Abstract
Objective
Mutations in GJB2, SLC26A4, and mitochondrial (mt)DNA 12S rRNA genes are the main cause of nonsyndromic hearing impairment. The present study analyzed these mutations in ethnic minority and Han Chinese patients with nonsyndromic hearing impairment from Qinghai, China.
Methods
The SNPscan assay was used to analyze mutation spectra and frequencies in the two patient groups.
Results
GJB2 mutations were detected in 9.5% (20/210) of minority patients and 20.88% (48/230) of Han Chinese patients. The most common Han Chinese GJB2 variants were c.235delC and c.299_300delAT, whereas c.235delC and c.109G > A were the most prevalent in minority patients. SLC26A4 mutations were detected in 5.71% (12/210) of minority patients and 14.35% (33/230) of Han Chinese patients, and mtDNA 12S rRNA mutations were detected in 4.28% (9/210) of minority patients and 9.13% (21/230) of Han Chinese patients.
Conclusions
These data indicate that the mutation frequencies of three deafness-associated genes were significantly higher in Han Chinese patients than in minority patients. Moreover, the GJB2 mutation spectrum was shown to differ between these two patient groups.
Introduction
Hearing impairment (HI) in humans is a genetically heterogeneous disorder. It has an incidence of approximately 1 in 1000 children worldwide, of which over half of cases can be attributed to a genetic cause. 1 Nonsyndromic hearing impairment (NSHI) accounts for approximately 70% of inherited HI, and is associated with more than 100 different genes with autosomal dominant (20%–25%), autosomal recessive (75%–80%), X-linked (1%–2%), and maternal inheritance (≥1%) patterns. 2 Previous reports have suggested that mutations in GJB2, SLC26A4, and mitochondrial (mt)DNA 12S rRNA genes are the main causes of HI.3–5
Qinghai Province is home to 43 ethnic groups, of which Tibetan, Hui, Tu, Mongolian, and Salar are native to Qinghai. Comprehensive genetic analyses of patients with hearing loss in different regions of China will provide epidemiological information that can inform effective genetic testing and accurate counseling. Data from the Qinghai Province Disabled Persons’ Federation show that 1.4% of all individuals in Qinghai Province have a HI. However, the molecular etiology of these patients has not been investigated systematically.
Currently, mutations in deafness-associated genes are detected using various methods, including direct sequencing, microarray analysis, PCR-restriction fragment length polymorphism analysis, and denatured high-performance liquid chromatography. Although direct sequencing is the gold standard approach for detecting mutations, it is expensive, time-consuming, and inefficient for the sequencing of large fragments. The single nucleotide polymorphism scan (SNPscan) technique is both high-throughput and cost-effective, and several studies have demonstrated its high accuracy, sensitivity, and specificity.6–8 Thus, it is considered a valid tool for the genetic diagnosis of inherited HI. The objective of this study was to investigate the molecular etiology of NSHI using SNPscan in patients with hearing loss from Qinghai Province. The information obtained from this study could provide a scientific basis for the diagnosis, intervention, and genetic counseling of patients with HI and their families.
Materials and methods
Patient selection
A total of 440 unrelated patients with NSHI from three special education schools in Qinghai Province were enrolled in this study. The cohort consisted of 215 male patients and 225 female patients ranging in age from 1 to 26 years, with an average age of 13.4 ± 3.5 years. As a control group, 200 age- and sex-matched healthy controls with no HI were recruited from the same region. The study protocol was approved by the Ethics Committee of the Second Hospital of Lanzhou University. Written informed consent was obtained from all subjects prior to blood sampling. The medical history of each patient was determined, including the age of HI onset, family history, mother’s health during pregnancy, previous history of infection, head trauma, and the use of aminoglycoside antibiotics. Patients received routine physical and otorhinolaryngological examinations, as well as age-appropriate audiological examinations including pure-tone audiometry or auditory brainstem response testing, immittance testing, and distortion product otoacoustic emissions testing. Patients with middle ear disorders or syndromic HI were excluded from this study. Patients with mutations in SLC26A4 were examined by temporal bone computed tomography (CT) scanning for the diagnosis of enlarged vestibular aqueduct (EVA) or inner ear malformation. EVA diagnosis was based on findings of a diameter of >1.5 mm at the midpoint between the common crus and the external aperture.
