Abstract
Objective
Long non-coding RNA (lncRNA) colon cancer-associated transcript 2 (CCAT2) plays oncogenic roles in several cancers, including esophageal squamous cell carcinoma (ESCC). However, the specific mechanism of how CCAT2 influences ESCC tumorigenesis is still unknown.
Methods
Using RT-qPCR, the mRNA expression levels of CCAT2 in 33 paired ESCC and adjacent non-cancer tissues and cell lines were measured. Lentiviral vector sh-CCAT2 was designed and transfected into TE10 cells. CCK-8 and transwell assays were employed to detect the effects of CCAT2 knockdown on cell proliferation and invasion, respectively. RT-qPCR and western blots were used to detect the effects of CCAT2 knockdown.
Results
CCAT2 was overexpressed in ESCC tissues compared with corresponding adjacent tissues. CCAT2 knockdown could suppress cell proliferation and invasion in vitro. Furthermore, knockdown of CCAT2 could suppress the mRNA and protein levels of β-catenin and Wnt-induced-secreted-protein-1 (WISP1), as well as the mRNA levels of their downstream targets VEGF-A, MMP2, and ICAM-1. High expression of CCAT2 and WISP1 were associated with poor prognosis of ESCC patients.
Conclusions
In conclusion, a novel CCAT2/β-catenin/WISP1 axis was revealed in ESCC progression and may provide a promising therapeutic target against ESCC. CCAT2 and WISP1 are potential molecular biomarkers for predicting prognosis of ESCC.
Keywords
Introduction
Esophageal carcinoma (EC) is the ninth most common cancer and the sixth most common cause of cancer-associated death globally. 1 Esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC) are the two major histological types of EC. In the highest-risk region from Iran through central Asia to north-central China, more than 90% of EC patients had the ESCC histological type. 1 Although continuous improvements to surgical techniques and chemoradiation strategies for ESCC have been achieved, the 5-year overall survival rate is still less than 20% worldwide. 2 This disheartening prognosis can be attributed to the high invasiveness and the propensity for metastasis. In recent years, target therapies, such as camrelizumab (PD-1 inhibitor) and pembrolizumab (PD-L1 inhibitor), have played increasingly important roles in the second-line treatments of advanced ESCC. 3 Thus, it is necessary to explore more novel targets to better understand the mechanisms of this cancer, predict the survival rates, and develop new drugs.
Long non-coding RNAs (lncRNAs) are a class of non-coding RNA that are more than 200 nucleotides in length. 4 Generally, lncRNAs were thought to have no protein-coding potential. 4 Conversely, other studies have indicated that some lncRNAs could encode functional micropeptides like myoregulin.5,6 LncRNAs were initially believed to be “noise” of genome transcription and have no biological function. In recent years, increasing evidence has suggested that lncRNAs actively participate in the development of diverse human diseases, especially various cancers, through regulation of specific target gene expression.4,7 Currently, many lncRNAs have been confirmed to function either as oncogenes or tumor suppressors in different cancer types.8–10
Colon cancer-associated transcript 2 (CCAT2) is a 1752 bp lncRNA located on chromosome 8q24.21. It was initially reported to be highly overexpressed in colon cancer and to promote tumor growth, metastasis, and chromosomal instability. 11 Subsequently, abnormal expression of CCAT2 was observed in a variety of human tumors, including breast, lung, gastric, liver, cervical, and esophageal cancers.12–17 CCAT2 could promote breast cancer growth by activating the Wnt/β-catenin signaling pathway. 13 Silencing of CCAT2 could inhibit the growth and metastasis of gastric cancer cells by blocking the mTOR signaling pathway. 18
Studies have shown that overexpression of CCAT2 is associated with smoking status 12 and poor prognosis of ESCC patients. 19 However, the specific molecular mechanism regarding how CCAT2 acts as an oncogene in ESCC has not been clearly elucidated. In the present study, the expression levels of CCAT2 were detected in both ESCC tissues and cell lines. The prognostic significance of CCAT2 in ESCC was also analyzed. Furthermore, the effect of CCAT2 knockdown on the proliferation and invasion of ESCC cells and the potential regulatory mechanism relating to the Wnt/β-catenin/Wnt-induced-secreted-protein-1 (WISP1) signaling pathway were investigated. These findings may enrich our knowledge about the underlying mechanism, as well as provide a potentially promising biomarker and novel therapeutic target for ESCC.
