Abstract
Objective
This cross-sectional study investigated the circulating strains of rotavirus and screened for noravirus in Ibadan, Nigeria as the country introduces the rotavirus vaccine into its national immunization program.
Methods
Sixty-five stool samples were collected from children younger than 5 years with clinically diagnosed diarrhea and screened for the presence of rotavirus and norovirus using RT-PCR. Rotavirus-positive samples were further analyzed to determine the G and P genotypes using semi-nested multiplex PCR.
Results
The rates of rotavirus and norovirus positivity were 30.8% and 10.8%, respectively, whereas the rate of rotavirus and norovirus mixed infection was 4.6%. G1 was the predominant VP7 genotype, followed by G2, G9, and G1G2G9, whereas the predominant VP4 genotype was P[4], followed by P[6], P[8], and P[9]. The mixed P types P[4]P[8] and P[4]P[6] were also detected. G1P[4] was the most common VP4 and VP7 combination, followed by G2P[4], G1[P6], G1P[8], G2P[6], G2P[9], G9P[6], G2G9P[4], G2P[4]P[6], G1P[4]P[8], G2G9P[8], G1G2G9P[8], and G1[non-typable] P[non-typable], which were detected in at least 5% of the samples. Four samples had a combination of non-typable G and P types.
Conclusions
It is essential to monitor the circulation of virus strains prior to and during the implementation of the immunization program.
Introduction
Rotavirus and norovirus remain the two most important etiologies of acute viral gastroenteritis in children younger than 5 years, resulting in severe morbidity and mortality in the developing world. 1 Globally, approximately 128,500 mortalities in children were associated with rotavirus infection in 2016. 2 Asia and sub-Saharan Africa account for approximately 80% of all rotavirus deaths globally.2–5 Despite the sharp decrease in rotavirus-associated mortalities from 450,000 deaths per year at the beginning of the millennium to 128,500 deaths in 2016, rotavirus remains the leading cause of acute viral infantile diarrhea globally.2,6 Norovirus is the second most important etiology of viral gastroenteritis in children, accounting for 12% of all cases of sporadic acute gastroenteritis in children younger than 5 years.7,8 Globally, an estimated 200 million cases of diarrhea and 50,000 deaths in children younger than 5 years are caused by norovirus infection each year, mostly occurring in developing countries.9,10 However, norovirus is well established as the leading cause of viral gastroenteritis across all ages globally. 8
Rotavirus belongs to the family Reoviridae. The virion is a triple-layered particle consisting of six structural proteins encoded by 11 segments of double-stranded RNA that can be separated on a polyacrylamide gel. 11 The outer layer consists of two neutralizing antigens, namely a protease-sensitive P type antigen (VP4) and glycoprotein G type antigen (VP7). The antigenic and molecular properties of these surface proteins have been used to classify rotavirus strains. 11
In Nigeria, studies have demonstrated that approximately 90% of all children are infected with rotavirus by 2 years of age.12,13 In addition, rotavirus is associated with an estimated 50 000 deaths annually in Nigeria, which has the second highest dead rate attributable to rotavirus globally. 14 At present, several rotavirus vaccines have been licensed, including Rotarix, a P[8]G1 monovalent human rotavirus vaccine, and RotaTeq, a [P5]G1–4, G9 human-bovine pentavalent vaccine.15–17 Rotarix is commercially available in Nigeria, but the country has not introduced the vaccine to its National Expanded Immunization Program.12,18–20 The introduction of the vaccine is still pending as at late 2020, although the vaccine alliance GAVI approved Nigeria’s application for support in early 2020.14,21
Noroviruses are single-stranded, positive-sense RNA viruses of the family Caliciviridae and genus Norovirus. 7 They are classified into genogroups, genotypes, and strains/variants. 22 Group two (GII) norovirus is the most frequently detected type in humans, with most strains belonging to the GII.4 genotype. 23 However, norovirus GII.17 is rapidly emerging as the most dominant strain in some parts of the world. 24 Osazuwa and Grobler 25 reported the presence of norovirus GII.17 Kawasaki among Nigerian children younger than 5 with diarrhea. Although the molecular epidemiology of these viruses in developed countries has been extensively described, the exact role of noroviruses in acute gastroenteritis in children in sub-Saharan Africa is unclear. 26
Some studies reported the molecular epidemiology of rotavirus in Nigeria.1,12,13,27,28 However, it is important to update our knowledge on rotavirus by conducting surveillance to determine the emergence of new strains or unusual genotypes, which can arise from the reassortment of co-circulating strains of different genotypes, prior to the official introduction of the vaccine into the country’s national immunization program.
