Abstract
Objective
Automated systems are needed for the rapid and accurate diagnosis of
Methods
This analytical study employed a cross-sectional approach. First, 44 clinical isolates of
Results
Compared with the results for VITEK®-MS, the congruence rates for TDR-300B and VITEK®2 were 80.76% (21/26) and 92.30% (24/26), respectively. Further, high sensitivity was observed for TDR-300B and VITEK®2 (95.45% and 100%, respectively). In addition, TDR-300B had a lower positive predictive value and accuracy than VITEK®2, albeit without significance.
Conclusions
Conclusively, there were no significant differences regarding the diagnostic efficiency of TDR-300B and VITEK®2 for
Introduction
An efficient disease management program relies on the rapid and accurate diagnosis of causative agents. Unfortunately, this is hampered by the complexity and variability of sample types and volumes and analytical processes involved in routine testing procedures. With the advent of advanced diagnostics, most microbiology work-up algorithms are automated and computerized.
12
These systems generate faster and more precise results. VITEK®2 (BioMerieux, Marcy-l’Étoile, France) and TDR-300B (Mindray, Shenzhen, China) are highly efficient and robust bacterial diagnostic systems. Although there is substantial evidence for the reliability of VITEK®2 for bacterial identification, scarce data on the performance of the TDR-300B biochemical test array have been reported.13–16 In the present study, we evaluated and compared the diagnostic capability of TDR-300B and VITEK®2 to VITEK®-MS for the accurate identification of
Materials and methods
Study outline and specimen
This analytical study, which employed a cross-sectional approach, was performed to correlate the identification capabilities of TDR-300B and VITEK®2 in comparison with VITEK®-MS (BioMerieux). Forty-four clinical samples were procured from patients, especially those admitted to the ICU of Atma Jaya Catholic University of Indonesia (North Jakarta, Indonesia), between 2015 and 2018. These samples mainly comprised sputum samples along with non-sputum samples, including those from the appendix, blood, feces, eye scab, pus, throat, and urine. The samples were stored at −20°C until further use. Bacterial isolation was performed in a fully equipped microbiology laboratory (Microbiology Department, School of Medicine and Health Science, Atma Jaya Catholic University of Indonesia) according to standard procedures and guidelines. 17
Research ethics and patient consent
Clinical specimens were obtained with the prior informed consent of each patient. 18 The present study was performed according to the standard guidelines and approval of the Departmental Ethical Committee of the School of Medicine and Health Science, Atma Jaya Catholic University of Indonesia (Ethical approval number: 11/05/KEP-FKUAJ/2017).
Microbiology work-up and bacterial isolation
Phenotype identification exploits the observable morphological (e.g, cell shape, size, and arrangement, colony features, cell motility), cultural (e.g., size, shape, surface, elevation, edged, opacity, color, hemolysis, consistency), and biochemical characteristics (metabolic properties) of a microorganism using a manual, semi-automated, or automated technique. Conventionally, these include Gram staining, assessments of growth characteristics on solid-nutrient agar medium (e.g., blood agar, MacConkey agar), and biochemical metabolic tests (e.g., indole production test, citrate utilization test, oxidase test). 19
In all cases, microbial growth was allowed at 35 ± 2°C for 18–24 hours. After phenotype characterization, a single colony was isolated for each isolate and processed for further analysis.