Materials and methods
Ethics statement and sampling
Paired EC tissue and normal adjacent esophageal tissue were acquired from 33 patients who had undergone surgical EC resection between August 2016 and August 2017 at Taizhou People’s Hospital. None of the patients had received preoperative anticancer treatment. According to the EC tumor-node-metastasis (TNM) criteria of the American Joint Committee on Cancer Classification (8th edition), 20 11 patients were classified as T1 or T2 stage, while 22 patients were T3 or T4 stage. Furthermore, 18 patients had lymph node metastasis, while the remaining did not. The normal adjacent tissue samples were at least 5 cm from the edge of the tumor. All samples were obtained from fresh surgical specimens, then immediately flash-frozen in liquid nitrogen and stored at −80°C until further use. The study was conducted with approval (Approval No. KY2019016) from the Human Research Ethics Committee of Taizhou People’s Hospital, and written informed consent was acquired from each patient.
Cell lines and cell culture
ESCC cell lines (KYSE-410, KYSE-150, TE10, TE11, and TE13) and human normal esophageal epithelial cell line HET-1A were purchased from Meixuan company (Shanghai, China). All cell lines were cultured in RPMI-1640 (Gibco, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Gibco), 100 U/mL penicillin, and 100 μg/mL streptomycin (Gibco). Cells were incubated at 37°C in a 5% CO2–95% air atmosphere.
Cell transfections
Knockdown of CCAT2 was performed using lenti-CCAT2 lentivirus purchased from Meixuan company. Cells transfected with sh-NC were considered as the negative control group and cells without transfection were considered as the blank control. Transfections were performed according to the manufacturer’s instructions. The multiplicity of infection (MOI) was 20:1. Following the transfections, TE10 cells were then harvested for RT-qPCR.
Quantitative reverse transcription polymerase chain reaction (RT-qPCR)
Total RNA was extracted from specific tissues or cells using TRIzol (Takara Bio, Inc., Tokyo, Japan). Then, cDNA was synthesized by using PrimeScript RT Master Mix (Takara Bio, Inc.). RT-qPCR was performed on an ABI7500 (Applied Biosystems, Foster City, CA, USA) instrument with SYBR Premix ExTaq™ Kit (Takara Bio, Inc.). Relevant primer sequences can be found in Table 1. RT-qPCR conditions were 95°C for 30 s and 40 cycles of 95°C for 5 s and 60°C for 31 s. All RT-qPCR experiments were performed in triplicate. RT-qPCR was performed to detect the mRNA expression levels of target genes. ΔCt values were used to determine their absolute expression and ΔΔCt values were used to determine their relative expression as fold changes occur. 21 By using the 2−ΔΔCt method, the relative expression levels of the target genes for each sample were normalized to those of the endogenous control GAPDH.
Relevant RT-qPCR primers.

