Abstract
H7N9 avian influenza virus (AIV) caused human infections in 2013 in China. Phylogenetic analyses indicate that H7N9 AIV is a novel reassortant strain with pandemic potential. We conducted a systemic review regarding virus-induced pathogenesis, vaccine development, and diagnosis of H7N9 AIV infection in humans. We followed PRISMA guidelines and searched PubMed, Web of Science, and Google Scholar to identify relevant articles published between January 2013 and December 2018. Pathogenesis data indicated that H7N9 AIV belongs to low pathogenic avian influenza, which is mostly asymptomatic in avian species; however, H7N9 induces high mortality in humans. Sporadic human infections have recently been reported, caused by highly pathogenic avian influenza viruses detected in poultry. H7N9 AIVs resistant to adamantine and oseltamivir cause severe human infection by rapidly inducing progressive acute community-acquired pneumonia, multiorgan dysfunction, and cytokine dysregulation; however, mechanisms via which the virus induces severe syndromes remain unclear. An H7N9 AIV vaccine is lacking; designs under evaluation include synthesized peptide, baculovirus-insect system, and virus-like particle vaccines. Molecular diagnosis of H7N9 AIVs is suggested over conventional assays, for biosafety reasons. Several advanced or modified diagnostic assays are under investigation and development. We summarized virus-induced pathogenesis, vaccine development, and current diagnostic assays in H7N9 AIVs.
Introduction
Influenza A is an enveloped virus belonging to the family Orthomyxoviridae that comprises a negative-sense segmented RNA genome. Influenza viruses are extremely variable owing to a lack of proofreading during genome replication; therefore, mutations are continuously accumulated.1–3 Antigenic drift and antigenic shift are ongoing processes that result in the existence of a large amount of influenza viruses whose natural reservoirs include wild waterfowl and shorebirds. 4 Reports indicate minimal evolution and apparent signs of disease in almost all natural reservoirs, but the occurrence of accumulated mutations in the genome or in certain fragments might benefit cross-species transmission.4–7 Influenza A viruses have the capacity to evolve rapidly and cause epidemics or even pandemics in domestic poultry, lower mammals, and humans. Avian influenza virus (AIV) transmission has been reported in several countries, with sporadic human infections owing to transfer of the virus from wild birds (e.g., migrating or wild aquatic birds) to domestic poultry.6,8 The first reported AIV infection in humans was caused by H5N1 virus and occurred in Hong Kong in 1997.6,9 Subsequently, AIVs have been continuously monitored and surveyed. Increasingly more new subtypes of AIVs have emerged in recent decades. 6
In February of 2013, a new atypical influenza virus, H7N9, was first reported in an 87-year-old male in Shanghai, China and the case was confirmed by the Chinese Center for Disease Control and Prevention on 29 March 2013.10–12 H7N9 is an emerging avian influenza A virus owing to genetic reassortment. 13 This virus was first identified as having low pathogenicity and was associated with asymptomatic or mild disease in poultry, causing sporadic human infections.12,14–17 A clinical survey of human infections with H7N9 viruses revealed characteristics of upper and lower respiratory tract illnesses that varied from mild to moderate (conjunctivitis or uncomplicated influenza-like illness) and could result in hospitalization.18–20 These novel H7N9 AIVs were reported to have gradually developed into highly pathogenic avian influenza (HPAI) viruses,21,22 indicating multiple increased polybasic amino acids at the hemagglutinin (HA) cleavage site (PEVPKRKRTAR/GL). 22 High mortality rates of about 50% are reported in human cases.18,22 Phylogenetic analysis of HA sequences in H7N9 isolates indicates that two major lineages of H7N9 have been established, including the Yangtze River Delta lineage and the Pearl River Delta lineage. 23 Reports indicate that the Yangtze River Delta lineage is broadly dispersed and was the original source of the H7N9 outbreaks in humans.23,24 China is currently facing its sixth epidemic of H7N9 human infection since 2018, and there has been a total of 1566 laboratory-confirmed human cases reported since 2013. 25 The World Health Organization (WHO) reports that three subtypes of AIVs are currently known to cause human infection, namely, H5, H7, and H9. 26 Among them, H5N1 and H7N9 have caused the most human infections. 27 This implies a challenge for global public health services in controlling emerging influenza viruses, even though the number of human infections with the H7N9 influenza virus has gradually decreased with treatment. 28 At present, there is no prophylactic vaccine against H7N9 AIV infection licensed for use in humans. The vaccine candidates currently under investigation include HA/NA peptide, plasmid-based, baculovirus-insect and mammalian protein expression system, and virus-like particle (VLP) vaccines.29–35 Some developed candidate vaccines are moving on to clinical trials. In addition, accurate and prompt diagnostic methods that have high sensitivity and specificity are essential for H7N9 influenza disease control and prevention. Several modified, advanced, conventional, and molecular diagnostic assays have been developed to enhance the capability of detecting H7N9 antigen or antibody.36–40 In this study, we conducted a systemic literature review of the emergent H7N9 AIVs in China, mainly focusing on pathogenesis, vaccine development, and diagnosis of human infection.
