Abstract
Objective
To explore underlying mechanisms that regulate hMSH2 expression and drug susceptibility in epithelial ovarian cancer (EOC).
Methods
Using data from the Cancer Genome Atlas (TCGA) we used bioinformatical analysis to predict transcription factors (TFs) that potentially regulate hMSH2. RT-qPCR, Western blot, and luciferase assays were undertaken using ovarian cancer cell lines to verify the identified TF. Expressions of the TF were modulated using overexpression or knockdown, and the corresponding cellular responses to cisplatin were examined.
Results
The TF, E2F1, was found to regulate the hMSH2 gene. The expression level of E2F1 correlated with cisplatin susceptibility in vitro. Kaplan-Meier analysis of 77 patients with EOC showed that low E2F1 expression was associated with worse survival.
Conclusions
To our knowledge, this is the first report of E2F1 regulated MSH2 expression playing a role in drug resistance of platinum-based treatments for patients with EOC. Further work is need to confirm our results.
Introduction
Epithelial ovarian cancer (EOC) ranks fifth in deaths caused by cancer in women and leads to more deaths than any other reproductive system cancer. 1 According to the American Cancer Society estimates for ovarian cancer, approximately 19,880 women will be new cases, and approximately 12,810 women will die in 2022 in the United States. (https://seer.cancer.gov/statfacts/html/ovary.html). Late diagnosis and platinum drug resistance are the main reasons for high mortality.2,3 Despite good initial responses to platinum-based chemotherapy, up to 80% women will relapse and progress to platinum resistance with a median progression-free survival (PFS) of 12–18 months.3,4 Therefore, primary or acquired resistance to platinum in ovarian cancer impedes treatment and is an important factor in cancer-associated mortality. 5
Platinum agents may inhibit DNA synthesis and affect its replication by changing the formation of inter- and intra-strand DNA cross-links. 6 Therefore, a significant determinant of the resistance to platinum agents may originate in DNA repair capacity. 7 As an important DNA repair mechanism, the mismatch repair (MMR) system plays an essential role in platinum susceptibility 7 . Cisplatin and carboplatin related DNA damage can be recognized by the human mismatch-repair protein, hMSH2 or by the hMSH2-hMSH6 complex.8–10 Indeed, hMSH2 is thought to have a notable role in the platinum resistance of ovarian cancer. 10 A study in bladder cancer cells has shown that as a mediator of cisplatin sensitivity, circRNA generated from the circularization of LIFR gene (circLIFR) could interact with hMSH2, and positively regulate the sensitivity to cisplatin in vivo and in vitro. 11 In a previous study, we found that hMSH2 low expression due to hypermethylation was associated with platinum resistance in EOC and we hypothesised that hMSH2 might be a target for epigenetic therapy in platinum-resistant patients.11 However, more regulatory mechanisms of hMSH2 need to be explored to study the MMR system in platinum resistance.
The E2F family is a group of genes that encode transcription factors that are widely expressed in many organs. 12 The eight members of E2F have different homology which affects their function; E2F1–3 are activators of transcription, whereas E2F4–8 act as repressors.12,13 Importantly, the deregulated expression of E2F proteins has been found to be a common phenomenon in a wide range of cancers.12,14 E2F1-regulated cellular functions include cell cycle control, DNA replication and apoptosis.15,16 In this study, bioinformatic analysis was used to explore underlying mechanisms that regulate hMSH2 expression and drug susceptibility in epithelial ovarian cancer (EOC).
Materials and Methods
Data source and online tools
The 2 kb upstream sequence from the hMSH2 start codon was obtained from the UCSC Genome Browser 17 (https://genome.ucsc.edu). PROMO database 18 (http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3) was used to seek and analyse the putative transcription factor binding sites (TFBSs) of the 2 kb sequence. After selection, the maximum matrix dissimilarity rate was set to 5% for target DNA sequence scanning.
The GEPIA 2 19 (http://gepia2.cancer-pku.cn/#index) was used to analyse gene expression profiles. The Cancer Genome Atlas (TCGA) dataset (http://xenabrowser.net) was collected to investigate the expression of E2F1 and hMSH2 in 33 cancers.
