Abstract
Objective
The activation of hepatic stellate cells (HSCs) is a key event in schistosome-induced liver fibrosis. Previous studies have shown that soluble egg antigens and the recombinant P40 protein from Schistosoma japonicum eggs inhibit HSC activation. In the present study, we observed the direct effect of the S. japonicum recombinant (r)SjE16 protein on HSCs.
Methods
The sequence of SjE16 was analyzed by bioinformatics. Then western blotting, quantitative PCR, and MTT assays were performed to observe the effects of rSjE16 on HSCs.
Results
The SjE16 protein has no signal peptide or transmembrane region. rSjE16 significantly inhibited expression levels of α-smooth muscle actin and collagen I protein in LX-2 cells. rSjE16 also significantly increased the expression levels of interleukin (IL)-6 and IL-8, and enhanced the expression of matrix metalloproteinase (MMP)-2, MMP-9, and peroxisome proliferator-activated receptor-γ in LX-2 cells. LX-2 cell viability was not inhibited by rSjE16.
Conclusion
rSjE16 may be involved in the progression of HSC activation via a complex molecular mechanism, which requires further study to fully understand.
Introduction
Schistosomiasis is a serious zoonotic parasitic disease that threatens human health and social economic development. 1 In China, Schistosoma japonicum is the causative agent of schistosomiasis. During the development of schistosomiasis, schistosome eggs lodged in the host liver or intestinal tissue were reported to induce granuloma formation and cause a series of pathological changes including liver fibrosis, portal hypertension, and ascites.2,3 Liver fibrosis manifests as excessive deposition of extracellular matrix (ECM), including α-smooth muscle actin (α-SMA) and type I collagen (collagen I), 4 and hepatic stellate cells (HSCs) located within the space of Dissé in the sinusoid are the main effector cells.5,6 HSCs are normally quiescent with lipid droplet retention and increased gene expression of peroxisome proliferator-activated receptor γ (PPARγ).6,7 However, in response to all types of liver damage, they undergo transdifferentiation from vitamin A-storing perisinusoidal cells into proliferative and fibrogenic myofibroblasts. 6
Anthony et al.8,9 found that eggs from both S. japonicum and S. mansoni suppressed the activation of HSCs and had an anti-fibrotic function. Additionally, they showed that S. japonicum eggs upregulated the expression levels of inflammatory factors interleukin (IL)-6 and C-C motif chemokine ligand (CCL2) in HSCs to induce a proinflammatory phenotype. 9 We also demonstrated that soluble egg antigens (SEA) from S. japonicum inhibited HSC activation via PPARγ and transforming growth factor (TGF)-β signaling pathways. 10 Because SEA is a complex of antigen proteins, we wanted to further investigate which individual SEA protein has this function.
E16, an egg-specific 16 kDa EF-hand protein, is expressed in the larvae, insects, eggs, and miracidium, and is characterized by four calcium-binding sites to belong in the calcium-binding protein family.11,12 Using crude extracts from S. japonicum eggs, Hu and Wang et al.11,13 showed that E16 was abundantly expressed in eggs and 92% homologous to the egg protein from S. mansoni (SmE16). Further study showed that the recombinant E16 protein from S. japonicum (rSjE16) has the potential for infection diagnosis and curative effect evaluation.11,13 SjE16 was also reported to recruit neutrophils and activate macrophages to induce inflammatory hepatic granuloma initiation.14,15 However, the direct effect of SjE16 on HSCs has not yet been investigated. Here, we observed the role of rSjE16 protein from S. japonicum in HSC activation, and determined the expression changes of various inflammatory factors in rSjE16-treated LX-2 cells. These data provide the basis for further study exploring key components of SEA with inhibiting effects in HSC activation.
Materials and methods
Reagents and antibodies
3,4,5-Dimethyl N-methylthiazol-2-yl-2,5-d-phenyl tetrazolium bromide (MTT) was obtained from Sigma (St Louis, MO, USA). Mouse monoclonal antibodies against α-SMA (dilution 1:200), PPARγ (dilution 1:400), and matrix metalloproteinase-9 (MMP-9, dilution 1:200) were obtained from Santa Cruz Biotechnology (Santa Cruz, CA, USA), and the mouse monoclonal antibody against His (dilution 1:200) was obtained from TIANGEN (Beijing, China). The polyclonal rabbit antibody against MMP-2 (dilution 1:100) was obtained from Santa Cruz Biotechnology, the primary mouse antibody against collagen I (dilution 1:200) was purchased from Abcam (Cambridge, MA, USA), and the primary rabbit antibody against glyceraldehyde phosphate dehydrogenase (GAPDH, dilution 1:1000) was obtained from Goodhere (Zhejiang, China). All secondary antibodies were obtained from Santa Cruz Biotechnology. Ovalbumin from chicken egg white (OVA) was purchased from Sigma.