CCAT2, colon cancer-associated transcript 2; WISP1, Wnt-induced-secreted-protein 1; VEGF, Vascular endothelial growth factor; ICAM-1, Intercellular cell adhesion molecule-1; MMP2, Matrix Metallopeptidase 2; GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Western blotting
Cells were lysed with RIPA buffer supplemented with protease inhibitor (Roche, Basel, Switzerland). Both nuclear and cytoplasmic proteins were extracted and separated using the Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime, Shanghai, China) according to the manufacturer’s instructions. Protein concentrations were determined by bicinchoninic acid assay (BCA) assay (Thermo Fisher Scientific, Waltham, MA, USA). Equal amounts (20 µg) of protein were subjected to 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to a polyvinylidene fluoride (PVDF) membrane. After blocking with 3% BSA-TBST for 1 hour, the membrane was incubated with primary antibody at 4°C overnight. The following primary antibodies were used: rabbit anti-β-catenin (1:1000, Cell Signaling Technology, Danvers, MA, USA), rabbit anti-WISP1 (1:1000, Abcam, Cambridge, UK), mouse anti-H2A (1:1000, Cell Signaling Technology), and mouse anti-GAPDH (1:5000, Abcam). The following secondary antibodies were used (1:10,000): horseradish peroxidase (HRP)-conjugated goat‐anti-mouse (Abcam) and HRP‐goat‐anti-rabbit (Abcam). Proteins were detected using an Odyssey infrared scanner (Li-Cor Biosciences, Lincoln, NE, USA). The band intensities were estimated using Photoshop Software.
CCK-8 assay
Cells (5 × 103 cells/well) were seeded in 96-well plates and maintained at 37°C overnight. Cell viability was assessed using a Cell Counting Kit (CCK)-8 (Beyotime) according to the manufacturer’s instructions. After the cells were treated as indicated, the CCK-8 solution was added into each well. After 4 hours of incubation, the absorbance at 450 nm was measured using a microplate reader (Multiskan Mk3, Thermo Fisher Scientific).
Transwell assay
The invasion ability of ESCC cells was evaluated by Transwell assay in a Matrigel‐coated invasion chamber (Corning, Corning, NY, USA). ESCC cells (5 × 104 cells/well) suspended in 200 µL of serum-free medium were seeded in the upper chamber and 600 µL of RPMI-1640 containing 10% FBS were placed in the lower chamber. The invading cells in the lower chamber were fixed in 1% formaldehyde and stained with 0.1% crystal violet after a 48-hour incubation. Positively stained cells were observed with an inverted microscope (Olympus, Tokyo, Japan) and counted at five randomly selected fields. Each experiment was independently repeated in triplicate.
Statistical analysis
All statistical analyses were performed with GraphPad Prism software (GraphPad Software, San Diego, CA). A two-tailed Student’s t test was used for statistical analysis unless otherwise indicated. For survival analysis, overall survival curves were plotted using the Kaplan–Meier method and were evaluated for statistical significance using a log-rank test. A P-value of less than 0.05 was considered statistically significant. Data are presented as mean ± standard deviation (SD).
Results
CCAT2 is upregulated in ESCC
We detected the expression levels of CCAT2 by RT-qPCR in 33 matched ESCC tissues. The results showed that CCAT2 expression levels in ESCC tissues were significantly higher than those in adjacent non-cancer tissues (t = 4.03, P < 0.001) (Figure 1a).The expression levels of CCAT2 in ESCC cell lines were also examined. Human normal esophageal mucosal cell line HET-1A was used as the control group. The results suggest that CCAT2 is highly expressed in all five cell lines (KYSE-150, 1.56-fold upregulated; TE11, 1.98-fold upregulated; TE13, 3.32-fold upregulated; KYSE-410, 6.82-fold upregulated; TE10, 16.95-fold upregulated) relative to HET-1A cells. CCAT2 expression in the TE10 cell line was the most significantly upregulated (P < 0.001) (Figure 1b). Therefore, TE10 cells were selected for further experiments to detect the biological function of CCAT2 in ESCC.

RT-qPCR assays were used to detect the mRNA expression levels of colon cancer-associated transcript 2 (CCAT2). (a) The expression levels of CCAT2 in esophageal squamous cell carcinoma (ESCC) tissues were significantly higher than those in adjacent non-cancer tissues (***: t = 4.03, P < 0.001). (b) CCAT2 was highly expressed in five ESCC cell lines, of which TE10 cells had the most significantly high CCAT2 expression (P < 0.001).
