Abstract
Objective
To develop a nomogram that discriminates lung cancer from benign lung nodules through metabolic profiling.
Methods
This was a retrospective cohort study that recruited 848 participants who were randomized into training and validation sets at a 7:3 ratio. Clinical characteristics and metabolic profiles were retrieved. Variables in the training set with statistically significant differences were selected for further least absolute shrinkage and selection operator (LASSO) regression. The nomogram was built from 13 variables identified by stepwise regression analysis. Receiver operating characteristic, calibration curve, and decision curve analyses were conducted to evaluate the performance of the nomogram by internal validation.
Results
Thirteen variables were selected through LASSO regression to build the nomogram: age, sex, ornithine, tyrosine, glutamine, valine, serine, asparagine, arginine, methylmalonylcarnitine, tetradecenoylcarnitine, 3-hydroxyisovaleryl carnitine/2-methyl-3-hydroxybutyrylcarnitine, and hydroxybutyrylcarnitine. The nomogram had good discrimination for the training set, with an area under the curve of 0.836 (95% confidence interval: 0.830–0.890). Moreover, the calibration curve with 1000 bootstrap resamples showed that the predicted value coincided well with the actual value. Decision curve analysis described a net benefit superior to baseline within the threshold probability range of 15% to 93%.
Conclusions
The nomogram constructed from metabolic profiling accurately predicted risk of lung cancer.
Introduction
Lung cancer is one of the most common malignancies worldwide and has the highest morbidity and mortality. 1 Although there are many methods of diagnosing lung cancer, the early identification of malignancy from pulmonary nodules remains a great challenge for physicians. Chest computed tomography (CT) has been recommended as an efficient screening tool for lung cancer, which has dramatically improved detection rates.2,3 However, CT remains to have high false-positive rates. Misdiagnosing patients with benign nodules can cause adverse effects such as anxiety, tension, and unnecessary clinical examinations such as further enhanced CT and tissue biopsy. Although other diagnostic methods such as serum tumor biomarkers and bronchoscopy have increasingly been used for lung cancer screening, these also have shortcomings. Using blood samples to detect tumor biomarkers is relatively simple and convenient, but the low sensitivity, especially for the diagnosis of early-stage lung cancer has limited its clinical application.4,5 CT-guided transthoracic needle biopsy is considered the gold standard for diagnosis, nevertheless, it comes with certain risks such as pneumothorax, pulmonary embolism, and trauma. Therefore, the procedure is hard to accept for most patients. Overall, there is great need for a more accurate, feasible, and effective diagnostic model to distinguish patients with early-stage lung cancer from those with benign nodules.
Metabolic reprogramming is considered one of the hallmarks of cancer.6,7 In the 1920 s, Otto Warburg showed that cancer cells preferentially use glycolysis even in the absence of hypoxic conditions in the growth medium (the Warburg effect). 6 Additionally, amino acid synthesis and catabolism play essential roles in the metabolic changes of tumors. 7 Amino acids perform many essential functions such as redox homeostasis, energy regulation, biosynthesis, and the maintenance of homeostasis in the body (e.g., the urea cycle). This wide range of metabolic activities has made amino acid metabolism increasingly relevant for cancer research. 8 For example, the function of carnitine is to bring long-chain fatty acids into the mitochondria for β-oxidation, followed by the formation of acylcarnitine, an excess of which is then released into the bloodstream. 9 In lung cancer, the acylcarnitine profile can reflect the metabolic status of fatty acids, as the de novo synthesis of fatty acids is closely related to the proliferation and metastasis of tumor cells. 10 Accordingly, amino acids and carnitines are potential biomarkers for diagnosing cancer, as they serve essential roles in cell physiology as both basic metabolites and metabolic regulators.11–13
Recently, the emergence of metabolomics has enabled researchers to obtain a deeper understanding of many diseases, especially cancers. Metabolomics measures low molecular weight metabolites (<1000 amu) including amino acids, carnitines, fatty acids, and lipids. These and their downstream metabolites potentially participate in lung carcinogensis. 14 Thus, amino acid- and carnitine-targeted metabolomic profiling has been used to construct predictive models for lung cancer.15,16 However, there are many limitations to these studies. For example, the sample sizes have been too small and early-stage patients are rarely enrolled.15,17 Most importantly, using amino acid- and carnitine-targeted metabolomic profiling to discriminate lung cancer from patients with lung nodules has not been explored.