Bioinformatic analysis of SjE16
The SjE16 signal peptide (GenBank: CAX76376.1) was predicted by the SignaIP-5.1 Server (http://www.cbs.dtu.dk/services/SignalP/) in the Center for Biological Sequence Analysis platform. Online software was used to predict if the transmembrane region existed in the SjE16 protein (https://embnet.vital-it.ch/software/TMPRED_form.html) as well as various physiological parameters including the theoretical isoelectric point, and estimated half-life (https://web.expasy.org/protparam/). The NetAcet 1.0 server (http://www.cbs.dtu.dk/services/NetAcet/) predicted substrates of N-acetyltransferase A, while the NetPhos 3.1 server (http://www.cbs.dtu.dk/services/NetPhos/) predicted serine, threonine, or tyrosine phosphorylation sites in eukaryotic proteins using ensembles of neural networks. Finally, the polarity, flexibility, and hydrophilicity of the SjE16 protein were predicted via expasy (https://web.expasy.org/protscale/).
Expression and purification of rSjE16
rSjE16 was produced by the prokaryotic expression system as previously described. 16 Briefly, based on the SjE16 sequence published in GenBank (FN320650.1), full-length SjE16 was acquired from S. japonicum eggs, cloned into the pET-28a (+) vector preserved in our laboratory, 16 and validated by restriction analysis (using Eco RI and Hind III) and sequencing. Then, pET-28a-SjE16 was constructed and transformed into Escherichia coli BL21 (DE3) preserved in our laboratory, 16 and recombinant SjE16 soluble protein was induced by isopropyl-beta-D-thiogalactopyranoside (IPTG) in E. coli BL21 (DE3). Harvested bacterial cells underwent ultrasonic treatment and were combined with Ni-NTA His•Bind Resin (Novagen, Madison, WI, USA). Then the protein was eluted with elution buffer containing imidazole at different concentrations (50 mM, 100 mM, 250 mM, 500 mM, and 750 mM). After dialysis with elution buffer containing a low concentration of imidazole, it was dialyzed three times with phosphate-buffered saline without imidazole. After identification by western blotting, rSjE16 was treated with polymyxin B-agarose beads (Sigma) to remove endotoxins, then its concentration was determined by the Bradford assay (Sangon Biotech, Shanghai, China).
Cell culture and treatment
Human LX-2 cells preserved in our laboratory 10 were cultured in Dulbecco’s modified Eagle’s medium (Invitrogen, Waltham, MA, USA) supplemented with 10% fetal bovine serum (Invitrogen) in a humidified incubator at 37°C with 5% CO2. During culture, LX-2 cells show characteristics of activated HSCs with increased expression of α-SMA and decreased expression of PPARγ. 17 In the present study, LX-2 cells were plated in six-well plates at a density of 1 × 105 cells/well and cultured for 24 hours. Then rSjE16 was added at various concentrations for 48 hours.
RNA isolation and quantitative PCR (qPCR)
Total RNA was extracted from LX-2 cells using TRIzol reagent (Invitrogen), then reverse-transcribed into cDNA using the RevertAid™ First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Waltham, MA, USA). qPCR was performed according to the protocol of the SYBR® Premix Ex Taq™ RT-PCR Kit (Takara Bio, Shiga, Japan) using the Eco Real-time PCR system (Illumina, San Diego, CA, USA). PCR conditions were: an initial denaturation step at 95°C for 30 s, then 40 cycles of 95°C for 5s, 60°C for 30 s, and 72°C for 30 s. A final cycle of 95°C for 15 s, 60°C for 30 s, and 95°C for 15 s was included for melting curve analysis. IL-6 primers were: F, 5′-GGCACTGGCAGAAAACAACC-3′ and R, 5′-GGCAAGTCTCCTCATTGAATCC-3′; and IL-8 primers were: F, 5′-TTTTGCCAAGGAGTGCTAAAGA-3′ and R, 5′-AACCCTCTGCACCCAGTTTTC-3′. The expression levels of all target genes were normalized to the internal control (GAPDH) of the same samples for standardization, and relative gene expression was measured by the 2(−ΔΔCt) method.