Bacterial identification
The Gram-negative, oxidase-positive rods selected on blood agar and MacConkey agar were subjected to the semi-automated TDR-300B and fully automated VITEK®2 methods of identification. In this study,
In the semi-automated TDR-300B instrument, there are separate devices for specimen insertion, inoculation, incubation, and analysis with a manual loading of the specimen in each device. Specifically, the inoculated cultures (on the non-fermentative [NF]-64 card) are incubated outside the instrument. On the contrary, VITEK®2 is a fully automated instrument that allows microbial analysis and antimicrobial sensitivity tests on the same device. Both instruments perform automated identification and antimicrobial sensitivity test analysis.20,21
Inoculum preparation
Single colonies on the nutrient agar slant were selected and suspended in 2.5 mL of 0.45% sterile saline to adjust the bacterial suspension to a 0.5 McFarland turbidity standard using a densitometer (BioMérieux) according to the manufacturer’s instructions. Each bacterial suspension was prepared within 30 minutes of inoculation into the NF-64 card/gram-negative (GN) cassette for identification by TDR-300B or VITEK®2 (Figure 1). 22
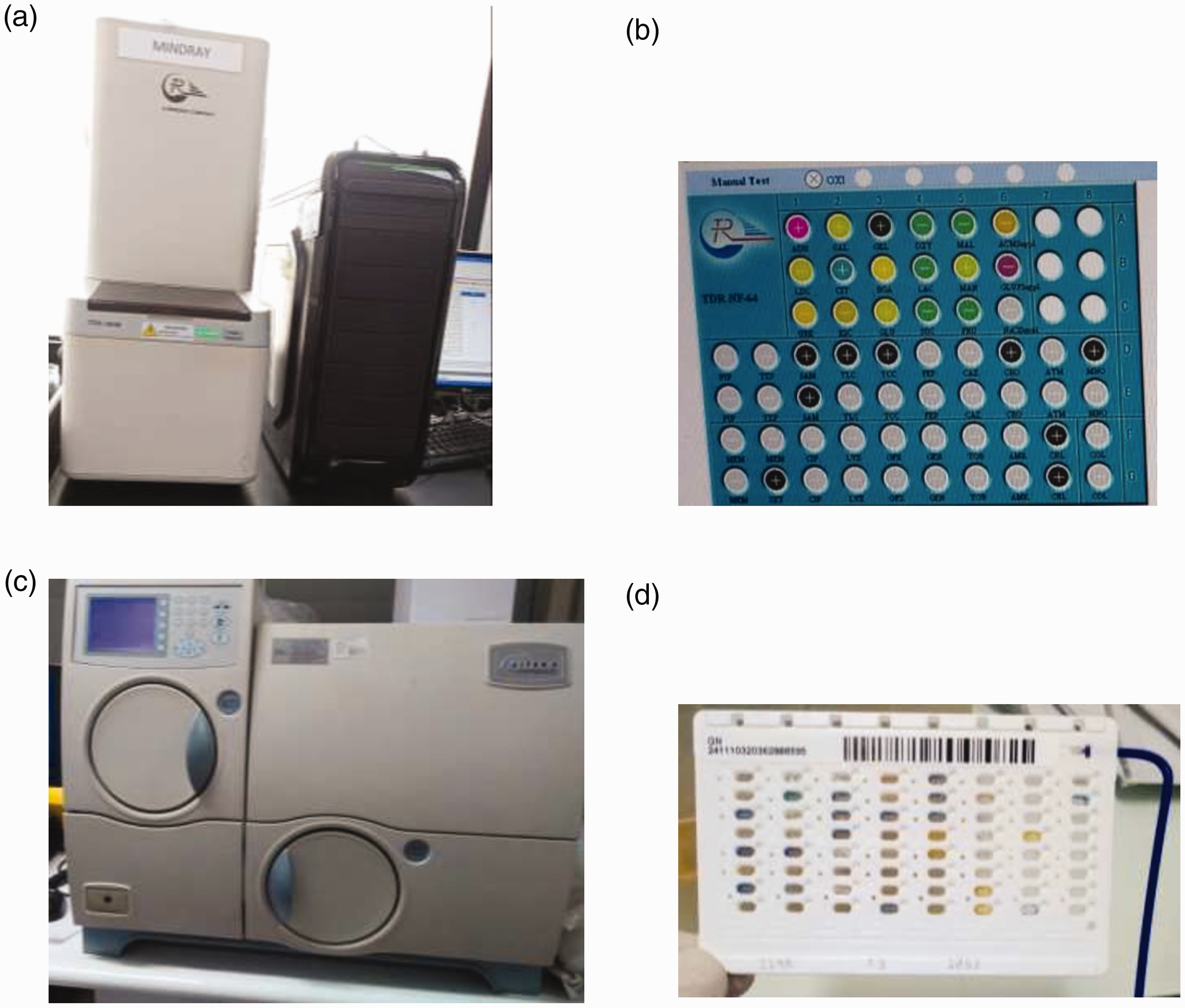

(a) TDR-300B and its (b) NF-64 card. (c) VITEK®2 and its (d) GN cassette
TDR-300B semi-automated identification method
Colonies on a nutrient agar slant were made suspended in a 0.5 McFarland solution and inoculated in the NF-64 card. The biochemical test array of the NF-64 card, which allows for 18 biochemical tests simultaneously, is presented in Table 1. The inoculated card was then incubated at 35 ± 2°C for 18 to 24 hours. TDR-300B Microorganism Analysis System version 1.0.0.7 software (Mindray) was used for reading.
