Abstract
Objective
This study explored the mechanism of squamous cervical cancer (SCC) progression.
Methods
Reverse transcription-quantitative polymerase chain reaction and western blotting were used to evaluate the expression of myosin light chain 9 (MYL9) in SCC tissues and cell lines. Furthermore, Transwell and Boyden assays were used to assess the function of MYL9 in SCC progression. In addition, the levels of lactate and aerobic glycolysis were used to explore the detailed mechanism of MYL9 in SCC.
Results
The mRNA and protein levels of MYL9 were elevated in SCC tissues, and MYL9 knockdown inhibited the migration and invasion of SCC cell lines. A mechanistic study demonstrated that MYL9 promotes SCC migration and invasion by enhancing aerobic glycolysis and increasing the activity of the Janus kinase 2 (JAK2)/signal transducer and activator of transcription 3 (STAT3) pathway.
Conclusions
MYL9 was upregulated in SCC, and it enhanced JAK2/STAT3 pathway activity and promoted metastasis and glycolysis in SCC.
Keywords
Introduction
Cervical cancer is the most common cancer in developing countries, and squamous cervical cancer (SCC) is the most common cancer globally.1–3 Although persistent high-risk human papillomavirus (HPV) infection is the main risk factor for SCC, 4 the detailed molecular mechanisms of migration/invasion in SCC are unclear. Thus, it is essential to study the detailed mechanisms driving the progression of SCC, which could facilitate the development of effective targeted therapeutic methods and decrease the morbidity and mortality of SCC.
Myosin light chain 9 (MYL9) encodes the regulatory light chain of myosin II (the most studied member of the myosin superfamily) and plays a crucial role in tumor progression and metastases.5–11 Accumulating evidence indicates that MYL9 is differentially expressed in various human tissues. Total MYL9 expression is lower in colon, gastric, bladder, non-small cell lung, and prostate cancers10,12–15 than in the corresponding normal tissues. Conversely, MYL9 is upregulated in ovarian cancer, 16 esophageal squamous cell carcinoma, 5 and glioblastoma, 8 and its overexpression is correlated with poor clinical outcomes. A previous study demonstrated that MYL9 induced aerobic glycolysis in endometrial fibroblasts. 17 However, to our knowledge, the detailed mechanism, expression, and function of MYL9 in SCC have not yet been reported.
When tumor cells have good oxygen availability, they utilize glycogen and undergo aerobic glycolysis (Warburg effect), thereby generating high levels of lactate. Glycolysis plays an important role in the occurrence and development of tumors, but the detailed molecular mechanisms remain to be clarified. Glucose transporter (GLUT1), a transmembrane protein, is responsible for the uptake of glucose into the cells of many tissues through facilitative diffusion. 18 Hexokinase II (HK2) is a predominant isoform in insulin-sensitive tissues, and it is also upregulated in many types of tumors associated with enhanced aerobic glycolysis. 19 Lactate dehydrogenase A (LDHA) is a glycolysis enzyme that converts pyruvate to lactate. 20
Our research demonstrated that MYL9 is upregulated in SCC, and it promotes migration and invasion by accelerating glycolysis. In addition, MYL9 activates the Janus kinase 2 (JAK2)/signal transducer and activator of transcription 3 (STAT3) signaling pathway, thereby promoting tumor progression in SCC.
Materials and methods
Cells and tissues
Human SCC cell lines (Siha, contains integrated copies of HPV-16; Caski, contains integrated copies of HPV-16 and HPV-18) were obtained from the Cell Resource Center, Shanghai Institute of Life Sciences, Chinese Academy of Sciences (Shanghai, China). Both cell lines were cultured in high-glucose Dulbecco’s Modified Eagle’s Medium (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Cytiva, Marlborough, MA, USA), and cells were cultured at 37°C in an atmosphere of 5% CO2. In addition, 36 pairs of matched SCC and healthy tissues were acquired from patients who underwent surgery at the Integrated Hospital of Traditional Chinese Medicine, Southern Medical University. Written informed consent was obtained from all patients before surgery, and this research was approved by the hospital’s ethics committee (No.: NFZXYEC-201906sb-K1). All procedures performed in this study followed with the ethical standards of the institutional research committee and the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
The mRNA sequences of clinical tissues or SCC cells were detected and quantified using RT-qPCR to better explore MYL9 gene expression. Total RNA was extracted from SCC tissues or cell lines using TRIzol (Takara Bio, Inc., Shiga, Japan). Complementary DNA (cDNA) was synthesized from 1 µg of total RNA using SuperScript III RT (Thermo Fisher Scientific). The PCR mix contained 1 µL of cDNA, 1× SYBR Green (Toyobo, Osaka, Japan), and specific primers. The primers were synthesized by IGE Co., Ltd (Guangzhou, China). The experiments were conducted more than three times using LightCycler 480 II (Roche, Basel, Switzerland). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA expression was detected as the internal control. The expression in each matched fresh peritumoral tissue sample was set as the control (the expression of MYL9 in all peritumoral tissues was set as 1), and the relative expression was determined by the 2−ΔΔCt method.