Western blotting analysis
LX-2 cells pretreated with or without rSjE16 for 48 hours were harvested and resuspended in protein lysis buffer to extract protein. The protein concentration was measured by the Bradford method. Equivalent amounts of protein were separated by 10% sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and electrotransferred onto polyvinylidene fluoride membranes. The membranes were blocked with 10% nonfat dry milk in Tris-buffered saline and Tween 20 (TBST), then probed with primary antibodies at 4°C overnight. Subsequently, the membranes were washed three times in TBST, then incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 hour, washed again with TBST, then visualized using a chemiluminescence kit (Merck, Darmstadt, Germany). Band quantification was performed by ImageJ software (National Institutes of Health, Bethesda, MD, USA). As an internal control, GAPDH was detected with the corresponding antibody.
MTT assays
The viability of LX-2 cells was detected by the MTT assay. Cells were plated at a density of 1 × 104 cells/well in 96-well plates for 24 hours, then cultured for a further 48 hours in the presence of 5 µg/mL or 10 µg/mL rSjE16. To evaluate the viability of rSjE16-treated cells, a working solution of MTT was added to each well and cells were incubated at 37°C for an additional 4 hours. Dimethyl sulfoxide was then added to dissolve the formazan products. The absorbance was measured at 490 nm with an ELISA reader (Bio-Tek, Winooski, VT, USA).
Statistical analysis
All data are presented as means ± standard error of the mean of three independent experiments. The independent samples t-test was used for analysis with SPSS software version 17.0 (IBM, Armonk, NY, USA). P values <0.05 were regarded as significant.
Results
rSjE16 was expressed and purified successfully in vitro
The amino acid sequence of SjE16 was first analyzed by bioinformatics and shown to contain no Sec signal peptide (Sec/SPI), no lipoprotein signal peptide (Sec/SPII), and no Tat signal peptide (Tat/SPI). This indicated that no signal peptide would need to be cleaved during construction of the SjE16 plasmid to express rSjE16 protein in the prokaryotic expression system. The SjE16 protein was also shown to lack a transmembrane region. The theoretical isoelectric point of the SjE16 protein is 4.88 and its theoretical molecular weight is 16725.73 Da. The formula is C731H1161N197O241S5. Its estimated half-life is 30 hours in mammalian reticulocytes in vitro, greater than 20 hours in yeast in vivo, and greater than 10 hours in E. coli in vivo.
The NetAcet 1.0 server revealed the existence of one acetylation site (position 2aa), eight casein kinase II kinase sites (positions 2aa, 23aa, 37aa, 39aa, 48aa, 57aa, 96aa, and 114aa), four protein kinase G kinase sites (positions 48aa, 57aa, 75aa, and 96aa), 10 UNSP kinase sites (positions 37aa, 57aa, 61aa, 76aa, 92aa, 96aa, 106aa, 133aa, 139aa, and 143aa), one protein kinase A kinase site (position 57aa), and three protein kinase C kinase sites (positions 76aa, 122aa, and 143aa) in the SjE16 protein. Predictions of SjE16 protein polarity, flexibility, hydrophilicity. and accessibility are shown in Figure 1. The polarity values (Figure 1a) are between 6.169 (position 37aa) and 33.501 (position 128aa), average flexibility values (Figure 1b) are between 0.394 (position 119aa) and 0.496 (position 89aa), and hydrophilicity values (Figure 1c) are between –0.900 (position 140aa) and 1.822 (position 128aa). The accessibility index (Figure 1d) which scales the free energy of a globular protein to transfer from inside to outside is between –0.711 (position 69aa) and 0.189 (position 11aa).
