Abstract
Objective
This study aimed to investigate the type and frequency of mutations in 10 genes in 85 colorectal cancer (CRC) patients in Huizhou and the guiding significance of targeted drug use.
Methods
The 10-gene panel next-generation sequencing (NGS) was used to assess genetic variants in 85 CRC patients from the Huizhou area combined with clinical information for a comprehensive analysis.
Results
Upon initial mutation testing, 68% (58/85) were positive. The mutation frequencies of these genes, including KRAS, PIK3CA, NRAS, ERBB2, BRAF, EGFR, and PDGFRA, were 51%, 20%, 5%, 4%, 4%, 1%, and 1%, respectively. Overall, 29 mutation types were detected from seven genes. More mutations were detected in more advanced cancers. There were three samples with multiple mutations of a single gene, including KRAS (n = 2) and ERBB2 (n = 2), 12 samples with multiple mutations of double genes, including KRAS/PIK3CA (n = 10), BRAF/PIK3CA (n = 1), and NRAS/PIK3CA (n = 1), and one sample with multiple mutations of three genes, including ERBB2/KRAS/PIK3CA (n = 1). Theoretically, 27 patients could receive targeted treatment. During the actual treatment, 10 patients received bevacizumab, cetuximab, or fruquintinib with no progression ranging from 12 to 24 months.
Conclusion
Gene mutations detected by a 10-gene panel were useful for targeting therapy of CRC in Huizhou.
Introduction
Colorectal cancer (CRC) is among the most lethal and prevalent malignancies worldwide, being responsible for nearly 881,000 cancer-related deaths in 2018. 1 CRC is the fifth most common cause of death in men and women in China. 2 However, more than half of all cases and deaths are attributed to modifiable risk factors, such as smoking, an unhealthy diet, high alcohol consumption, physical inactivity, and excess body weight, and are thus potentially preventable. 3 The 5-year survival rate for CRC is ∼64%, but drops to 12% for metastatic CRC. Further investigation is still required to develop effective approaches for medical intervention. 4 Nearly a quarter of CRCs are diagnosed at an advanced stage with metastases and 20% of the remaining cases may develop metachronous metastases, which result in difficulties in curative surgical control and subsequent tumor-related death.5–7
The ideal CRC treatment is to completely remove the tumor and any metastases, most of which require surgical intervention. 8 Several targeted therapies have been approved for CRC treatment. Identification of gene mutations that can indicate potential response to therapy is rapidly processing. The anti-epidermal growth factor receptor (EGFR) antibody cetuximab is approved together with panitumumab for the treatment of patients with metastatic CRC with tumors lacking KRAS and BRAF mutations. 9 KRAS and BRAF mutations are associated with poor survival, independent of microsatellite instability status, in patients with curatively resected CRC. 10
Next generation sequencing (NGS) has been an invaluable tool to involve genomic sequencing in clinical practice. It has become a new strategy for identifying relevant tumor-driving genes and related signal transduction pathways as potential therapeutic targets. NGS technology has facilitated many important findings in cancer research and treatment. Additionally, the incorporation of clinically-relevant target sequences into NGS-based gene panel tests has generated practical diagnostic tools that can enable individualized cancer patient care. 11
To demonstrate the applicability of this sequencing panel in clinical samples, we used a 10-gene panel in the present study to profile a set of CRC samples from Huizhou (Guangdong Province, China). We aimed to investigate the mutation types and frequencies of these 10 genes in CRC patients from Huizhou and compared the results with data from The Cancer Genome Atlas (TCGA). This research can help contribute to the clinical management of CRC patients in Huizhou.
Materials and methods
Ethics statement
Investigations were conducted in accordance with ethical standards, the Declaration of Helsinki, and national and international guidelines. The study was also approved by Ethics Committee of Huizhou Municipal Center Hospital (Approval No. [2018]01).