Knockdown of CCAT2 suppresses ESCC cell proliferation and invasion in vitro
To further study the biological function of CCAT2 in ESCC, lentivirus sh-CCAT2 was designed and transfected into TE10 cells. As shown in Figure 2a, compared with the sh-NC group, the expression level of CCAT2 was significantly decreased after sh-CCAT2 transfection. We constructed a cell growth curve using the CCK-8 method to detect the effect of CCAT2 knockdown on TE10 cell proliferation in vitro. The experimental results showed that knockdown of CCAT2 effectively inhibited the proliferation of TE10 cells in vitro compared with the sh-NC group (P < 0.001) (Figure 2b). Furthermore, transwell assays were performed to investigate the effect of CCAT2 knockdown on cell invasion. The results indicated that the invasiveness of TE10 cells transfected with sh-CCAT2 was significantly decreased compared with the sh-NC group (P < 0.001) (Figure 2c). Taken together, these data demonstrate that knockdown of CCAT2 suppresses cell proliferation and invasion of ESCC cells in vitro.
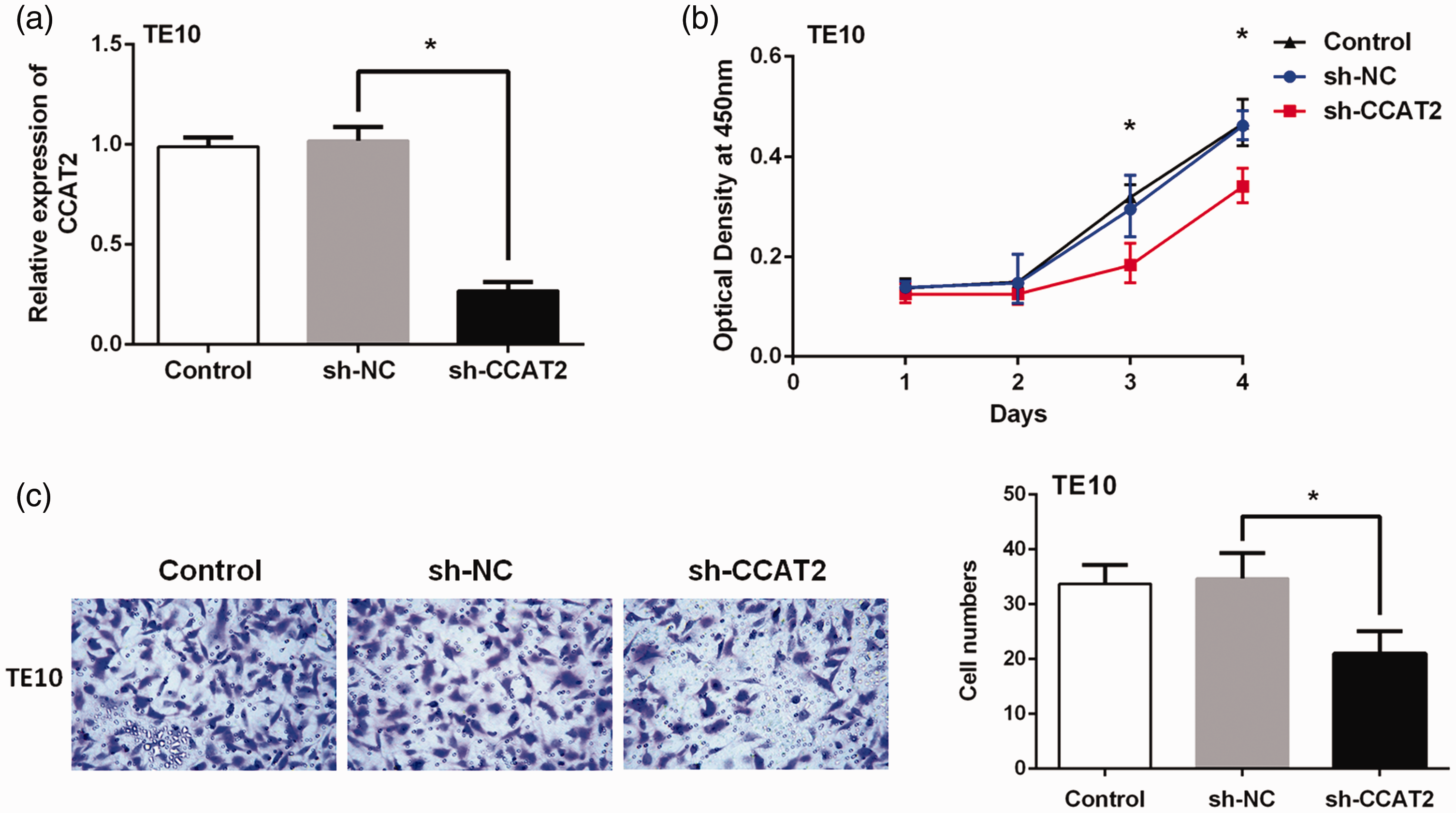
(a) The knockdown efficiency of colon cancer-associated transcript 2 (CCAT2) was measured by RT-qPCR in TE10 cells transfected with short hairpin (sh)-RNAs targeting CCAT2, as well as a negative control (sh-NC) (*P < 0.01). (b) Cell Counting Kit (CCK)-8 assays were performed to investigate the effects of CCAT2 knockdown on esophageal squamous cell carcinoma (ESCC) cell proliferation; (n = 9; *P < 0.01). (c) A transwell assay was performed to investigate the effects of CCAT2 knockdown on ESCC cell invasion; (n = 9; *P < 0.01).
CCAT2 can regulate the β-catenin/WISP1 signaling pathway
