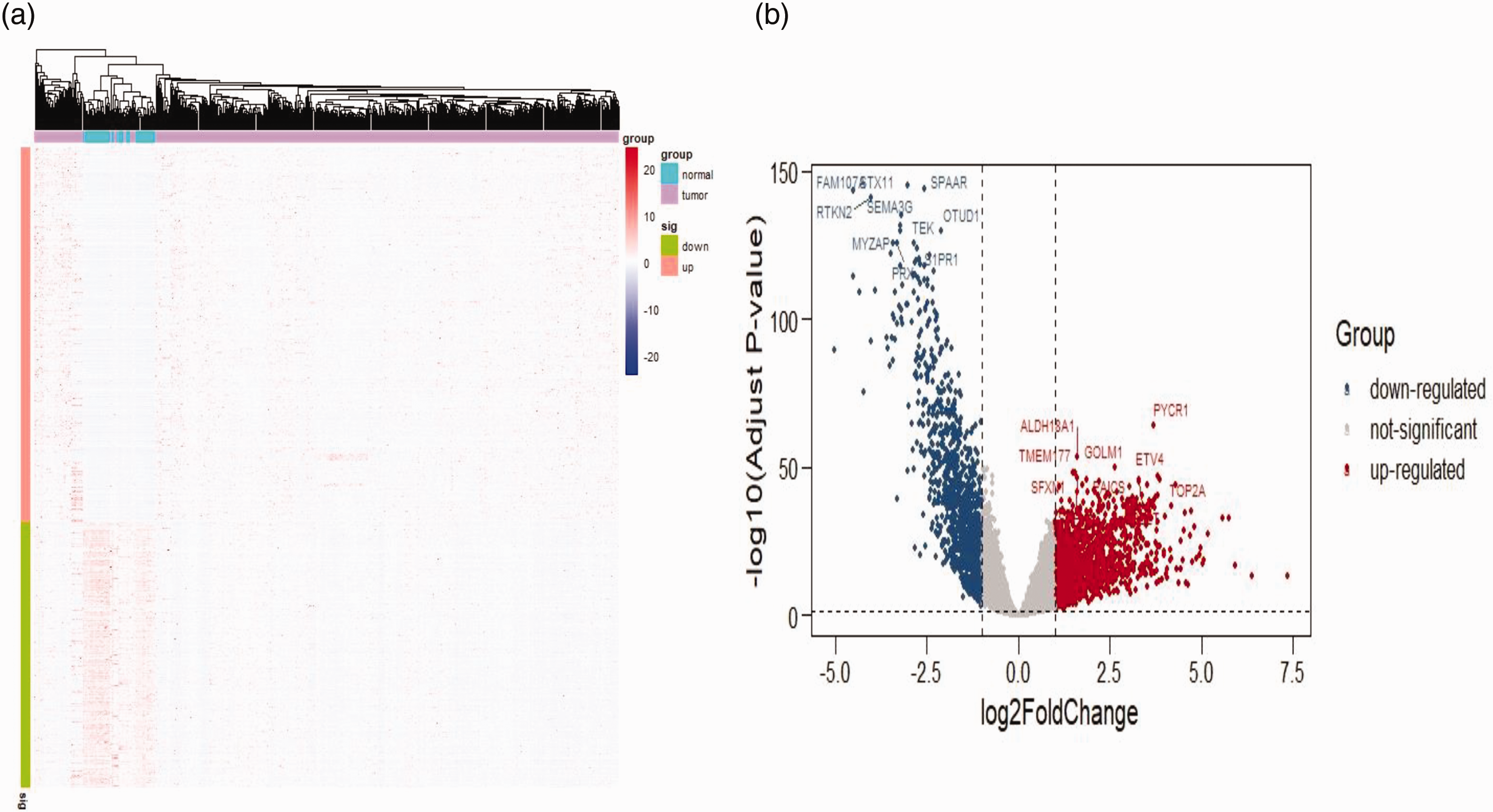
Levels of gene expression in lung adenocarcinoma: (a) heatmap of peroxiredoxin 6 (PRDX6) gene in lung adenocarcinoma and (b) volcano plot of lung adenocarcinoma. The colour version of this figure is available at: http://imr.sagepub.com.