Patients and samples
Written consent was obtained from all 85 patients included in the study. Patients who were referred to genetic counseling at the Huizhou Central People's Hospital between 2018 and 2019 with a history of CRC were included in the study. Information about age at diagnosis, histotype, grading, stage, tumor invasiveness, and receptor status was collected. Tissue and peripheral blood samples were collected once informed consent was obtained.
Library construction and sequencing
Genomic DNA (gDNA) was extracted from tissue and blood samples using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) and quantified using the Qubit dsDNA BR Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). gDNA samples were collected and then the fragments were detected by ultrasonic interruption and gel electrophoresis. The DNA-seq library preparation was followed by Oncomine Focus Assay (Thermo Fisher Scientific). Ultra-deep sequencing was performed on a Nextseq500 sequencer (Illumina, San Diego, CA, USA) according to the manufacturer's standard process (average sequencing depth >500X).
Mutation analysis
After obtaining the raw sequencing data, the Trim Galore program was used to filter the data by removing any low-quality reads at both ends of sequencing. Burrows-Wheeler Aligner (BWA) was adopted to align the cleaned sequences to the human reference genome (GRCH37/hg19). Picard (https://broadinstitute.github.io/picard/) was performed for duplicate removal, local realignment, and base quality recalibration. Variant and genotype calling were performed using the Genome Analysis Toolkit (GATK). Variants were annotated using the ANNOVAR software tool (v2016-02-01). The fusion genes were confirmed by the Integrative Genomics Viewer (IGV) tool. The 10-gene panel included AKT1, BRAF, EGFR, ERBB2, KIT, KRAS, MET, NRAS, PDGFRA, and PIK3CA.
Comparison of mutation frequencies of related genes in our cohort and TCGA cohort
We included the TCGA data from cBioPortal (https://www.cbioportal.org/). This cohort included 276 CRC samples. We downloaded the mutation data, counted the frequency of the 10 genes of interest, and compared them with our cohort.
Sanger sequencing
The mutations in this study were validated using Sanger sequencing.
Statistical analysis
Statistical analysis was conducted using SPSS Statistics 22.0 software (IBM Corp., Armonk, NY, USA), and P < 0.05 was considered statistically significant.
Results
Patient specimen characteristics
The clinical characteristics of the patients are listed in Table 1. There were 85 patients in this study, including 49 men (57.6%) and 36 women (42.4%). The samples were all from resection/tissue excision. For pathological stage, there were 24 stage II (28.2%), 36 stage III (42.4%), and 25 stage IV (29.41%) samples. According to histopathological type, we had 2 patients with signet ring cell carcinoma (2.4%) and 83 patients with adenocarcinoma (97.6%). For tumor grade, there were 1 well-differentiated (1.2%), 58 moderately-differentiated (68.2%), 15 poorly-differentiated (17.7%), and 11 other cases (12.9%). Patient ages ranged from 36 to 87 years, with a median of 60 years.
Clinical characteristics of colorectal cancer (CRC) patients.

Characteristics of sequencing data
The 85 CRC tissue specimens were sequenced at an average sequencing depth of 1000X. The parameters of quality control included DNA in target rate (>90%) and homogeneity (>90%).
Analysis of variation types of genes
The genes in the present study, including BRAF, EGFR, ERBB2, KRAS, NRAS, PDGFRA, and PIK3CA, had different types of mutations. The results are shown in Table 2. There were 43 patients (50.6%) with KRAS mutations, including c.34G > A, c.34G > C, c.35G > T, c.35G > C, c.35G > A, c.38G > A, c.99T > G, c.176C > G, and c.436G > A. There were four patients (4.7%) with NRAS mutations, including c.34G > T, c.35G > T, c.38G > A, and c.38G > T. There were 17 patients (20%) with PIK3CA mutations, including c.253G > A, c.277C > T, c.1592T > C, c.1633G > A, c.1634A > G, c.1636C > G, c.1637A > G, c.2176G > A, and c.3140A > G. There were three patients (3.5%) with ERBB2 mutations, including c.2033G > A, c.2264T > C, c.2305G > T, and c.2329G > C. There were three patients (3.5%) with a BRAF mutation (c.1799T > A). There was one patient (1.2%) with an EGFR mutation (c.1799T > A) and one patient (1.2%) with a PDGFRA mutation (c.2458G > A). Mutations in the AKT1, KIT, and MET genes were not found in the present study.