To better understand the carcinogenic mechanism of CCAT2 in ESCC, we first detected the effect of CCAT2 knockdown on Wnt/β-catenin signaling pathway activity. The RT-qPCR assays showed that knockdown of CCAT2 resulted in decreased mRNA levels of β-catenin (P < 0.01) (Figure 3a). Using western blots, we found that knockdown of CCAT2 also downregulated protein levels of β-catenin both in the cytoplasm (P < 0.01) and nucleus (P < 0.01) of ESCC cells (Figure 3b–d).

(a) The mRNA expression levels of β-catenin were detected by RT-qPCR; (n = 3). Knockdown of colon cancer-associated transcript 2 (CCAT2) decreased the mRNA levels of β-catenin in TE10 cells (P < 0.01). (b) Western blot assays were performed to detect the protein expression of β-catenin; (n = 3). Knockdown of CCAT2 could downregulate the protein levels of β-catenin in both (c) the cytoplasm (P < 0.01) and (d) nucleus (P < 0.01) of esophageal squamous cell carcinoma (ESCC) cells.
We further examined the correlation between CCAT2 and Wnt/β-catenin pathway downstream factor WISP1, which acts as an important oncogene in EC. Surprisingly, the mRNA expression levels of WISP1 in ESCC tissues of the 33 patients were also significantly higher than in adjacent non-cancer tissues (t = 3.65, P < 0.001) (Figure 4a). There was a positive correlation between the expression levels of CCAT2 and WISP1 in these ESCC samples (r = 0.73, P < 0.001) (Figure 4b). Furthermore, we found that knockdown of CCAT2 reduced both the mRNA (P < 0.01) (Figure 5a) and protein (P < 0.01) (Figure 5b) expression levels of WISP1 in ESCC cells via qRT-PCRs and western blots, respectively.

(a) RT-qPCR assays were used to detect the mRNA expression levels of Wnt-induced-secreted-protein-1 (WISP1). The mRNA expression levels of WISP1 in the 33 esophageal squamous cell carcinoma (ESCC) tissues were also significantly higher than in the adjacent non-cancer tissues (***: t = 3.65, P < 0.001). (b) The expression levels of colon cancer-associated transcript 2 (CCAT2) were positively correlated with the expression levels of WISP1 in these ESCC samples (r = 0.73, P < 0.001).

