Abstract
Objective
This study aimed to perform an integrated pan-cancer analysis to characterize the expression patterns, prognostic value, genetic alterations, and immunologic roles of transforming growth factor beta 1 (TGFB1) across diverse human cancer types.
Methods
Bioinformatics analyses were conducted using multiple public databases including The Cancer Genome Atlas, Genotype-Tissue Expression, Clinical Proteomic Tumor Analysis Consortium, TIMER2, GEPIA2, cBioPortal, StringDB, and others. Differential expression, survival, immune correlation, and protein interaction network analyses were performed.
Results
TGFB1 was overexpressed in several tumor types compared with that in normal tissues. High TGFB1 expression was associated with an advanced stage and poorer prognosis in certain cancers. TGFB1 mutations occurred in 1.3% of 10,967 cases surveyed. TGFB1 expression correlated with tumor-infiltrating immune cells and immunotherapy-related genes.
Conclusions
This comprehensive multi-omics analysis revealed the complex expression and prognostic landscape of TGFB1 across cancers. TGFB1 is emerging as a potential immunotherapeutic target in certain contexts. Further research should elucidate its multifaceted tumor-promoting and tumor-suppressive mechanisms. Our pan-cancer analysis provides new insights into TGFB1 as a prognostic biomarker and immunotherapeutic target in human cancers, and our findings may guide future preclinical and clinical investigations of TGFB1-directed therapies.
Keywords
Introduction
Both the incidence of cancer and cancer mortality are growing rapidly worldwide, and cancer is a leading cause of death in the 21st century and a major obstacle in improving life expectancy worldwide.1–3 In 2018, there were 18.1 million new cases and 9.6 million cancer deaths worldwide. By 2040, the global burden of cancer is expected to increase to 27.5 million new cases and 16.3 million cancer deaths, solely because of the growth of the aging population. 1 Given the aging population, it is anticipated that the number of cancer-related deaths will continue to rise both globally and in the United States, posing a significant public health burden. 4 Given the substantial health burden posed by cancer globally, an urgent need exists to advance the understanding of molecular drivers of tumor progression and identify new therapeutic targets. Traditional cancer treatments, including surgery, chemotherapy, and radiotherapy, continue to serve as the cornerstone of current therapeutic strategies for cancer. Immunotherapy, however, has emerged as a promising approach for treating patients with a wide spectrum of malignancies.5,6 Notably, immunotherapy based on immune checkpoint inhibitors (ICIs) has demonstrated remarkable therapeutic efficacy across various malignancies.7,8 It is important to acknowledge that the effectiveness of immunotherapy can vary significantly among different types of cancer and even within individual tumors of the same cancer type owing to the inherent heterogeneity of tumors. 9 Transforming growth factor beta 1 (TGFB1) has become a prominent target in the realm of antitumor immunotherapy. 10 An increasing body of research has unveiled the pivotal role of TGFB1 in cancer pathogenesis, rendering it a promising candidate for the development of novel drugs. 11
TGFB1, also called CED, LAP, DPD1, TGFB, IBDIMDE, TGF-beta, and TGF-beta1, is a member of the TGF-beta superfamily. This multifaceted protein is intricately involved in processes such as cell proliferation, differentiation, and growth. Moreover, TGFB1 exerts regulatory control over the expression and activation of other growth factors, including interferon-gamma and tumor necrosis factor-alpha.12–14 When activated, TGFB1 binds to ubiquitously expressed cell-surface TGFB1 type I receptors (TGFBR1) and type II receptors (TGFBR2). These receptors are transmembrane serine/threonine kinases that have been implicated in tumorigenesis. 15 TGFB1 is synthesized by both healthy epithelial cells and tumor cells, and it plays a dual role in the context of cancer. It can either promote cancer development by inhibiting immunosurveillance, encouraging epithelial-mesenchymal transition, and facilitating metastases, or alternatively, it can suppress tumor growth by directly hindering cell cycle progression, leading to growth arrest and activating apoptotic pathways.13,16,17 Although TGFB1 has emerged as a key molecule involved in cancer, a comprehensive pan-cancer analysis regarding the oncogenic role of TGFB1 in relation to cancer is currently lacking. Therefore, we conducted a pan-cancer analysis using public databases to investigate the association of TGFB1 with tumorigenesis and immunotherapy.
Our study aimed to conduct an integrated multi-omics analysis of the oncogenic properties of TGFB1 across 33 different tumor types to provide insights into its potential as a prognostic biomarker and immunotherapeutic target. Findings from this pan-cancer characterization of TGFB1 may help guide future preclinical and clinical investigations of TGFB1-directed therapies.
Materials and methods
Ethics statement
This research utilized publicly available databases and did not involve any human or animal studies or experiments. Consequently, ethical approval was not required for this study.
Multi-omics analysis
To obtain comprehensive information regarding the gene function of TGFB1, we performed multi-omics analysis using several databases. First, the chromosome localization of TGFB1 was obtained from the GeneCards database (https://www.genecards.org/), which integrates gene-centric data from 150 web sources. Furthermore, oncogene information and protein sequence data for TGFB1 were gathered from the National Center for Biotechnology Information (NCBI) Gene and Protein databases (https://www.ncbi.nlm.nih.gov/). Conserved protein domains and the three-dimensional (3D) structure of TGFB1 were acquired from the UniProt database (https://www.uniprot.org/). The Illustrator for Biological Sequences tool 18 (version 1.0, http://ibs.biocuckoo.org/) was then used to visualize the conserved domains in the TGFB1 amino acid sequence.
To identify cross-species conservation of the TGFB1 protein, a HomoloGene search was performed in NCBI, and a phylogenetic tree was constructed using the Constraint-based Multiple Alignment Tool (COBALT, https://www.ncbi.nlm.nih.gov/tools/cobalt/) from NCBI. Finally, to determine the subcellular localization of TGFB1, the Human Protein Atlas (HPA) (https://www.proteinatlas.org/) database was used.