PrognoScan database 20 (http://www.abren.net/PrognoScan/) was used to analyse the correlation between survival in pan-cancer and E2F1 expression using Cox proportional hazard model. The prognostic value of E2F1 mRNA levels for patients with EOC was evaluated using the Kaplan-Meier plotter 21 (http://www.kmplot.com).
Patients and specimens
Tissue samples were collected from 77 women with histologically confirmed EOC who were enrolled into the study from May 2017 to May 2018. All patients had undergone cytoreductive surgery followed by platinum-based chemotherapy at the affiliated Xingtai People Hospital of Chengde Medical University. No other treatments had been applied before surgery. Participants were grouped into platinum-sensitive (n = 56) or platinum-resistant (n = 21) patients according to a platinum-free interval (PFI), which was calculated as the interval from the date of the last platinum treatment to the date of disease progression. Platinum-resistant patients had a PFI of <6 months, and platinum-sensitive patients had a PFI of >6 months. 22 Patients were followed-up every three months for more than 36 months. Survival status was evaluated using overall survival (OS) and PFS.
Prior to study commencement, patients were informed about its objectives and they provided written informed consent. The study was approved by the local Ethics Committee of Xingtai People Hospital of Chengde Medical University (Approval Letter NO. 2017 (036).
Quantification of E2F1 and hMSH2 transcript levels in tissue samples
TRIzol reagent (ThermoFisher Scientific, US) was used to isolate total RNA from EOC tissues, according to the manufacturer's protocol. Revert Aid First Strand cDNA Synthesis Kit (ThermoFisher Scientific, US) was used to synthesize the total cDNA. Custom primers for E2F1 (forward:
Immunohistochemistry
In total, 36 tissue samples were obtained from the 77 patients and immunohistochemistry (IHC) staining was performed for hMSH2 and E2F1 using the hMSH2 antibody (ab212188, Abcam, 1:10000) and E2F1 antibody (ab288369, Abcam, 1:50). Nuclear staining without cytoplasmic staining was considered positive. E2F1 and hMSH2 protein expressions were evaluated based on previous studies.23,24 Specifically, staining intensity and percentage of positive staining area of the IHC results were set as judgment criteria.
Cell culture
OVCAR-3 and SKOV-3 cells were purchased from iCell Bioscience Inc. (Shanghai, China) and were cultured at 37°C with 10% CO2 in RPMI-1640 or DMEM/F-12 (Gibco; Thermo Fisher Scientific, Inc.), respectively.
RNA interference and E2F1 overexpression
E2F1 plasmids and their negative controls (NC) were purchased from the Gene Pharmaceutical Technology Company (Shanghai, China). The specific sequences for the interference RNAs were as follows
F: GGGAGAAGUCACGCUAUGA, R: UCAUAGCGUGACUUCUCCC (siE2F1-1); and F: CCUGAUGAAUAUCUGUACU, R: AGUACAGAUAUUCAUCAGG (siE2F1-2). The E2F1 expressing (E2F1-overexpression, OE) and the negative control lentivirus (NC) were constructed by the Gene Pharmaceutical Technology Company (Shanghai, China). The Lipofectamine 3000 reagent (Invitrogen, Carlsbad, USA) was used for transfections.
Western blot assay
Western blot assay was performed according to a previous protocol. 25 Antibodies were hMSH2 (Abcam, ab212188, 1:1,000), E2F1 (Abcam, ab137415, 1:1,000), GAPDH (Abcam, ab9485, 1:2,500), and the anti-rabbit secondary antibody (Abcam, ab6721, 1:2,000). The GAPDH was used as the loading control.
Luciferase Reporter Assay
The luciferase assay was performed as previously described. 26 Briefly, a hMSH2 promoter region covering −1500 ∼ +500 bp of the genomic loci of hMSH2 was subcloned into a reporter vector. The luciferase activity was examined when the E2F1 gene was up or down regulated. The assay was performed in ovarian cell lines SKOV-3 and OVCAR-3. Cells were seeded into 96-well plates using 5000 cells per well. After 24 h, 5 ng of pRL-TK, 100 ng of pGL3.0-basic or pGL3.0-MSH2, and 5 pmol of siRNA or 100 ng overexpressing plasmid were transfected into cells. After 48 hours, Firefly and Renilla luciferase activities were detected using the dual-luciferase reporter assay system (Promega, WI, USA).