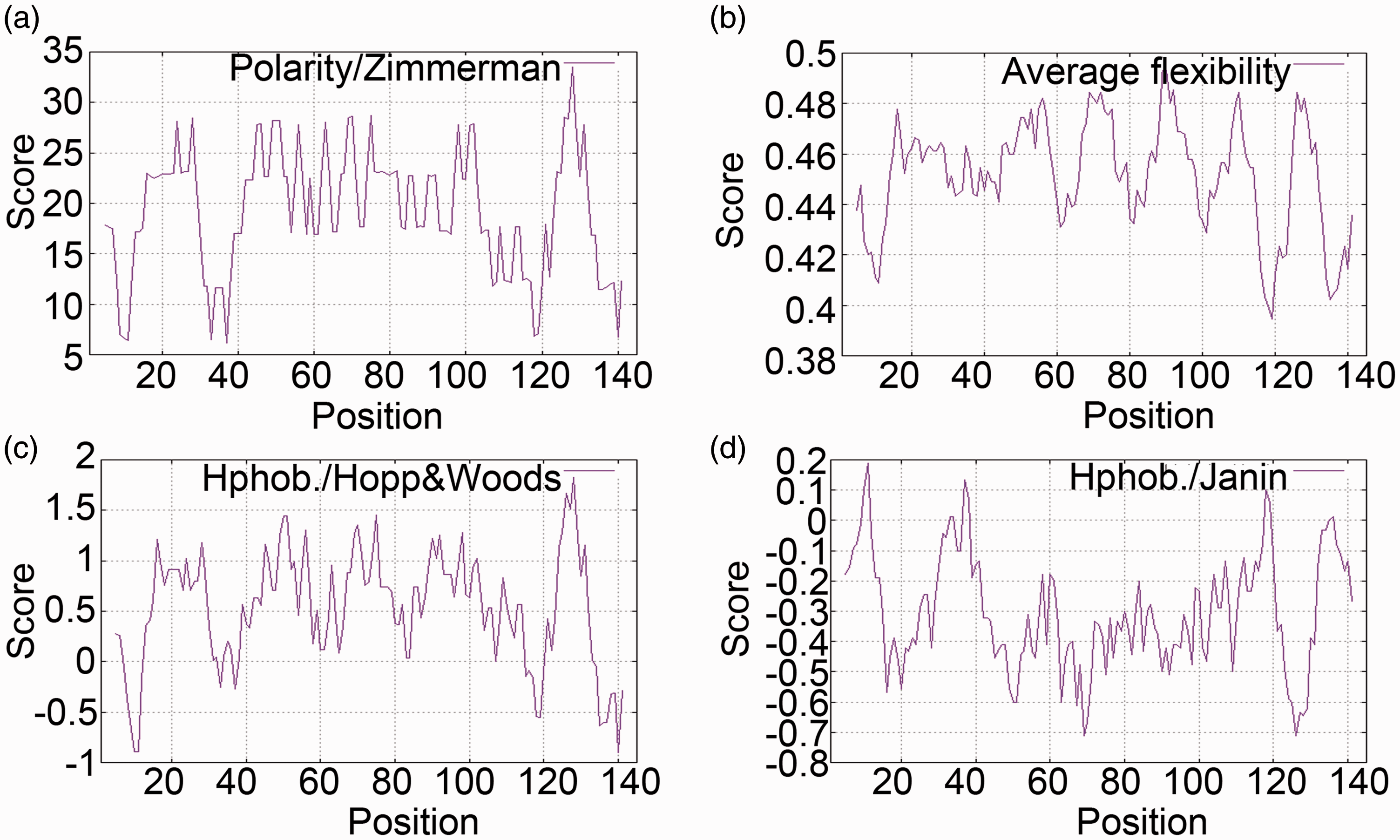
Bioinformatic analysis of the SjE16 protein. Polarity (a), flexibility (b), hydrophilicity (c), and accessibility (d) of the SjE16 protein were predicted via the expasy platform. The x axis represents amino acid positions and the y axis represents scores computed by each algorithm.
The amplified full-length cDNA sequence of SjE16 was cloned into the prokaryotic expression vector pET-28a (+) and the recombinant pET-28a-SjE16 plasmid was verified by double restriction endonuclease reaction and DNA sequencing (Figure 2a). It was then transformed into E. coli BL21 (DE3) to express the rSjE16 prokaryotic protein (Figure 2b) which was purified using Ni-NTA His•Bind Resin (Figure 2c). The obtained rSjE16 protein was also verified by western blotting using an anti-His monoclonal antibody (Figure 2d). All results showed a specific reaction at a molecular weight of ∼20 kDa.