Analysis of variation types of genes by next-generation sequencing (NGS) in the present study.

As shown in Table 3, the mutation frequencies of these 10 genes are AKT1 (2.10%), BRAF (11.60%), EGFR (2.80%), ERBB2 (3.70%), KIT (4.90%), KRAS (40.80%), MET (3.60%), NRAS (6.20%), PDGFRA (5.10%), and PIK3CA (27.50%). The mutation frequencies of the six genes, including BRAF, EGFR, ERBB2, NRAS, PDGFRA, and PIK3CA, in the present study were lower than those from TCGA database. The mutation frequencies of the RAS genes (KRAS and NRAS) in the present study were higher than those from TCGA database. The mutations of these genes were useful for targeted drug guiding.
Mutation frequency of variation types of genes by next-generation sequencing (NGS) in the present study.
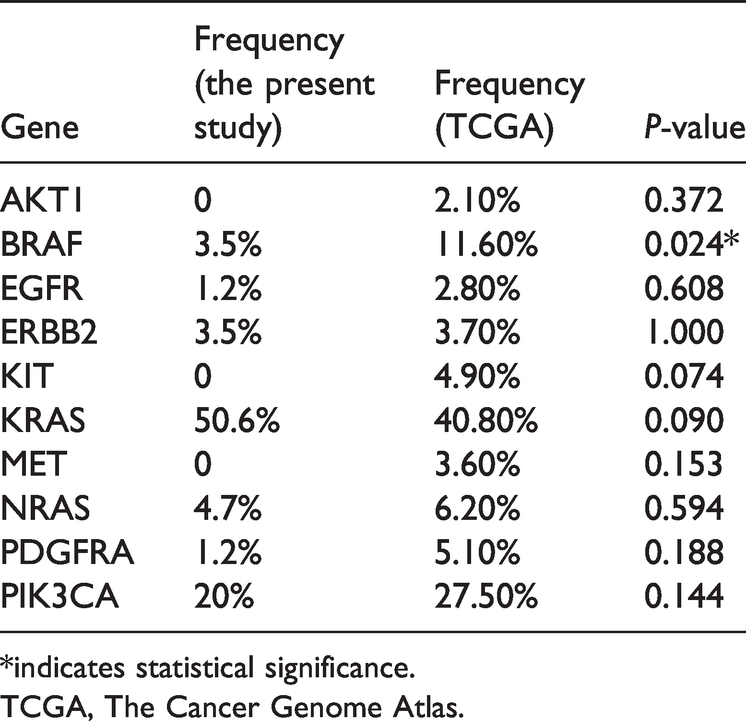
*indicates statistical significance.
TCGA, The Cancer Genome Atlas.
Types of mutations in the various stages of CRC
Fifty-eight patients in this study had genetic variants of the 10 genes in the panel, including 16 patients (stage II), 26 patients (stage III), and 16 patients (stage IV) with 15, 15, and 16 variants, respectively. The results show that more mutations were detected in higher stage tumors. As shown in Table 4, mutations shared by stage II patients, stage III patients, and stage IV patients included c.35G > T, c.35G > A, c.35G > C, and c.38G > A of KRAS and c.1633G > A of PIK3CA. Mutations shared by stage II patients and stage III patients included c.3140A > G of PIK3CA, while mutations shared by stage III patients and stage IV patients included c.1799T > A of BRAF and c.34G > A of KRAS.
Types of mutations in the various stages of colorectal cancer (CRC).