SNPscan for mutation detection
A mutation database6,8 was generated by the direct sequencing of exons from GJB2 and SLC26A4 in more than 7000 patients with HI. SNPscan genotyping was performed using a custom-designed 2 × 48-plex SNPscan™ Kit (Genesky Biotech Inc., Shanghai, China) that was developed according to SNP genotyping technology based on double ligation and multiplex fluorescence PCR, as described previously. 8 A SNPscan assay was conducted according to the manufacturer’s recommendations, as described elsewhere. 8 Raw data were analyzed using GeneMapper v4.0 software (Applied Biosystems, Waltham, MA, USA), and the genotypes at each locus were determined based on the dye colors generated by labeling and by the fragment sizes of allele-specific ligation PCR products. For quality control, the assay was randomly repeated for 4% of the samples and concordant results were obtained. Genomic DNA was extracted from the peripheral blood leukocytes of 440 patients with NSHI and 200 controls with normal hearing using a DNA extraction kit (Axygen Scientific, Inc., Union City, CA, USA).
Statistical analysis
Statistical analysis was performed using SPSS 21.0 software (IBM Corp., Armonk, NY, USA). Intergroup differences in rate or frequency were compared using the two-tailed chi-square test. A P value of <0.05 was considered statistically significant.
Results
Of the 440 patients included in this study, the majority were Han Chinese (n = 230, with an average age of 12.5 ± 3.4 years; female/male ratio 122/108), followed by Tibetan (n = 59, with an average age of 11.4 ± 2.8 years; female/male ratio 26/33), Hui (n = 53, with an average age of 10.0 ± 3.1 years; female/male ratio 27/26), Tu (n = 44, with an average age of 12.8 ± 3.2 years; female/male ratio 24/20), and Mongolian (n = 44, with an average age of 12.3 ± 2.7 years; female/male ratio 21/23). Other ethnicities were observed at much lower frequencies (Table 1). Pure-tone audiometry showed that 422 probands experienced severe to profound levels of hearing impairment, while only 18 were identified as having moderate hearing impairment probands (Table 1). Mutation frequencies for the three deafness-associated genes in 440 patients with hearing loss are shown in Table 2. A total of 115 mutations in GJB2 (n = 36), SLC26A4 (n = 77), and mtDNA12S rRNA (n = 2) genes were identified in at least two individuals selected from our mutation database. All SNPs were genotyped successfully with a call rate above 98%.
Clinical characteristics of 440 patients with NSHI.

HI, hearing impairment; NSHI, nonsyndromic hearing impairment.
Prevalence of three deafness-associated genes in Qinghai minority and Han Chinese patients.

GJB2
Forty-two patients (29 homozygotes and 13 compound heterozygotes) carried two confirmed pathogenic mutations. Twenty-six patients carried monoallelic mutations in the heterozygous form. Thus, the detection rate of GJB2 mutations was 15.45% (68/440). GJB2 mutant alleles accounted for 12.5% (110/880) of all alleles in the 440 patients with NSHI. Of these patients, 9.5% (42/440) had confirmed molecular etiology, and 5.9% (26/440) carried one pathogenic mutation. In control individuals, seven (3.5%, 7/200) were heterozygous carriers of GJB2 variants, including three with c.235delC, one with c.299_300delAT, and three with c.109G > A. A significant difference in the detection rate of GJB2 mutations was found between patients and controls (χ2 = 18.994, p = 0.001).
Nine variants were identified in the patient cohort, as shown in Table 3, and all were pathogenic. The frequency of c.235delC was 7.84% (69/880), followed by 2.05% (18/880) for c.109G > A, 1.93% (17/880) for c.299_300delAT, 0.45% (4/880) for c.35delG, 0.34% (3/880) for c.257C > G, 0.23% (2/880) for c.427C > T, 0.23% (2/880) for c.176_191del16, 0.11% (1/880) for c.512insAAC, and 0.11% (1/880) for c.232 G > A. The most prevalent variant was c.235delC, accounting for 65.45% (72/110) of all GJB2 mutant alleles in this population. Among the 40 patients carrying the c.235delC variant (9.1%, 40/440), 22 were homozygous for this variant, 10 had a compound heterozygous state with another pathogenic variant, and eight were heterozygous.
GJB2 genotypes of 440 patients with NSHI.