(a) The mRNA expression of Wnt-induced-secreted-protein-1 (WISP1) was detected by RT-qPCR; (n = 3). Knockdown of colon cancer-associated transcript 2 (CCAT2) decreased the mRNA levels of WISP1 in TE10 cells (P < 0.01). (b) Western blot assays were performed to detect the protein expression of WISP1; (n = 3). Knockdown of CCAT2 downregulated the protein levels of WISP1 (P < 0.01). (c-e) The expression of β-catenin/WISP1 downstream targets vascular endothelial growth factor A (VEGF-A), matrix metallopeptidase 2 (MMP2), and intercellular cell adhesion molecule-1 (ICAM-1) were examined by qRT-PCR; (n = 3). Knockdown of CCAT2 decreased the mRNA levels of VEGF-A (P < 0.05), MMP2 (P < 0.001), and ICAM-1 (P < 0.01).
The effects of CCAT2 knockdown on the β-catenin/WISP1 signaling pathway downstream factors VEGF-A, MMP2, and ICAM-1 were also examined. The RT-qPCR results showed that following CCAT2 knockdown in TE10 cells, the mRNA levels of the abovementioned three factors significantly decreased (VEGF-A, P < 0.05; MMP2, P < 0.001; ICAM-1, P < 0.01) (Figure 5c–e). Thus, these results indicate that CCAT2 regulates the Wnt/β-catenin/WISP1 signaling pathway.
Correlation between CCAT2 expression and overall survival of ESCC patients
To further investigate the correlation between CCAT2 expression and survival of patients with ESCC, Kaplan–Meier analyses were performed. As shown in Figure 6, the overall survival rates of ESCC patients in the high CCAT2 expression group were markedly decreased compared with those of the low CCAT2 expression group (P < 0.05). Furthermore, the survival of ESCC patients with high WISP1 expression was obviously shorter than that of patients with low WISP1 expression (P < 0.05). Patients with high expression levels of both CCAT2 and WISP1 had the worst prognosis (P < 0.05).

Kaplan–Meier survival curves for esophageal squamous cell carcinoma (ESCC) patients according to their expression levels of (a) colon cancer-associated transcript 2 (CCAT2), (b) Wnt-induced-secreted-protein-1 (WISP1), and (c) CCAT2 combined with WISP1 (no risk factors: low expression of CCAT2 and WISP1, one risk factor: high expression of CCAT2 or WISP1, two risk factors: high expression of CCAT2 and WISP1).
Discussion
In recent years, many lncRNAs, such as HOTAIR, 22 UCA1, 23 and MALAT-1, 24 have been found to have abnormal expression patterns in ESCC and participate in multiple processes of carcinogenesis. These discoveries provided a new research area for exploring the pathogenesis of ESCC and searching for novel targets to treat this disease. In this study, we confirmed that CCAT2 is highly expressed in ESCC tissues and cell lines. Among these cell lines, CCAT2 expression was the highest in TE10 cells. In addition, the results of the CCK-8 and transwell experiments showed that CCAT2 knockdown could significantly inhibit the proliferation and invasion of TE10 cells.
Studies have discovered that CCAT2 affects cancer progression by regulating the Wnt/β-catenin pathway.11,13,25,26 The Wnt/β-catenin pathway is a classical signaling pathway involved in multiple processes of carcinogenesis, including proliferation, apoptosis, differentiation, and metastasis. 27 β-catenin is a key downstream factor of Wnt/β-catenin signaling that interacts with various lncRNAs to regulate tumorigenesis in different cancers. 28 In colorectal cancer, CCAT2 has been shown to physically interact with the Wnt-regulated transcription factor TCF7L2, increasing its transcriptional activity and the subsequent recruitment of β-catenin. 11 In prostate cancer, knockdown of CCAT2 reduced Wnt/β-catenin signaling by inhibiting the expression of TCF7L2. 29 Cai et al. reported that suppressing the expression of CCAT2 in breast cancer cells decreased the translational levels of β-catenin both in the cytoplasm and nucleus. 13 Guo et al. reported that knockdown of CCAT2 in glioma cells suppressed β-catenin translocation from the cytoplasm to the nucleus, while there was little effect on the total cellular β-catenin levels. 26 Wang et al. showed that a CCAT2-specific short interfering RNA (siRNA) and FH535 (an inhibitor of Wnt signaling) had similar effects on cell proliferation, apoptosis, migration, and invasion of EC cells. Their immunocytochemistry results also indicated that silencing of CCAT2 could decrease the expression of β-catenin. 25 In this study, we found that knockdown of CCAT2 in TE10 cells significantly decreased both mRNA and protein levels of β-catenin, which was consistent with previous results. Similar to breast cancer cells, 13 the expression levels of β-catenin, both in the cytoplasm and nucleus of ESCC cells, were decreased after CCAT2 knockdown.