Nomograms are statistically-based tools that provide the overall probability of a certain result, and then combine multiple independent variables to assign proper weights to each variable on the basis of their predictive ability. Moreover, nomograms have been widely applied to estimate cancer diagnosis, risk of distant metastasis, and prognosis.18,19 In this study, we developed a diagnostic nomogram for lung cancer on the basis of amino acid- and carnitine-targeted metabolomic profiling by liquid chromatography-mass spectrometry (LC-MS/MS) in large-scale samples. The goal of this nomogram is for physicians to be able to predict early-stage lung cancer risk and assess whether pulmonary nodules are benign or malignant, thus maximizing benefits patients with malignancies and minimizing damage to those with benign lesions.
Materials and methods
Study population
We recruited participants who were screened for lung nodules by chest CT between April 2018 and December 2020 at the Second Affiliated Hospital of Dalian Medical University (Dalian, China). The patients enrolled in the study were selected randomly. We collected metabolic data before surgery. The subjects were randomly divided into training and validation sets at a 7:3 ratio. The inclusion criteria for patients with lung cancer were as follows: (1) patients with stage I/II lung cancer according to the 8th edition American Joint Committee on Cancer (AJCC) Tumor-Node-Metastasis (TNM) staging system; and (2) patients naïve to antineoplastic therapy, radiotherapy, or chemotherapy before surgery or cancer diagnosis. Owing to significant relationships between the profiles of circulating metabolites and certain diseases (autoimmune diseases; serious heart, liver, and kidney diseases; metabolic syndrome; other cancers),20–24 patients with the above conditions were excluded from this study.
This was a retrospective cohort study that was approved by the Ethics Committee of the Second Affiliated Hospital of Dalian Medical University (permit number: 2018-048). We have de-identified all patient details. All methods in the study were performed in accordance with the Declaration of Helsinki. Verbal informed consent was obtained from all participants prior to study participation, and the patients provided written informed consent for publication of this study. This study was designed and reported in accordance with the transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD) statement. 25
Measurement of tumor markers and metabolic profiling
Blood samples were collected from all participants after overnight fasting. For the amino acid- and carnitine-targeted metabolomic profiling, LC-MS/MS was used to test serum samples for 20 amino acids (alanine [Ala], arginine [Arg], asparagine [Asn], aspartate [Asp], citrulline [Cit], glutamine [Gln], glutamic acid [Glu], glycine [Gly], histidine [His], leucine [Leu], lysine [Lys], methionine [Met], ornithine [Orn], phenylalanine [Phe], proline [Pro], serine [Ser], threonine [Thr], tryptophan [Trp], tyrosine [Tyr], and valine [Val]) and 27 carnitines (carnitine [C0], acetylcarnitine [C2], propionylcarnitine [C3], butyrylcarnitine [C4], hydroxybutyrylcarnitine [C4OH], methylmalonylcarnitine [C4DC], isovalerylcarnitine/3-methylbutyrylcarnitine [C5], 3-hydroxyisovaleryl carnitine/2-methyl-3-hydroxybutyrylcarnitine [C5OH], glutarylcarnitine [C5DC], penenylcarnitine [C5:1], hexanoylcarnitine [C6], adipylcarnitine [C6DC], octanoylcarnitine [C8], decanoylcarnitine [C10], dodecanoylcarnitine [C12], tetradecanoylcarnitine [C14], hydroxytetradecanoylcarnitine [C14OH], myristylcarnitine [C14DC], tetradecenoylcarnitine [C14:1], hexadecanoylcarnitine [C16], hydroxyhexadecanoylcarnitine [C16OH], 3-hydroxypalmitoleoylcarnitine [C16:1OH], octadecanoylcarnitine [C18], eicosylcarnitine [C20], docosacarnitine [C22], twenty-four carbonyl carnitine [C24], and carnosylcarnitine [C26]), respectively. LC-MS/MS was performed on an API 3200 quadrupole mass spectrometer (Applied Biosystem, Waltham, MA, USA) equipped with an electrospray ionization (ESI) probe, Chemo View 1.4.2, and Agilent 1200 high-performance liquid chromatography (Agilent Technologies, Santa Clara, CA, USA). LC-MS/MS was performed at the Dalian Institute of Chemical Physics, Chinese Academy of Sciences within 48 hours of sampling.