Spearman’s correlation coefficient analysis between the levels of expression of the peroxiredoxin 6 (PRDX6) gene and 20 major genes in lung adenocarcinoma.
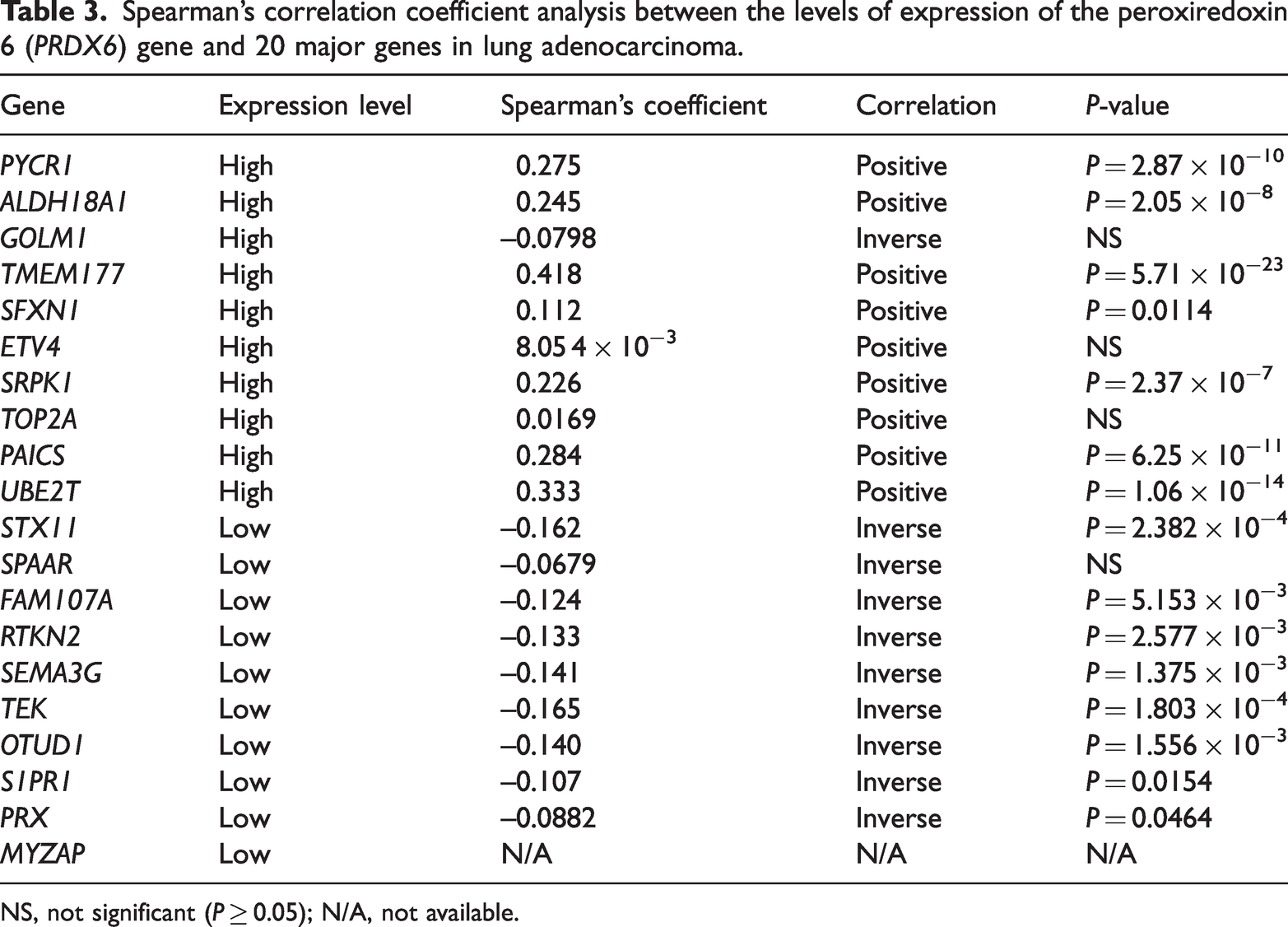
NS, not significant (P ≥ 0.05); N/A, not available.