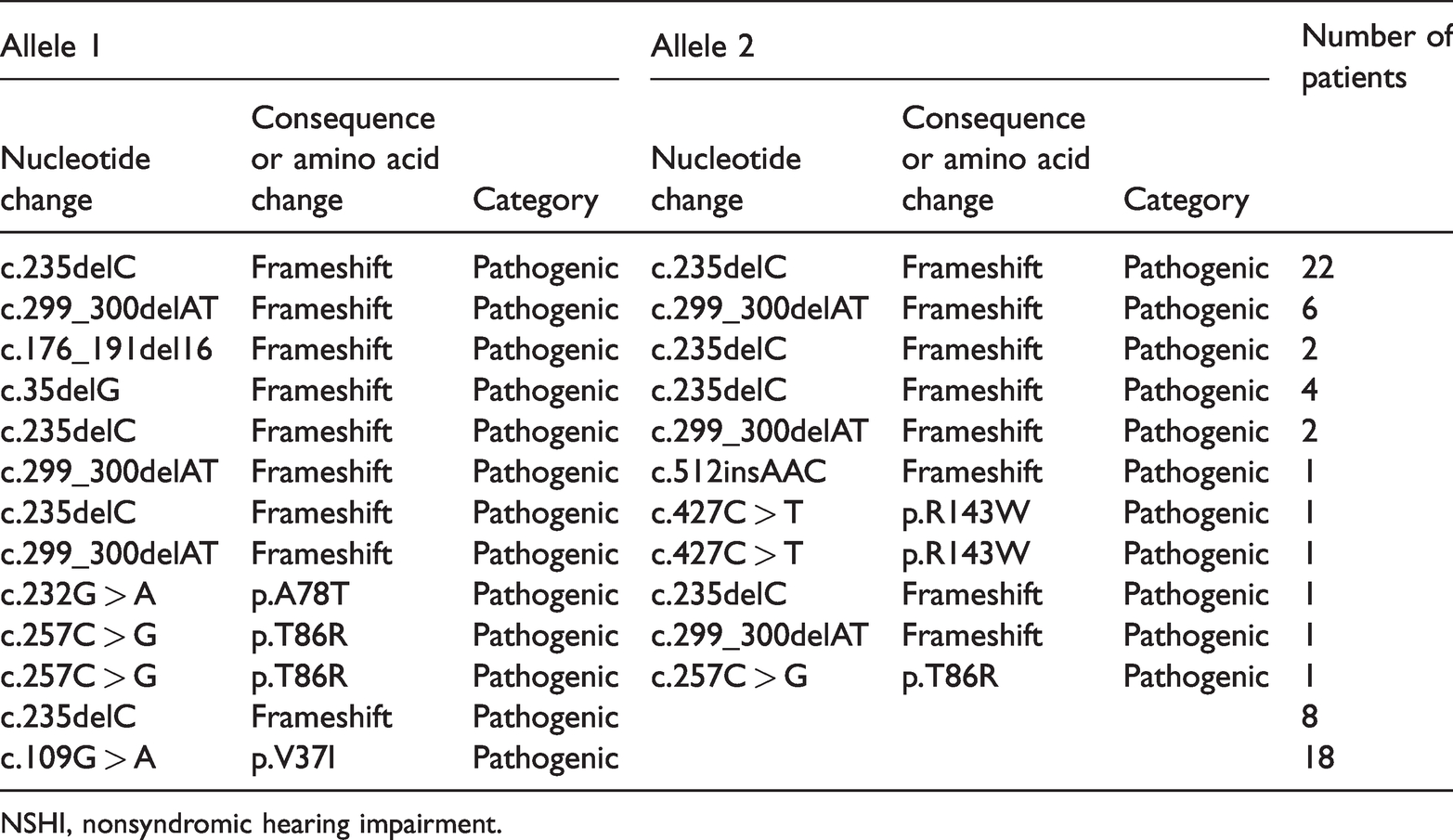
NSHI, nonsyndromic hearing impairment.
The detection rate of GJB2 mutations was 9.5% (20/210) in minority patients; six variants were detected: c.235delC, c.109G > A, c.257C > G, c.299_300delAT, c.35delG, and c.176_191del16, with allele frequencies of 3.81% (16/420), 1.67% (7/420), 0.71% (3/420), 0.48% (2/420), 0.48% (2/420), and 0.23% (1/420), respectively. Three Hui and two Tu patients were homozygous for the c.235delC variant. Three Hui patients carried c.35delG/c.235delC, one Hui patient was homozygous for c.257C > G, and another Hui patient carried c.235delC/c.299_300delAT. One Tu patient carried c.176_191del16/c.235delC. 109G > A monoallelic mutations were found in one Hui, one Man, one Tibetan, two Tu, and two Mongolian patients. One Tibetan patient was heterozygous for c.235delC.
