Abstract
We report a case of a woman with diabetes mellitus and choledocholithiasis who had a low fever with chills and severe weakness for 7 days. The patient’s abdominal tenderness was positive. Computed tomography and magnetic resonance imaging showed a giant abscess in the liver. The production of extended-spectrum β-lactamases by hypervirulent Klebsiella pneumoniae was found in the purulent fluid of the liver by nanopore-based metagenomic third-generation sequencing combined with an antibiotic susceptibility test. The patient recovered after intravenous antibiotic therapy and percutaneous drainage. Patients with a history of diabetes mellitus and choledocholithiasis should be aware of the possibility of pyogenic liver abscesses caused by hypervirulent Klebsiella pneumoniae. To rapidly control the development of this disease, nanopore-based metagenomic third-generation sequencing plays an important role not only in rapidly identifying pathogens, but also in guiding the use of antibiotics.
Keywords
Introduction
Since 1986 when the first case of pyogenic liver abscess (PLA) caused by hypervirulent Klebsiella pneumoniae (hvKp) was reported in Taiwan, China, PLA has gradually become a focus of global attention. 1 Infections from hvKp have spread throughout the world, causing various infections. 2 Underlying diseases, such as diabetes and bile duct stones, increase the risk of hvKp infections. 3 Metagenomic third-generation sequencing technology plays an important role in rapidly assisting the diagnosis of pathogens, and the detection results of drug-resistant genes have resulted in positive effects on the treatment and prognosis of patients. In this report, we describe a patient who was admitted with a giant liver abscess caused by hvKp.
Case report
The reporting of this study conforms to the CARE guidelines. 4 We obtained consent for treatment and signed consent to publish from the patient. The publication of the study has been approved by the Ethics Committee of the Third People's Hospital of Changzhou (approval number: O2A-A20210011).
A woman in her early 90s presented with chills and fever, accompanied by severe general weakness and a loss of appetite, which lasted for 7 days. She had a history of diabetes mellitus and choledocholithiasis, as well as hypertension for 30 years, which was under medical control. The patient had no special travel history of visiting areas with specific pathogens.
At a physical examination, there was swelling in the two transverse fingers below the xiphoid process. The swelling was clearly bordered and tender. Additionally, percussion pain in the liver area was observed. The patient had no clinical symptoms of eye or mental system diseases.
The results of a peripheral blood analysis are shown in Table 1. Multi-row computed tomography of the chest showed huge hypodense foci in the left lobe of the liver (Figure 1a). Abdominal ultrasound demonstrated a mixed echogenic mass in the left liver, which measured approximately 136.6 × 78.4 mm with dotted flocculent echogenicity, and was considered a liver abscess (Figure 1b).
Peripheral blood analysis results of the patient.


(a) Computed tomography scan of the patient shows huge hypodense foci in the left lobe of the liver on the first day of admission. (b) Abdominal color ultrasound findings of the patient on the first day of admission. A mixed echogenic mass in the left liver, measuring approximately 136.6 × 78.4 mm with dotted flocculent echogenicity, was observed. This mass was considered as a liver abscess and (c) Contrast-enhanced ultrasound of the liver on the third day of admission shows a space-occupying lesion in the left liver (148 × 116 mm)
Upon admission, because of the large size of the lesion, the patient received an intravenous infusion of the broad-spectrum carbapenem antibiotic biapenem as antibiotic therapy and albumin as nutritional support to rapidly control the development of the disease. On the third day after admission, contrast-enhanced ultrasound of the liver revealed a space occupying lesion in the left liver (148 × 116 mm) (Figure 1c). Magnetic resonance imaging confirmed the diagnosis of the abscess (Figure 2). On the seventh day, 310 mL of thick purulent fluid in the liver abscess was collected using ultrasound-guided puncture aspiration for identifying pathogenic microbes. Microbes were identified using metagenomic third-generation sequencing. Metagenomic third-generation sequencing is based on the third-generation single-molecule sequencing (nanopore) platform. After efficient genome nucleic acid extraction and library construction, the sample was sequenced by the Nanopore GridION platform (Oxford Nanopore Technologies, Shanghai, China). The sequencing results were compared and analyzed with professional medical microbial databases. We found that K. pneumoniae was positive with 3394 reads (Figure 3). Moreover, virulence genes (rmpA, iutA, and iroB) and drug resistance genes (CTX-M, tetA, and qnrS) were identified (Table 2). These results indicated that K. pneumoniae might be multidrug-resistant. On the 10th day, a bacterial culture of the liver drainage further confirmed the existence of K. pneumoniae, which was in accordance with the results of metagenomic third-generation sequencing. However, a blood culture was negative. The antibiotic susceptibility test was automated via the minimal inhibitory concentration using VITEK®2 COMPACT (Bio Mérieux, Lyon, France). The antibiotic susceptibility test suggested that the strain efficiently produced extended-spectrum β-lactamases (ESBLs) (Table 3). The combination of metagenomic third-generation sequencing and antibiotic susceptibility findings indicated ESBL-producing hvKp.

