Abstract
Objective
This meta-analysis aimed to update knowledge about the association between the SLC4A7 variant rs4973768 and breast cancer incidence.
Methods
Studies were identified from relevant digital databases. Fixed- or random-effects models were used to calculate odds ratios and 95% confidence intervals. Statistical Q and I2 tests and sensitivity analyses were used to detect interstudy heterogeneity and test the statistical stability of overall estimates, respectively. Egger’s tests were applied to detect publication bias among included studies. In silico analysis was used to ascertain increased expression of SLC4A7 mRNA in rs4973768 with the mutant allele. Trial sequential analysis was used to calculate the study’s sample size.
Results
The overall odds ratios reflected a positive correlation between the SLC4A7 rs4973768 polymorphism and susceptibility to breast cancer in five genetic comparisons of alleles T and C, and tests revealed significant heterogeneity in the allele comparison. After stratification by ethnicity, heterogeneity in Asian and White populations substantially decreased (Ph = 0.984, I2 = 0%) and remained stable (Ph = 0.083, I2 = 46.3%), respectively. The mutant allele was associated with increased expression of SLC4A7 mRNA in rs4973768. The cumulative z curve indicated that our conclusions were robust.
Conclusions
Our updated consequence shows that the SLC4A7 rs4973768 polymorphism is associated with increased breast cancer risk.
Introduction
Breast cancer is one of the most common cancers in women worldwide and accounts for 30% of female cancers in the USA, ranking first among female malignant tumors. 1 In its 2020 statistics, China’s National Cancer Center reported 416,371 new cases of female breast cancer, ranking first in the number of new cases of female cancer in China. 2 Breast cancer is mostly sporadic, and some breast cancers are hereditary. The pathogenesis of breast cancer is not completely clear. 3 A variety of gene changes produce abnormal gene products or functional changes that can lead to breast cancer. Therefore, identifying breast cancer susceptibility genes is very important. 4
As a possible substrate of tyrosine kinase, the SLC4A7 protein was shown to be downregulated in cell lines and neoplastic tissues.5,6 SLC4A7 rs4973768 exists in the 3′-UTR, whose specific function remains unclear.5,6 Because the variant was first associated with susceptibility to breast cancer in 2009, 7 a large number of studies have successively investigated the association between rs4973768 and breast cancer risk.8–17 However, subsequent studies have yielded inconsistent results. A meta-analysis on the relationship between SLC4A7 rs4973768 and breast cancer was conducted in 2012. 18 However, several new studies have examined the relationship between the rs4973768 variant and breast cancer risk since 2012.13–17 Therefore, to update the knowledge of the relationship between rs4973768 and breast cancer risk, we conducted a meta-analysis incorporating relevant studies published before 2022 and assessed the robustness of our conclusions using trial sequential analysis (TSA).
Methods
This study was approved by the ethics committee of the First Affiliated Hospital of Xi’an Jiaotong University and was conducted in accordance with the Declaration of Helsinki. The systematic review and meta-analysis were designed and reported in accordance with the 2020 preferred reporting items for systematic reviews and meta-analysis guidelines. 19 In error, we did not prospectively register this trial; however, we have now registered the trial retrospectively in INPLASY (registration number 202320013).
Search strategy
A comprehensive literature search was conducted in academic databases including PubMed, Embase, Medline, Google Scholar, the Cochrane Library, the Chinese National Knowledge Infrastructure, and Wanfang. Combinations of keywords, such as “BC, breast carcinoma or breast neoplasm” and “rs4973768 or SLC4A7,” were used in our search strategy. After reviewing the included articles, further relevant articles were assessed for incorporation into our research.
Selection criteria
The inclusion criteria for the meta-analysis were the following: 1) studies with both case and control groups; (2) studies assessing the relationship between the SLC4A7 rs4973768 polymorphism and susceptibility to breast cancer; (3) studies with sufficient information such as genotype frequency to allow the generation of odds ratios (ORs) and 95% confidence intervals (95% CIs); and (4) full-text articles of studies with human participants. Studies that met any of the following criteria were excluded: (1) conference abstracts, comments, reviews, case reports, or editorials; (2) studies with inadequate data for OR calculations; (3) studies without control groups; and (4) animal studies.