Electrophoretic Mobility Shift Assay
Biotin labelled 146 bp DNA fragment in the hMSH2 promoter region containing predicted E2F1 binding site was synthesized (Integrated DNA Technologies, Coralville, IA, USA). Corresponding sequence is as below:
5′-
The synthesized DNA fragment at 2 × 10−7 M concentration was mixed with recombinant human E2F1 protein (ab82207; Abcam) in a buffer containing 50 mM Tris –HCl pH 7.0 at 20°C, 100 mM NaCl, 5% v/v glycerol, 0.01 mg/mL bovine serum albumin, 0.1 mMDTT in 20 µl volume at 37°C for 30 min. The amount of E2F1 proteins were 50, 100, and 150 ng respectively. After binding, the mixtures were run in a 8% polyacrylamide native gel casted in the same buffer as the binding step at 10 V/cm. After electrophoresis was completed, DNA was transferred from gel to a zeta-probe nylon membrane (BIO-RAD) using standard capillary transferring. The DNA was crosslinked to the membrane by exposure to an UV transilluminator for 3 min. The membrane was blocked in 0.1 M Tris –HCl pH 7.5, 0.1 M NaCl, 2 mM MgCl2, and 3% BSA for 1 h at room temperature. The membrane was incubated in the washing buffer (0.1 M Tris –HCl pH 7.5, 0.1 M NaCl, 2 mM MgCl2) containing streptavidin alkaline phosphatase (Promega, V5591, 1:1000) at room temperature for 10 min and then washed in the wash buffer twice, 10 min each time. Additional 10 min wash was performed in 0.1 M Tris –HCl pH 9.5, 0.1 M NaCl, 50 mM MgCl2. The membrane was then developed with the BCIP/NBT colour development substrate (Promega, S3771) according to the manufacturer’s instructions. The reaction was stopped by soaking the membrane I the TE buffer (10 mM Tris –HCl pH 8.0, 1 mM EDTA pH 8.0).
Cisplatin cytotoxicity assay
Culture media containing different concentrations of cisplatin ranging from 5–30 μM were applied to E2F1-overexpression cells and negative controls for 24 h. Cell proliferation was checked using a Cell Counting Kit-8 (ab228554; Abcam) according to the manufacturer's protocol. Absorbance was measured using EL800 microplate reader (BioTek Instruments, Inc., Winooski, VT) at 492 nm. Each experiment was performed three times.
Apoptosis assay
Apoptosis was performed using an Annexin V Apoptosis Detection kit I (BD Biosciences, Franklin Lakes, NJ, USA). E2F1-overexpression and negative control cells were treated with 10 μM cisplatin (Sigma-Aldrich, St., Louis, MO, USA) for 24 h. Phosphate buffered saline (PBS) was used as control. After treatment, 5 µl propidium iodide and annexin V were added to the cell suspension, followed by a 30 min incubation (dark, room temperature). The volume was then made up to 500 μl; cells were counted using FACSAriaTM II (BD Biosciences). Each experiment was performed three times.
Statistical analysis
All statistical analyses were performed using SPSS software (version 21 for Windows®; IBM Corp, Armonk, NY, USA). P value <0.05 was considered statistically significant. Cell apoptosis and viability assays were analysed using Student’s t-test. Wilcoxon Rank Sum test was used for E2F1 and hMSH2 mRNA expression, and Spearman's correlation analysis was used for their co-relationship analysis. Associations of patients' prognosis with E2F1 expression was analysed using Kaplan-Meier analysis.
Results
Patient characteristics
Median age of the patients was 56 years (range 23–77). Histologically, 55 (71%) patients were diagnosed with serous adenocarcinoma and 22 (29%) with endometrioid carcinoma (Table 1). According to International Federation of Gynaecology and Obstetrics (FIGO) staging, 29 cases (38%) were stages I–II, and 48 (62%) were stages III–IV. There were no statistically significant differences in age, stage, grade, pathological classification, or tumour size between patients in the platinum-sensitive (n = 56) and platinum-resistant (n = 21) group (Table 1).