Analysis of the levels of PRDX6 gene expression in lung adenocarcinoma showed that the levels were significantly higher in lung adenocarcinoma compared with normal tissues (P < 0.05) (Figure 3a). This finding was also confirmed by immunohistochemistry (P < 0.01) (Figures 3b and 3c). Five different mutation sites in different domains of the PRDX6 gene were identified using the cBioPortal database (Figure 3d) (Table 4). Knockdown of the PRDX6 gene using siRNA-PRDX6 resulted in a significant decrease in cell growth as measured using the CCK-8 assay at 24 and 48 h compared with the siRNA-NC group (P < 0.05) (Figure 3e).
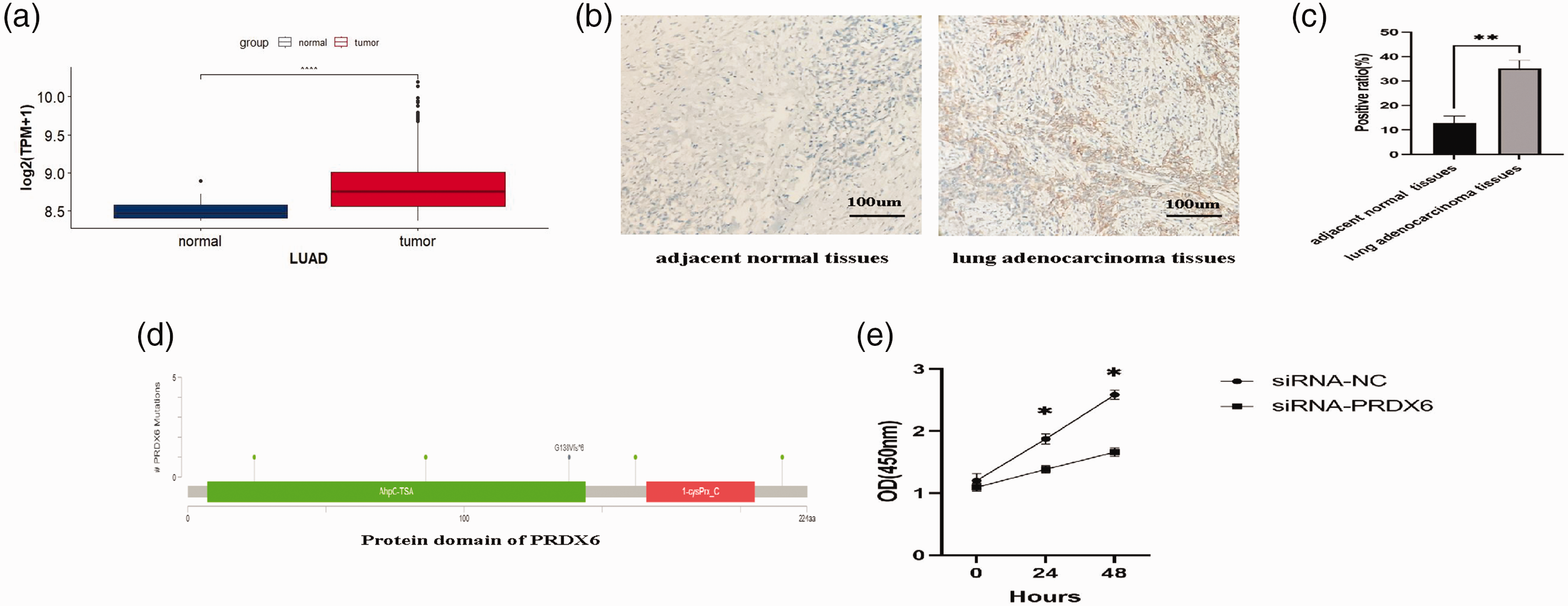
Levels of expression and biological function of the peroxiredoxin 6 (PRDX6) gene in lung adenocarcinoma: (a) box-whisker plots of PRDX6 in lung adenocarcinoma (LUAD) (the central horizontal line is the median; the extremities of the box are the 25th and 75th percentiles; the error bars represent the minimum and maximum outliers; and the circles above the error bar represent extreme outliers); (b) representative photomicrographs of immunohistochemical staining of PRDX6 protein in adjacent normal tissues and lung adenocarcinoma (scale bar, 100 µm); (c) quantitative analysis of the immunohistochemical staining levels of PRDX6 protein in lung adenocarcinoma tissues compared with adjacent normal tissues; (d) mutation sites in the PRDX6 gene in lung adenocarcinoma and (e) a cell proliferation assay was used to measure cell proliferation in the siRNA-NC and siRNA-PRDX6 groups at 0, 24 and 48 h. *P < 0.05, **P < 0.01, ****P < 0.0001; unpaired Student’s t-test. The colour version of this figure is available at: http://imr.sagepub.com.