Materials and methods
Study population and sample collection
For this cross-sectional study, 65 samples were randomly collected from children younger than 5 years presenting with acute diarrhea (defined as the passage of loose, watery, or blood-tinged stool three or more times within a 24-hour period) at Oni and Sons Memorial Hospital (Ibadan, Nigeria) and Adeoyo General Hospital (Ibadan, Nigeria) over a period of 7 months (April–November 2017). A structured questionnaire was used to obtain the basic demographic data, medical histories, and clinical information of the children from their parents/caregivers, who provided consent for the use of these data. The samples were transported on ice to the Department of Virology, University College Ibadan, University of Ibadan (Ibadan, Nigeria), where they were stored at −20°C until screened for rotavirus and norovirus using commercially available molecular detection kits. Before RNA extraction, samples were processed by thawing at room temperature. A small amount of each stool sample was then dispersed into a 10 mL centrifuge tube containing 5 mL of sterile normal saline. The emulsified stool sample was then centrifuged at 2000 × g for 10 minutes, and the supernatant was decanted into a fresh centrifuge tube for RNA extraction.
RNA extraction and cDNA synthesis
A suspension of 10% (v/v) phosphate-buffered saline (pH 7.0) was prepared for RNA extraction. Viral RNA was extracted from stool suspensions of samples using a total RNA purification kit (Jena Bioscience® GmbH, Frankfurt, Germany). Extracted RNA was reverse-transcribed into cDNA using a SCRIPT cDNA synthesis kit (Jena Bioscience® GmbH) according to the manufacturer’s instructions.
Rotavirus detection and VP4 and VP7 genotyping by semi-nested multiplex PCR
Rotavirus detection was performed using the VP4F and VP4R primers, whereas the VP7F and VP7R primers were used for VP7 amplification. The methods described by Simmonds et al. 29 and Rodriguez-Daiz et al. 30 were adopted for VP4 (P[4], P[6], P[8], P[9], P[10]) and VP7 (G1–G4, G8–G10) genotyping, respectively. For product amplification, 4 µL of cDNA were added to a 21 µL reaction mixture consisting of 5 µL of Red Load Taq Master/high yield mix (Jena Bioscience® GmbH), 0.2 µM each of the first-round primers (Table 1), and 14 µL of nuclease-free water. The set-up for the second round of amplification consisted of 1 µL of each of the genotype specific primers and the reverse primer of the first-round reaction with the same volume of the Red Load PCR Mix. The resulting amplicons from second-round PCR were visualized by agarose gel electrophoresis using an ultraviolet gel documentation system (Roche, Basel, Switzerland). The gel concentration was set to 2% agarose, and the gel was run in a 1 × dilution of Tris borate EDTA buffer in a gel tank at 120 V for 30 minutes. Table 1 presents the primer positions and sequences. Cycling conditions were adopted from the protocol of Simmonds et al., 29 Rodriguez-Diaz et al., 30 and Motayo et al. 31 The control set-up for second-round PCR included nuclease-free water for the negative (no template) control and PCR-positive archived samples with the GenBank accession numbers (genotype) KU866451 (G1), KU866454 (G3), MN781164 (G8), KY964455 (P[4]), and MN781170 (P[8]). All samples were tested in duplicate before genotypes were assigned.