Demographic baseline characteristics of patients with epithelial ovarian cancer.

Data are expressed as n.
FIGO: International Federation of Gynaecology and Obstetrics; ns, not significant.
hMSH2 expression was associated with the E2F1 gene
A previous study7,27 suggested that hMSH2 could be a modulator of platinum susceptibility. Therefore, we decided to explore potential regulators of hMSH2. In total, 49 transcription factors were screened with PROMO software. The transcription factor E2F1, was found to be a strong candidate that binds the −162 to −155 and the −124 to −117 regions of the hMSH2 gene (Figure 1a). A positive correlation was found between expression of hMSH2 and E2F1 in samples of EOC from TCGA database (r = 0.4, P < 0.05, Figure 1b). Additionally, significant positive correlations were found in the co-expression of hMSH2 and E2F1 in a pan-cancer analysis (Table 2). A positive correlation was also found between hMSH2 and E2F1 mRNA levels in samples from 77 patients with EOC (r = 0.73, P < 0.001, Figure 1c).

E2F1 is predicted to regulate hMSH2. (a) Two predicted binding sites of E2F1 upstream of transcription start site (TSS) of hMSH2. (b) Correlation between expression of E2F1 and hMSH2 in samples of epithelial ovarian cancer (EOC) from the Cancer Genome Atlas (TCGA) database. TPM, transcript per million and (c) Correlation of E2F1 and hMSH2 mRNA levels in samples from 77 patients with EOC.
Correlation of E2F1 and hMSH2 across 33 cancer types from the Cancer Genome Atlas (TCGA) database.

E2F1 regulates hMSH2 expression in ovarian cancer cell lines
Although a strong co-expression pattern was found between E2F1 and hMSH2 genes, the hypothesis that E2F1 regulates hMSH2 needed to be validated. Therefore, the E2F1 gene was either over-expressed exogenously or was knocked down endogenously in SKOV-3 and OVCAR-3 ovarian cancer cell lines. The mRNA and protein