Gene mutations of the peroxiredoxin 6 (PRDX6) gene in lung adenocarcinoma.

With regard to the diagnostic efficacy of the PRDX6 gene in lung adenocarcinoma, ROC curve analysis showed that the areas under the curve (AUC) for PRDX6 in the overall cohort of patients with lung adenocarcinoma was 0.5082 (Figure 4a). The cut-off value was 8.160, the specificity was 0.932 and the sensitivity was 0.375. Patients were then analysed on the basis of smoking history, sex and race and the AUCs were 0.5788, 0.5648 and 0.5812, respectively (Figures 4b, 4c and 4d). The cut-off values for smoking history, sex and race were 8.610, 8.165 and 8.303, respectively. The specificities for smoking history, sex and race were 0.800, 0.406 and 0.615, while the sensitivities were 0.373, 0.706 and 0.571, respectively.
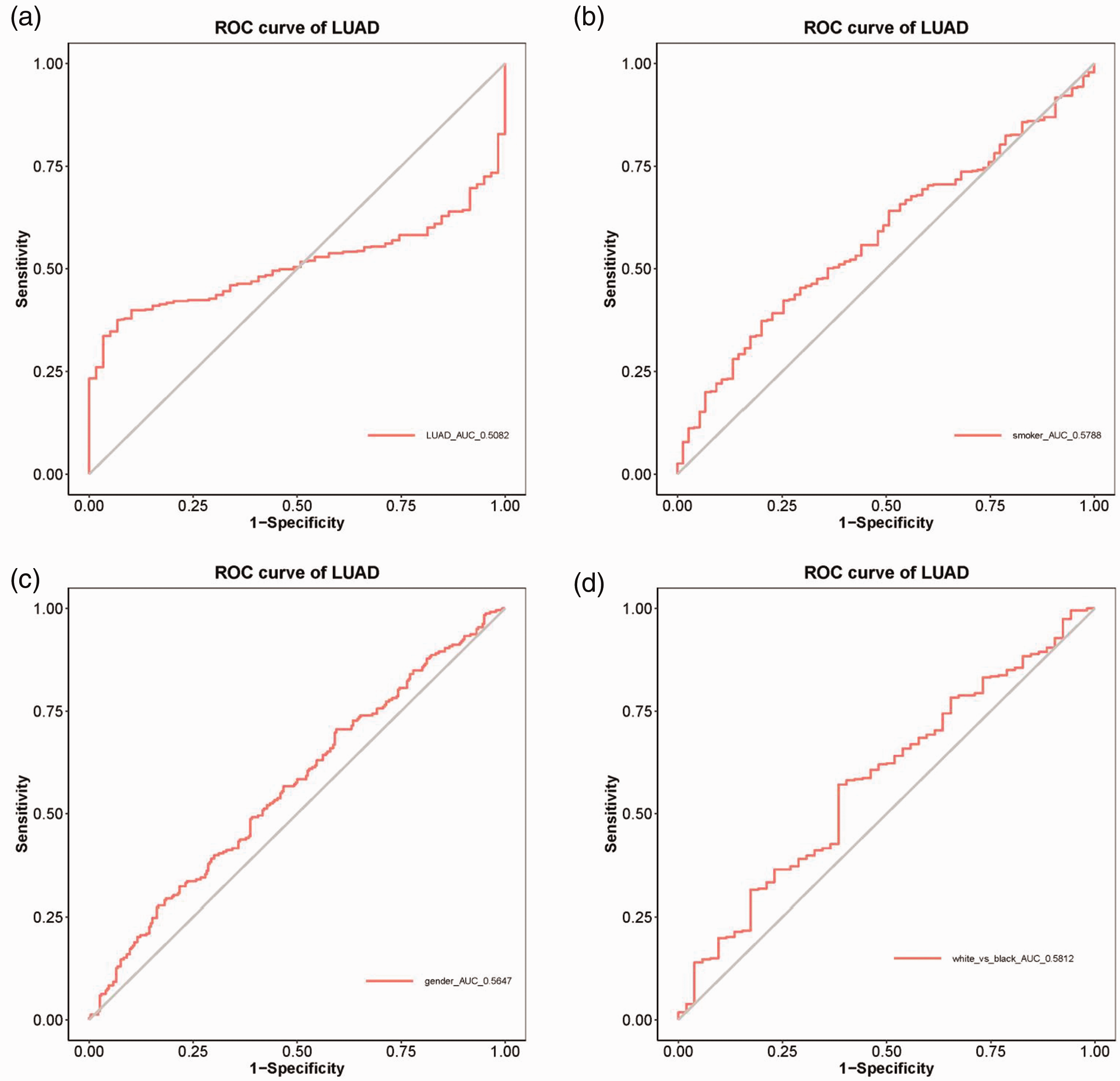
Diagnostic efficacy of the peroxiredoxin 6 (PRDX6) gene in lung adenocarcinoma (LUAD) patients: (a) overall cohort; and in those analysed according to smoking history (b), sex (c) and race (d). ROC, receiver operating characteristic. The colour version of this figure is available at: http://imr.sagepub.com.
With regard to the impact of the PRDX6 gene on survival in lung adenocarcinoma, the survival curve showed that patients with a high level of PRDX6 gene expression had a significantly shorter survival duration compared with those with low PRDX6 expression levels (P < 0.05) (Figure 5a). Among patients with or without a smoking history, the survival time of patients