Primers used to screen and genotype rotavirus and norovirus from stool samples.
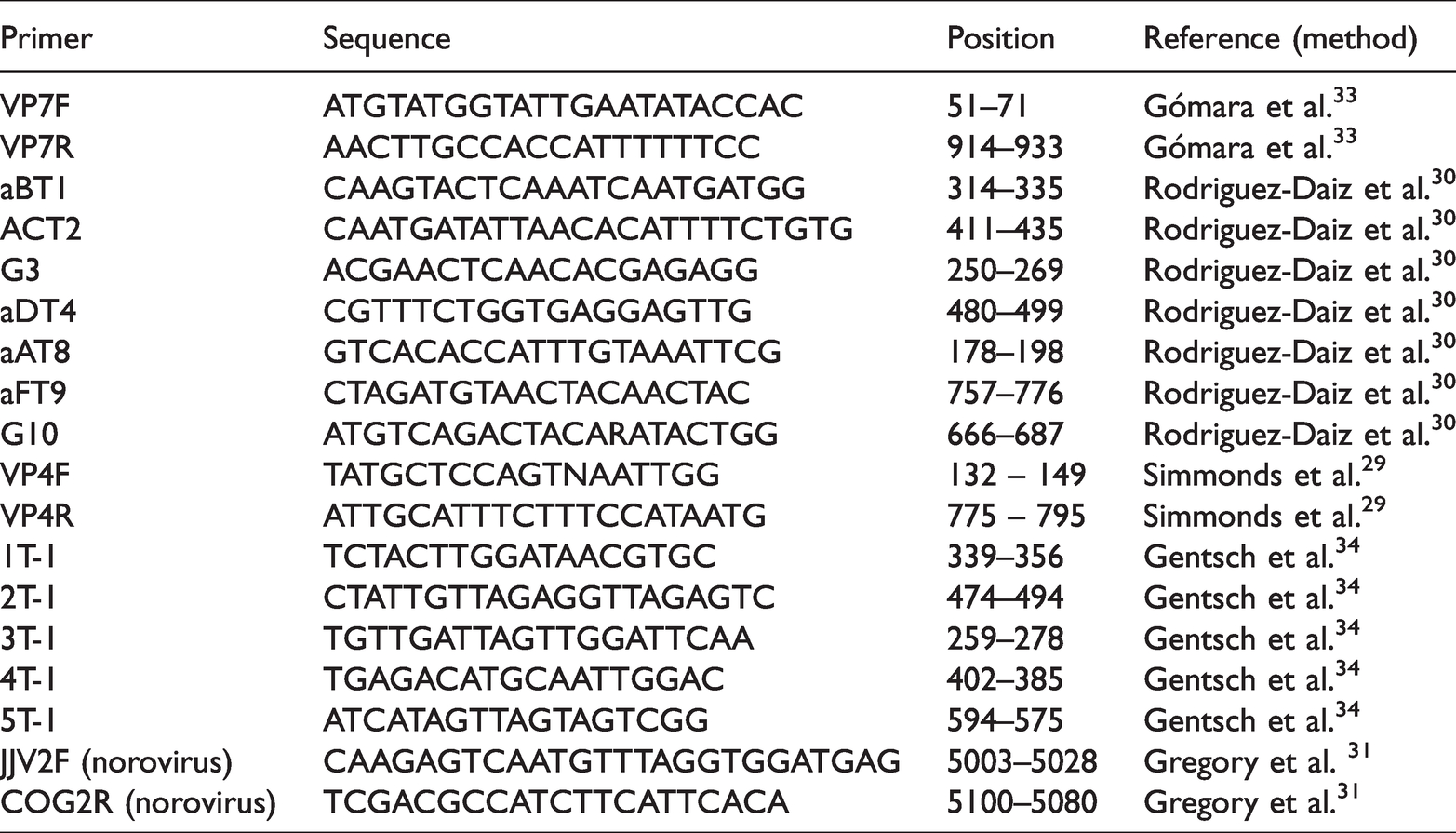
Norovirus detection
The methods described by Gregory et al. 31 were adopted for endpoint PCR detection of norovirus using the JJV2F and COG2R primers. The cycling protocol was 95°C for 5 minutes and 42 cycles of 95°C for 30 s, 55°C for 45 s, 72°C for 1 minute, and 70°C for 5 minutes.