Statistical analysis
Clinical indicators including sex, age, 20 amino acids, and 27 carnitines were collected and compared. Continuous variables are expressed as mean (interquartile range), while categorical variables are expressed using relative frequencies and proportions. The Mann–Whitney U test was used to compare continuous variables, and the chi-square test was used to compare categorical variables.
Variables with statistically significant differences were selected for further logistic least absolute shrinkage and selection operator (LASSO) regression. LASSO regression was performed to identify relevant variables and avoid overfitting the models based on the 30 variables using the ‘glmnet’ package of R software (www.r-project.org). To fit the model, the penalty term λ (i.e., the minimum binomial deviance) was used to select variables for the model. A logistic regression analysis was then performed. The nomogram was developed upon logistic regression model coefficients using the ‘rms’ package of R. A receiver operating characteristic (ROC) curve was constructed from the model. Areas under the curve (AUC) and 95% confidence interval (CIs) of the nomogram were calculated. The predictive performance of the nomogram was measured by the concordance index and calibrated with 1000 bootstrap resamples to decrease overfit bias. We also evaluated AUC results of each single index in the nomogram and compared with the final model to demonstrate the superiority of combining the indexes. Furthermore, calibration curves were plotted separately for the training and validation sets using 1000 bootstrap resampling. Finally, decision curve analysis (DCA) was used to assess net clinical benefit. Statistical analyses were performed using SPSS 23.0 software (IBM Corp., Armonk, NY, USA) and R 4.0.3 software (www.r-project.org). All tests were designated to have statistical significance at P < 0.05.
Results
Participant characteristics
In total, we recruited 848 participants who had been screened for lung nodules by chest CT. Among the enrolled cases, there were 478 lung cancer patients and 370 subjects with benign nodules. The cases were then divided at a 7:3 ratio into the training and validation sets. In the training set, 334 cases were diagnosed with lung cancer by biopsy. Clinical features of lung cancer patients in the training and validation sets are presented in Supplementary Table 1. Among the lung cancer patients, 125 were male and 209 were female. Lung cancer patients tended to be older than subjects with benign lesions (median age at diagnosis: 62.50 years vs. 56.00 years, respectively, P < 0.001). In the validation set, the median age of the lung cancer patients was 64.00 years, and 42.36% were male. Characteristics of amino acid- and carnitine-targeted metabolomic profiling are listed in Table 1.
Demographic information and metabolic profiling of subjects in training and validation sets.

NA, not applicable.
Nomogram development
A comparison of clinical indicators between the 334 early-stage lung cancer patients and 260 subjects with benign pulmonary nodular lesions identified 30 variables with statistically significant differences between the groups including age, sex, Orn, Thr, Asp, Tyr, Cit, Glu, Val, Pro, Ser, Asn, Trp, Gln, Met, Arg, C4OH, C4DC, C5DC, C5OH, C5:1, C6DC, C8, C12, C14, C14:1, C16, C22, C24, and C26.
Next, LASSO regression was performed to identify relevant variables and avoid overfitting the models based on the 30 variables using the “glmnet” package of R software. To fit the model, the penalty term λ (see Methods) was used to select the variables for the model. In total, 25 variables were screened out including age, sex, Ser, Arg, Pro, Glu, Val, Thr, Tyr, Asn, Met, Gln, Orn, C4OH, C4DC, C5OH, C5DC, C5:1, C8, C12, C14:1, C16, C22, C24, and C26 (λ = 0.005612, M = 0.9985, SE = 0.01982) (Figure 1).

Cross validation plot for the penalty term.