levels of hMSH2 were analysed by RT-qPCR and Western blot analysis. Results showed that when E2F1 expression was knocked down, the hMSH2 mRNA and protein levels (siE2F1 group) were correspondingly decreased compared to the random control (siNC) (Figure 2a). Results from the overexpression experiment agreed with the knockdown experiment. When the E2F1 mRNA or protein levels increased, the hMSH2 mRNA and protein levels were upregulated (Figure 2b, OE-E2F1). These results indicated that the expression of hMSH2 was regulated by E2F1.
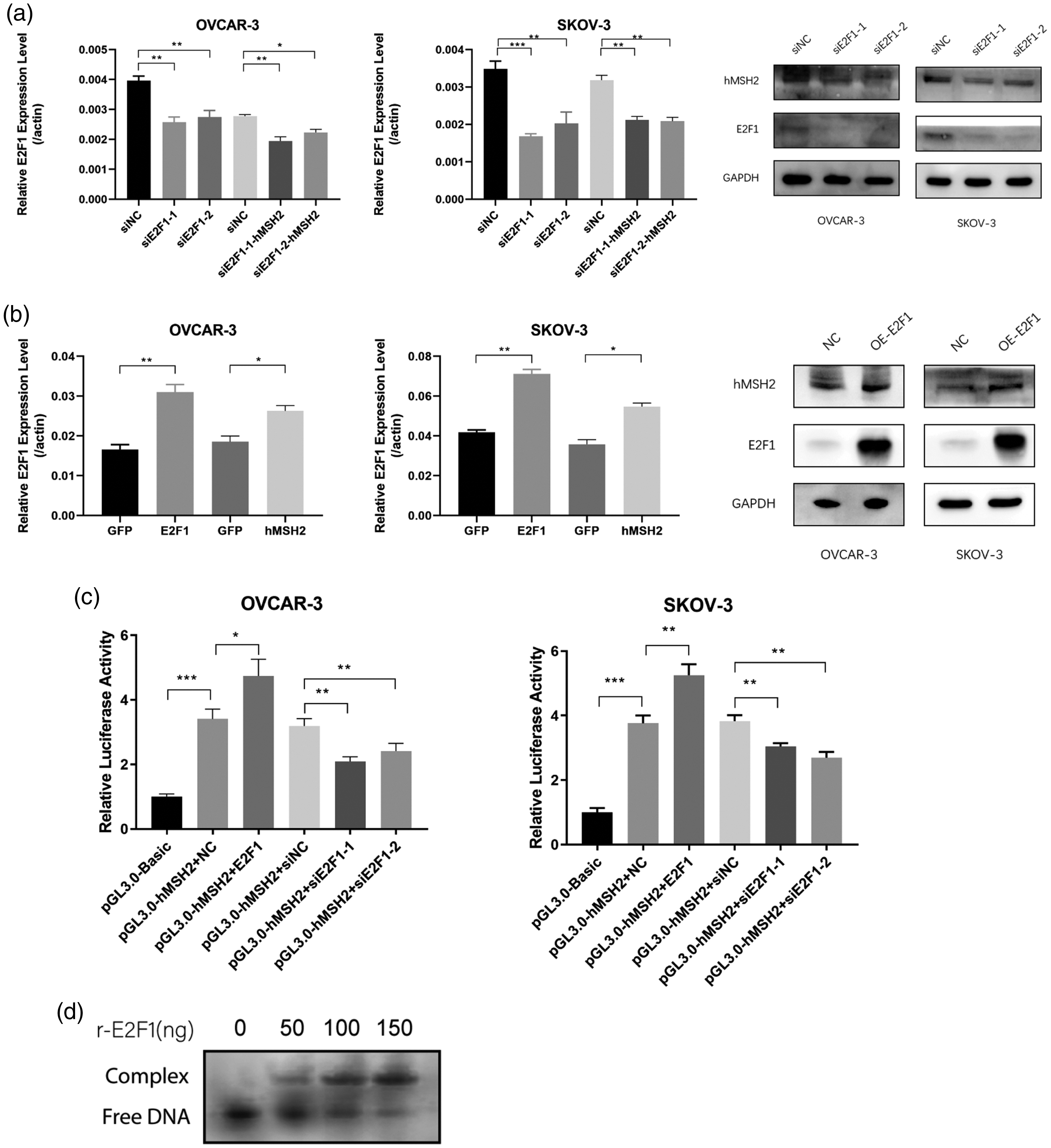
E2F1 regulates the expression of hMSH2 in ovarian cancer cells. (a) E2F1 knockdown resulted in hMSH2 under-expression. E2F1 was knocked down in OVCAR-3 and SKOV-3 cells and the mRNA (left, middle) and protein (right) levels of hMSH2 were also downregulated correspondingly. GADPH, glyceraldehyde 3-phosphate dehydrogenase; si-NC, random control; si-E2F1, E2F1 knockdown. *P < 0.05, **P < 0.01, and ***P < 0.001. (b) E2F1 overexpression resulted in hMSH2 upregulation. E2F1 was overexpressed in OVCAR-3 and SKOV-3 cells, and hMSH2 was also upregulated correspondingly at mRNA (left) and protein (right) levels. GFP, green fluorescent protein; GADPH, glyceraldehyde 3-phosphate dehydrogenase; NC: empty vector control; OE-E2F1, E2F1 overexpression. *P < 0.05 and **P < 0.01. (c) Luciferase showed the regulation of E2F1 on the hMSH2 promoter. The promoter region of hMSH2 was subcloned into a reporter vector and the luciferase signal intensity altered in coordination with the E2F1 level. *P < 0.05, **P < 0.01, and ***P < 0.001 and (d) Electrophoretic Mobility Shift Assay showing binding of E2F1 to the promoter region of hMSH2. Synthesized 146 bp promoter region of hMSH2 was incubated with 50, 100, and 150 ng recombinant E2F1 and separated in native gel. Upper bands show the shifted mobility of DNA/E2F1 complex.