Methods
Literature retrieval and inclusion criteria
In this review, we summarized the latest information the pathogenesis of H7N9 influenza virus in humans, vaccine development, and current laboratory assays for the diagnosis of infection with H7N9 AIV. This review was conducted according to the PRISMA guidelines, using systematic methods to identify and select eligible articles from PubMed, Web of Science, and Google Scholar databases. We used search strings containing a combination of terms including H7N9 influenza virus, diagnosis, vaccine, and pathogenesis. The systemic search covered publication dates between January 2013 and December 2018. Search results were limited to articles published in English language. Articles related to pharmacodynamics, clinical trials, and meta‐analyses were excluded. Full‐text articles were reviewed to assess their relevance and quality of the methodology. Articles with the following were excluded: insufficiently detailed materials and methods; small sample size (n ≤ 20); irrelevant or insufficient information related to the review objectives; articles reporting H7N9 influenza and other microbes as a combined outcome, use of bioinformatics or mathematical models; and no quantitative data could be retrieved. We also searched gray literature such as WHO websites and announcements, and local ministries of health and centers for disease control and prevention. A flow chart of the systemic literature search and article selection is shown in Figure 1.

Flow chart of criteria and selection guidelines for literature included in this systemic review
Literature analysis
Two independent reviewers assessed the level of data quality in the selected articles. Disagreements were resolved by joint discussion and consensus
Ethics statement
In this study, we conducted a literature review regarding the current status of avian H7N9 influenza virus. Human clinical trials, case reports, and studies correlated to human ethical issues were excluded from this review. Ethics approval and informed consent were not required for this study.
Results
Pathogenesis of H7N9 influenza infection
Emergence of a novel H7N9 virus in China
H7N9 AIV first appeared in China in 2013 as a new atypical strain arising as a result of either mutations or genetic reassortment of whole genome segments, including neuraminidase (NA) genes from wild birds in Korea, HA genes from duck H7 viruses, and internal genes derived from two distinct lineages of H9N2 viruses.41,42 More specifically, the new H7N9 virus is derived from multiple and sequential reassortment among HxN9, H7N3, and H9N2 viruses where the H7 and N9 subtypes are from H7N3 and HxN9 AIVs, respectively, which were subsequently transmitted to and began circulating in poultry.41,43 The H7N9 virus is known to evolve at a rapid rate. Currently, several emerging genotypes of H7N9 AIVs have been identified, including cocirculating low pathogenic avian influenza (LPAI) and HPAI, causing infections in poultry and humans. Previous reports have indicated that in the first wave of the H7N9 influenza outbreak, a total of 27 genotypes were observed and documented between 2013 and 2014.12,44 The G0 genotype is predominantly found in human H7N9 infection whereas the G2, G4, and G5 genotypes are mainly detected in poultry. Most H7N9 genotypes were detected in the Yangtze River Delta area in China. More recent genotypic studies have identified dynamic reassortment of some internal genes among LPAI H7N9/H9N2/H6Ny and HPAI H7N9, as well as surface genes between the Yangtze River Delta and Pearl River Delta lineages, resulting in at least 36 genotypes identified in 2017.45,46 Among them, three main genotypes predominate, including G1 (A/chicken/Jiangsu/SC537/2013-like strain), G3 (A/chicken/Zhongshan/ZS/2017-like strain), and G11 (A/Anhui/40094/2015-like strain). 46
Most human cases of H7N9 AIV infection have been found in areas endemic for the virus.18,19,42,47 Closure of live poultry markets effectively controlled H7N9 epidemics in Shanghai, Nanjing, Hangzhou, and Huzhou during the first wave of the H7N9 outbreak in China, 48 as well as in Shenzhen, Guangzhou, Hangzhou, Foshan, and Ningbo during the second outbreak wave. 47 This suggests that cases of human H7N9 AIV infection resulted from direct contact with infected poultry. Although animal-to-human transmission is the main route of infection, some studies have reported the possibility of human-to-human transmission using clusters of human cases and tracing data; however, these studies found very low basic reproduction number (R) estimates.9,49–53 