Quality assessment
All included studies were assessed for quality using the Newcastle–Ottawa Scale. The scale measures three aspects—selection, comparability, and exposure— and has a total score of 9. The studies could therefore be categorized based on their final score as high (score higher than 6), medium (score between 4 and 6), or low (score lower than 4) quality.
Data extraction
Data extraction was conducted by two reviewers separately. Incompatible opinions were resolved through discussion to reach a final agreement. The information acquired from each suitable study was the first author’s name, publication year, patient ethnicity, disease type, the sum of cases and controls, genotype and/or allele frequencies in case and control groups, and evidence of Hardy–Weinberg equilibrium in controls.
Statistical analysis
All data analyses were calculated using STATA software version 17.0 (StataCorp LP, College Station, TX, USA). The strength of the association between the SLC4A7 rs4973768 variant and breast cancer susceptibility was determined by calculating pooled ORs and 95% CIs. The interstudy heterogeneity assumption was examined using χ2-based q-statistics and I2 tests. If heterogeneity was deemed significant (P < 0.05 in the Q test or I2 > 50%), the random-effects model was used to obtain summarized OR estimates using the DerSimonian and Laird method; otherwise, the fixed-effects model was applied using the Mantel–Haenszel method. Additionally, when the heterogeneity between studies was statistically significant, subgroup analysis was conducted to identify potential causes of heterogeneity. Egger’s linear regression tests were used to assess probable publication bias among the studies. The stability of the joint consequences was inspected by applying a sensitivity analysis in which each of the studies in the analysis was sequentially removed, and then summary ORs were recalculated to observe changes between the original and recalculated ORs. P < 0.05 was considered statistically significant.
In silico analysis and trial sequential analysis
To explore the influence of polymorphisms on SLC4A7, we replicated the association between the rs4973768 variant and SLC4A7 expression levels using the GTEx Analysis Release V6 (dbGaP Accession phs000424.v7.p2) database. 20
TSA was conducted in accordance with the guideline of a former publication. 21 We set a significance of 5% for type I error and a 20% significance for type II error to calculate the required sample size and built the TSA monitoring boundaries.
Results
Study characteristics
In the online databases, 83 potentially relevant papers were identified through our search strategy. After identification and screening, 10 eligible articles covering independent studies on rs4973768 were ultimately included in our meta-analysis (Figure 1).8–17 Among these studies, seven and three offered information on White8–10,13,14,16,17 and Asian populations11,12,15, respectively. Table 1 provides more comprehensive information on the studies included in the meta-analysis.

Flowchart for literature search and selection.
Main characteristics of the studies included in the meta-analysis.

NOS: Newcastle–Ottawa Scale; HWE: Hardy–Weinberg equilibrium; Y signifies conforming to HWE.
Meta-analysis results
Data on the association between the SLC4A7 rs4973768 variant and breast cancer risk are summarized in Table 2. The joint results proved that the SLC4A7 rs4973768 polymorphism increased susceptibility to breast cancer in five genetic comparisons of allele T versus allele C (OR = 1.1, 95% CI = 1.05–1.14, P < 0.001), TT + CT versus CC (OR = 1.12, 95% CI = 1.06–1.18, P < 0.001), TT versus CC + CT (OR = 1.1, 95% CI =1.06–1.14, P < 0.001), TT versus CC (OR = 1.17, 95% CI = 1.09–1.25, P < 0.001), and CT versus CC (OR = 1.1, 95% CI = 1.04–1.16, P = 0.001).
Result of meta-analysis of the SLC4A7 rs4973768 polymorphism and breast cancer risk.