Types of multiple co-occurring mutated genes
Before reporting of variants, all non-coding or synonymous variants were filtered out of the dataset. We also conservatively excluded any potential population polymorphisms (those with population allele frequency ≥0.1% from the dataset [1000 Genomes]). After exclusion of these population polymorphisms, 68.3% of the specimens (58/85) had at least one mutation detected by the 10-gene assay (Table 5).
Types of multiple co-occurring mutated genes. Percentage of specimens with one, two, or three genes mutated in a single colorectal cancer (CRC) tumor specimen. No tumors harbored more than five mutated genes in a single specimen.

Prognosis information of patients
The data show that 78 of the 85 CRC patients had surgery performed, while 7 did not. Additionally, 76 CRC patients were administered first-line chemotherapy, including 55 with the XELOX regimen, while 8 patients were given second-line chemotherapy, including 3 with first-line chemotherapy. Currently, 24 of 85 patients have progression, ranging from 1 to 37 months, and 61 patients have no progression, ranging from 7 to 24 months. Twenty-six of the 85 patients had metastases, including 4 patients with lung metastases, 3 patients with liver metastases, and 6 patients with hepatopulmonary metastases.
The 10-gene panel includes CFDA/FDA approved drugs, such as panitumumab, regorafenib, and cetuximab. The drugs in clinical trials include nintedanib, nimotuzumab, and sorafenib. The CFDA/FDA approved drugs in other cancer species include cobimetinib, afatinib, apatinib, icotinib, osimertinib, dabrafenib, dasatinib, erlotinib, vandetanib, gefitinib, cabozantinib, crizotinib, lapatinib, pertuzumab, trametinib, trastuzumab, sunitinib, temsirolimus, vemurafenib, imatinib, and everolimus. The CRC patients with KRAS, NRAS, or BRAF wild-type genotypes (without mutation) are suitable for treatment with the tumor-targeting drugs cetuximab or panitumumab (NCCN guideline v2.2019 for CRC).
Cetuximab has been approved by the CFDA. From Table 2, 55.3% (47/85) of the patients with a RAS (NRAS or KRAS) wild-type genotype cannot benefit from cetuximab drugs. The mutation positive rate of BRAF (c.1799T > A, p.V600E) was 4%. The BRAF mutation (c.1799T > A, p.V600E) is a common mutation in CRC, which is not sensitive to cetuximab or panitumumab and can only be treated with BRAF inhibitors. Theoretically, 27 patients could receive targeted treatment. During the actual treatment, six patients received bevacizumab, three patients received cetuximab, and one patient received fruquintinib. Ten patients had no progression, ranging from 12 to 24 months.
Discussion
CRC is a highly heterogeneous disease, the treatment of which has been relegated to clinicopathological staging for decades. Better individualized treatment plans can be achieved using genetic testing. 12 The addition of targeted therapies has further improved the reported overall survival (OS) rates of metastatic CRC to 21 to 24 months. 13
The improvement in progression-free survival produced by panitumumab monotherapy was significantly greater in patients with non-mutated (wild-type) KRAS than in those with mutant KRAS (in whom no benefit from panitumumab was observed). 14 Molecular tumor profiling is becoming important in the management of advanced cancer. 15 The presence of NRAS mutations helped predict a poor outcome for CRC patients with metastases. 16 RAS mutations are found in nearly half of patients with CRC, most of whom also harbor KRAS or NRAS mutations (36% for KRAS and 3% for NRAS). 17 In the present study, 55.3% of the patients harbored a RAS (KRAS and NRAS) mutation. During the actual treatment, these patients were treated with XELOX adjuvant chemotherapy after surgery, and only one patient has died.