Gene expression analysis of TGFB1
Baseline TGFB1 mRNA expression in normal tissues was retrieved from the HPA. Gene expression values were normalized using the transcripts per million. Subsequently, the “Gene_DE of Exploration” module of the TIMER2 (http://timer.comp-genomics.org/) database was explored using “TGFB1” as input. The differences in TGFB1 mRNA expression between tumor and paracancerous tissues were explored. To examine the expression level of the TGFB1 gene among different tumor and normal tissues, the “Expression Analysis-Expression DIY-Box Plots” module of the GEPIA2 (http://gepia2.cancer-pku.cn/) database and the “Differential analysis of gene expression” module of Sangerbox 3.0 (http://vip.sangerbox.com/) were used. The UALCAN (http://ualcan.path.uab.edu/) database was used to perform protein expression analysis of the Clinical Proteomic Tumor Analysis Consortium dataset. The expression levels of total TGFB1 protein were compared between primary tumor and normal tissues.
Clinical pathological analysis of TGFB1
The “Pathological Stage Plot” module of GEPIA2 was applied to assess the relationship between TGFB1 gene expression and cancer stages of patients in The Cancer Genome Atlas (TCGA). We also used the UALCAN database to explore the relationship between the TGFB1 mRNA expression level and clinicopathological cancer stage, including a within-stage correlation exploration.
Prognostic analysis of TGFB1
To further analyze the potential relationship between TGFB1 expression and clinical prognosis, we used the “Survival Map” and “Survival Analysis” modules of GEPIA2 to construct a survival map and overall survival (OS) and disease-free survival (DFS) Kaplan–Meier curves. The patients were divided into high- and low-expression groups based on the median expression of TGFB1. Furthermore, the Sangerbox 3.0 web tool was also used to explore the association between TGFB1 expression and progression-free interval (PFI) as well as disease-specific survival (DSS).
Genomic alterations of TGFB1
The cBioPortal (https://www.cbioportal.org/) database is an online resource used to explore multidimensional cancer genomics information. TCGA pan-cancer atlas studies, including 32 tumor studies and 10,967 samples, were selected for further analysis. cBioPortal was used to explore TGFB1 alterations across all samples from TCGA studies. Total genomic alteration information for various tumors was summarized using the “Cancer Types Summary” module. The “Comparison/Survival” module was used to obtain the survival differences, including OS, DFS, progression-free survival, and DSS for tumor samples with or without genetic alterations of TGFB1. In addition, the specific mutation locations and type and frequency information of TGFB1 were displayed in the schematic diagram of the protein structure or the 3D structure using the “Mutations” module of the cBioPortal database.
Immune correlation analysis
The TIMER2 database was used to quantify the associations between TGFB1 expression in diverse cancer types and the abundance of immune infiltrates, including cancer-associated fibroblasts, CD8+ T cells, CD4+ T cells, T regulatory cells, macrophages, and B cells. Scatterplots were used to visualize the association between TGFB1 expression and cancer-associated fibroblast infiltration, which was adjusted by purity. The Tumor Immune System Interaction Database (http://cis.hku.hk/TISIDB/index.php) was used to evaluate the correlation of TGFB1 expression with ICI-related genes. The tumor mutational burden (TMB), and microsatellite instability (MSI) are important biomarkers of the tumor microenvironment (TME). Sangerbox was used to analyze the associations of TGFB1 expression with the TMB and MSI.
Methylation analysis of TGFB1
The promoter methylation expression level of TGFB1 in various human cancers was assessed using ULCAN. DNA sights with the highest methylation rate were predicted using MethSurv (https://biit.cs.ut.ee/methsurv). Meanwhile, we also characterized the effect of methylation on OS. The prognostic predictive value of the CpG sites with the highest DNA methylation rate in various human cancers was analyzed using the MethSurv database.
Protein–protein interaction (PPI) network and functional and pathway enrichment analysis
The TGFB1-binding proteins were obtained using the STRING database (https://string-db.org/) with a query consisting of a single protein name (“TGFB1”) and organism (“
Statistical analysis
Differential expression was determined by a log fold change >1 and p < 0.05. Correlations were assessed using the Pearson coefficient. Survival differences were calculated using the log-rank test, with p < 0.05 considered statistically significant.
Results
Multi-omics analysis of TGFB1
This study aimed to investigate the oncogenic role of TGFB1 in human cancers. TGFB1 (Gene ID: 7040, updated on 6 February 2022) is located at chromosome 19q13.2 and contains seven exons (Figure 1a). The secondary structure of the TGFB1 protein is displayed in Figure 1a. TGFB1 has two conserved domains including the “straightjacket” domain and the “arm” domain, as predicted by the UniProt database (Figure 1b). The straitjacket and arm domains encircle the TGFB1 monomers and are linked by strong bonding between Lys-56 and Tyr-103/Tyr-104. The predicted 3D structures of TGFB1-encoded proteins from the UniProt database are displayed in Figure 1c. We also found that the functional domains of TGFB1 were highly conserved across different species (Figure 1d). The multispecies phylogenetic tree also revealed the conserved nature of TGFB1 among

Gene structure, protein structure, and conserved domain of transforming growth factor beta 1 (TGFB1). (a) Chromosome localization and secondary structure of the human TGFB1 protein. (b) The conserved domains of the TGFB1 protein structure. (c) The three-dimensional structure of the TGFB1 protein. (d) Conservation of the TGFB1 protein among different species. (e) The phylogenetic tree of TGFB1 in different species. (f) Location of the TGFB1 protein in cells and (g) TGFB1 protein sites in the Golgi apparatus of HEL (human erythroleukemic cell line) cells and extracellular secretion.