Ethical clearance
Ethical approval for this study was obtained from the Department of Planning, Research and Statistics Division, Ministry of Health, Oyo state, Nigeria (reference number: AD 13/479/578, approval date: 13 February 2014). Verbal informed consent was given by the parents/caregivers of the children before enrolment into the study. Participants’ data were de-identified.
Statistical analysis
Data from the questionnaires administered and others generated in the study were analyzed using descriptive statistics with SPSS version 20.0 (IBM, Armonk, NY, USA). The reporting in this study conforms to the STROBE guidelines. 32
Results
High prevalence of rotavirus in children younger than 20 months
Study samples were obtained from 36 boys (55.4%) and 29 girls (44.65%). The rates of rotavirus and norovirus infection were 30.8% and 10.8%, respectively. The prevalence of rotavirus and norovirus was high in children younger than 24 months (Fig 1) and 25 months (Table 2), respectively. Rotavirus and norovirus mixed infection was observed in three samples (4.6%).
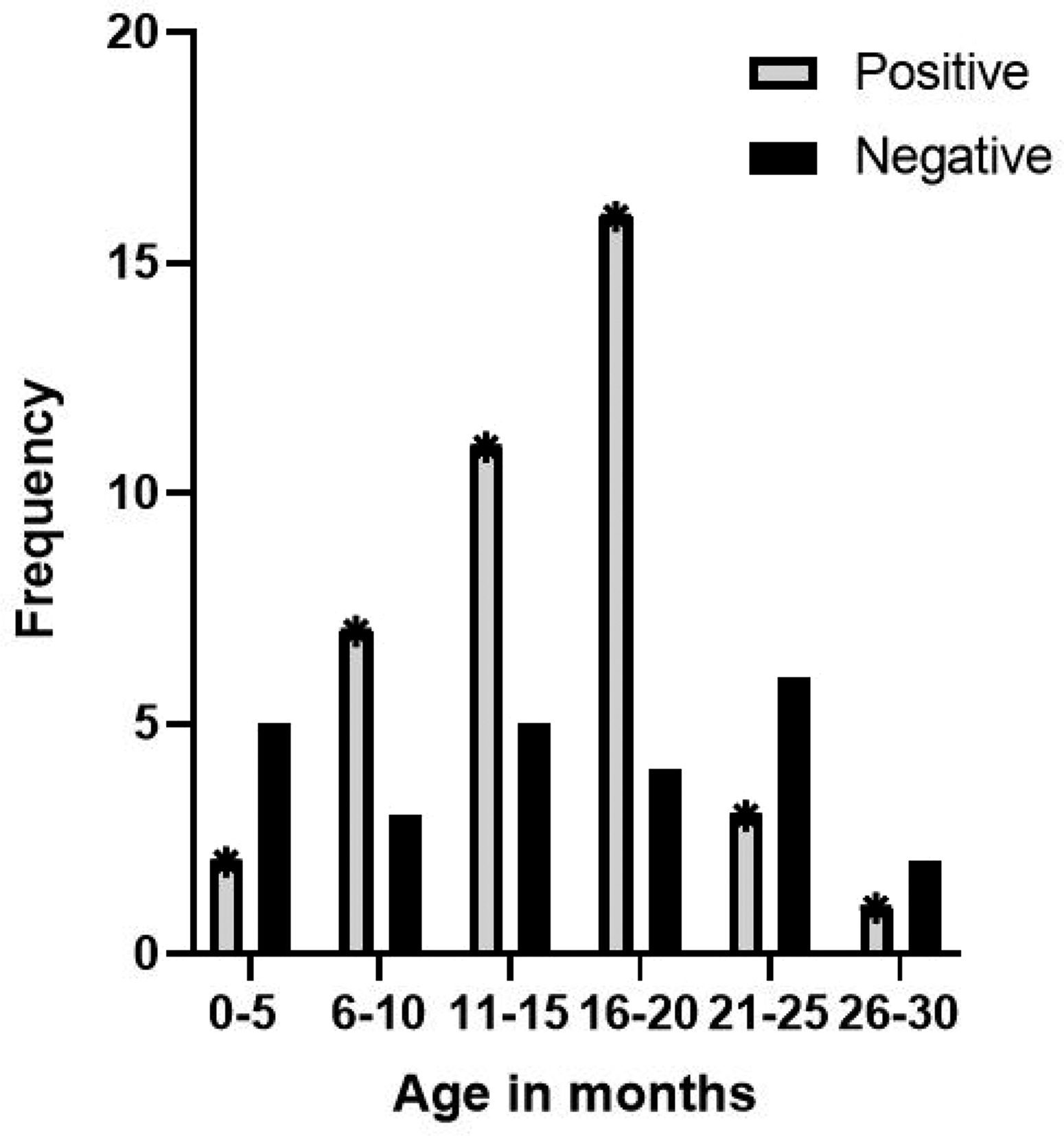
Age distribution of children with rotavirus infection in Ibadan, Nigeria.
Distribution of norovirus infection by age.

VP7 (G) genotyping and distribution
VP7 (G) genotypes were assigned to 80% of the rotavirus-positive samples, whereas 20% were non-typable. Of the seven VP7 serotypes subjected to multiplex PCR, three serotypes were detected including G9. Genotype G1 was most dominant (30%), closely followed by G2 (25%) and G9 (5%). Mixed infection with two or three VP7 genotypes was also detected. G1G2 mixed infection was predominant (10%), whereas G1G2G9 was detected in 5% of the tested samples. Mixed infection with G1 and a non-typable strain was also detected in 5% of samples, whereas 20% of the samples were non-typable.