The recombinant protein rSjE16 was expressed and purified successfully in vitro. (a) Construction of the recombinant pET-28a (+)-SjE16 plasmid. M1: λ-Hind III DNA marker; M2: DL2000 DNA marker; lane 1: pET-28a (+) vector; lane 2: the recombinant pET-28a-SjE16 plasmid following mono-restriction enzyme digestion; lane 3: the recombinant pET-28a-SjE16 plasmid following dual-restriction enzyme digestion; lane 4: PCR products. (b) The recombinant rSjE16 prokaryotic protein expressed in Escherichia coli BL21 (DE3) separated by SDS-PAGE. M: standard protein marker; lane 1: pET-28a (+) with no IPTG induction; lane 2: pET-28a (+) induced with IPTG; lane 3: pET-28a-SjE16 with no IPTG induction; lane 4: pET-28a-SjE16 induced with IPTG. (c) The recombinant rSjE16 prokaryotic protein was purified using Ni-NTA His•Bind Resin. M: standard protein marker; lanes 1 and 2: the fusion protein unsuccessfully combined with Ni-NTA His•Bind Resin; lanes 3–7: the fusion protein successfully combined with Ni-NTA His•Bind Resin and eluted using imidazole at different concentrations (50 mM, 100 mM, 250 mM, 500 mM, and 750 mM); lane 8: supernatant of whole bacterial proteins after sonication; lane 9: the fusion protein after purification and dialysis. (d) The purified fusion protein rSjE16 was verified using antibodies against His by western blotting. Lane 1: whole proteins from bacterial solution transformed with pET-28a (+) and induced with IPTG; lane 2: whole proteins from bacterial solution transformed with pET-28a-SjE16 and induced with IPTG; lane 3: the purified fusion protein rSjE16.
rSjE16 suppressed the expression of α-SMA and collagen I proteins in LX-2 cells
To explore the role of rSjE16 in activated HSCs, LX-2 cells were treated with different concentrations of rSjE16 (0, 5, and 10 µg/mL) for 48 hours and then harvested for western blotting. Significant decreases in the expression levels of α-SMA and collagen I proteins were detected in LX-2 cells treated with rSjE16 compared with nontreated cells (P<0.05, Figure 3a). The negative control OVA had no direct effect on α-SMA or collagen I protein expression (Figure 3b). Additionally, the viability of LX-2 cells was not inhibited by rSjE16 at any of the concentrations tested (Figure 3c). A concentration of 5 µg/mL rSjE16 for 48 hours was chosen as the best working condition for follow-up experiments.

rSjE16 suppressed the expression of α-SMA and collagen I proteins in LX-2 cells. LX-2 cells were treated with rSjE16 at concentrations ranging from 0 to 10 μg/mL for 48 hours. (a) Expression levels of α-SMA and collagen I proteins were detected by western blotting using GADPH as the internal standard for normalization. *P < 0.05 compared with each control group (with no stimulus). (b) OVA was used as the negative control. (c) LX-2 cell viability was detected by the MTT assay (P > 0.05, compared with the control group of LX-2 cells with no stimulus).
rSjE16 regulated the expression of other fibrotic associated proteins in LX-2 cells
PPARγ has been identified as an adipogenic transcription factor involved in the activation of quiescent HSCs and phenotype alteration,18,19 so we examined its expression in rSjE16-treated LX-2 cells. As expected, LX-2 cells treated with rSjE16 showed significantly enhanced PPARγ protein expression compared with nontreated LX-2 cells (P<0.05, Figure 4). rSjE16 at the effective stimulus concentration of 5 µg/mL also significantly enhanced the expression of MMP-9 and MMP-2, which both play an important role in homeostatic regulation of the extracellular environment, including the degradation and reestablishment of ECM in various physiological and pathological conditions (P<0.05, Figure 4). Collectively, these findings indicated that rSjE16 regulated the expression of fibrotic proteins in LX-2 cells.