The dual luciferase assay is a technology that is widely used to study gene transcription and regulation.28,29 We performed a luciferase assay to demonstrate that the promoter region of hMSH2 was regulated by E2F1. We subcloned the hMSH2 promoter region before the firefly luciferase gene as the reporter, and used a non-E2F1 regulated promoter (SV40) combining renilla luciferase gene as the basal comparison. When E2F1 was overexpressed (pGL3.0-hMSH2 + E2F1, OVCAR-3 and SKOV-3 cell lines), the relative firefly luciferase activity (firefly/renilla) increased compared to the negative control (NC, no E2F1 overexpression) (Figure 2c). Correspondingly, when the endogenous E2F1 was knocked down, the relative firefly luciferase activity also decreased (pGL3.0-hMSH2 + siE2F11-1/2). These results showed E2F1 regulates the promoter of hMSH2.
We confirmed the direct binding of E2F1 to the promoter of hMSH2 using electrophoretic mobility shift assay (EMSA). The synthesized DNA corresponding the hMSH2 promoter region covering predicted E2F1 binding site was incubated with recombinant E2F1 and the DNA/protein mixtures were analysed with EMSA. The DNA showed an apparent mobility shift after the incubation with the recombinant E2F1 (Figure 2d). The result reaffirmed the regulatory function of E2F1 on hMSH2.
E2F1 overexpression reduces cisplatin resistance in ovarian cancer cells
Apoptotic analysis and cell viability assays were performed to validate the effect of E2F1 on cisplatin susceptibility in SKOV-3 and OVCAR-3 cells. Cells overexpressing E2F1 (OE group) were exposed to various concentrations of cisplatin for 24 h, and cell viabilities were compared with controls via CCK-8 assays. A moderate but statistically significant decrease of viable cells was found in the OE group (P < 0.05, Figure 3a). The flow cytometry assay also showed substantially increased apoptotic cells in the E2F1 OE group when 10 μM cisplatin was applied for 24 h (P < 0.05, Figure 3b and 3c).

Overexpression of E2F1 increased susceptibility of ovarian cancer cells to cisplatin. (a) Percentage of viable cells (OVCAR-3 and SKOV-3) with or without E2F1 overexpression incubated with cisplatin (5–30 μM) after 24 h and (b and c) Percentage of apoptotic cells with or without E2F1 overexpression counted by flow cytometry after cisplatin treatment. Con, control; OE, over expressed. *P < 0.05 and **P < 0.01.
E2F1 associated with platinum resistance in EOC
With the above supporting evidence, we explored the expression of E2F1 and hMSH2 in platinum-sensitive and resistant EOC tissues. The average E2F1 mRNA level in sensitive patients was higher than in resistant patients and reached statistical significance (Figure 4a). Similarly, the hMSH2 mRNA expression displayed the same pattern as E2F1 and was consistent with previous findings. 10 . Immunohistochemistry also confirmed that higher E2F2 and hMSH2 protein levels were associated with platinum sensitivity (Figure 4b).

E2F1 levels in tissues from patients with ovarian cancer correlated with prognosis following platinum-based therapy. (a) mRNA levels of E2F1 and hMSH2 in platinum sensitive or resistant patients were statistically significantly different. (b) Immunohistochemistry confirmed that higher E2F2 and hMSH2 protein levels were associated with platinum sensitivity and (c) The progression-free survival (PFS) and overall survival (OS) curves of patients with high or low levels of E2F1. The median PFS of patients with E2F1high was 29 months, compared with 9.5 months for patients with E2F1low. Patients with E2F1high had a more favourable median survival (34 months) compared with patients with E2F1low (23 months). The differences between the two groups were statistically significant.
Finally, we studied the clinical significance of E2F1 in respect of patients’ prognosis following platinum-based therapies. Patients were categorized into high and low groups according to the E2F1 mRNA level, and their PFS and OS were determined. Substantial differences were observed in survival; the median PFS of patients with E2F1high was 29 months, compared with 9.5 months for patients with E2F1low (Figure 4c). Similarly, the OS profile also showed that patients with E2F1high had a much more favourable median survival (34 months) compared with patients with E2F1low (23 months). The differences between the two groups in PFS and OS were statistically significant (P = 0.0034 and P = 0.0032, respectively).