21
The thermocycling conditions
17
were 95°C for 10 minutes to activate DNA polymerase, followed by 45 cycles of 95°C for 15 s, 60°C for 15 s, and 72°C for 10 s. The specificity of amplification products was confirmed by melting curve analysis. Independent experiments were performed in triplicate. The primer sequences used in the polymerase chain reaction (PCR)
16
were as follows: MYL9 forward, 5′-
Transfection
siRNA targeting MYL9 (siMYL9) was designed by Guangzhou Ribobio Co., Ltd. (Guangzhou, China). The specific sequence of siMYL9 was 5′-GCAACATCGTCTTCAAGAA-3′. SiRNA (concentration: 50 nM, 10 µL) was transfected into Siha and Caski cells using Lipofectamine® 2000 (Thermo Fisher Scientific) according to the manufacturer’s protocol.
Transwell and Boyden assays
SCC cell migration and invasion assays were performed using Transwell chambers (8 μm, 24-well insert; Corning Inc., Corning, NY, USA). For the Transwell assay, cells (1 × 105) in serum-free medium were added to the upper chamber, and medium containing 20% FBS was added to the lower chamber. Cells were incubated for 12 hours at room temperature. For the Boyden assay, diluted Matrigel (BD Biosciences, Franklin Lakes, NJ, USA) was used to pre-coat the insert membrane. Cells (1 × 105) were cultured for 36 hours under the same conditions. Finally, the cells that migrated to or invaded the lower chambers were fixed with methanol, stained with Giemsa or crystal violet, and counted in six random fields under a microscope (BDS200, Olympus, Tokyo, Japan).
Lactate production assay
A lactate assay kit (Abcam, Cambridge, UK) was used to detect lactate production colorimetrically per the manufacturer’s protocols. siRNA-transfected SCC cells were seeded into a 96-well cell culture plate and incubated overnight at 37°C. The cells were subsequently starved for 12 hours, and the supernatant was obtained and used to detect lactate levels using a microplate reader (Biotek, Winooski, VT, USA) at 450 nm.
Western blotting
Flasks containing SCC cells were washed three times with ice-cold phosphate-buffered saline, and total protein was harvested using RIPA lysis buffer (Beyotime Institute of Biotechnology, Jiangsu, China) containing phenylmethanesulfonyl fluoride (Bio-Rad Laboratories, Inc., Hercules, CA, USA) and phosphatase inhibitors (Bio-Rad Laboratories) at a ratio of 100:1:1. A bicinchoninic acid protein assay kit (Beyotime Institute of Biotechnology) was used to determine the protein concentrations. Proteins (30 µg) were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis on 10% gels and transferred to a polyvinylidene fluoride membrane (Beyotime Institute of Biotechnology). The membranes were blocked with 5% bovine serum albumin in TBS containing Tween-20 (TBST) for 1 hour at 37°C and incubated with primary antibodies against MYL9 (mouse, 60233-1-Ig, Proteintech, Rosemont, IL, USA), GLUT1 (rabbit, 21829-1-AP, Proteintech), HK2 (mouse, 66974-1-Ig, Proteintech), LDHA (rabbit, 19987-1-AP, Proteintech), β-tubulin (mouse, Proteintech), and GAPDH (mouse, CW0100M, CWBIO, Beijing, China) overnight at 4°C. Following three washes with TBST, the membrane was incubated with horseradish peroxidase-conjugated goat anti-rabbit or anti-mouse IgG antibody at 37°C for 1 hour. The membranes were visualized using an electrochemiluminescence chromogenic kit (Beyotime Institute of Biotechnology) in a dark room, and images were captured with a ChemiDoc™ Molecular Imager (Bio-Rad Laboratories). The experiments were repeated at least three times.
Statistical analysis
All experiments were performed in sets of three or more, and all data were presented as the mean ± standard deviation. The differences between the two groups were analyzed using Student’s t-test, and graphs were plotted using GraphPad Prism 7 (GraphPad Software, Inc., San Diego, CA, USA). P < 0.05 was considered statistically significant.