with a high level of PRDX6 gene expression was significantly shorter compared with those with low PRDX6 expression levels (P < 0.05 for both comparisons) (Figure 5b). Among male and female patients, the survival time of patients with a high level of PRDX6 gene expression was significantly shorter compared with those with low PRDX6 expression levels (P < 0.05 for both comparisons) (Figure 5c). Among patients stratified according to race (white or black ethnic background), the survival time of patients with a high level of PRDX6 gene expression was significantly shorter compared with those with low PRDX6 expression levels (P < 0.05 for both comparisons) (Figure 5d).

Kaplan–Meier survival curve analysis of the peroxiredoxin 6 (PRDX6) gene in lung adenocarcinoma (LUAD) patients: (a) overall cohort; and in those analysed according to smoking history (b), sex (c) and race (d). ROC, receiver operating characteristic. The colour version of this figure is available at: http://imr.sagepub.com.
Discussion
At present, despite the achievements in the research of the pathogenesis of lung adenocarcinoma and the development of new treatment methods, lung adenocarcinoma is still one of the most aggressive and deadly cancer types with a total survival period of <5 years.3,18,19 Therefore, it is very important to investigate the potential of molecular targeted therapy for lung adenocarcinoma. This current study aimed to explore the expression levels and clinical role of the PRDX6 gene in lung adenocarcinoma.