TGFB1 expression
To explore the mechanism underlying the oncogenic role of TGFB1, we determined whether TGFB1 expression is related to tumorigenesis. First, the TGFB1 mRNA expression level was compared in normal tissues based on the HPA database (Supplementary Figure 1a). The bar charts showed that TGFB1 had the highest expression in the spleen, followed by the bone marrow, lungs, and esophagus, indicating that TGFB1 was mainly expressed in the bone marrow and lymphoid tissues. Additionally, TGFB1 expression was high in most normal tissues, indicating the low tissue specificity of TGFB1 mRNA expression. Second, the expression levels of TGFB1 in normal tissues and tumor tissues were evaluated using the TIMER database. As shown in Figure 2a, a comparison between paracancerous tissues and tumor tissues revealed that TGFB1 mRNA was significantly differentially expressed in breast invasive carcinoma (BRCA), cholangiocarcinoma (CHOL), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), kidney renal clear cell carcinoma (KIRC), lung adenocarcinoma (LUAD), thyroid carcinoma (THCA), uterine corpus endometrial carcinoma (UCEC) (p < 0.001); esophageal carcinoma (ESCA), liver hepatocellular carcinoma (LIHC), lung squamous cell carcinoma (LUSC) (p < 0.01); and prostate adenocarcinoma (PRAD) (p < 0.05). Similar results were also observed using the Sangerbox 3.0 database (Supplementary Figure 1b). We used GEPIA2 to combine data from TCGA and the Genotype-Tissue Expression database to verify the expression levels of TGFB1. The results were similar to those obtained using the TIMER database in that TGFB1 was differentially expressed in adrenocortical carcinoma (ACC), CHOL, GBM, HNSC, KICH, KIRC, brain lower grade glioma (LGG), pancreatic adenocarcinoma (PAAD), stomach adenocarcinoma (STAD), testicular germ cell tumors (TGCT), thymoma (THYM), UCEC, and uterine carcinosarcoma (UCS) (Figure 2b), but there was no significant difference for other tumors, including bladder urothelial carcinoma (BLCA), BRCA, cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), colon adenocarcinoma (COAD), lymphoid neoplasm diffuse large B-cell lymphoma, ESCA, kidney renal papillary cell carcinoma (KIRP), acute myeloid leukemia (LAML), LIHC, LUAD, LUSC, ovarian cancer, pheochromocytoma and paraganglioma (PCPG), PRAD, rectum adenocarcinoma (READ), sarcoma (SARC), skin cutaneous melanoma (SKCM), and THCA (Supplementary Figure 1c). Further evaluation of TGFB1 protein expression levels across different tumors was performed using the UALCAN database, and the results revealed that the TGFB1 protein was distinctly expressed in PAAD, HNSC, LUAD, UCEC, clear cell renal cell carcinoma, BRCA, and GBM (Figure 2c). These results indicated that TGFB1 was differentially expressed in most tumors, which implied that TGFB1 expression may be involved in the occurrence of these tumors.
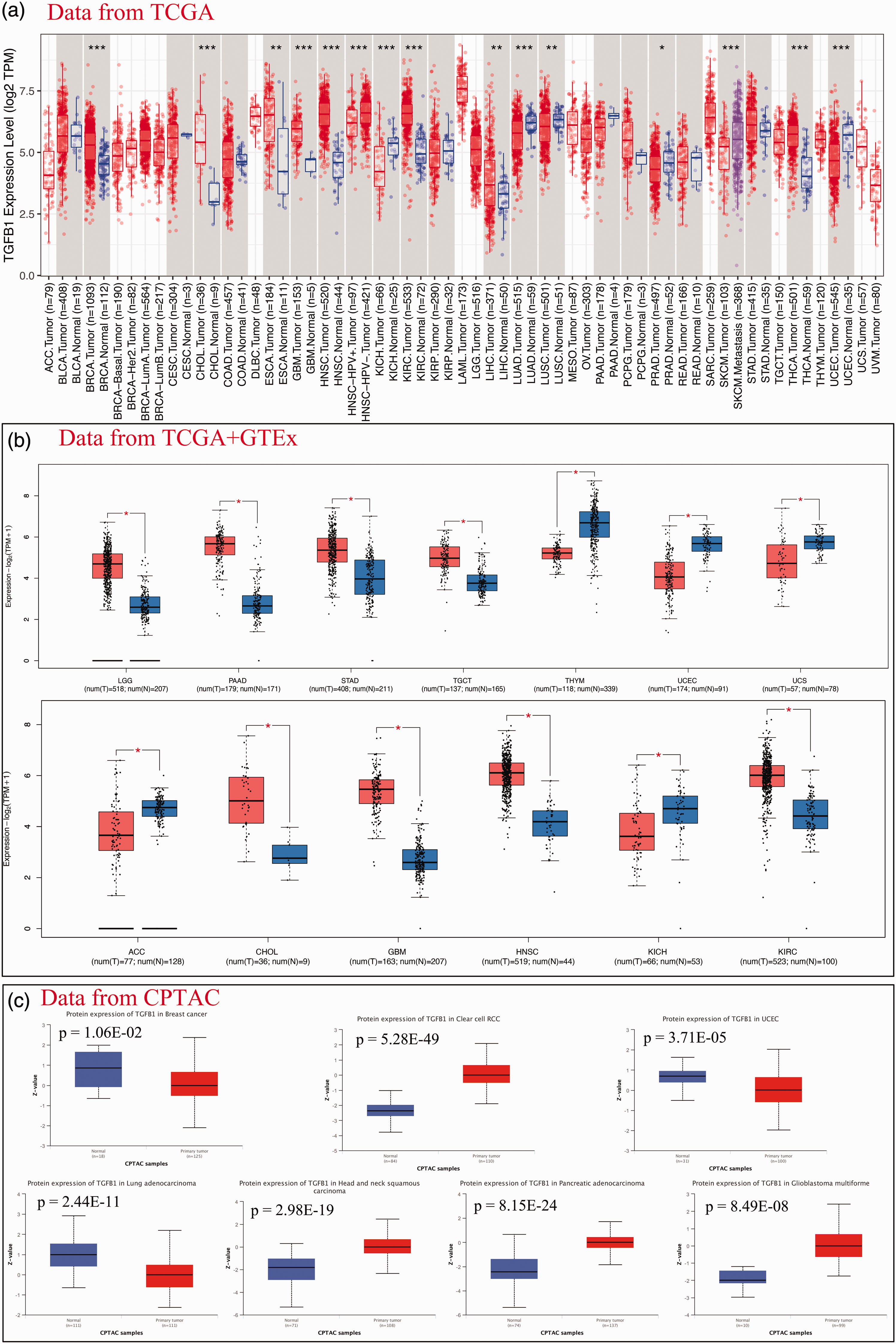
The expression levels of transforming growth factor beta 1 (TGFB1) in tumor tissues and normal tissues. (a) The mRNA expression levels of TGFB1 in a pan-cancer analysis using the TIMER database.Continued.*p < 0.05, **p < 0.01, ***p < 0.001. (b) Significantly different expression levels of TGFB1 between normal and tumor tissues and (c) Protein expression levels of TGFB1 in different tumor and normal tissues.