The detection rate of GJB2 mutations was 20.88% (48/230) in Han Chinese patients; eight variants were observed: c.235delC, c.299_300delAT, c.109G > A, c.427C > T, c.35delG, c.176_191del16, c.512insAACG, and c.232G > A, with allele frequencies of 10% (46/460), 3.26% (15/460), 2.39% (11/460), 0.43% (2/460), 0.22% (1/460), 0.22% (1/460), 0.22% (1/460), and 0.22% (1/460), respectively. Variations in the mutational spectrum of GJB2 were found between minority and Han Chinese patients. Hotspot variants were c.235delC and c.109G > A in minority patients and c.235delC and c.299_ 300delAT in Han Chinese patients. A significant difference in the detection rate of GJB2 mutations was found between minority and Han Chinese patients (χ2 = 10.815, p = 0.001).
SLC26A4
Thirty patients (16 homozygotes and 14 compound heterozygotes) carried two confirmed pathogenic mutations. Fifteen patients carried monoallelic mutations in the heterozygous form. Thus, the detection rate of SLC26A4 mutations was 10.23% (45/440). SLC26A4 mutant alleles accounted for 8.5% (75/880) of all alleles in the 440 patients with NSHI. Of these patients, 6.82% (30/440) had confirmed molecular etiology, and 3.41% (15/440) carried one pathogenic mutation. Three control individuals (1.5%, 3/200) were heterozygous carriers of SLC26A4 c.919-2A > G. A significant difference in the detection rate of SLC26A4 mutations was found between patients and controls (χ2 = 15.096, p = 0.001).
Thirteen variants were identified in the patient cohort, as shown in Table 4, and all were pathogenic. The allele frequency of c.919-2A > G was 4.77% (42/880), followed by 1.93% (17/880) for c.2168A > G, 0.45% (4/880) for c.1226G > A, 0.23% (2/880) for c.2027T > A, 0.23% (2/880) for c.249G > A, 0.11% (1/880) for c.1229C > T, 0.11% (1/880) for c.1174A > T, 0.11% (1/880) for c.1517T > G, 0.11% (1/880) for c.1343C > T, 0.11% (1/880) for c.1336C > T, 0.11% (1/880) for c.1520delT, 0.11% (1/880) for c.754T > C, and 0.11% (1/880) for c.170C > A. The most prevalent variant was c.919-2A > G, accounting for 68% (51/75) of all SLC26A4 mutant alleles in this population. Among the 31 probands carrying the c.919-2A > G variant (7.04%, 31/440), 11 were homozygous, nine had a compound heterozygous state with another pathogenic variant, and 11 were heterozygous.