Biochemical test array of the TDR-300B NF-64 card.

NF, non-fermentative.
VITEK®2 automated identification method
Bacterial suspensions (0.5–0.63 McFarland) were prepared and inoculated in the GN cassette. The biochemical test array of the GN cassette is presented in Table 2. The analysis was performed using VITEK®2 software version 07.01 (BioMérieux).
Biochemical test array of the VITEK®2 GN cassette.

GN, gram-negative.
VITEK®-MS standard automated identification method
VITEK®-MS was used as a gold standard for the identification of
Statistical analysis
In this study, the sample size (n) was calculated as follows:

The accuracy interval applied was equal to 95%, and then Zα was 1.96. Using these values, the target sample size was calculated as follows:
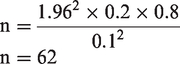
The congruence between TDR-300B and VITEK®2 was analyzed using the interrater reliability test. Results were reported as the percent of agreement (Pa) or kappa value. Fisher’s exact test and Bayes’ formula were applied to compare the results obtained using TDR-300B and VITEK®2 with those obtained using VITEK®-MS.
The final identification results for TDR-300B and VITEK®2 were classified as (a) correct identification of an organism at the genus and species levels, (b) misidentified because of an incorrect identification, and (c) not identified when an acceptable level of identification was not achieved.
Results
Forty-four clinical isolates labeled as
Performance of TDR-300B in the identification of P. aeruginosa
The accuracy of TDR-300B in the identification of
Identification of

Performance of VITEK®2 in the identification of P. aeruginosa
VITEK®2 correctly identified
TDR-300B versus VITEK®2
The congruency values for TDR-300B and VITEK®2 at the genus and species levels were 83.87% (26/31 isolates) and 80.64% (25/31 isolates), respectively (Table 3). This incongruence at the species level occurred because of a mismatch for two isolates, namely
Agreement for nine biochemical tests between TDR-300B and VITEK®2.
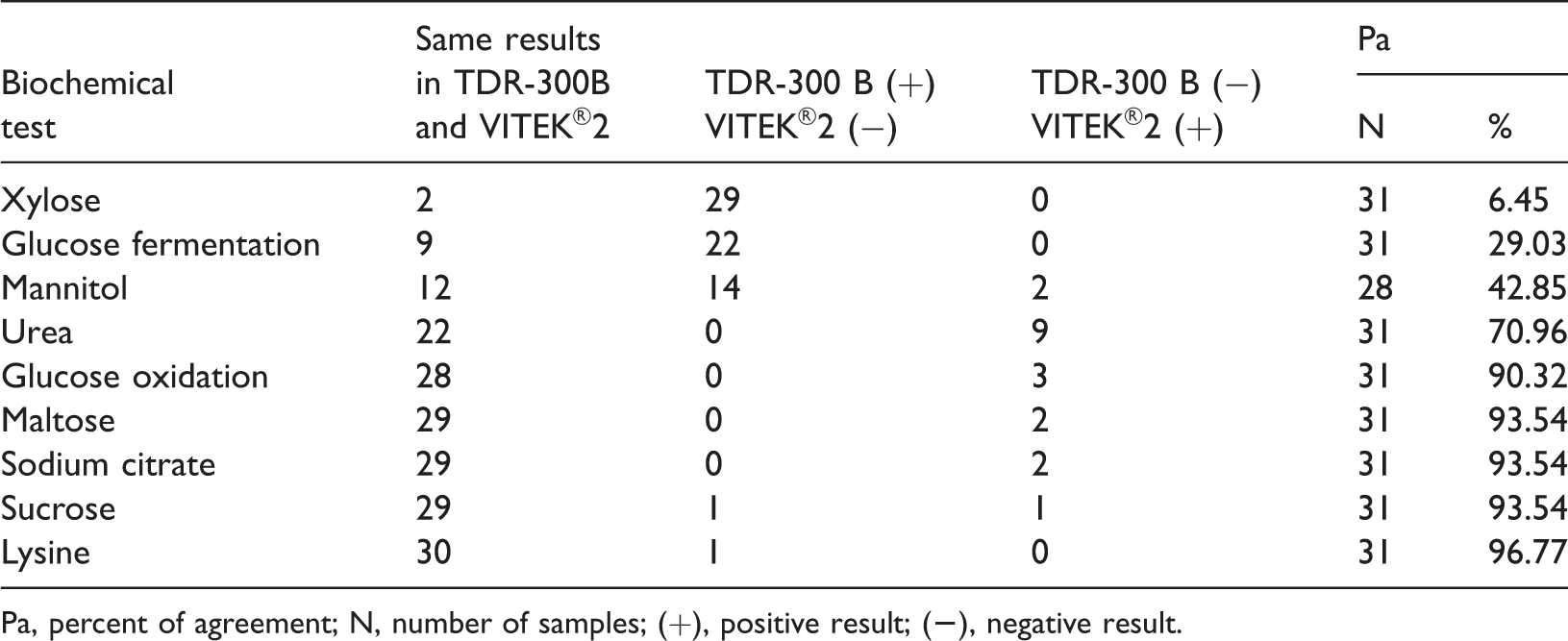
Pa, percent of agreement; N, number of samples; (+), positive result; (
VITEK®-MS was also employed for the identification of the retrieved clinical isolates. Of the 31 clinical isolates, 25 samples were identified as
Relative to VITEK®-MS, TDR-300B exhibited congruence for 84.61% (22/26) and 80.76% (21/26) of the samples at the genus and species levels, respectively. The incongruence at the species level resulted from the misidentification of five isolates:
