Abstract
Objective
To identify genes associated with the clinicopathological features of colorectal cancer (CRC).
Methods
Gene expression profiles were downloaded and preprocessed by GEOquery and affy R packages, respectively. The limma package was applied to identify the differentially expressed genes (DEGs) in CRC. Gene Ontology and Kyoto Gene and Genome Encyclopedia (KEGG) pathway enrichment analyses for the DEGs were carried out using the clusterProfiler package. Protein–protein interaction (PPI) and weighted gene co-expression (WGC) networks were constructed using the STRING database and WGCNA package, respectively.
Results
A total of 523 DEGs (283 downregulated and 240 upregulated genes) in CRC tissues were identified. These DEGs were mainly enriched in 111 biological processes, 16 cellular components and 40 molecular functions, such as proteinaceous extracellular matrix, extracellular structure organization and chemokine-mediated signalling pathway. PPI and WGC networks showed that four upregulated genes (
Conclusions
The study provides new insights into understanding the pathogenesis of CRC. These identified genes may act as potential targets for CRC diagnosis and treatment.
Introduction
Colorectal cancer (CRC) is not only one of the most common malignancies, but also one of the leading causes of cancer-related death. 1 In recent decades, studies that have focused on the diagnosis, prognosis and treatment of CRC have made great progress, but the global burden of CRC is still increasing and more than two million new cases and one million deaths are expected by 2030. 2 Therefore, there is an urgent need to find effective ways to obtain new promising biomarkers and therapeutic targets for CRC.
With the emergence and wide application of microarray and RNA-sequencing technology, more and more gene expression data have been recently generated and deposited in publicly available databases such as Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA). Reanalysis of these data will contribute to a better understanding of the mechanism of the occurrence and development of diseases; and should lead to the identification of new disease-related molecules. For example, a previous study found several hub genes and pathways related to anaplastic thyroid carcinoma by mining public databases, including the
This current study identified the differentially expressed genes (DEGs) between CRC tissues and adjacent normal tissues via analysing microarray data in the GEO database. Integrated analysis of the DEGs based on bioinformatic methods was then undertaken, which aimed to identify the molecular mechanisms involved in CRC occurrence and development and to provide biomarker targets for future research.
Materials and methods
Microarray data
The gene expression profiles of 17 paired CRC and adjacent normal tissues (GSE110224) were obtained from the GEO database (https://www.ncbi.nlm.nih.gov/geo/) using the GEOquery package in R (www.r-project.org). All data were generated based on Affymetrix Human Genome U133 Plus 2.0 Array (GPL570). 6 Raw data were preprocessed by the affy R package with Robust Multichip Averaging (RMA) algorithm. The probeset IDs were converted into gene symbols using the annotation package hgu133plus2.db. If multiple probesets corresponded to the same gene, the mean value of those probesets expression was used.
Identification of the DEGs in CRC
The limma package of R (www.r-project.org) was used to identify the DEGs in CRC.
7
The
GO and KEGG pathway enrichment analyses
To further evaluate the functions of the DEGs and understand the biological processes (BPs), cellular components (CCs), molecular functions (MFs) and pathways closely related to CRC, the clusterProfiler package in R (www.r-project.org) was used to identify and visualize gene ontology (GO) terms and Kyoto Gene and Genome Encyclopedia (KEGG) pathways enriched by the DEGs. 8 The adj.P.Val <0.05 was set as the significant enrichment.
Construction of biological networks of the DEGs
In order to identify key genes involved in CRC, the protein–protein interaction (PPI) and weighted gene co-expression (WGC) networks of the DEGs were constructed by STRING database (www.string-db.org) and weighted gene co-expression network analysis (WGCNA) R package, 9 respectively. The minimum required interaction score in the PPI network was set as 0.7. In the WGC network, the soft-threshold power and the minimum threshold of interaction weight were set as 12 and 0.6, respectively. Subsequently, biological networks were visualized using Cytoscape software (https://cytoscape.org/). In the WGC network, the modules associated with colorectal cancer were identified by the MCODE plugin. 10 The key nodes (proteins/genes) in each biological network were selected based on degree centrality. The nodes with degree >23 and >6 were defined as key nodes in the PPI network and WGC network, respectively. Key nodes that overlapped in the two networks were screened out as key genes using a Venn diagram.
Validation of the expression levels of key genes
To validate the expression levels of key genes, a gene expression profiling interactive analysis (GEPIA) tool (http://gepia.cancer-pku.cn/) was used to explore the related data in TCGA and Genotype-Tissue Expression databases, and to analyse the expression levels of key genes in CRC tissues compared with normal tissues.
11
Furthermore, the UALCAN tool (http://ualcan.path.uab.edu/) was used to assess the expression of key genes in CRC based on individual cancer stages and histological subtypes.
12
A
Results
There were 523 DEGs in CRC tissues, including 240 upregulated and 283 downregulated genes (Figure 1). The