SLC26A4 genotypes of 440 patients with NSHI.
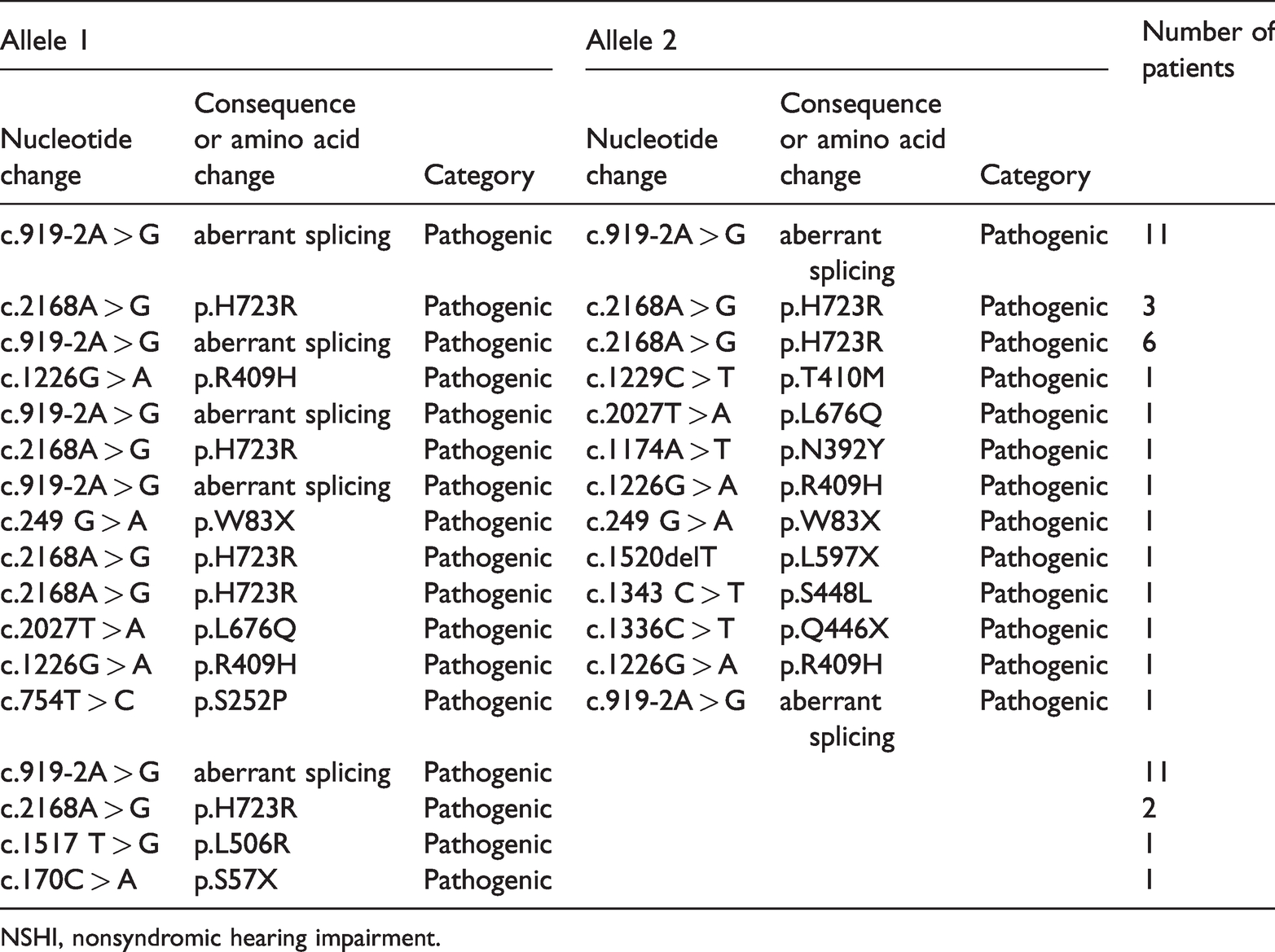
NSHI, nonsyndromic hearing impairment.
The detection rate of SLC26A4 mutations was 5.71% (12/210) in minority patients; four variants were observed: c.919-2A > G, c.2168A > G, c.1226G > A, and c.754T > C. One Salar, one Hui, and one Mongolian patient were homozygous for c.919-2A > G. One Hui and one Tibetan patient carried c.919-2A > G/c.2168A > G, one Tibetan patient was homozygous for c.1226G > A, one Tu patient carried c.754T > C/c.919-2A > G, and three Tibetan, one Hui, and one Mongolian patient were heterozygous for c.919-2A > G. The allele frequency of c.919-2A > G was 3.33% (14/420), followed by 0.48% (2/420) for c.2168A > G, 0.48% (2/420) for c.1226G > A, and 0.24% (1/420) for c.754T > C.
The detection rate of SLC26A4 mutations was 14.35% (33/230) in Han Chinese patients; 12 variants were identified: c.919-2A > G, c.2168A > G, c.1226G > A, c.1229C > T, c.2027T > A, c.249 G > A, 1174A > T, c.1517T > G, c.1343C > T, c.1336C > T, c.1520delT, and c.170C > A. Allele frequencies were 6.09% (28/460) for c.919-2A > G, 3.26% (15/460) for c.2168A > G, 0.43% (2/460) for c.1226 G > A, 0.43% (2/460) for c.2027T > A, 0.43% (2/460) for c.249G > A, and 0.22% (1/460) for each of the others. No variations in the SLC26A4 mutation spectrum were found between minority and Han Chinese patients, and c.919-2A > G was the most prevalent variant. The allele frequency of c.919-2A > G in minority patients was not significantly different from that in Han Chinese patients (χ2 = 3.663). A significant difference in the detection rate of SLC26A4 mutations was found between minority and Han Chinese patients (χ2 = 8.912, p = 0.003).