Abdominal magnetic resonance findings of the patient on the third day of admission. The volume of the left liver is increased, and lumpy mixed signal shadows can be seen in the left liver. There is a multilocular cyst in the left liver and a liver abscess can be seen. (a) A T1-weighted image shows an equal and low signal. (b) A T2-weighted image shows an equal and high signal. (c–e). Contrast enhanced scanning shows enhancement of the intralesional reticular septum and different sizes of cystic cavities. (c) Enhanced arterial phase; (d) enhanced venous phase and (e) delayed enhancement phase.

Results of nanopore-based metagenomic next-generation sequencing for pathogen detection in thick purulent fluid of a liver abscess (date: 16 January 2022). Klebsiella pneumoniae was identified as the predominant pathogen.
Metagenomic third-generation sequencing results of Klebsiella pneumoniae isolated from liver abscess drainage fluid.

Antibiotic susceptibility test results of Klebsiella pneumoniae isolated from liver abscess drainage fluid.

On the 20th day, the general condition of the patient gradually improved. The inflammatory indicators were mostly restored to normal levels. Moreover, color ultrasound showed that the liver abscess had shrunk (109.6 × 57.7 mm) (Figure 4a, b). Therefore, according to the drug resistance results, de-escalation was performed in which biapenem was replaced with piperacillin/sulbactam by intravenous infusion. Unfortunately, 1 week after changing the treatment regimens, the patient had a high fever (38.8°C). Therefore, biapenem was resumed until discharge. On the 52nd day, a reexamination suggested that the liver abscess had improved (Figure 4c). The patient was discharged and subsequently took levofloxacin orally.