Results
MYL9 is upregulated in SCC
Using 36 pairs of SCC and peritumoral tissues (Table 1), we observed a significant difference in the mRNA expression of MYL9 between tumor and healthy tissue (P < 0.0001, Figure 1a, b). Additionally, MYL9 protein was highly expressed in SCC cells (Figure 1c).
The detailed information of 36 patients with SCC.

SCC, squamous cervical carcinoma.
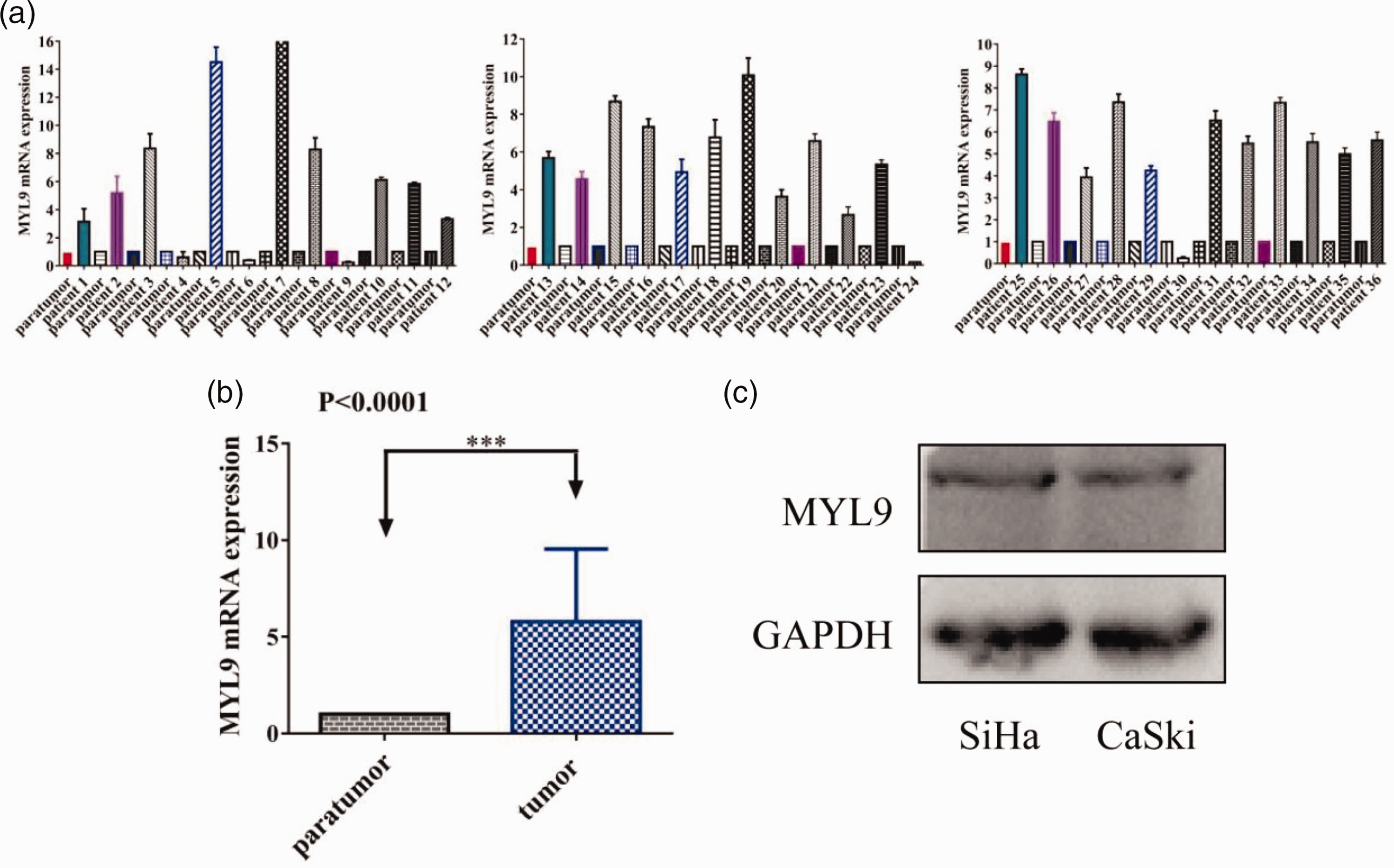
MYL9 mRNA and protein expression in SCC tissues and cell lines. (a, b) Reverse transcription-quantitative polymerase chain reaction revealed higher MYL9 mRNA expression in SCC tissues than in peritumoral tissues and (c) Western blotting illustrated that MYL9 protein was expressed in SCC cells. MYL9, myosin light chain 9; SCC, squamous cervical cancer.