WISP1 is a member of the Cyr61-CTGF-Nov (CCN) growth factor family and a downstream target gene of the Wnt/β-catenin pathway. 30 WISP1 is widely expressed in a variety of human organs and plays an important role in many pathophysiological processes, including embryonic development, inflammation, injury repairs, and tumor progression. 31 In recent years, studies have found that WISP1 is abnormally expressed in a variety of cancers and promotes tumor growth, metastasis, and epithelial-to-mesenchymal transition (EMT) through regulation of specific signaling pathways.31,32 Chiang et al. reported that the high expression of WISP1 promoted breast cancer cell proliferation, migration, and invasion, and induced EMT, through the downregulation of E-cadherin and upregulation of N-cadherin, Snail, and β-catenin. 33 Ge et al. reported that inhibition of WISP1 inhibited proliferation, migration, and adhesion, and enhanced apoptosis of hepatocellular cancer cells through downregulation of phosphorylated protein kinase B (pAKT), glycogen synthase kinase (GSK)-3β, and matrix metallopeptidase (MMP)-2. 34
Overexpression of WISP1 was closely associated with tumor size, lymph node metastasis, and poor prognosis of ESCC patients. 35 Additionally, the high expression of WISP1 may induce radiotherapy resistance of ESCC. 36 Inhibiting WISP1 could reverse EMT-associated radiotherapy resistance and lead to the death of ESCC cells. 37 In our study, high expression of WISP1 was observed in the ESCC samples and positively correlated with CCAT2 expression. Knockdown of CCAT2 downregulated β-catenin and WISP1 expression levels, suggesting that CCAT2 is a regulator of the β-catenin/WISP1 pathway. In addition, inhibition of CCAT2 decreased the expression levels of β-catenin/WISP1 downstream targets VEGF-A, MMP2, and ICAM-1, which play vital roles in cancer angiogenesis, migration, and invasion.38–40 Targeting the CCAT2/β-catenin/WISP1 axis may inhibit metastasis and increase radiotherapy sensitivity in ESCC patients by reversing the EMT-associated progression. Moreover, patients with high expression levels of both CCAT2 and WISP1 had the worst prognosis, suggesting that CCAT2 combined with WISP1 may be prognostic biomarkers for ESCC.
In conclusion, our findings demonstrate that lncRNA CCAT2 is overexpressed in ESCC tissues and cells and acts as an oncogene in this cancer. Furthermore, a novel CCAT2/β-catenin/WISP1 axis was revealed in ESCC progression and is a potentially promising therapeutic target for treating ESCC. In addition, CCAT2 combined with WISP1 are possible molecular biomarkers for predicting the prognosis of ESCC patients. However, there are still two shortcomings of this study, namely that only in vitro experiments were conducted and that a small number of patient tissues were used. Further investigation into targeting the CCAT2/β-catenin/WISP1 axis in animal models and expanding the number of patient tissues are still needed.
Footnotes
Author contributions
CY drafted the manuscript. CY, FL, and WZ performed the experiments and data analysis. CY and FL collected the samples. JH edited the manuscript and contributed to the research conception and funding acquisition. All authors reviewed the manuscript and gave final approval of the submitted version.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by the Jiangsu Provincial Medical Innovation Team (Grant No. CXTDA2017042).