TGFB1 and tumor clinicopathological staging
The expression levels of TGFB1 were correlated with tumor stage, among which, the TGFB1 expression level was significantly (p < 0.05) correlated with tumor clinicopathological staging in BLCA, KIRC, SKCM, STAD, and THCA (Figure 3a–e). However, there was no significant correlation for other tumors. Further correlation analyses of TGFB1 expression with different clinicopathological stages was conducted for each tumor using the UALCAN database. The results also demonstrated that TGFB1 expression may be a critical factor for tumor progression. Additionally, the different clinicopathological stages of seven tumors revealed high TGFB1 expression compared with that in paracancerous tissues, including BRCA, CHOL, ESCA, HNSC, KIRC, THCA, and LIHC (Figure 3f–l). In contrast TGFB1 showed lower expression levels than those in normal tissues in UCEC, SKCM, PAAD, LUSC, LUAD, KICH, and COAD (Figure 3m–s). Furthermore, TGFB1 expression was associated with the clinicopathological staging of STAD, uveal melanoma (UVM), and BLCA (Figure 3t–v). These findings suggest that TGFB1 may serve as a valid and reliable biomarker for the clinical diagnosis of these tumors.

Association analysis of transforming growth factor beta 1 (TGFB1) expression with clinicopathological stages. (a–e) The association between TGFB1 expression and pathological clinical stage. (a) thyroid carcinoma (THCA), (b) kidney renal clear cell carcinoma (KIRC), (c) stomach adenocarcinoma (STAD), (d) bladder urothelial carcinoma (BLCA), (e) skin cutaneous melanoma (SKCM). (f–v) Correlations between TGFB1 expression and normal as well as different pathological clinical stages. (f) breast invasive carcinoma (BRCA), (g) esophageal carcinoma (ESCA), (h) THCA, (i) cholangiocarcinoma (CHOL), (j) head and neck squamous cell carcinoma (HNSC), (k) KIRC, (l) liver hepatocellular carcinoma (LIHC), (m) lung adenocarcinoma (LUAD), (n) lung squamous cell carcinoma (LUSC), (o) pancreatic adenocarcinoma (PAAD), (p) kidney chromophobe (KICH), (q) SKCM, (r) uterine corpus endometrial carcinoma (UCEC) and (s) colon adenocarcinoma (COAD), (t) STAD, (u) uveal melanoma (UVM) and (v) BLCA.
TGFB1 and prognosis
The prognostic value of TGFB1 was explored in the GEPIA2 database. As shown in Figure 4a, stronger expression of TGFB1 was related to poor prognosis for OS in COAD (p = 0.04, hazard ratio (HR) = 1.7), LAML (p = 2.20E−02, HR = 1.9), LGG (p = 5.1E−03, HR = 1.7), mesothelioma (MESO) (p = 9.20E−04, HR = 2.3), and STAD (p = 2.20E−02, HR = 1.4). However, high expression of TGFB1 was associated with greater OS in SKCM (p = 6.70E−03, HR = 0.69). In terms of DFS, the results demonstrated a significant correlation between high TGFB1 expression and poor prognosis in KIRC (p = 4.10E−03, HR = 1.7), LGG (p = 8.10E−03, HR = 1.5), and PRAD (p = 1.90E−04, HR = 2.3) (Figure 4b). Cox proportional hazard regression analysis was also used to determine the correlation of TGFB1 expression and PFI and DSS. The forest plots indicated that higher TGFB1 expression is linked to worse PFI in COAD, KIRC, KIRP, LGG, LUSC, MESO, PRAD, STAD, and UVM (Figure 4c). Additionally, the forest plots indicated that higher TGFB1 expression is linked to worse DSS in BLCA, CESC, COAD, HNSC, KIRC, KIRP, LGG, LUSC, MESO, STAD, TGCT, and UVM (Figure 4d). Meanwhile, we presented the significant results for each tumor in Kaplan–Meier curves (Supplementary Figure 2), which indicate that TGFB1 might serve as a potential prognostic biomarker for several cancer types.

Correlation between transforming growth factor beta 1 (TGFB1) expression and survival prognosis of cancers. (a) Overall survival. (b) Disease-free survival. (c) Progression-free survival and (d) Disease-specific survival. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute myeloid leukemia; LGG, lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, ovarian cancer; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
TGFB1 and genetic alteration
Malignant tumor is a disease of genetic alterations, and these mutated genes offer potential molecular therapeutic targets.19,20 We analyzed the TGFB1 mutation status in all tumor samples from TCGA datasets using the cBioPortal database. The results revealed that TGFB1 was mutated in 141 (1.3%) of 10,967 tumor cases. Amplification was the predominant mutation type of TGFB1, followed by missense mutations and deep deletion (Figure 5a). As shown in Figure 5a, TGFB1 mutations were detected in 22 tumor types, and no mutations were detected in 10 tumor types. Furthermore, we found that TGFB1 had a high alteration frequency (gene altered in 7.02% of 57 cases) in patients with UCS (Figure 5a). Amplification as the only genetic alteration type occurred in CESC (1.68% of 297 cases) and MESO (1.15% of 87 cases). Similarly, mutation as the only genetic alteration type occurred in SKCM (1.35% of 444 cases), GBM (0.34% of 592 cases), and KIRC (0.35% of 283 cases) (Figure 5a). The sites, types, and case number of the mutated TGFB1 (NM_000660 | ENST00000221930) were analyzed using the “mutation” module of the cBioPortal database. Missense mutation of TGEB1 was the predominant genetic mutation type, and the R180* mutation type in the TGFb-propeptide domain was detected in one case of GBM and two cases of UCEC (Figure 5b). The R180* site is displayed in the 3D structure of the TGFB1 protein (Figure 5c). In addition, an evaluation of TGFB1 mutations and their association with disease prognosis in pan-cancer samples was assessed. A worse prognosis for DFS (p = 4.40E−3), but not for OS (p = 0.766), progression-free survival (p = 0.174), and DSS (p = 0.597), was observed for patients with tumors lacking TGFB1 alterations (Figure 5d) compared with those for patients with TGFB1 alterations (Figure 5e–g). These results suggest that mutated TGFB1 may be involved in the pathogenesis and progression of cancer.