Of the 30 patients with HI with SLC26A4 mutations, 23 were examined by temporal bone CT scanning, and 20 were found to have bilateral EVA and/or other inner ear malformations (Figure 1). Of these, nine were homozygous for c.919-2A > G, seven were compound heterozygotes (c.919-2A > G/c.2168A > G [n = 5], c.919-2A > G/c.2027T > A [n = 1], and c.2168A > G/c.1520 delT [n = 1]), and four carried only one pathogenic variant (c.919-2A > G, c.2168A > G, and c.2027T > A). Temporal bone CT scanning results were normal in the remaining patients and control groups. Two minority patients (one Hui and one Salar) presented with EVA with genotypes of c.919-2A > G/c.919-2A > G and c.919-2A > G/c.2168A > G, respectively.

Temporal bone computed tomography scanning of patients showing the bilateral enlarged vestibular aqueduct (black arrows).
mtDNA 12S rRNA
Twenty-eight patients (6.36%, 28/440) carried the mtDNA 12S rRNA A1555G variant, of whom 21 were Han Chinese and seven were minority patients (five Tu, one Tibetan, and one Mongolian). Of these 28 patients, 17 had a clear history of aminoglycoside use, and 16 of these were Han patients. Two Tibetan patients (9.52%, 2/210) carried the mtDNA 12S rRNA C1494T variant (Table 5). All mtDNA mutations were homogeneous. The mtDNA 12S rRNA A1555G variant rate was 9.13% (21/230) in Han Chinese patients and 3.33% (7/210) in minority patients, representing a significant difference (χ2 = 6.191, p = 0.013). Overall mtDNA 12S rRNA mutation rates differed significantly between minority and Han Chinese patients (4.28% [9/210] vs 9.13% [21/230], respectively; χ2 = 4.055, p = 0.044). None of the 200 control individuals carried the mtDNA 12S rRNA A1555G or mtDNA 12S rRNA C1494T variants.
mtDNA 12S rRNA variants in Qinghai minority and Han Chinese patients with NSHI.

NSHI, nonsyndromic hearing impairment.
Discussion
In this study, mutation analysis using the SNPscan technique was performed to evaluate 440 patients with NSHI with moderate to profound sensorineural hearing impairment. A total of 32.5% of the patients with hearing loss showed evidence of genetic involvement. The detection rate for mutations in three deafness-associated genes was significantly higher in Han Chinese patients than in minority patients. Moreover, a significant difference in the mutational spectrum of GJB2 was found between minority patients and Han Chinese patients.
GJB2
Previous reports have suggested that mutations in GJB2 are the most common cause of NSHI in many populations.9,10 However, their prevalence varies among different ethnic groups. Epidemiological studies of deaf populations reported GJB2 mutation detection rates of 5% in Korea, 14% in Australia, 17% in Tunisia, 20% in Japan, and 43% in Israel. 11 Additionally, Dai et al. 12 obtained a GJB2 mutation detection rate of 20.94% (432/2063) in patients with NSHI from different regions of China. The present study found a GJB2 mutation detection rate of 15.45%, which differs significantly from that reported by Dai et al. (χ2 = 6.827, p = 0.009). GJB2 c.35delG, c.167delT, and c.235delC were shown to be the most prevalent variants in Caucasians, Ashkenazi Jews, and Asians, respectively. 11 We found that GJB2 c.235delC and c.109G > A were hotspots in minority patients, while c.235delC and c.299_300delAT were hotspots in Han Chinese patients, representing variations in the mutational spectrum of GJB2. Similarly, Dai et al. 12 showed that c.235delC and c.299_300delAT were hotspots in Chinese patients with NSHI, accounting for 16.7% (345/2063) and 4.4% (90/2063) of variants, respectively.