However, Kucharski et al. 43 reported finding no evidence of reduced human-to-human transmission in the two outbreak waves associated with the closure of live poultry markets, indicating that higher basic R estimates were reported for the human-to-human transmission component. Despite the high susceptibility and occurrence of transmission among household or family members, it has not been well demonstrated that the H7N9 virus is capable of human-to-human spread.54,55 Poultry exposure remains the main risk factor for human H7N9 infection.43,55,56 In addition, travelers also represent a potential risk factor in the dissemination of H7N9 AIVs from epidemic regions to non-epidemic areas.3,57–60 The WHO has suggested that the health status of international travelers returning from H7N9 epidemic areas should be assessed on arrival, and these individuals should be monitored or observed by health care workers for any apparent symptoms indicative of possible infection . 26 Although imported cases of human H7N9 infection have been identified, the opportunity for spread at community level is unlikely as the virus has not adapted to humans.3,57–60 Furthermore, health advisories and communications should warn travelers to avoid exposure to poultry and raw poultry products while visiting affected areas. In people with a history of travel to affected regions, screening should begin within 2 weeks of departure, prior to the onset of acute respiratory illness. Patients are encouraged to disclose recent travel histories, and health care workers should obtain information regarding travel to affected areas.26,52,61 Public health and laboratory partners are suggested to notify suspected cases during the diagnostic work-up so as to receive guidance in a timely manner. 61
Receptors and infection
The novel H7N9 influenza virus has been reported to be efficiently transmitted from avian species to humans.62–64 Previous studies isolated two distinct human H7N9 viruses, A/Shanghai/1/2013 and A/Anhui/1/2013, carrying Gln-226 and Leu-226 on the HA proteins, respectively, 65 which present different receptor-binding properties. The HA molecule of the A/Anhui/1/2013 strain has dual-receptor specificity, with the ability to bind to human-type α2,6-linked sialic acid and avian-type α2,3-linked sialic acid receptors.65,66 HA receptor binding affinity and specificity are required for efficient virus transmission between individuals and between species.67,68 Reducing the binding affinities of HA to avian receptors is a critical factor in human-to-human transmission. 66
The receptor-binding tropism of H7N9 changing to human receptors is possibly owing to the acquisition of two hydrophobic residues by substitution of Gln226Leu and Gly186Val. These substitutions were found in influenza pandemic viruses H2N2 and H3N2 in 1957 and 1968, respectively, as well as aerosol-transmissible H5N1 mutants. 68 The receptor binding properties of the human H7N9 virus are diverse, comprising different lineages of avian H7N9 isolates, but the cleavage activity of its NA is clearly low for human-type α2,6-linked sialic acid receptors. Regarding the pathogenicity of H7N9 viruses, disease caused by virus infection is usually asymptomatic and mortality is rare in chickens, ducks, and other birds. Similarly, various studies have demonstrated that H7N9 viruses replicate well in ferrets, 69 pigs,70,71 and mice,71,72 but are rarely lethal. Since it first emerged in China in 2013, there have been five reported outbreak waves of H7N9 AIV, with the majority caused by H7N9 LPAI strains. 3 More recently, HPAI H7N9 viruses have been isolated from humans in Guangdong, with high fatality rates.22,73 These viruses contain an insertion of basic amino acids at the HA proteolytic cleavage site.46,74 Highly pathogenic H7N9 human isolates were found to possess HA with a preference for human-type receptors and NA with inhibitor resistance;22,75 furthermore, these strains retained a series of genetic features contributing to human infection, such as 186V in the HA protein, according to H3 numbering.22,75 Sequence analysis has indicated that human HPAI H7N9 clinical isolates, such as A/Guangdong/17SF006/2017 and A/Taiwan/1/2017 possess mutations in the 627K domain of the PB2 subunit, conferring efficient replication in mammals. 22 In addition, amino acid substitutions associated with drug-resistance have been detected on NA in these HPAI H7N9 isolates. 75
Clinical symptoms