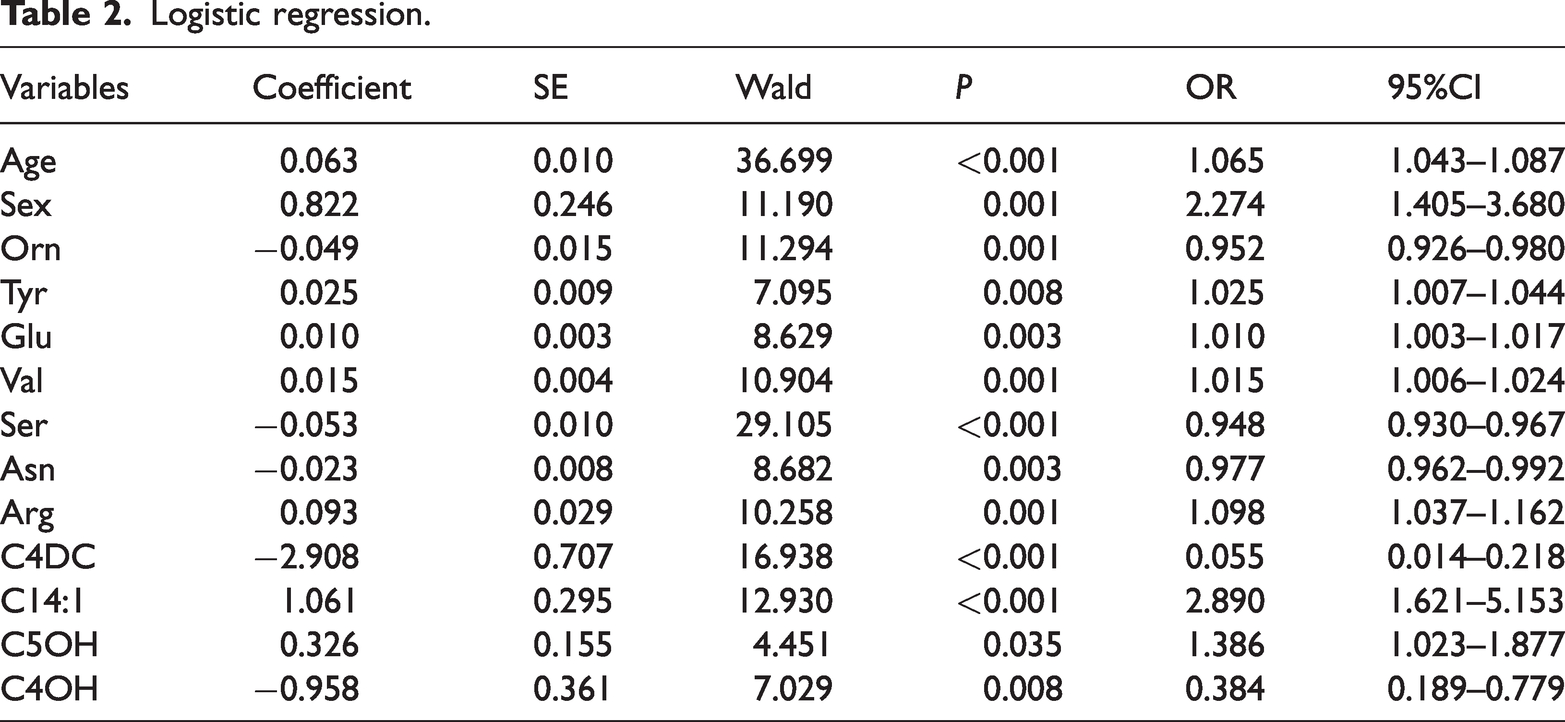
Finally, 13 independent predictive factors for early-stage lung cancer were found by multivariate logistic regression: age (odds ratio [OR]: 1.065, 95%CI: 1.043–1.087, P < 0.001), sex (OR: 2.274, 95%CI: 1.405–3.680, P = 0.001), Orn (OR: 0.952, 95%CI: 0.926–0.980, P = 0.001), Tyr (OR: 1.025, 95%CI: 1.007–1.044, P = 0.008), Glu (OR: 1.010, 95%CI: 1.003–1.017, P = 0.003), Val (OR: 1.015, 95%CI: 1.006–1.024, P = 0.001), Ser (OR: 0.948, 95%CI: 0.930–0.967, P < 0.001), Asn (OR: 0.977, 95%CI: 0.962–0.992, P = 0.003), Arg (OR: 1.098, 95%CI: 1.037–1.162, P = 0.001), C4DC (OR: 0.055, 95%CI: 0.014–0.218, P < 0.001), C14:1 (OR: 2.890, 95%CI: 1.621–5.153, P < 0.001), C5OH (OR: 1.386, 95%CI: 1.023–1.877, P = 0.035), and C4OH (OR: 0.384, 95%CI: 0.189–0.779, P = 0.008) (Table 2). A nomogram containing these 13 variables was established in accordance with coefficients of the regression model (Figure 2). Each variable was assigned a score ranging from 0 to 100 points. By calculating the total scores of various covariates and placing the total score, the probability of lung cancer could be efficiently estimated.