SLC26A4
Mutations in SLC26A4 are responsible for both syndromic and NSHI, and are the second most common cause of NSHI in China. The prevalence of SLC26A4 mutations differs among ethnic groups. We identified SLC26A4 mutations in 13.48% (38/282) of patients with HI, which differs significantly from the rate of 14.54% (342/2352) reported by Yuan et al. 13 in patients with NSHI from different regions of China (χ2 = 5.776, p = 0.016). The mutational spectrum of SLC26A4 also differs among ethnic groups. Previous studies have indicated that the most common SLC26A4 variants are p.T416P and IVS8 + 1G > A in northern European populations and c.2168A > G in Japanese or Korean populations;14,15 however, the most common variants in Chinese populations are c.919-2A > G and c.2168A > G.16,17 In support of this, we found c.919-2A > G to be the most prevalent in our cohort, accounting for 8.87% (25/282). Dai et al. 17 reported a frequency for this variant of 12.5% (408/3271) in patients with NSHI from different regions of China, which differs significantly from our findings (χ2 = 10.955, p = 0.01). We identified c.2168A > G as the second most common variant, at 2.95% (13/440), which does not differ significantly from the 3.52% (10/284) reported by Yuan et al. 4 (χ2 = 0.18).
EVA is the most common form of inner ear malformation associated with prelingual or postlingual sensorineural hearing impairment. Defects in SLC26A4, which encodes pendrin, may cause nonsyndromic enlarged vestibular aqueduct and pendrin syndrome. 18 We detected SLC26A4 mutations in 86.9% (20/23) of patients with EVA, similar to the 89.74% (35/39) seen by Yuan et al. 4 (χ2 = 0.112). Previous studies documented regional and ethnic diversity in the incidence of nonsyndromic EVA in NSHI populations, with reported ratios of 92% in Koreans, 78.1% in Japanese, 40% in Caucasians, 16 30% in Italians, 19 13.6% in Czechs, 20 and 16.7% in Australians. 21 SLC26A4 mutations are seen in a much higher percentage of Asian patients with EVA compared with Caucasian populations, 16 although other genetic and/or environmental factors may play a role in the development of EVA in Asian populations. For example, Qinghai Province in northwest China is in a high-altitude region with a reduced ambient oxygen tension, increased solar radiation, extreme diurnal ranges in temperature, an arid climate, and poor soil quality. High-altitude hypoxia exerts severe physiological stress on the body, including the development of an embryonic auditory organ. 22 Therefore, hypoxia and high-altitude environments may be responsible for the development of EVA.
mtDNA12S rRNA
mtDNA 12S rRNA A1555G and C1494T variants have been associated with both aminoglycoside-induced HI and NSHI in many families of different ethnicities. A1555G is more commonly detected, although its prevalence varies according to ethnicity, with reported rates of 1.6% to 8.56% in Asian populations and 0.6% to 2.5% in Caucasian populations.23–26 We found that 6.36% (28/440) of our patients with NSHI carried A1555G, and that it was present in significantly more Han Chinese than minority patients. This may be because of a founder effect: northern Han Chinese individuals largely derive from the Mongolian lineage, which differs from the Caucasian ancestry of European countries. Alternatively, the widespread use of aminoglycoside antibiotics in northwest China plays an important role in the occurrence of HI. Our results indicate that 16 Han Chinese patients with the A1555G variant had a history of aminoglycoside use, accounting for 72.73% (16/22) of all patients with A1555G. The C1494T variant, and especially the A1555G variant in the highly conserved decoding site of mtDNA 12S rRNA, are the main ototoxic targets of aminoglycoside antibiotics. These variants increase sensitivity to aminoglycoside ototoxicity and lead to permanent and profound deafness. 27 C1494T exhibits a lower carrier rate in Chinese patients with NSHI compared with A1555G. We observed a C1494T variant rate of 0.71% (2/282), which is in line with that reported by Li et al. 28 of 0.4% (13/3133) in patients with NSHI from different regions of China (χ2 = 0.014).
Conclusions
Our data demonstrate that mutation screening using the SNPscan assay is a powerful and effective method for evaluating a large deaf cohort. A total of 32.5% of the patients with hearing loss in our study showed evidence of genetic involvement, and 15.48%, 10.23%, and 6.82% of patients had inherited HI caused by GJB2, SLC26A4, and mtDNA 12S rRNA mutations, respectively. Our results revealed significant differences between minority and Han Chinese patients with respect to deafness-associated gene mutational frequencies. This information will help the design of genetic testing for deafness and enable accurate molecular diagnoses to be achieved. A combination of active genetic counseling, intervention, and the avoidance of aminoglycoside use in patients with HI and their matrilineal relatives should prevent the occurrence of HI.
Footnotes
Acknowledgments
We would like to thank all the patients and their parents.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Cuiying Graduate Supervisor Applicant Training Program of Lanzhou University Second Hospital (grant no. 201804).