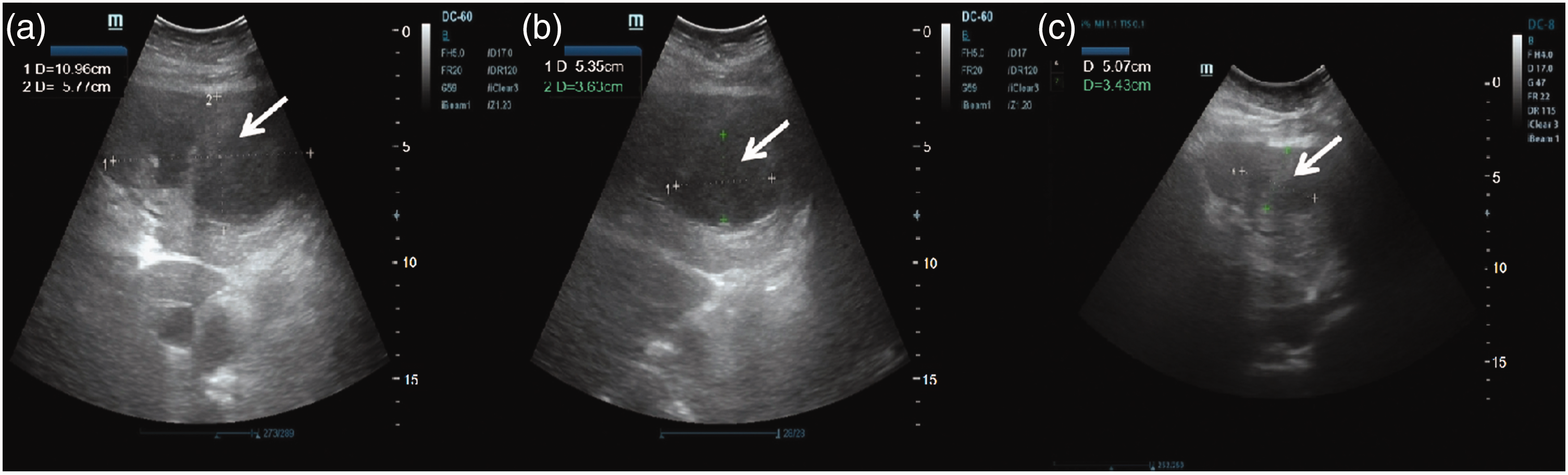
Abdominal color ultrasound of the liver. (a, b) Abdominal color ultrasound findings of the patient on the 20th day of admission. (a) The liver volume is normal. A mixed echo mass can be seen in the left liver, with a diameter of 109.6 × 57.7 mm. (b) A necrotic area can be seen in the left liver, with a diameter of 53.5 × 36.3 mm. (c) Abdominal color ultrasound findings of the patient on the 52nd day of admission. The liver has a normal volume and shape, smooth surface, soft medium texture, and diffuse, fine, enhanced, and scattered echo in the parenchyma. A parenchymal heterogeneous light mass can be seen in the left liver, which is 50.7 × 34.3 mm in size, and the boundary is not clear.
Discussion
We report a case of a giant liver abscess caused by ESBL-producing hvKp with no other metastatic infection. We also performed a systematic review of hvKp-infectious PLA cases. The literature search was performed with English language restriction using the electronic database PubMed (https://pubmed.ncbi.nlm.nih.gov/) from 1 January 1983 to 21 September 2022. The search terms were as follows: “hvKp”, “liver abscess”, and “case report”. We found that an hepatic abscess caused by drug-resistant hvKp is rare (Table 4).
Cases of liver abscess caused by hypervirulent Klebsiella pneumoniae.

ESBLs, extended-spectrum β-lactamases.
Since the first case of a liver abscess caused by K. pneumoniae was reported in 1986, 1 PLA caused by K. pneumoniae has become a new type of infectious disease worldwide. 5 K. pneumoniae is a major pathogen, and it also causes pneumonia, 6 endophthalmitis, 7 meningitis, 8 and sepsis. 9 Some of these conditions are metastatic infections occurring after a liver abscess. 10 PLA is usually caused by a mixed infection of multiple bacteria. 11 A case of PLA caused by a single bacterial infection (K. pneumoniae), without any other metastatic infection, occurred in our patient.
Compared with invasive PLA, the pathogenesis of cryptogenic PLA is unclear. K. pneumoniae colonizes the human gastrointestinal tract. In a previous report, a liver abscess associated with K. pneumoniae was related to a gastrointestinal tumor. 12 In addition, inflammation caused by bile duct stones is one source of a liver abscess. 13 These conditions may result in PLA. Interestingly, diabetes mellitus is associated with the occurrence and recurrence of PLA caused by K. pneumoniae infection to various degrees. 14 The possibility of PLA should be considered in patients with a history of these diseases. In our case, the patient had diabetes and choledocholithiasis. Our findings suggest that more attention should be paid to the occurrence and recurrence of PLA.