Logistic regression.

The nomogram for predicting the risk of lung cancer.
Performance of the nomogram
We evaluated the performance of the nomogram from three aspects. First, we plotted ROC curves and calculated AUC values to assess discrimination ability. Using the training set, the nomogram showed an AUC of 0.836 (95%CI: 0.830–0.890, P < 0.05) and of 0.860 after 1000 bootstrap resamples (Figure 3a). The AUC of the nomogram in the validation set was 0.781 (95%CI: 0.722–0.841, P < 0.05) and remained 0.781 after 1000 bootstrap resamples (Figure 3b). We also evaluated AUC results of each individual index of the nomogram including age (AUC: 0.667), sex (AUC: 0.561), Orn (AUC: 0.731), Tyr (AUC: 0.663), Glu (AUC: 0.644), Val (AUC: 0.631), Ser (AUC: 0.616), Asn (AUC: 0.580), Arg (AUC: 0.552), C4DC (AUC: 0.690), C14:1 (AUC: 0.562), C5OH (AUC: 0.607), and C4OH (AUC: 0.674) (Supplementary Table 2). Compared with each individual index, the nomogram demonstrated the superiority of combining these 13 indexes.

Receiver operating characteristic (ROC) curve for the nomogram (a) in the training set and (b) in the validation set.
Ni. et al. previously used metabolomics to build a model for screening lung cancer patients that included Gly, Val, Met, Cit, Arg, and C16-carnitine. 15 Next, we performed calibration curves with 1000 bootstrap resamples, which showed that the predicted value closely coincided with the actual value (Figure 4). Moreover, DCA reported a net benefit superior to baseline within the threshold probability range of 15% to 93% (Figure 5). These results confirmed that the nomogram had good performance for screening early-stage lung cancer.

Calibration plot for the nomogram (a) in the training set and (b) in the validation set.

The decision curve analysis (DCA) curve for the nomogram.
Discussion
In this study, we developed a novel nomogram to diagnose early-stage lung cancer from benign lung nodules. The nomogram, which includes two clinical factors and 11 metabolic factors, showed superior predictive power.
Liquid biopsy has become an emerging tool for tumor management, and among serum markers, tremendous progress has been made with regard to circulating tumor DNA (ctDNA). Liquid biopsy not only plays an important role in tumor diagnosis but also in assessing tumor prognosis, monitoring treatment efficacy and resistance, and treatment selection. 26 Additionally, there are many commercial kits that are currently in use. 26 Metabolic reprogramming is considered one of the hallmarks of cancer. 7 In recent years, metabolic fingerprints have been used as early diagnostic markers of tumors, including for lung adenocarcinoma, 27 colorectal cancer, 28 breast cancer, 29 and early-stage gastric cancer. 30 The emergence of advanced tools such as metal-organic framework hybrids, 28 mass spectrometry, 29 plasmonic alloys, 30 and bionanoparticles 31 have greatly accelerated the use of metabolic fingerprints. Various models for cancer management that use metabolic fingerprinting have also shown excellent diagnostic efficacy.26–31 It is believed that the advancement of analytical techniques will also increase screening for early-stage cancers.