MYL9 knockdown suppresses in vitro SCC cell migration and invasion
We performed Transwell and Boyden assays to detect the role of MYL9 in SCC. Our results indicated that SCC cell migration and invasion were significantly suppressed by MYL9 knockdown (Figure 2).

Transwell and Boyden assays demonstrated that MYL9 knockdown suppresses SCC cell migration and invasion. (a) Knockdown of MYL9 reduced the migration of SCC cells in the Transwell assay and (b) Knockdown of MYL9 reduced cell invasion in SCC cells in the Boyden assay (magnification, ×200; scale bar = 50 µm). MYL9, myosin light chain 9; SCC, squamous cervical cancer.
MYL9 promotes JAK2/STAT3 pathway activity and SCC migration and invasion by enhancing aerobic glycolysis
Owing to the close correlation between the progression of cancer cells and aerobic glycolysis, lactate levels were determined, and western blotting was performed. Interestingly, our results demonstrated that lactate levels were lower in the siMYL9 group (Figure 3a), and GLUT1, HK2, and LDHA protein expression was significantly inhibited after MYL9 knockdown (Figure 3b). To further explore the mechanism of MYL9 in aerobic glycolysis in SCC, western blotting was performed, and the phosphorylation of JAK2 and STAT3 was significantly reduced in the siMYL9 group (Figure 3c).
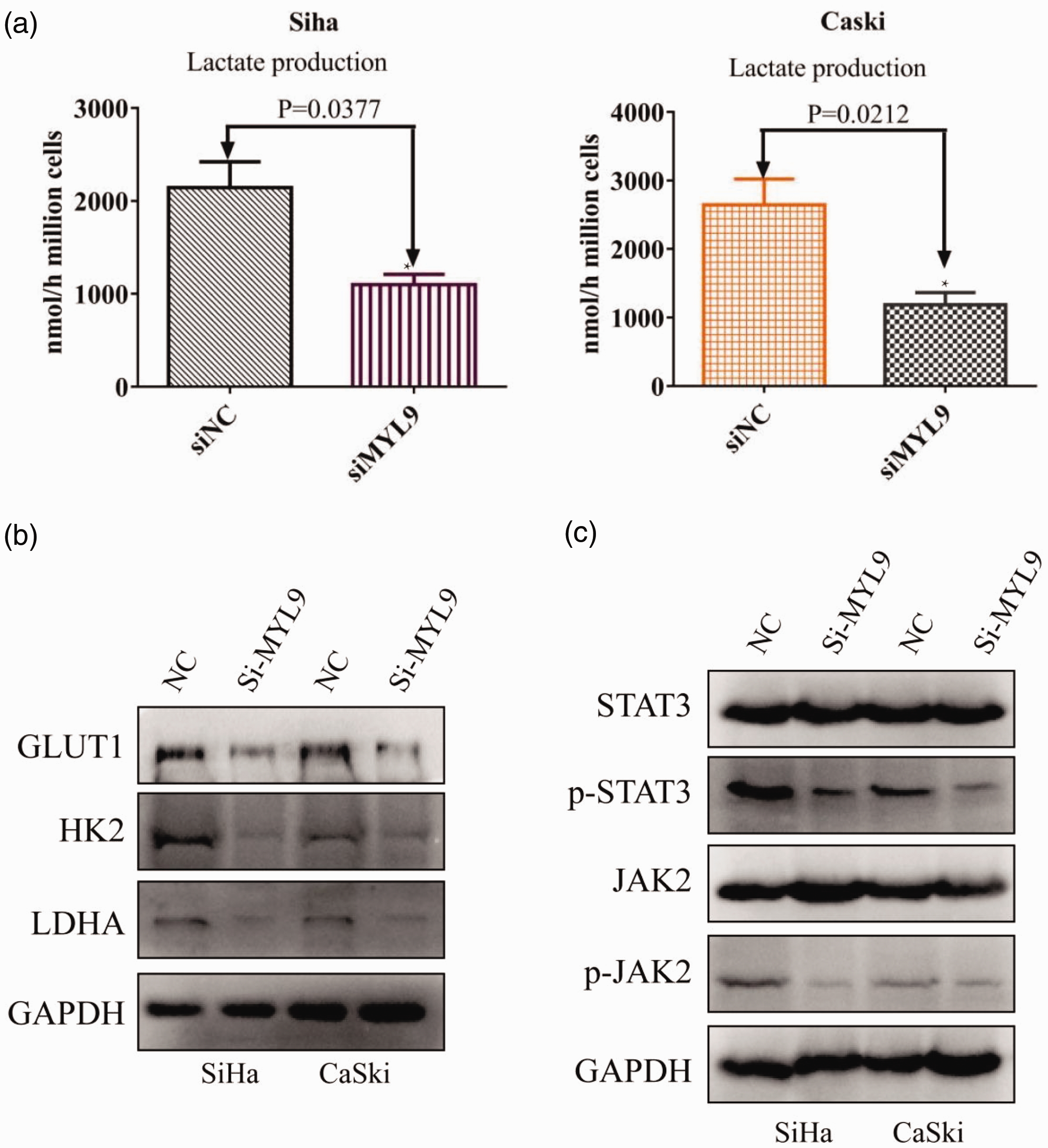
MYL9 promoted the progression of SCC by enhancing aerobic glycolysis. (a) MYL9 knockdown decreased the Lactate production of SCC. (b) MYL9 knockdown inhibited the expression of GLUT1, HK2, and LDHA and (c) MYL9 knockdown inactivated the JAK2/STAT3 pathway. MYL9, myosin light chain 9; SCC, squamous cervical cancer; GLUT1, glucose transporter 1; HK2, hexokinase II; LDHA, lactate dehydrogenase A.