Transforming growth factor beta 1 (TGFB1) mutation landscape in various cancer types. (a) Summary of alteration frequency according to mutation type of TGFB1 in different tumors. (b) Mutation diagram of TGFB1 in pan-cancer analysis. (c) Some TGFB1 mutations are shown on the three-dimensional structure of the protein. (d–g) Correlation between TGFB1 mutation and survival prognosis in pan-cancer analysis. (d) Disease-free survival, (e) Disease-specific survival, (f) Overall survival and (g) Progression-free survival. BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; COAD, colon adenocarcinoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LGG, lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; PAAD, pancreatic adenocarcinoma; PRAD, prostate adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; OV, ovarian cancer.
TGFB1 and immune cell infiltration
The TME, which is a complex immune network consisting of tumor-infiltrating immune cells, tumor cells, and stroma cells, contributes to tumor biology and the therapeutic response to ICIs. 21 Thus, successful immunotherapeutic strategies in cancer treatment require effective infiltration of the tumor by tumor-suppressive immune cells. Using the TIMER2 tools, we evaluated the association between TGFB1 expression and tumor-infiltrating immune cells. We observed a significantly positive correlation of TGFB1 expression and the tumor-associated immune cell infiltration of cancer-associated fibroblasts for TCGA tumors including BRCA, COAD, ESCA, KIRP, LGG, LIHC, LUAD, PAAD, PRAD, READ, SKCM, STAD, TGCT, THYM, UCEC, and UCS (p < 0.05) (Figure 6). Furthermore, partial correlation analysis between TGFB1 expression and infiltration of other immune cells was conducted using the TIMER2.0 database. The results demonstrated a remarkable correlation between TGFB1 expression and the infiltration of CD8+ T cells, CD4+ T cells, T regulatory cells, and B cells (Supplementary Figure 3). The results revealed that the level of TGFB1 expression was significantly (p < 0.05) correlated with immune markers of various immune cells.
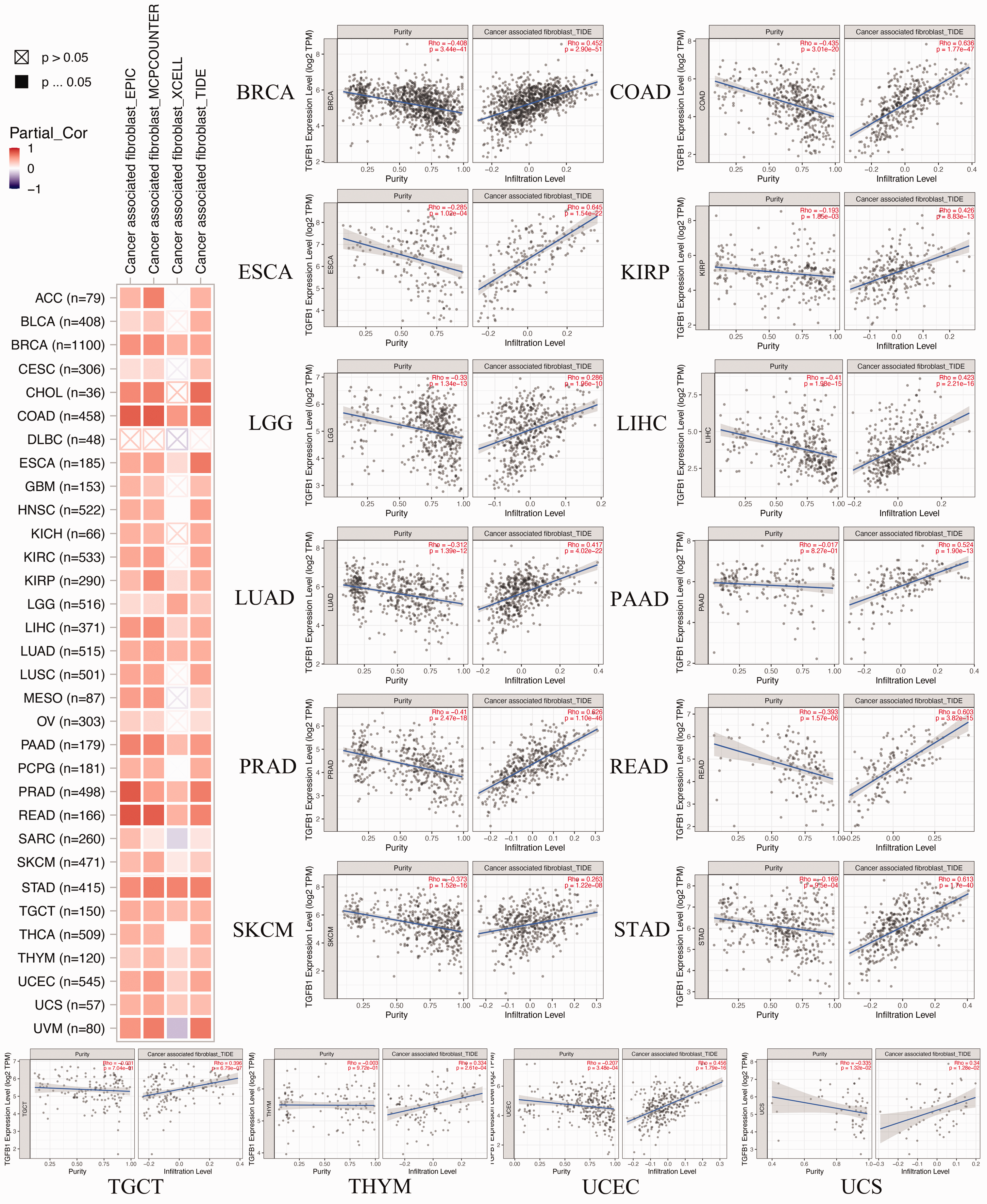
Correlation analysis between transforming growth factor beta 1 (TGFB1) expression and immune infiltration of cancer-associated fibroblasts. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute myeloid leukemia; LGG, lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, ovarian cancer; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
TGFB1 and immunotherapy
ICIs represent a significant breakthrough that has positively impacted the treatment outcomes for cancer patients. 22 Our findings demonstrate a substantial positive correlation between TGFB1 expression and ICI-associated genes, particularly immunoinhibitory genes, across various cancer types in TCGA (Figure 7a). PD-1 (PDCD1), PD-L1 (CD274), and CTLA-4 antibodies are considered the cornerstones of cancer immunotherapy. Notably, in LIHC, TGFB1 expression exhibited a significant association (p < 0.05) with PD-1 and CTLA-4, independent of PD-L1 (Figure 7b). The relationship between TGFB1 expression and two important markers, the TMB and MSI, varied considerably among different cancer types within TCGA dataset. TGFB1 expression was negatively correlated with the TMB in CHOL, ESCA, KIRP, BLCA, and stomach and esophageal carcinoma. Conversely, a positive correlation was found with the TMB in COAD, SARC, colorectal adenocarcinoma, KIRC, and a combined category of kidney cancers (Figure 7c). Furthermore, the expression of TGFB1 was positively correlated with MSI in COAD, SARC, colorectal adenocarcinoma, and CESC, while it exhibited a negative correlation with MSI in glioma, combined kidney cancers, stomach and esophageal carcinoma, and STAD (Figure 7d). These results collectively suggest that TGFB1 expression may represent a promising new target for future immunotherapeutic interventions.