VP4 (P) genotyping and distribution
Regarding VP4 (P) genotyping, 55% of the 20 rotavirus-positive samples were assigned individual P types, whereas 10% had mixed P types. Analysis of the P type illustrated that P[4] was predominant (30%), followed by P[6] and P[8] (15% each), whereas P[9] was detected in 5% of the samples. Mixed P type infection was observed with P[4]P[8] and P[4]P[6] being present in 5% of samples each. Five of the samples could not be assigned a P genotype.
VP4 and VP7 genotype combination
Analysis of combined P and G types revealed that G1P[4] was the most common combination (15%), whereas G2P[4] was detected in 10% of the tested samples. In addition, G1[P6], G1P[8], G2P[6], G2P[9], G9P[6], G2G9P[4], G2P[4]P[6], G1P[4]P[8], G2G9P[8], G1G2G9P[8], and G1NTP[NT] were each present in 5% of the samples. Four of the genotyped samples had a combination of non-typable G and P types (Table 3).
Rotavirus mixed G and P genotypes circulating among children with diarrhea in Ibadan, Nigeria.

Discussion
Vaccines are unarguably the superior prophylaxis for viral diseases. Over time, vaccines have proven effective for both preventing and eradicating these diseases. However, to assess vaccine efficacy, it is critical to continually monitor the emergence of unusual strains resulting from reassortment. Although rotavirus vaccines are available and they caused a sharp reduction in rotavirus-associated childhood mortalities,35–37 the virus remains an important cause of severe childhood acute gastroenteritis in countries that have not included the vaccine in their national routine vaccination programs.38–40 As of July 2021, Nigeria is one of nine sub-Saharan African countries that have yet to introduce the rotavirus vaccine in their national immunization programs 41 even though the vaccine is commercially available for people who can afford it.