In recent years, there has been an increasing number reports of multidrug-resistant K. pneumoniae infection or hvKp infection, and drug resistance of bacteria is becoming increasingly more severe. 15 The strains of hvKp are typically capsular type K1 or K2. The expression of iron uptake system-related genes is positively related to the high virulence and drug resistance of K. pneumoniae. 16 Many studies have shown that hvKp is not necessarily a mucus phenotype, and some virulence genes were identified through highly virulent strains, such as iuc, rmpA, and rmpA2. 17 In our patient, the results of an antimicrobial susceptibility test of hvKp only confirmed that it produced ESBLs, but metagenomic third-generation sequencing showed that hvKP carried a drug-resistant gene to aminoglycosides, cephalosporins, and tetracyclines. The reason for the discrepancy between the genotype and the phenotype may result from transcription or translation. Because this study was a retrospective analysis, we did not retain the original strain. Generally, hvKp rarely has a drug-resistant phenotype. In our case, hvKp carried multiple drug resistance genes. A rapid and accurate diagnosis and rational administration of drugs are required to urgently control the development of this disease. The infection of multidrug-resistant hvKp will become a growing concern in the future.
Overall, monomicrobial cryptogenic non-invasive PLA secondary to K. pneumoniae remains a rare condition worldwide. The clinical symptoms of this condition are fever, chills, weakness, abdominal tenderness, and other non-specific symptoms. 18 Compared with traditional bacterial culture, metagenomic third-generation sequencing can be used to rapidly and accurately diagnose one or more different pathogenic microorganisms under the premise of unknown pathogens. There may be some differences between genotypes and phenotypes of drug resistance owing to an insufficient sample size, which suggests that metagenomic third-generation sequencing cannot completely replace bacterial culture. However, this method is especially suitable for diagnosing acute and critical cases and difficult infections. Currently, metagenomic next-generation sequencing is mostly performed on the Illumina platform. However, compared with the Illumina platform, nanopore-based metagenomic third-generation sequencing does not require polymerase chain reaction pre-processing, it has a more abundant database of drug resistance genes, and the turn-around time is approximately 5 to 7 hours. In our case, we used imaging and nanopore-based metagenomic third-generation sequencing to rapidly identify the pathogen of infection. DNA sequencing results of resistance genes are still important for the prognosis of patients.
Percutaneous drainage is the primary treatment for a liver abscess caused by K. pneumoniae. After performing an antimicrobial susceptibility test and gene sequencing, appropriate antibiotic treatment should be selected to avoid the abuse of antibiotics.
PLA caused by multidrug-resistant hvKp has rarely been reported (Table 4).19–26 The strain in our case showed a trend of multidrug resistance. There needs to be awareness of infection caused by hypervirulent strains with drug resistance. Clinically, in infected patients with a history of diabetes mellitus and gallstone disease, we should pay more attention to PLA. Additionally, with regard to cryptogenic liver abscess with an unknown origin, nanopore-based metagenomic third-generation sequencing is especially suitable for rapidly identifying the pathogen and analyzing the virulence or drug resistance genes within the effective range. This knowledge is necessary to guide clinical precise drug use, effectively improve the patient's condition, alleviate the form of bacterial drug resistance, and reduce an unnecessary economic burden.
In conclusion, the possibility of PLA should be considered in patients with a history of diabetes mellitus and gallstone disease. With regard to cryptogenic liver abscess with an unknown origin, nanopore-based metagenomic third-generation sequencing is especially suitable for rapidly identifying the pathogen and predicting drug resistance.
Footnotes
Acknowledgement
We thank the patient for agreeing to submit her case.
Author contributions
MMC conceived the study, treated the patient, and helped to draft the manuscript. XYH performed the statistical analysis and drafted the manuscript. TZ treated the patient, reviewed the medical records, and collected the data. TMX treated the patient. SL helped to review the manuscript. All authors read and approved the final manuscript.
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sector.