Correlation analysis between transforming growth factor beta 1 (TGFB1) expression and immune checkpoint inhibitor (ICI)-related genes, microsatellite instability (MSI), and tumor mutational burden (TMB). (a) The relationship of TGFB1 expression and ICI-related genes. Red indicates positive correlations; Blue indicates negative correlations. (b) The relationships of TGFB1 expression with CD274, PDCD1, CTLA4, LAG3 in liver hepatocellular carcinoma (LIHC). (c) Analysis of the correlation between TGFB1 expression and TMB (c) and MSI (d). ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute myeloid leukemia; LGG, lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
Methylation analysis of TGFB1
Abnormal methylation plays an important role in the regulation of expression levels; therefore, we further analyzed the correlation between the methylation level and TGFB1 expression level. We first compared the different levels of TGFB1 promoter methylation between tumors and normal tissues based on the UALCAN database. The results showed that the promoter methylation level of TGFB1 was significantly down-regulated in BLCA, CHOL, KIRC, LUSC, PCPG, STAD, THCA, and TGCT samples compared with that in normal tissues (p < 0.05) (Figure 8a–h). In contrast, the promoter methylation level of TGFB1 was significantly up-regulated in UCEC, PRAD, BRCA, and COAD (p < 0.05) (Figure 8i–l). Furthermore, the DNA methylation levels of TGFB1 compared with the prognostic power of each single highest methylated CpG site was explored using the MethSurv database. The results confirmed that cg09926389 of TGFB1 was the highest methylated CpG site in most tumor tissues, including LAML, ACC, BLCA, LGG, BRCA, ESCA, GBM, HNSC, KICH, KIRC, LUAD, LUSC MESO, PAAD, SARC, SKCM, STAD, and UCS (Supplementary Figure 4). cg11037750 of TGFB1 was the highest methylated CpG site in CESC and KIRP tissues, and cg27540367 was the highest methylated CpG site in COAD, LIHC, READ, UCEC, and UVM tissues (Supplementary Figure 4). Meanwhile, cg09926389 of TGFB1 was associated with survival in MESO, PAAD, ACC, BRCA, LAML, LUSC, HNSC, and KIRC (Figure 8m–t). cg11037750 of TGFB1 was associated with a significant prognosis in KIRP (Figure 8u). cg27540367 was associated with a significant prognosis in UVM (Figure 8v). These results indicated that methylation of TGFB1 may be a key biological process that is involved in the development of cancers.
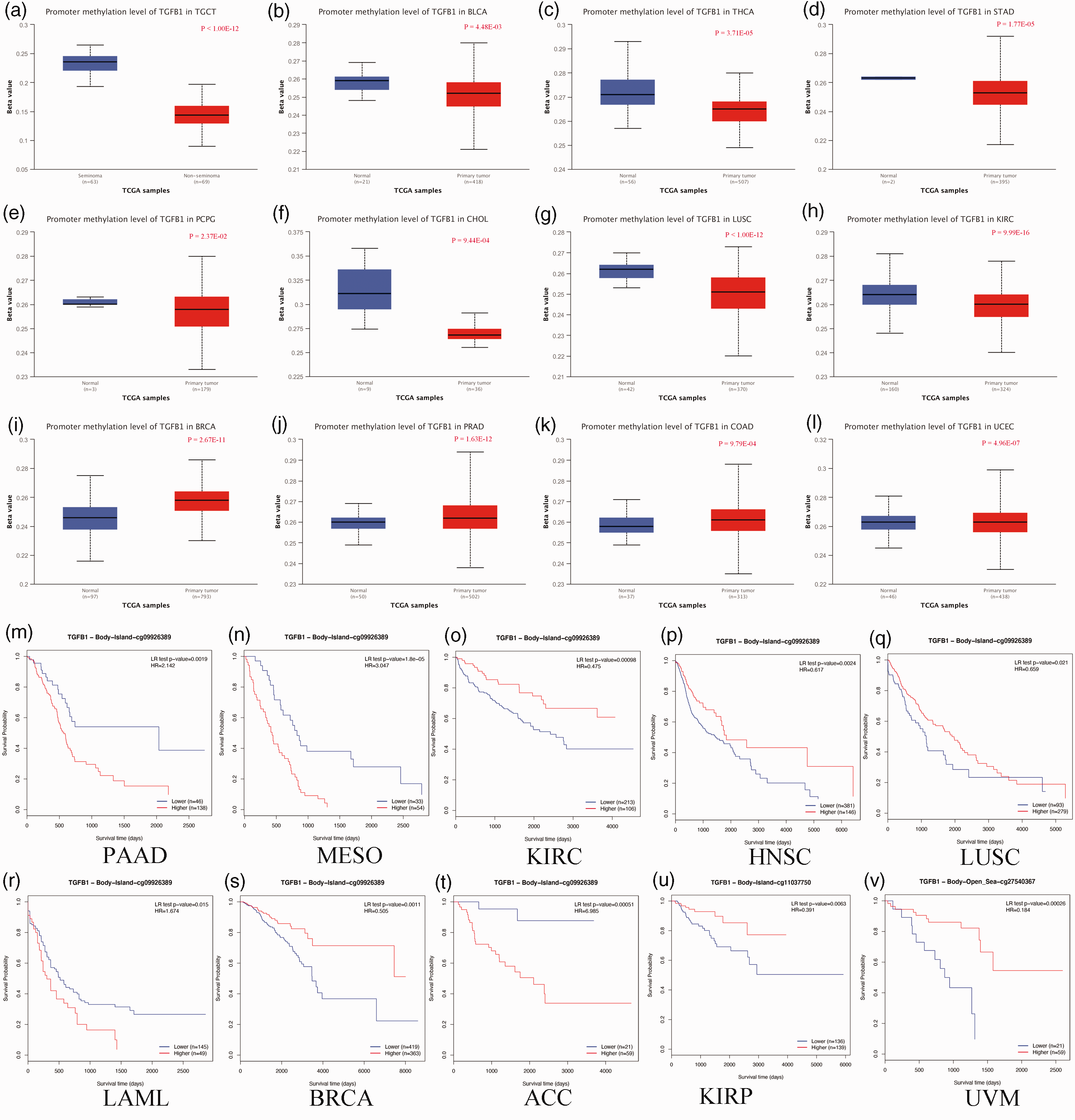
DNA methylation analysis of transforming growth factor beta 1 (TGFB1). (a–l) Differential DNA methylation levels of the TGFB1 promoter in various tumor types. (a) Testicular germ cell tumors (TGCT), (b) bladder urothelial carcinoma (BLCA), (c) thyroid carcinoma (THCA), (d) stomach adenocarcinoma (STAD), (e) pheochromocytoma and paraganglioma (PCPG), (f) cholangiocarcinoma (CHOL), (g) lung squamous cell carcinoma (LUSC), (h) kidney renal clear cell carcinoma (KIRC), (i) breast invasive carcinoma (BRCA), (j) prostate adenocarcinoma (PRAD), (k) colon adenocarcinoma (COAD), (l) uterine corpus endometrial carcinoma (UCEC). (m–v) The prognosis value of single CpG sites in TGFB1 in various tumor types. (m) Pancreatic adenocarcinoma (PAAD), (n) mesothelioma (MESO), (o) KIRC, (p) head and neck squamous cell carcinoma (HNSC), (q) LUSC, (r) acute myeloid leukemia (LAML), (s) BRCA, (t) adrenocortical carcinoma (ACC), (u) kidney renal papillary cell carcinoma (KIRP) and (v) uveal melanoma (UVM).
PPI network