Volcano plots showing the upregulated and downregulated genes in colorectal cancer tissues. Blue dots represent significant mRNAs with an adjusted
The top five upregulated and downregulated genes in colorectal cancer tissues compared with adjacent normal tissues.

The GO analysis showed that these DEGs were significantly enriched in 111 BPs, 16 CCs and 40 MFs. The top five enriched CCs were proteinaceous extracellular matrix, extracellular matrix, microvillus membrane, apical part of the cell and endoplasmic reticulum lumen (Figure 2a). The top five enriched BPs were extracellular matrix organization, chemokine-mediated signalling pathway, positive regulation of response to external stimulus, extracellular structure organization and leukocyte chemotaxis (Figure 2b). The top five enriched MFs were cytokine activity, receptor ligand activity, CXCR chemokine receptor binding, chemokine activity and chemokine receptor binding (Figure 2c). Furthermore, KEGG pathway enrichment analysis indicated that these DEGs were mainly involved in cytokine–cytokine receptor interaction, interleukin (IL)-17 signalling pathway, bile secretion, nitrogen metabolism and chemokine signalling pathway (Figure 2d).
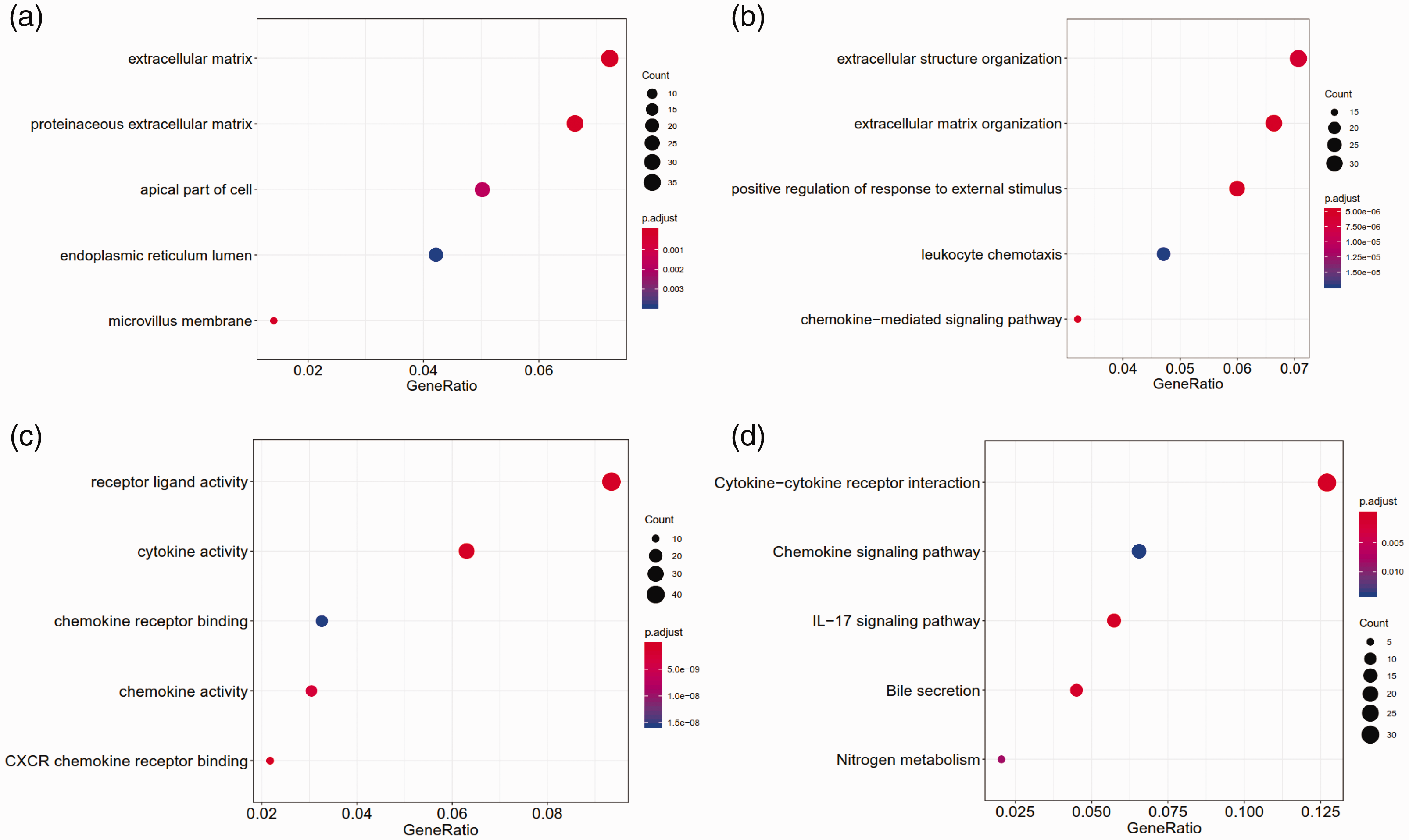
The top five gene ontology terms and Kyoto Gene and Genome Encyclopedia (KEGG) pathways enriched by the differentially expressed genes: (a) cellular components; (b) biological processes; (c) molecular functions; (d) KEGG pathways. IL-17, interleukin-17. The colour version of this figure is available at: http://imr.sagepub.com.
The WGC network showed 95 nodes and 161 edges (Figure 3a). Eight modules associated with colorectal cancer were identified in the WGC network (Table 2).