Several studies, such as the Mayo Clinic model 32 and the Pan-Canadian Early Detection of Lung Cancer Study (PanCan) risk prediction model 33 have explored models to discriminate early-stage lung cancer from pulmonary nodules on the basis of imaging features. However, these models have some limitations. The Mayo Clinic model underestimates the risk of lung nodules 34 and has a bias towards solid-predominant nodules. 35 The PanCan risk prediction model was established from PanCan data, and its efficacy has not been ideal through external validation.33,36 Therefore, the PanCan risk prediction model requires further study before it can be applied in the clinic. As mentioned above, both of these models used radiological features, which can involve some speculation. Unfortunately, different imaging physicians may have different observations, and the deviation caused by different imaging physicians can reduce the accuracy of these models. Therefore, a risk model established using objective laboratory-based factors may be more reliable. Notably, previous studies have demonstrated that metabolites are relatively strong objective predictors of lung cancer.15,37 In early-stage tumors, due to increased anabolism, synthetic raw materials such as amino acids and carnitines have been to be greatly altered compared with healthy tissue. Thus, we applied changes in amino acids and carnitines to construct a risk nomogram model for lung cancer screening.
Our model included four carnitines: C4DC, C4OH, C5OH, and C14:1. The function of carnitines is to bring long-chain fatty acids into the mitochondria for β-oxidation, followed by the formation of acylcarnitine, excess of which is subsequently released into the bloodstream. 9 In lung cancer, the de novo synthesis of fatty acids is closely related to the proliferation and metastasis of cancer cells. 10 Therefore, the acylcarnitine profile in lung cancer can reflect the metabolic status of fatty acids. In this study, C4DC, C4OH, C5OH, and C14:1 were included in a prediction model for early-stage lung cancer for the first time.
Our model also included seven amino acids (Orn, Asp, Glu, Val, Ser, Tyr, and Arg) that have been shown to be closely related to biological functions that are active during carcinogenesis. Orn is a non-essential amino acid that is produced as an intermediate molecule in the urea cycle and is a key substrate for the synthesis of Pro, polyamines, and Cit, which have significant regulatory roles in cancer. 38 Kim et al. found that Orn was significantly elevated in lung cancer by comparing plasma free amino acid levels between lung cancer and healthy subjects, 39 which is consistent with our findings. Asp is formed from aspartic acid as a substrate, using the amino group produced by transamination during Gln formation to Glu, which is bound to ATP by the action of asparagine synthetase. In a mouse model of breast cancer, lowering Asp levels inhibited tumor metastasis. 40 In acute lymphoblastic leukemia, tumor cells cannot synthesize Asp themselves and need exogenous supplementation, but normal cells can synthesize Asp. Therefore, if Asp, an essential substrate for protein synthesis in tumor cells, is depleted, it can inhibit tumor growth without affecting normal cells. 41 Glu is an acidic amino acid that is secreted by tumor cells and is one of the causes of the acidic tumor microenvironment, which provides space for tumor invasion and inhibits T lymphocyte immunity. In breast cancer, normal cells do not readily secrete Glu, whereas invasive tumor cells do. 42 Glu metabolism is an important factor in tumor growth and invasion, and in tumor cells, increased Glu secretion and dysregulated Glu metabolic signaling are thought to be possibly associated with a malignant tumor phenotype.43,44 Ni et al. found that glutamate levels were decreased in lung cancer patients compared with in normal subjects. 15 This is contrary to our results, probably because our study population was primarily patients with early-stage lung cancer. Moreover, the role of Glu in lung cancer needs to be further explored. Val is one of the branched-chain amino acids and can activate various signaling pathways and participate in tumor growth and metastasis. Additionally, Val can also serve as a nitrogen donor to produce macromolecules such as nucleotides, which are involved in tumor growth. 45 Ser is also a non-essential amino acid that supports a variety of metabolic processes for cell growth and survival, including the synthesis of proteins, amino acids, and glutathione. As an important single-carbon donor in the folate cycle, Ser is involved in nucleotide synthesis, methylation reactions, and NADPH production for antioxidant defense.46,47 Many rapidly proliferating cells are dependent on exogenous Ser. Both in vivo and ex vivo, Ser deficiency significantly inhibits the growth of some tumor cells. 48 Tyr is an important amino acid for protein synthesis and an important nutrition substance for some tumor cells, with inhibition of Tyr uptake inhibiting growth via nutrient depletion in these cells. 49 Arg is a semi-essential amino acid that can serve as an important building block for protein synthesis and as a precursor for a variety of metabolites, including polyamines and nitric oxide, which have significant immunomodulatory effects in cancer.50,51 Furthermore, elevated Arg levels in tumor cells induce strong metabolic changes including a shift from glycolysis to oxidative phosphorylation in activated T cells; Arg also promotes the generation of central memory-like cells with longer viability and, in a mouse model, higher antitumor activity. 52 Thus, intracellular Arg concentrations directly interfere with T cell survival and metabolic functions, which are essential for antitumor responses. 53
To the best of our knowledge, this study represents the first diagnostic nomogram established on the basis of amino acid- and carnitine-targeted metabolomic profiling and without subjective factors such as imaging features. Our study had significant innovations in three aspects. First, we focused on new molecular markers to screen early-stage lung cancer among patients with lung nodules by metabolomic profiling. Second, a visual model was constructed with R language for clinical practice (i.e., discriminating benign/malignant lung nodules), and our application of the mathematical model to estimate lung nodules showed good performance. Third, the nomogram was based on data from a small amount of peripheral blood detected by LC-MS/MS. Together, these suggest similar methods could also provide the basis for establishing diagnostic models in other neoplasms.