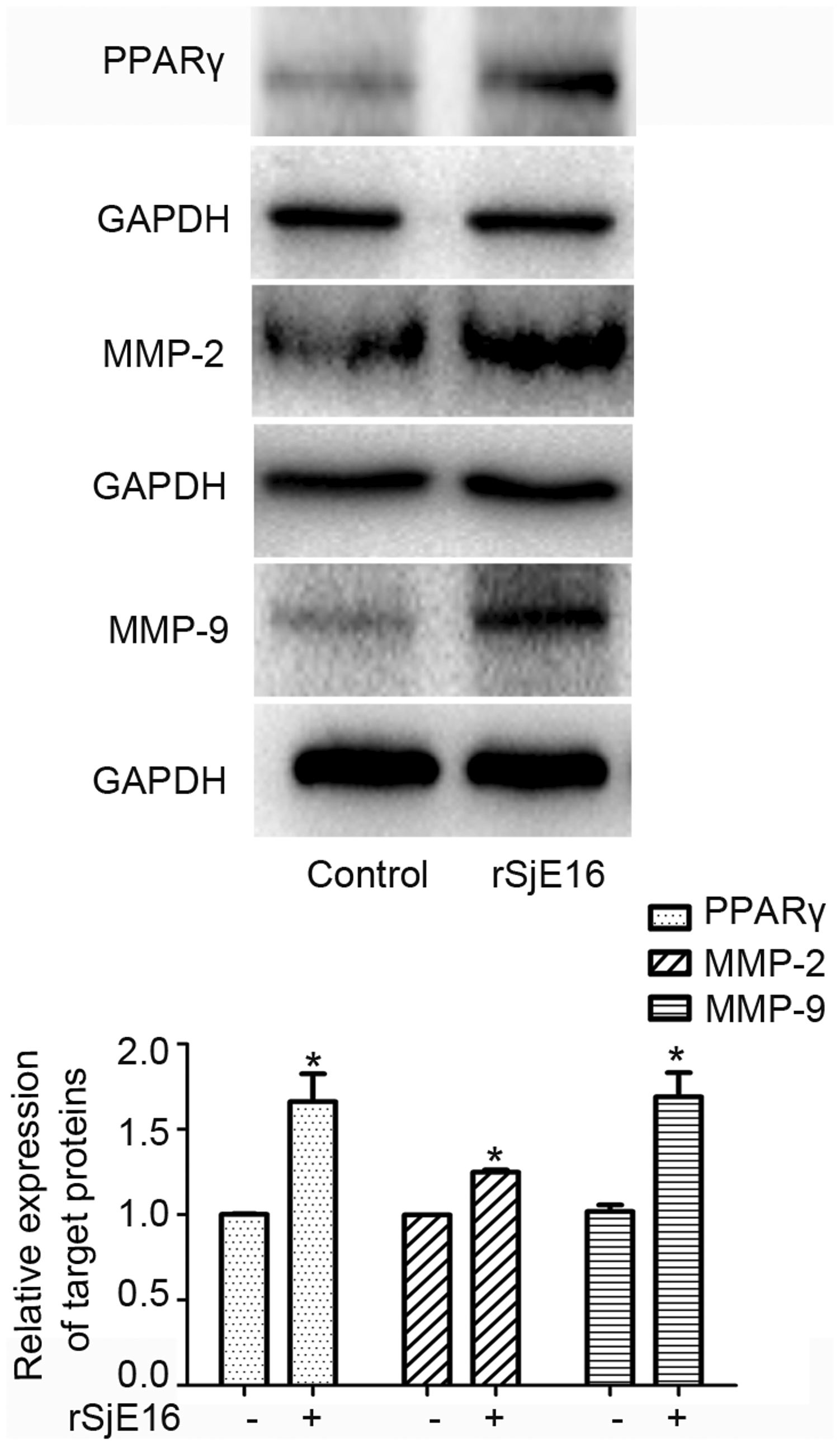
rSjE16 regulated the expression levels of other fibrotic-associated proteins in LX-2 cells. LX-2 cells were treated with 5 μg/mL of rSjE16 for 48 hours. Expression levels of PPARγ, MMP-2, and MMP-9 proteins were detected by western blotting. *P < 0.05 compared with each control group
rSjE16 upregulated IL-6 and IL-8 mRNA expression levels in LX-2 cells
HSCs produce cytokines and chemokines in an autocrine manner, and these are thought to be key events in fibrogenesis. Because S. japonicum eggs simultaneously upregulate CCL2 and IL-6 mRNA expression levels and induce a proinflammatory, anti-fibrogenic phenotype in HSCs, 9 we next investigated whether rSjE16 could regulate the expression of some inflammatory factors. As shown in Figure 5, IL-6 and IL-8 mRNA expression levels were both significantly upregulated in LX-2 cells treated with rSjE16 for 48 hours (P<0.05). These results indicate that rSjE16 promotes the generation of some inflammatory cytokines in LX-2 cells.