In recent times, Nigeria has witnessed the emergence of unusual rotavirus genotypes, including recombinants from different species, providing evidence of interspecies transmission, 42 as well as mixed rotavirus and norovirus infections, which have been implicated in the development of severe childhood gastroenteritis. 1 Therefore, as an update to our current knowledge on the circulating strains of rotavirus in Nigeria, this study revealed a rotavirus positivity rate of 30% in Ibadan, similar to the rate of 34.5% reported by Japhet et al. 1 in Ile-Ife. Our study focused on children who presented to the hospital for gastrointestinal illness, which significantly limited the sample size, with only 65 stool samples collected during the study period. However, Omatola et al. 43 previously reported a rotavirus prevalence of 18.5% in Ibadan, and the different rates might be related to differences in the sensitivity of the different diagnostic approaches used. The present study also recorded a GII norovirus positivity rate of 10.8% using stool samples obtained from children younger than 5 years, whereas the rate of rotavirus and norovirus co-infection was 4.6%. These data are consistent with previous findings by Babalola et al., 44 who reported a norovirus prevalence of 8% in Ibadan. However, Japhet et al. 45 reported norovirus positivity rates of 23.6% and 5.1% in Osun and Ogun states, respectively. Rotavirus and norovirus co-infection was observed in 4.6% of the samples screened. This is important because it is suggested that co-infection by rotavirus and other enteric viruses results in gut competition, which could result in reduced vaccine efficacy. This might be one of the factors responsible for the weaker and waning efficacy of the rotavirus vaccine in low-income countries, as similarly observed with the oral polio vaccine in resource-limited settings.46,47
This study identified G1 as the most dominant circulating VP7 genotype, in line with previous findings.12,13,18,48,49 However, studies by Lekana-Doukiet al., 50 Boni-Cisseet al., 51 Mokomane et al., 52 and Simwaka et al. 53 reported G6, G12, G9, and G2 as the predominant VP7 genotypes circulating in Gabon, Côte D’Ivoire, Botswana, and Zambia respectively. This highlights the diversity in the predominant circulating genotypes in different parts of Africa, which has diverse implications for determining which available vaccines will be most effective in different areas. Genotype G2 was detected as the second most important VP7 genotype among children in Ibadan. Similarly, G2 was reported as the second most dominant genotype among children in northwestern Nigeria 54 and in sewage. 48 Worthy of note is the detection of the G9 genotype, which has gained epidemiological and clinical relevance globally. Similarly as reported in previous studies, different mixed VP7 genotypes and untypable strains were also identified in this study. Adah et al. 13 reported the predominance of G1G2 mixed infection; however, a mosaic rotavirus infection by G2G9 was reported in this study. In addition, the combination of G1, G2, and G9 was present in a single infection. Similarly, the combination of G1, G2, G8, and G9 in a single infection was previously reported in Nigeria. 12 However, it is important to further study these mixed infections, although there is evidence that this is a common feature of rotavirus infection in Nigeria. Analysis of the VP4 (P) genotypes revealed the predominance of the P[4] genotype. This is similar to the findings of other studies.19,55–58 The next most prevalent genotypes were P[6] and P[8]; however, Adah et al., 13 Aminu et al., 12 and Ayolabi et al. 18 previously identified P[6] as the predominant P type from samples collected in Maiduguri, Lagos, and Northwestern Nigeria, respectively. P[9] was also detected, whereas P[4]P[6] and P[4]P[8] mixed genotypes were also present. Interestingly, these P combinations were also reported in a previous study in Lagos. 18
Globally, G1P[8], G2P[4], G3P[8], G4P[8], G9P[8], and G12P[8] are the six most common rotavirus VP7 (G) and VP4 (P) combinations associated with human infections.59–62 However, the evolution of the virus resulting from the accumulation of point mutations, reassortment and recombination events, and interspecies transmission has resulted in a new and diverse panel of unusual G and P combinations. In this study, the predominant G and P combination was G1P[4], which was not consistent with the globally dominant G/P combinations. Motayo et al. 19 reported G1P[4] as the common genotype in sewage isolates in Nigeria. In addition, a previous study in Detroit, MI reported G1P[4] as the dominant circulating strain in children with severe diarrhea, 55 and a review by Almalki and colleagues 63 identified G1P[4] as the dominant strain in Yemen. The emergence of G1P[4] might have resulted from reassortment with G2P[4]. G2P[4] was detected in this study, in line with findings from other studies.18,62 Other combinations detected in this study included G1[P6], G1P[8], G2P[6], G2P[9], G9P[6], G2G9P[4], G2P[4]P[6], G1P[4]P[8], G2G9P[8], G1G2G9P[8], and G1[non-typable] P[non-typable]. The emergence of unusual G and P combinations is common with rotavirus infection. This might be the result of infection by multiple strains or zoonotic transmission, which could lead to reassortments and subsequently affect vaccine efficacy.