Despite these advancements, this study also had several limitations. First, differences in pathological types of lung cancer were not discussed, and the candidate indicators included in the study were limited, which may omit important indicators such as nodule size, nodule morphology, and smoking history, which may influence the accuracy of the model. Second, external verification has not been performed, and further verification and optimization are needed in the future, such as large-sample multi-center studies. Third, the sensitivity and specificity of the nomogram were not ideal. Nevertheless, it provides new insights into the features of amino acid and acylcarnitine profiles that could contribute to the distinct separation between lung cancer and benign lung nodules.
In summary, this study focused on diagnosing the early-stage lung cancer by using metabolic profiling. The nomogram we developed included age, sex, and amino acid and carnitine profiles and exhibited a strong ability to distinguish patients with lung cancer from controls. The included metabolites may be important functional substances involved in the pathogenesis of lung cancer. This nomogram could become a novel screening tool for lung cancer.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231161204 - Supplemental material for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules
Supplemental material, sj-pdf-1-imr-10.1177_03000605231161204 for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules by Chenwei Li, Zhuo Chen, Hui Zhao, Cuicui Wang, Shujun Yu, Hengde Ma, Qi Wang and Xiaohui Du in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605231161204 - Supplemental material for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules
Supplemental material, sj-pdf-2-imr-10.1177_03000605231161204 for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules by Chenwei Li, Zhuo Chen, Hui Zhao, Cuicui Wang, Shujun Yu, Hengde Ma, Qi Wang and Xiaohui Du in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605231161204 - Supplemental material for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules
Supplemental material, sj-pdf-3-imr-10.1177_03000605231161204 for A nomogram based on metabolic profiling to discriminate lung cancer among patients with lung nodules by Chenwei Li, Zhuo Chen, Hui Zhao, Cuicui Wang, Shujun Yu, Hengde Ma, Qi Wang and Xiaohui Du in Journal of International Medical Research
Footnotes
Acknowledgements
We thank all the patients who kindly donated samples and the staff at the Second Hospital of Dalian Medical University. We acknowledge the supports of Dalian Institute of Chemical Physics, Chinese Academy of Sciences.
Author contributions
Chenwei Li and Hui Zhao conceived and designed the study. Chenwei Li, Zhuo Chen, and Cuicui Wang collected the data. Chenwei Li, Hengde Ma, Shujun Yu, and Qi Wang analyzed the data. Chenwei Li and Zhuo Chen drafted the manuscript and figures. Hui Zhao, Xiaohui Du, and Qi Wang edited and revised the manuscript. All authors contributed to the article and approved the submitted version.
Data availability statement
All data generated and analyzed during this study are included in the supplementary files.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (81903185 and 81703087), the Talent Innovation Support Plan of Dalian (2021RQ008), the United Fund of the Second Hospital of Dalian Medical University and Dalian Institute of Chemical Physics, Chinese Academy of Sciences (UF-ZD-202011), the Project of Education Department of Liaoning Province (LZ2020009), and the Science and Technology Innovation Fund Project of Dalian (2021JJ13SN67).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