The diagnostic capabilities of TDR-300B and VITEK®2 were compared with those of VITEK®-MS. Although TDR-300B had high sensitivity (95.45%), the value was 100% for VITEK®2 (Table 5). In addition, TDR-300B had a numerically lower positive predictive value (PPV) and accuracy (84% and 80.77%, respectively) than VITEK®2 (92% and 92.31%, respectively). However, the differences were not significant according to Fisher’s exact test. Thus, there were no significant differences observed in the diagnostic capabilities of TDR-300B and VITEK®2 regarding the identification of
Diagnostic capability of TDR-300B and VITEK®2 compared with VITEK®MS.

PPV, positive predictive value; CI, confidence interval; SD, standard deviation.
Discussion
Paim
Certain limitations of this study must be noted. This pilot study was conducted using limited available samples (sputum samples along with non-sputum samples such as the appendix, blood, feces, eye scab, pus, throat, and urine) procured from patients, particularly those admitted to the ICU of Atma Jaya Catholic University of Indonesia, between 2015 and 2018. To our dismay, only 44 clinical isolates (labeled as
The correct identification of microbial species relies on a combination of approaches including phenotypic and molecular techniques. In this pilot study, the isolates were identified using TDR-300B and VITEK®2 biochemical test-based diagnostic systems, and the results were compared with those of VITEK®-MS, an automated MALDI-TOF mass spectrometry method. Although VITEK®-MS is a robust, rapid, accurate, and cost-effective diagnostic platform, molecular tests are entrusted for the final confirmation of the identified species.27–29 Unfortunately, in a developing country such as Indonesia, it is not feasible to adopt a combination of various approaches because of economic constraints and thus, biochemical diagnostics are preferred in routine microbial diagnosis. Finally, employing molecular tests for the confirmation of all isolates used for the comparison of the diagnostic systems is emphasized, which is a limitation of this study.
Conclusion
There were no significant differences observed between TDR-300B and VITEK®2 in the identification of
Footnotes
Acknowledgements
We thank the School of Medicine, Atma Jaya Catholic University of Indonesia for providing all necessary support. We are grateful to Dr. Lely Saptawati of Moewardi Hospital (Surakarta, Indonesia) for experimental support.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author contributions
LHM, ET, and HP: Conceptualization, visualization, methodology, validation, and data curation. SJ: Formal analysis, data curation, original draft preparation, review, and editing. DE and TR: Methodology, data curation, and formal analysis. LHM: Supervision. All authors contributed significantly to the study, and all authors have approved the final manuscript.
Data availability
All relevant data is present in the manuscript, and the same is accessible for the peer-review process.