Rotarix (RV1) is the planned vaccine formulation to be introduced into Nigeria’s national immunization program. 41 The vaccine is a monovalent vaccine derived from an attenuated human G1P[8] strain. RotaTeq, a live pentavalent G1, G2, G3, G4, G9, and P[8] human-bovine rotavirus reassortant vaccine, has also been licensed by the World Health Organization for introduction or use in the national immunization programs of different countries. Although these vaccines have proven high and durable efficacy against circulating strains that are homotypic and heterotypic to the vaccine strains in low-mortality settings, weaker and waning efficacy have been reported in South Asia and Africa.64–67 Similar findings were reported for ROTAVAC, ROTASIL, and RV3-BB.68–71 The reason for these variations in efficacy is not fully understood; however, it might primarily be the result of greater strain diversity with several rotavirus types co-circulating, whereas maternal antibody interference, malnutrition, and other enteric infections could also play a role. However, it is important to note that despite these variations, the vaccines have greatly reduced the mortality and morbidity associated with rotavirus gastroenteritis in both high- and low-income settings. 72
Conclusion
The major limitations of this study included the low number of samples collected because it was a hospital-based study and our inability to confirm the detected genotypes of isolated viruses by sequencing. However, the results from this study revealed the genetic diversity of rotavirus strains circulating in Ibadan, Nigeria and the presence of mixed enteric viral infections. This underscores the need for continuous surveillance to monitor circulating strains as Nigeria prepares to fully introduce the vaccine in its national immunization program.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605221121956 - Supplemental material for Human rotavirus VP4 and VP7 genetic diversity and detection of GII norovirus in Ibadan as Nigeria introduces rotavirus vaccine
Supplemental material, sj-pdf-1-imr-10.1177_03000605221121956 for Human rotavirus VP4 and VP7 genetic diversity and detection of GII norovirus in Ibadan as Nigeria introduces rotavirus vaccine by Meshach Maunta Maina, Adedayo Omotayo Faneye, Babatunde Olanrewaju Motayo, Ntung Nseabasi-Maina and Adekunle Johnson Adeniji in Journal of International Medical Research
Footnotes
Data availability
All raw data are available from the corresponding author on request.
Author contributions
MMM planned and performed the experiments, conducted data analysis, and prepared and edited the manuscript. AOF designed experiments and aided in preparing and editing the manuscript. BOM performed experiments, aided in data analysis, and edited the manuscript. NNM aided in data analysis and prepared and edited the manuscript. AJA conceived and designed the experiments, aided in data analysis, and edited the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interest
The authors declare no conflict of interest.
Funding
This research did not receive external funding.
Acknowledgements
We are grateful to the technical staff of the Department of Virology, University College Hospital Ibadan for support, helpful advice, and discussions. We thank Nurse Ojo of Oni and Sons Memorial Hospital Ibadan and the staff of the Outpatient Department of Oni and Sons Memorial Hospital and Adeoyo General Hospital for support and helpful advice during sample collection.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