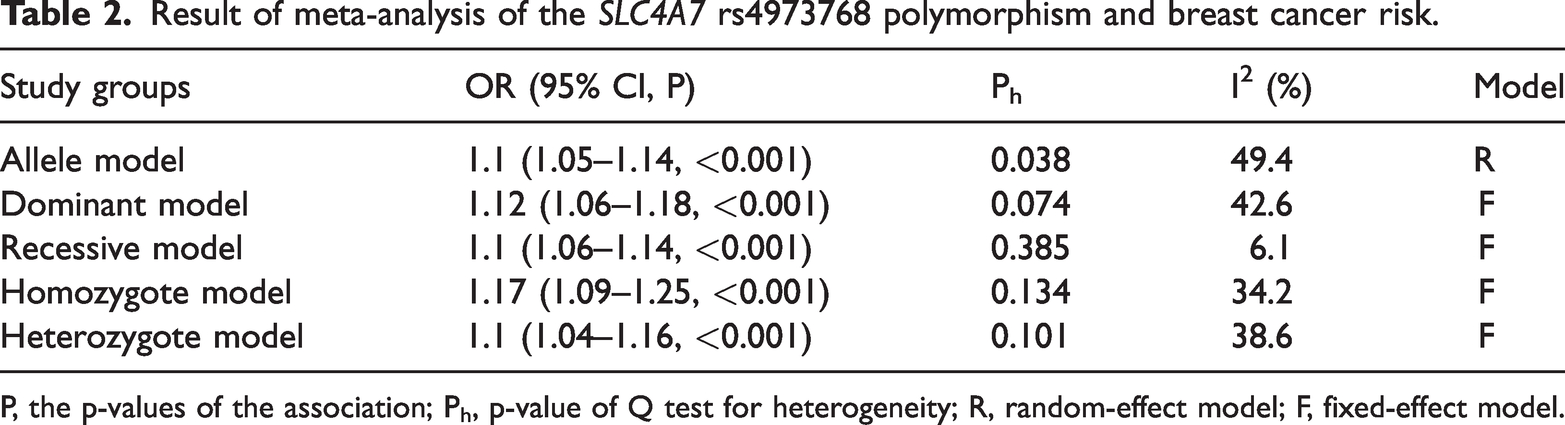
P, the p-values of the association; Ph, p-value of Q test for heterogeneity; R, random-effect model; F, fixed-effect model.
Heterogeneity test
The statistical results of the Q and I2 tests indicated obvious heterogeneity in the allele T versus allele C comparison (Figure 2). Therefore, the random-effects model was applied to calculate ORs in this comparison. The fixed-effects model was applied for genetic comparisons of the remaining four types of heterogeneity, which can be ignored (Supplementary Figure 1).

Forest plot for the association between the SLC4A7 rs4973768 polymorphism and breast cancer risk (including ethnicity subgroup analysis) in allele T vs allele C. The squares and horizontal lines correspond to the study-specific odds ratio (OR) and 95% confidence interval (CI), respectively. The area of the squares reflects the weight (i.e., inverse of the variance). The diamond represents the summary OR and the 95% CI.
Subgroup analysis was conducted when significant heterogeneity was observed in the comparisons. The results demonstrated that racial differences explained the significant heterogeneity of most or all sources. After stratification by race, the heterogeneity in the Asian population substantially decreased (Ph = 0.984, I2 = 0%); however, the heterogeneity in the White population remained (Ph = 0.083, I2 = 46.3%). All details are summarized in Supplementary Figure 2.
Sensitivity analysis
Results of sensitivity analysis indicated no material differences between original ORs and ORs recalculated after deleting any single eligible study (Figure 3) in all the comparisons, implying that our results were dependable and stable.

Sensitivity analysis for testing the stability of the overall estimate in allele T vs allele C (a), TT + CT vs CC (b), TT vs CC + CT (c), TT vs CC (d), and CT vs CC (e) comparisons.
Publication bias
Potential publication bias can be assessed with a visual inspection of Egger’s funnel plots and has statistical consequences. In our study, plot and P value (P > 0.05) results of Egger’s linear regression tests (Figure 4) revealed no significant publication bias among included studies in all comparisons.