Biological networks and Venn diagram of the differentially expressed genes: (a) protein–protein interaction network; (b) the weighted gene co-expression network; (c) Venn diagram of key nodes that overlapped in two biological networks. The colour version of this figure is available at: http://imr.sagepub.com.
Identification of modules associated with colorectal cancer in the weighted gene co-expression network.

The GEPIA tool was applied to confirm the expression levels of key genes (
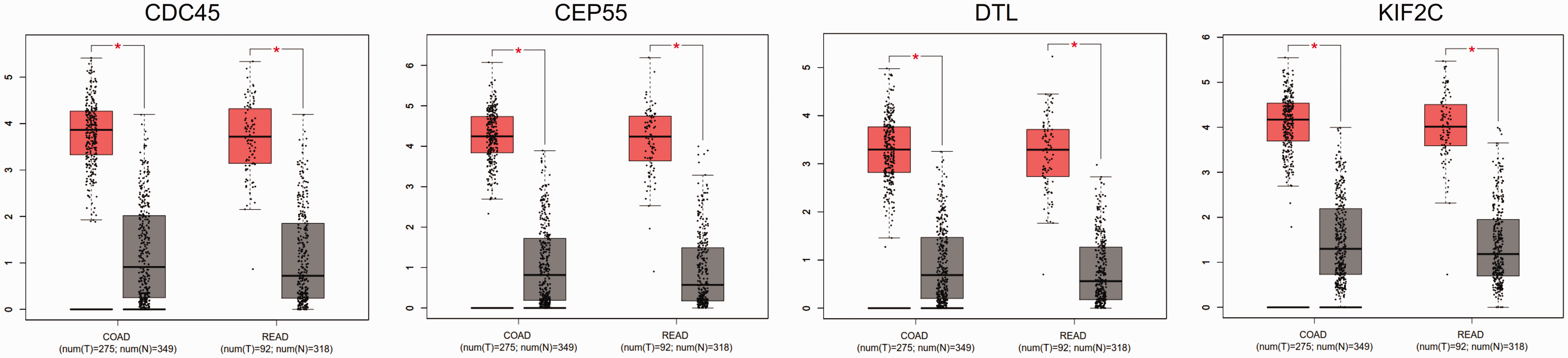
The expression of key genes in colorectal cancer and normal tissues. COAD, colon adenocarcinoma; READ, rectum adenocarcinoma. The pink and grey colours indicate tumour and normal tissues, respectively. *