The PIK3CA (c.3140A > G, p.H1047R) mutation is a common driving mutation in CRC. In CRC, the mutation of PIK3CA exon 21 decreased the response rate of cetuximab, while the mutation of PIK3CA exon 10 had no effect. 18 In CRC patients with a PIK3CA mutation, aspirin therapy can effectively prolong their survival. 19 In the present study, 20% of the patients harbored a PIK3CA mutation. During the actual treatment, 2 out of 17 patients underwent targeted therapy, 1 patient (BRAF: c.1799T > A & PIK3CA: c.1592T > C) with bevacizumab + XELOX dosing and 1 patient (PIK3CA: c.253G > A) with cetuximab + FOLFOX6 dosing. These two patients are currently surviving well.
Mutations in the BRAF gene are found in about 10% of CRC patients. 20 BRAF-mutated CRCs are characterized by a dismal prognosis and resistance to standard therapies, with a median OS of about 12 months. 21 Approximately 96% of all BRAF mutations are a T1799A transversion in exon 15, which results in a valine amino acid substitution: V600E. 22 Regardless of disease stage, the presence of this mutation appears to be correlated with greater chemoresistance and worse prognosis. 23 In the present study, three patients had the BRAF mutation (c.1799T > A, p. V600E). During the actual treatment, two of the three patients underwent targeted therapy, one patient (BRAF: c.1799T > A and PIK3CA: c.1592T > C) with bevacizumab + XELOX and one patient (BRAF: c.1799T > A) with bevacizumab. Both patients are currently surviving well.
Cetuximab, an EGFR-directed antibody, is an effective treatment alone or in combination with chemotherapy for patients with CRC. 24 Activation of EGFR signaling induces resistance to the ERBB2-directed therapeutic antibody trastuzumab in breast cancer cell lines in vitro and in vivo, suggesting a common mechanism for drug resistance to therapeutic antibodies in ERBB-driven cancers. 25 Generally, CRC is associated with overexpression of platelet-derived growth factor receptors (PDGFRs) in tumors and/or tumor-associated stromal cells.26,27 In the present study, 3.5% of the patients harbored an ERBB2 mutation, 1.2% of the patients harbored an EGFR mutation, and 1.2% of the patients harbored a PDGFRA mutation. During the actual treatment, one patient (ERBB2: c.2264T > C) underwent bevacizumab +Xeliri dosing. The patient is currently in good condition.
Cancer and diabetes are closely related to poorly regulated AKT activity.28,29 MET amplification and exon 14 mutations are extremely rare events in CRC. 30 The KIT gene can be targeted with the clinically approved inhibitor imatinib in the growth and maintenance of tumor initiating cells in CRC.31,32 In the present study, no mutations of PDGFRA, AKT, or MET were found.
In this study, we investigated the significance of 10-gene panel testing in guiding targeted drug use in CRC patients in Huizhou, China. Genetic test results were available for 68% (58/85) of the patients in this study. Twenty-nine mutation types in seven genes were detected. The results showed that a higher number of mutations were detected in more advanced tumors. Theoretically, 27 patients were eligible for targeted therapy. In practice, 10 patients were treated with bevacizumab, cetuximab, or frusintinib without progression for 12 to 24 months.
The limitations of this study include the small number of CRC specimens and the lack of comparison with germline testing as a mean to confirm the somatic status of the variants that were detected. Further studies with larger sample sizes and follow-up treatment data will provide a better understanding of this panel.
Conclusion
We developed a 10-gene panel detection method for CRC and examined the mutation landscape of 85 patients with CRC in Huizhou. Furthermore, 68.2% of the included CRC patients harbored a gene mutation. More mutations were detected in higher stage cancers. Specifically, 55.3% of the patients harbored a RAS (KRAS or NRAS) mutation. The single mutation and co-mutation with the highest frequency were KRAS (35.3%) and KRAS/PIK3CA (11.8%). The patients who had undergone genetic testing benefited from matched targeted drugs. Here, we demonstrated the guiding role of targeted sequencing in the clinical management of CRC in Huizhou.
Footnotes
Acknowledgement
The authors thank MyGene Diagnostics Co., Ltd. for technology support.
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and publication of this article: This study was funded by Huizhou Science and Technology Program (No. 20161128152026).