Next, we constructed a PPI network using the STRING database. As shown in Figure 9a, the PPI network, including 50 TGFB1-bound proteins verified experimentally, had 51 nodes and 142 edges. Additionally, the GeneMANIA results revealed that TGFB1-bound proteins were associated with the transmembrane receptor protein serine/threonine kinase signaling pathway, growth factor binding, regulation of the transmembrane receptor protein serine/threonine, the kinase signaling pathway, the response to transforming growth factor beta, the cellular response to transforming growth factor beta stimulus, cytokine binding, and regulation of the cellular response to growth factor stimulus (Figure 9b). The results were consistent with the Metascape gene analysis data (Figure 9c–d). Subsequently, the top six core genes, namely, amyloid beta precursor protein (APP), matrix metallopeptidase 2 (MMP2), TGFB2, TGFBR1, TGFBR2, and thrombospondin 1 (THBS1), were screened based on the calculation of the degree of connectivity in the PPI information network. According to the heatmap results, we concluded that TGFB1 was positively correlated with these genes in the majority of detailed cancer types (Figure 10a). As shown in Figure 10b–g, the TGFB1 expression level was positively correlated with those of the APP (R = 0.16), MMP2 (R = 0.38), TGFB2 (R = 0.19), TGFBR1 (R = 0.38), TGFBR2 (R = 0.37), and THBS1 (R = 0.35) genes (all p < 0.001).

Protein–protein interaction (PPI) network analysis. (a) PPI network of transforming growth factor beta 1 (TGFB1) protein and TGFB1-related proteins based on the STRING database. (b) The gene–gene network analysis of TGFB1 and its related genes. (c) Network of enriched terms, colored according to p-value and (d) Network of enriched terms, colored according to cluster ID.

Correlation analysis of transforming growth factor beta 1 (TGFB1) and six main TGFB1-related genes. (a) Heatmap of selected TGFB1-related genes and cancer types. (b–g) Correlation between TGFB1 expression and selected target genes, including amyloid beta precursor protein (APP) (b), matrix metallopeptidase 2 (MMP2) (c), TGFB2 (d), TGFBR1 (e), TGFBR2 (f), thrombospondin 1 (THBS1) (g).
GO and KEGG analysis
We performed functional and pathway enrichment analysis to explore the potential biological functions of TGFB1-bound genes. According to Figure 11a, among biological processes, these genes were mainly enriched in the transmembrane receptor protein serine/threonine kinase signaling pathway, extracellular matrix organization, and extracellular structure organization. Among cellular components, these genes were mainly enriched in the collagen-containing extracellular matrix, endoplasmic reticulum lumen, and external side of the plasma membrane. Among molecular functions, these genes were mainly enriched in growth factor binding, TGF-beta binding, and cytokine binding. According to the KEGG enrichment results, these genes were mainly enriched in proteoglycans in cancer, the PI3K-Akt signaling pathway, and the TGF-beta signaling pathway (Figure 11b). These results indicated that TGFB1 plays crucial roles in various cellular and physiological processes and exerts functional effects on tumor progression.

Gene Ontology (GO) functional enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway for transforming growth factor beta 1 (TGFB1) and its related genes. (a) GO analysis of TGFB1-related genes and (b) KEGG analysis of TGFB1-related genes.
Discussion
Studies have increasingly indicated an association of TGFB1 with the pathogenesis and progression of a wide range of complex human diseases, with a particular emphasis on various types of tumors.12,23,24 Aberrant expression or mutations in TGFB1 may play a role in the initiation and development of many tumors through shared pathways, although this aspect requires further investigation. 25 To the best of our knowledge, there have been no previous reports of a pan-cancer analysis of TGFB1 oncogenes. In this study, we employed bioinformatic methods in conjunction with publicly available databases, leveraging omics data encompassing expression patterns, gene mutations, disease prognosis, and associations with immune cell infiltration. Our analysis encompassed 33 different tumor types, facilitating a comprehensive evaluation of the TGFB1 gene. The results obtained from our analysis may provide valuable insights for prioritizing specific cancer types for subsequent clinical trials and experimental studies.