The expression of key genes in colorectal cancer based on individual cancer stages. COAD, colon adenocarcinoma; READ, rectum adenocarcinoma. *
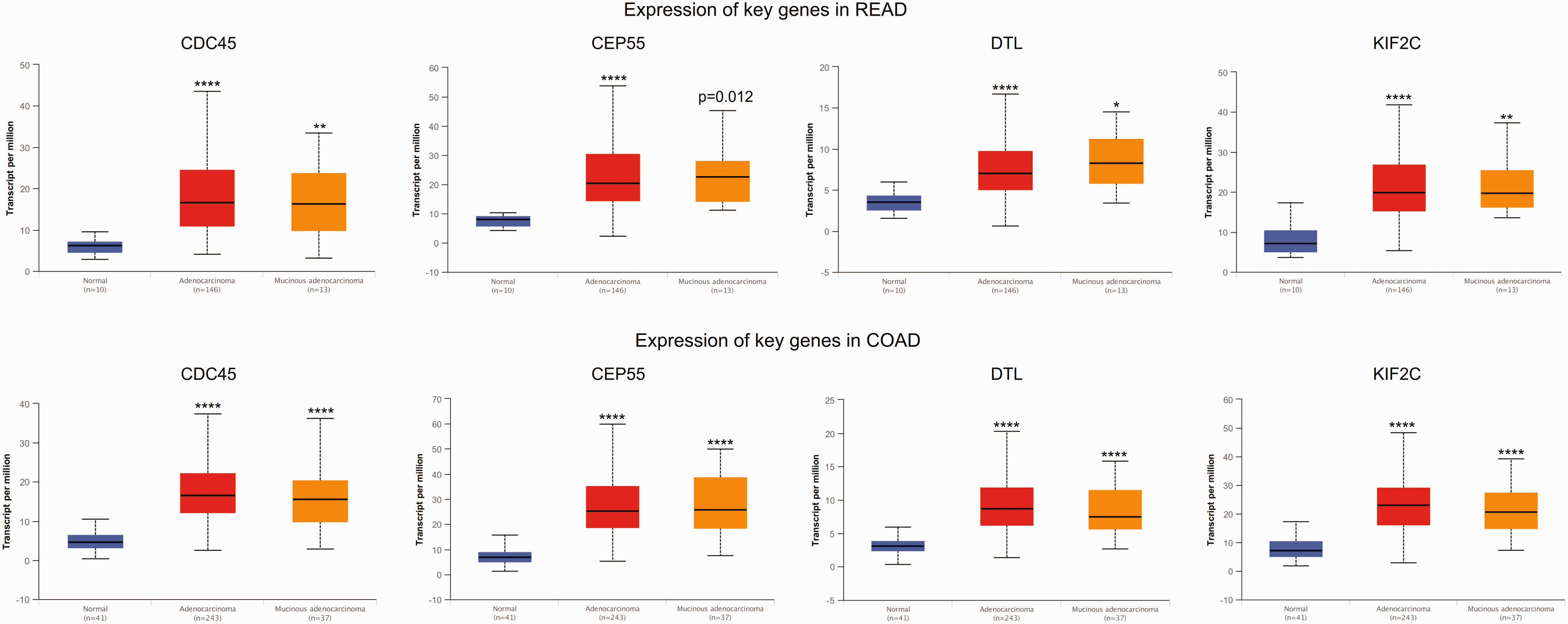
The expression of key genes in colorectal cancer based on histological subtypes. COAD, colon adenocarcinoma; READ, rectum adenocarcinoma. *

The expression of key genes in colorectal cancer based on sex. COAD, colon adenocarcinoma; READ, rectum adenocarcinoma. *

The expression of key genes in colorectal cancer based on age. COAD, colon adenocarcinoma; READ, rectum adenocarcinoma. *
Discussion
The present study investigated the DEGs in CRC based on 17 paired CRC and adjacent normal tissues from the GSE110224 dataset and found 523 DEGs, including 240 upregulated and 283 downregulated genes in CRC. Enrichment analyses indicated that these DEGs were significantly enriched in 111 BPs, 16 CCs and 40 MFs, such as proteinaceous extracellular matrix, leukocyte chemotaxis, cytokine activity and the chemokine signalling and IL-17 signalling pathways. Biological networks of the DEGs showed that the
The
In conclusion, the present study identified a series of CRC-related genes and pathways using bioinformatics analysis. Among them, the expression of the