This current study demonstrated that the PRDX6 gene expression levels in lung adenocarcinoma was higher than that in half of the range of cancers analysed and the mutation frequency of PRDX6 in lung adenocarcinoma ranked fourth in the same range of cancers. These findings indicated that it was meaningful to further explore the expression levels and clinical role of the PRDX6 gene in lung adenocarcinoma. The results of the heat map and volcano map showed that there were genes with high and low levels of expression in lung adenocarcinoma. The current study then focused on labelling the top 10 genes with high expression and the 10 genes with low expression genes. The PRDX6 gene was positively correlated with nine out of 10 high expression genes, of which seven were statistically significant; and it was inversely correlated with nine out of 10 low expression genes, of which eight were statistically significant. A previous study found that PRDX6 gene expression was markedly higher in non-small cell lung cancer tissues than in adjacent tissues, which seems consistent with the current findings. 20 Therefore, the findings of the current study suggest that the PRDX6 gene might be highly expressed in lung adenocarcinoma.
The current study examined the levels of expression of the PRDX6 gene in lung adenocarcinoma via the TCGA database. The results showed that PRDX6 was highly expressed in lung adenocarcinoma and immunohistochemistry was used to verify this result. Analyses also found five mutations at different sites in the PRDX6 gene in lung adenocarcinoma. A previous study found that when there are mutations in PRDX6 at C47S, D140A and H26A, the three endothelial activities of PRDX6 are affected. 21 The K122/142R mutation is the main Sumo1 binding site in the PRDX6 gene, which can increase the abundance of glutathione peroxidase. 22 These findings suggest that PRDX6 gene mutations in lung adenocarcinoma have considerable research potential.
This current study undertook preliminarily in vitro studies to explore the biological role of PRDX6 in lung adenocarcinoma were undertaken in NCI-H1395 cells and inhibition of PRDX6 gene expression inhibited cell proliferation. Several studies have found similar results in lung cancer.20,23,24 Analysis of the diagnostic value of PRDX6 for lung adenocarcinoma demonstrated that it had poor diagnostic efficacy; and this was also the case for patients with different demographic characteristics. These current findings suggest that PRDX6 could not be used as a clinical diagnostic indicator for lung adenocarcinoma patients. In order to explore the impact of PRDX6 gene expression on survival time of patients with lung adenocarcinoma, Kaplan–Meier survival curve analyses were undertaken. The results showed that patients with a high level of PRDX6 gene expression had a significantly shorter survival duration compared with those with low PRDX6 expression levels. This finding was not affected by different demographic characteristics of lung adenocarcinoma patients (smoking history, sex, race). These current findings suggest that the PRDX6 gene might be related to the poor prognosis of patients with lung adenocarcinoma.
This current study had several limitations. First, the number of patients collected in this study was too small. In the future, the sample size will be expanded to make the conclusions more universal and objective. Secondly, this study did not investigate the mechanism of action of the PRDX6 gene in lung adenocarcinoma, so will require further exploration in the future.
In conclusion, this current study found that the PRDX6 gene is highly expressed in lung adenocarcinoma and that it can promote the proliferation of NCI-H1395 cells. It appears to be associated with a poor prognosis in patients with lung adenocarcinoma, which might provide a new target for clinical treatment.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605241236276 - Supplemental material for Expression and clinical role of PRDX6 in lung adenocarcinoma
Supplemental material, sj-pdf-1-imr-10.1177_03000605241236276 for Expression and clinical role of PRDX6 in lung adenocarcinoma by Zixin Chen, Junjun Lin, Huifang Wang, Jing Wang and Zhou Zhang in Journal of International Medical Research
Footnotes
Author contributions
Z.Z. and Z.C. were responsible for the conception of the study; Z.C. was responsible for the bioinformatics analysis; Z.C., J.L. and J.W. wrote the original draft; Z.Z. and H.W. revised the manuscript; Z.Z., Z.C. and H.W. responded to the reviewers; Z.Z. reviewed and edited the manuscript. All authors have read and approved the manuscript.
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
This research received no specific grant from funding agency in the public, commercial, or not-for-profit sectors.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