rSjE16 upregulated the mRNA expression levels of IL-6 and IL-8 in LX-2 cells. LX-2 cells were treated with 5 μg/mL of rSjE16 for 48 hours. mRNA expression levels of IL-6 and IL-8 were measured by qPCR and normalized against that of GAPDH. Results are shown as the mean ± SEM of three independent experiments. *P < 0.05, relative to the control group.
Discussion
HSCs have been identified as a major effective cell type in liver fibrosis. During this process, activated HSCs synthesize large quantities of ECM, including α-SMA, collagen, and tissue inhibitors of metalloproteinases (TIMPs)/MMPs. 20 HSCs also release various cytokines, such as tumor necrosis factor (TNF)-α, IL-6, and TGF-β1, to induce the occurrence and development of liver fibrosis.21–23 The inhibition of HSC activation is a main strategy for blocking liver fibrosis. Previous studies showed that some schistosome components can return activated HSCs with a myofibroblastic phenotype back into quiescent lipid-storing cells and exhibit anti-fibrogenic activity on TGF-β1-activated HSCs.8–10,16 For example, SEA from S. japonicum suppressed the activation of HSCs through PPARγ and TGF-β signal pathways, 10 and rSjP40, a major component of schistosome eggs, attenuated TGF-β1-induced HSC activation in vitro. 16 Recently, we found that the recombinant T2 RNase protein from S. japonicum significantly diminished the expression levels of α-SMA and Smad4 in LX-2 cells. 24 However, the recombinant T2 RNase protein was unable to inhibit the expression of procollagen I in LX-2 cells. 24 In this study, we obtained another protein (rSjE16) from S. japonicum eggs using the prokaryotic expression system and observed its direct effects on HSC activation.
SjE16, a calcium-binding protein with two EF-hand motifs from S. japonicum eggs, modulates host immune responses during host–parasite interactions and can be used as a diagnostic indicator of schistosomiasis.14,25 The EF-hand motif is found in a large number of protein families, possessing diverse functions.26,27 In our study, we found that rSjE16 significantly reduced the expression of α-SMA and collagen I and significantly increased that of PPARγ. rSjE16 also induced significantly high expression levels of MMP-9 and MMP-2. MMPs, a family of ECM degradative enzymes, are promptly expressed in HSCs in response to a diverse range of hepatic toxins. They also block liver fibrosis through the cleavage of fibrillar ECM and the promotion of apoptosis in activated HSCs. 28 MMP-9 is one of the major MMPs shown to regulate ECM remodeling and it is also involved in the inflammatory response,29,30 while MMP-2 mainly inhibits type I collagen synthesis and promotes the apoptosis of HSCs through N-cadherin.31,32 Together, these results indicate that rSjE16 suppresses HSC activation and plays an anti-fibrotic role in HSCs.
Anthony et al. reported that S. japonicum eggs promoted the mRNA expression of CCL2 and IL-6 in LX-2 cells, accompanied by a reduction in α-SMA and collagen I expression. 9 Similarly, we found that rSjE16 induced a significant increase in IL-6 and IL-8 gene expression in LX-2 cells. IL-6 is a pleiotropic cytokine involved in inflammation, hematopoiesis, and immune regulation and is strongly expressed at the onset of schistosome egg laying. 33 Following long-term schistosome infection, IL-6 induces IL-10 production and negatively regulates the T-helper 1 response. 34 Furthermore, both exogenous IL-6 and IL-10 block carbon tetrachloride-induced liver fibrosis.35,36 The cytokine IL-8 produced by a variety of neutrophil-activating cells is a proinflammatory factor that participates in the development of several inflammatory diseases. Chuah et al. 37 showed that S. japonicum eggs induce an inflammatory phenotype of neutrophils by upregulating proinflammatory cytokines (IL-1α and IL-8) and MMP-9 expression. However, Robert et al. 38 reported that TNF-α and IL-8 have partial anti-fibrotic effects in LX-2 cells, which oppose those of TGF-β1. Based on these views, we speculate that the proinflammatory effect of rSjE16 partially induces the blocking of HSC activation in LX-2 cells.
In conclusion, we showed that rSjE16 inhibited the expression levels of α-SMA and collagen I and promoted those of PPARγ, MMP-9, MMP-2, and the inflammatory factors IL-6 and IL-8 in LX-2 cells. Further study is needed to understand the complex relationship between inflammation and fibrosis in rSjE16-treated LX-2 cells and its underlying molecular mechanisms.