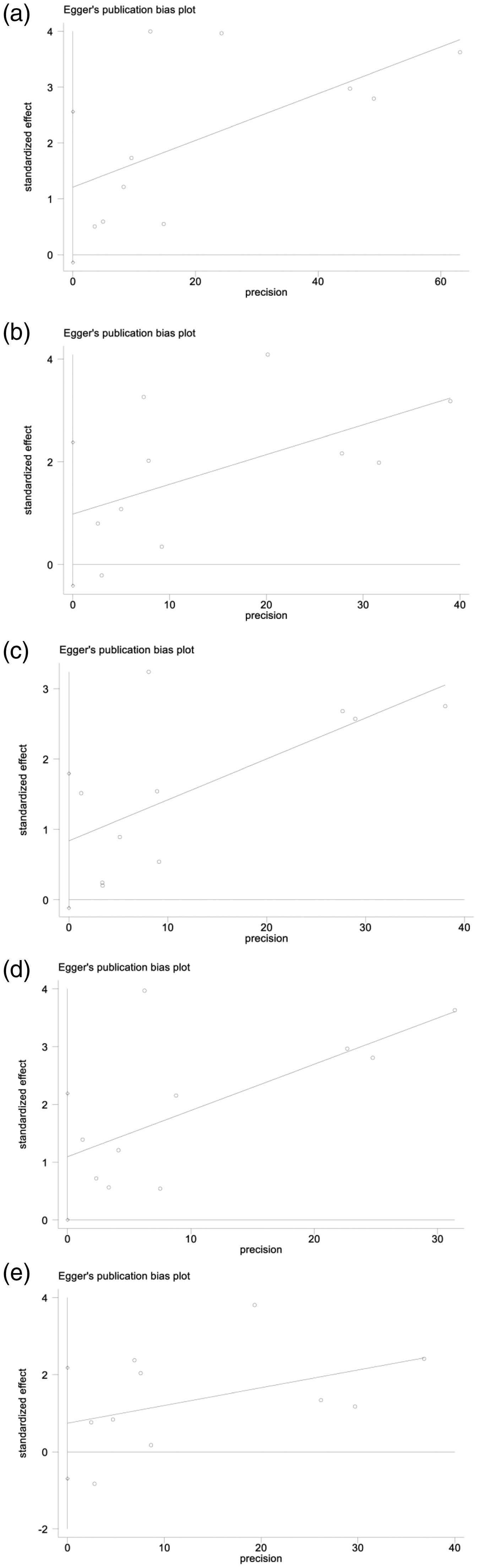
Egger’s publication bias plot and p-value in allele T vs allele C (a), TT + CT vs CC (b), TT vs CC + CT (c), TT vs CC (d), and CT vs CC and (e) comparisons. Each point represents a separate study for the indicated association.
In silico analysis and trial sequential analysis
The results of the GTEx portal database analysis indicated that the presence of the mutant allele was associated with the increased expression of SLC4A7 mRNA in the rs4973768 variant (P = 0.00069, Figure 5).

In silico analysis of SLC4A7 expression in rs4973768 and breast cancer SLC4A7 mRNA expression by eQTL analysis in human tissues based on the GTEx database.
The TSA result is shown in Figure 6. The required sample size was 109,971 samples, and the cumulative z curve crossed the trial sequential monitoring boundary before reaching the required sample size, signifying that our conclusions were robust.

Trial sequential analysis of SLC4A7 rs4973768 polymorphism under the allele contrast model.
Discussion
Breast cancer tumors are some of the most common malignant tumors among women, and targeted therapy of breast cancer is the hot spot of precision treatment.22–24 Targeted therapy refers to the delivery of a specific combination of drugs to carcinogenic sites in the body to cause the death of tumor cells without affecting normal tissues and cells around the tumor.25–27 this meta-analysis aimed to update the knowledge of the relationship between the rs4973768 single nucleotide polymorphism of the SLC4A7 gene and breast cancer to potentially provide support for future research on targeted gene therapy for breast cancer.
SLC4A7 is essential for encoding the sodium bicarbonate cotransporter, 28 influences the adjustment of intracellular pH drawn into visual and auditory sensory transfer and may be associated with susceptibility to cancer and cancer severity.5,6,29 However, genome-wide association studies have extensively examined the association between the SLC4A7 rs4973768 polymorphism and breast cancer risk and reported inconsistent results.17,30–32 Therefore, we used a meta-analysis to more precisely explore the correlation.