In recent years, advancements in high-throughput sequencing and mass spectrometry have enabled the detection of an increasing number of abnormally expressed genes related to ICIs across various cancer types. 26 TGFB1 is one of these common ICIs and is closely linked to cancer tumorigenesis and disease pathogenesis. 27 Previous studies have underscored the role of TGFB1 in the initiation and progression of clear cell renal cell carcinoma. 28 Additionally, existing evidence suggests that TGFB1 can serve as a prognostic biomarker in carcinomas.28,29 Our present study aligns with these findings. Comparative analysis revealed significant differences in TGFB1 expression between tumor tissue and normal paracancerous tissue across various cancer types, including BRCA, CHOL, GBM, and HNSC. Furthermore, our results indicate that elevated TGFB1 expression is associated with prognosis in several cancers, including COAD, LAML, LGG, MESO, STAD, SKCM, KIRC, and PRAD.
Increasing attention has been directed toward the TME. The TME encompasses stromal cells, immune cells, the extracellular matrix, the lymphatic and vascular systems, and bioactive molecules, collectively playing pivotal roles in cancer development, immune evasion, and therapeutic responses. 30 Multiple studies have highlighted the intricate and interconnected nature of the relationship between TME-infiltrating immune cells and TGFB1 expression. 31 In our present study, we also observed a significant association between TGFB1 expression and the extent of tumor-associated immune infiltration across various cancer types, particularly with regard to the infiltration of cancer-associated fibroblasts.
Currently, ICIs such as anti-CTLA-4 and anti-PD-L1/PD-1 antibodies have demonstrated enduring and effective responses against specific solid tumors. 32 However, treatment responses vary among individual patients, with differences in both efficacy and side effects. Hence, the identification of patients likely to benefit from ICI therapy is of paramount importance. In our study, we noted that TGFB1 expression increased concurrently with elevated expression of immune checkpoints on tumor cells. Consequently, heightened TGFB1 expression may serve as a valuable enhancer of immunotherapy outcomes in clinical practice. However, the precise mechanisms underpinning the relationship between TGFB1 expression and ICIs warrant further exploration.
TGFB is a prominent instigator of matrix remodeling and has garnered increasing interest in recent decades. This is primarily because of its pivotal role in the regulation of both normal and pathological processes in various organs and tissues, including functions such as immune tolerance, organogenesis, and synthesis of mucosa-restricted immunoglobulins (IgA) and their implication in cancer. Following GO and KEGG analysis of TGFB1 and TGFB1-related genes, our findings reveal that these genes are predominantly associated with the transmembrane receptor protein serine/threonine kinase signaling pathway, collagen-containing extracellular matrix, TGF-beta binding, proteoglycans in cancer, and the PI3K-Akt signaling pathway. These outcomes strongly suggest that TGFB1 plays critical roles in various cellular and physiological processes and exerts functional effects on tumor progression.
This study has several limitations. It provides initial data regarding the linkage between TGFB1 and various aspects of tumor progression, but further experimental investigations are necessary to elucidate the precise molecular mechanisms and processes through which TGFB1 contributes to carcinogenesis. Validation of our findings necessitates additional research conducted at the molecular and cellular levels. Simultaneously, the precise mechanisms governing the modulation of immunity by TGFB1 remain enigmatic. Furthermore, the availability of specific and comprehensive case data for drawing inferences regarding the efficacy of diverse medications in impeding tumor progression is currently lacking. In a forthcoming study, we will delve deeper into the intricacies of TGFB1 resistance mechanisms. Additionally, in our study, we have employed differentially expressed genes as a means to identify potential candidate genes associated with a specific condition or disease. However, it is crucial to recognize that differentially expressed genes alone do not always indicate causality and that they can be secondary to various factors.
Conclusions
This study showed that TGFB1 was elevated in multiple types of human cancer, and high expression correlated with poor prognosis. We also showed that abnormal expression of TGFB1 impacted tumor-related immune cell infiltration, the TME, ICI-related gene expression, and immunotherapy responses. These findings may contribute to clinical decision-making and cancer immunotherapy.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231221361 - Supplemental material for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer
Supplemental material, sj-pdf-1-imr-10.1177_03000605231221361 for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer by Zhitao Chen, Chenchen Ding, Jun Chen, Shusen Zheng and Qiyong Li in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605231221361 - Supplemental material for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer
Supplemental material, sj-pdf-2-imr-10.1177_03000605231221361 for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer by Zhitao Chen, Chenchen Ding, Jun Chen, Shusen Zheng and Qiyong Li in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605231221361 - Supplemental material for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer
Supplemental material, sj-pdf-3-imr-10.1177_03000605231221361 for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer by Zhitao Chen, Chenchen Ding, Jun Chen, Shusen Zheng and Qiyong Li in Journal of International Medical Research
Supplemental Material
sj-pdf-4-imr-10.1177_03000605231221361 - Supplemental material for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer
Supplemental material, sj-pdf-4-imr-10.1177_03000605231221361 for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer by Zhitao Chen, Chenchen Ding, Jun Chen, Shusen Zheng and Qiyong Li in Journal of International Medical Research
Supplemental Material
sj-pdf-5-imr-10.1177_03000605231221361 - Supplemental material for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer
Supplemental material, sj-pdf-5-imr-10.1177_03000605231221361 for Pan-cancer analysis revealing the multidimensional expression and prognostic and immunologic roles of TGFB1 in cancer by Zhitao Chen, Chenchen Ding, Jun Chen, Shusen Zheng and Qiyong Li in Journal of International Medical Research
Footnotes
Author contributions
All authors read and approved the final manuscript. Zhitao Chen and Chenchen Ding collected and analyzed the data. Zhitao Chen, Qiyong Li, and Shusen Zheng designed the study. Zhitao Chen and Jun Chen drafted the manuscript. Chenchen Ding, Qiyong Li, and Shusen Zheng reviewed and revised the manuscript. Chenchen Ding and Jun Chen edited the figures in the article.
All authors made substantial contributions to the study conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submission to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