Thus, these findings indicated that MYL9 induces JAK2/STAT3 pathway activity and promotes SCC migration and invasion by enhancing aerobic glycolysis.
Discussion
Recent studies illustrated that MYL9 is weakly expressed in non-small cell lung, gastric, bladder, colon, and prostate cancers.10,12–15 However, other reports illustrated that MYL9 is highly expressed in ovarian cancer, esophageal squamous cell carcinoma, and glioblastoma.5,8,16 To our knowledge, our study is the first to validate that MYL9 is highly expressed in SCC compared with its expression in peritumoral tissues, and MYL9 protein expression was also high in SCC cell lines, thus indicating MYL9 upregulation in SCC. However, additional studies are required to demonstrate its exact role in cancer.
Previous reports illustrated that MYL9 participates in many physiological and pathological functions including cell adhesion, polarity, and motility, and it plays a crucial role in some human cancers.5,8,12–16 Previous studies also reported that MYL9 plays a role in tumor suppression in gastric and colon cancers.10,12 On the contrary, other studies indicated that MYL9 has an oncogenic role in melanoma, glioblastoma, and breast cancer.8,9,11
GLUT1, HK2, and LDHA are key factors of aerobic glycolysis, and they play important roles in metabolic reprogramming and cancer progression.22–26 In this study, our results revealed that lactate levels were reduced in the MYL9 knockdown group compared with that in the control group, and GLUT1, HK2, and LDHA expression in SCC cells was decreased after the MYL9 gene was suppressed. MYL9 promoted SCC cell migration and invasion in vitro by enhancing aerobic glycolysis. The JAK2/STAT3 signaling pathway has been reported to produce a marked effect on all types of tumors.27–31 However, one limitation of the present study was that normal epithelial cells were not used as a control. Future studies are warranted to further demonstrate the detailed role of MYL9 in vivo.
Collectively, our results revealed that MYL9 was upregulated in SCC tissues compared with that in peritumoral samples, and MYL9 knockdown inhibited the migration and invasion of SCC cells by regulating aerobic glycolysis and its downstream factors (including GLUT1, HK2, and LDHA) via the JAK2/STAT3 pathway. Thus, MYL9 serves as a metastasis-related gene in SCC, and it is a potential biomarker for targeted treatment.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231208582 - Supplemental material for MYL9 promotes squamous cervical cancer migration and invasion by enhancing aerobic glycolysis
Supplemental material, sj-pdf-1-imr-10.1177_03000605231208582 for MYL9 promotes squamous cervical cancer migration and invasion by enhancing aerobic glycolysis by Bin Wen, Limei Luo, Zhaoyang Zeng and Xiping Luo in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605231208582 - Supplemental material for MYL9 promotes squamous cervical cancer migration and invasion by enhancing aerobic glycolysis
Supplemental material, sj-pdf-2-imr-10.1177_03000605231208582 for MYL9 promotes squamous cervical cancer migration and invasion by enhancing aerobic glycolysis by Bin Wen, Limei Luo, Zhaoyang Zeng and Xiping Luo in Journal of International Medical Research
Footnotes
Data availability statement
The datasets generated/analyzed during the present study are available.
Declaration of conflicting interest
The authors declare no conflicts of interest.
Funding
The present study was supported by the President funds of the Integrated Hospital of Traditional Chinese Medicine, Southern Medical University (No. 1201902001).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