Interestingly, findings on the role of SLC4A7 rs4973768 in the occurrence of breast cancer are inconsistent. Some previous studies have reported no overall association between SLC4A7 rs4973768 and breast cancer risk, 33 while other researchers have found that the allele C of SLC4A7 rs4973768 reduces the risk of breast cancer. 18 As mentioned earlier, SLC4A7 rs4973768 was not even previously linked to breast cancer. A meta-analysis in 2012 reported that SLC4A7 rs4973768 was associated with breast cancer but did not use TSA to address the random error observed in the repeated updating of traditional meta-analyses and did not calculate the required sample size. 18 Moreover, the authors identified the small sample size as a limitation of the study. 18 Between 2012 and 2022, many new published studies have reported the risk relationship between SLC4A7 rs4973768 and breast cancer.13–17 To further update out knowledge of the relationship between SLC4A7 rs4973768 and breast cancer occurrence, we implemented a systematic review and meta-analysis that included all the relevant literature published before 20228–17 and assessed the robustness of our conclusions using TSA. We systematically reviewed the association between the SLC4A7 polymorphism rs4973768 and breast cancer risk using 10 case–control studies including a total of 37,128 cases and 43,640 controls. Our meta-analysis concluded that SLC4A7 rs4973768 correlated with the occurrence of breast cancer, and allele C of the SLC4A7 rs4973768 mutation played a protective effect against breast cancer. The result of TSA showed the cumulative z curve crossed the trial sequential monitoring boundary before reaching the required sample size, confirming that our results were robust. In addition, we found that the mutant allele was associated with an increased expression of SLC4A7 mRNA in the rs4973768 variant (P = 0.00069), according to the results from the GTEx portal database. Overall, our findings provide evidence to support future research on SLC4A7 gene therapy targeting breast cancer.
This study had some limitations. First, because gene-with-gene and gene-with-environment interactions may affect the risk of breast cancer, assessing the interaction between a genotype polymorphism and possible factors of breast cancer such as psychology and environment is difficult. Such a study can only estimate the risk relationship between one abnormal genotype and breast cancer. Second, our meta-analysis included only Chinese and English studies, thereby introducing language bias. Third, only one polymorphic site of rs4973768 was included in our study; the polymorphic sites linked to other genes were not considered. The polymorphic sites linked to other genes may have affected the pathogenesis of breast cancer. More high-quality studies with large sample sizes must be included in the future.
Conclusion
Our meta-analysis indicated that the rs4973768 polymorphism of the SLC4A7 gene is markedly correlated with the occurrence of breast cancer, and allele C is protective against breast cancer. These findings provide preliminary research data for assessing the risk of breast cancer associated with the rs4973768 single nucleotide polymorphism and are expected to lay the foundation for future research in personalized medicine targeting breast cancer.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231166517 - Supplemental material for Update on the relationship between the SLC4A7 variant rs4973768 and breast cancer risk: a systematic review and meta-analysis
Supplemental material, sj-pdf-1-imr-10.1177_03000605231166517 for Update on the relationship between the SLC4A7 variant rs4973768 and breast cancer risk: a systematic review and meta-analysis by Yuhui Zhou, Xiaoxia Ma and Jinglan Sun in Journal of International Medical Research
Footnotes
Author contributions
(I) Conception and design: YZ, XM; (II) Administrative support: YZ, JS; (III) Provision of study materials or patients: YZ; (IV) Collection and assembly of data: YZ; (V) Data analysis and interpretation: YZ; (VI) Manuscript writing: all authors; and (VII) Final approval of manuscript: all authors.
Data availability statement
The data used to support the findings of this study are available from the corresponding author upon request.
Declaration of conflicting interests
All authors have completed the ICMJE uniform disclosure form. The authors have no conflicts of interest to declare.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Research & Development Plan of Shaanxi Province (2021SF-212).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