Discussion
Patients with advanced EOC have a 5-year survival rate below 40% due to late diagnosis and primary or acquired platinum resistance. 30 DNA repair deficiency may affect a patient's response to chemotherapy, and so is an important target for many chemotherapeutic agents. 7 Indeed, MMR dysfunction is recognised as a major factor in platinum resistance. 31 A previous study showed that hypermethylation of the MMR gene, hMSH2, was associated with platinum-resistance. 10 In addition to the epigenetic downregulation of hMSH2 expression, traditional regulatory mechanisms also play essential roles in controlling MMR genes.
Using bioinformatic analysis, we identified the transcription factor, E2F1, as having an association with hMSH2 in EOC tissues. Previous studies have also shown a regulatory role of E2F1 on MMR genes; for example, E2F1 has been shown to regulate MSH6 and MSH2 in rat cells. 32 In addition, the hMSH2 promoter was reported to respond to E2F1 in human lung carcinoma cells H1299. 13 Most work to-date, has focused on the importance of E2F1 in DNA replication and repair.15,16Furthermore, the understanding of drug resistance and role of E2F1 in cancer is limited. A recent study demonstrated that targeting the Yes-associated protein (YAP)-E2F1 DNA damage response pathway leads to an improvement in the sensitivity of EOC cells to a chemotherapeutic drug (WEE1 kinase inhibitor AZD1775). 33 These findings substantiate our hypothesis that E2F1 may have a role in regulating hMSH2 and platinum resistance in EOC.
To explore underlying mechanisms that regulates hMSH2 expression and drug susceptibility in EOC we investigated the connection between E2F1 and hMSH2 in tissues from patients with EOC. The regulatory function of E2F1 on the hMSH2 gene was confirmed in in vitro studies. Moreover, we found that expressional level regulation affected cisplatin susceptibility in EOC cells. Importantly, we believe that this is the first report showing a correlation between E2F1 and prognosis of platinum-based therapies. However, previous reports on the correlation between clinical prognosis and E2F1 in EOC have been contradictory. For example, one study proposed a positive association between favourable survival rate and increased activity of E2F1 in ovarian cancer. 34 Whereas, two other studies concluded that the overexpression of E2F1 was associated with an unfavourable outcome in patients with ovarian cancer.35,36 In addition, high malignancy (i.e., advanced histopathologic grade and high mitotic index) have been reported to be associated with an increased E2F1 expression in ovarian cancer. 37 Similar contradictory reports have been found in hepatocellular cancer. 38 Although one study in prostate cancer found E2Fs may act as promising biomarkers. 39 Limitations of this study include the lack of in vivo testing and the small sample size. A large controlled study is needed to verify our results.
In conclusion, we found an association between E2F1 and hMSH2 via bioinformatics and experiments such as knockdown, overexpression, and luciferase assay. To our knowledge, this is the first report of E2F1 regulated MSH2 expression playing a role in drug resistance of platinum-based treatments for patients with EOC. Our data suggested E2F1 may be a prognostic biomarker in EOC. Further research is required to confirm our findings.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231163780 - Supplemental material for hMSH2 coordinated with the expression of E2F1 promotes platinum response in epithelial ovarian cancer
Supplemental material, sj-pdf-1-imr-10.1177_03000605231163780 for hMSH2 coordinated with the expression of E2F1 promotes platinum response in epithelial ovarian cancer by Xiao-qian Hu, Bao-ying Zhang and Tian Hua in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605231163780 - Supplemental material for hMSH2 coordinated with the expression of E2F1 promotes platinum response in epithelial ovarian cancer
Supplemental material, sj-pdf-2-imr-10.1177_03000605231163780 for hMSH2 coordinated with the expression of E2F1 promotes platinum response in epithelial ovarian cancer by Xiao-qian Hu, Bao-ying Zhang and Tian Hua in Journal of International Medical Research
Footnotes
Acknowledgments
The authors greatly acknowledge Dr Miao Jie, Department of Pathology, Affiliated Xingtai People Hospital of Hebei Medical University for the assistance in collecting tissue samples.
Declaration of Conflicting Interests
The authors declare that there are no conflicts of interest.
Funding
This study was supported by the science and technology project of Xingtai City [grant number 2021ZC135].
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