H7N9 AIV has jumped the species barrier, causing sporadic human infections since 2013. H7N9 human influenza virus A/Anhui/1/2013 has sporadically infected humans, with a high mortality rate of about 40%. However, earlier H7N9 isolates lack the multiple basic amino acids at the HA cleavage site that are usually present on highly pathogenic AIV strains.76,77 Clinical observations show that patients with confirmed H7N9 avian influenza present with high fever (≥39°C), productive or nonproductive cough, dyspnea, hypoxia, lower respiratory tract inflammation, shortness of breath, and infiltrates have also been noted on chest imaging.3,78 Complications in human cases of H7N9 AIV infection include respiratory failure, acute respiratory distress syndrome, septic shock, refractory hypoxemia, rhabdomyolysis, encephalopathy, acute renal dysfunction, multiple organ dysfunction, ventilator-associated pneumonia, and bacteremia.3,78,79 These features differ from those of previous human cases of infection with H7 subtype AIV, which generally presents as mild flu-like illness or conjunctivitis. 6 The three first human H7N9 viruses (A/Shanghai/1/2013, A/Shanghai/2/2013, and A/Anhui/1/2013) isolated from patients were found to be identical in all eight viral gene segments. However, A/Shanghai/1/2013 has been found to be phylogenetically distinct from the other two strains mentioned above. 80 Compared with human cases of infection with HPAI H7N9 (e.g., A/Guangdong/17SF003/2016 or A/Guangdong/17SF006/2017), most patients infected with A/Shanghai/1/2013 have viral pneumonia and some patients have displayed acute respiratory distress syndrome (ARDS), especially elderly people, who also have high fatality rates.22,81 Overall, the clinical presentation of HAPI H7N9 infection is similar to that of LPAI H7N9 infection; 22 however, fewer patients with HPAI than LPAI H7N9 infection have underlying conditions. 82
Host immune response against H7N9 influenza virus
The pathogenesis of human H7N9 virus that leads to severe outcomes remains unclear. Previous studies of HPAI H5N1 virus and pandemic (H1N1) 2009 virus have suggested that poor prognosis in human disease caused by these influenza viruses might be attributed to a cytokine storm or hypercytokinemia in the body.83–85 Some studies of H7N9 AIV have analyzed the cytokine and chemokine profiles in human serum samples from H7N9-infected patients. Reports have showed the presence of high levels of cytokines and chemokines, such as interleukin (IL)-6, IL-17, IL-2, monocyte chemotactic protein (MIP)-1, the monokine induced by gamma interferon (MIG), and macrophage inflammatory protein (MIP)-1β, in the serum of patients infected with H7N9 AIV.86,87 Moreover, in patients with severe H7N9 infection, the expressed levels of IL-6 and IP-10 molecules are significantly higher than in asymptomatic patients with H7N9 infection. 86 However, a previous study using BALB/c and C57BL/6 mouse models assessed the pathogenicity of H7N9 virus, to examine the association of proinflammatory cytokines in lung injury and viral clearance. The findings of that study showed that TNF-α and IFN-γ suppress viral gene expression and increase viral clearance whereas IL-6 and MCP-1 contribute to lung injury in A/H7N9-infected mice. 88 In summary, H7N9 viruses potently induce the expression of proinflammatory cytokines (IL-2, IL-6, and IL-7) and chemokines (IP-10), which contributes to the regulation of cellular immune responses. Alternatively, H7N9 infection also induces strong inflammatory responses, leading to tissue destruction.
Clinical observation indicates that patients with severe H7N9 AIV infection have high serum levels of IP-10 and IL-6, which may reflect the severity of disease. 86 In addition, the PB1-F2 protein of the influenza virus has been implicated in the regulation of polymerase activity, immunopathology, susceptibility to secondary bacterial infection, and the induction of apoptosis. 89 Studies suggest that the PB1-F2 protein may assist in NLRP3-associated hyperinflammation in human H7N9 influenza virus infections via the formation of inflammasome in host cells. New treatment strategies for H7N9 PB1-F2-induced lung inflammation and cellular influx specifically target NLRP3 inflammasome formation, which may provide effective therapy to reduce the disease severity level and mortality rate in pathogenic H7N9 infection.
Drug resistance
The newly emerged avian H7N9 influenza virus is known to cause human infection with high mortality. Antiviral drugs are critical tools for the control, treatment, and prevention of H7N9 virus infection as humans lack immunity against H7 subtype influenza viruses. Both M2 ion channel blockers (adamantanes, amantadine, and rimantadine) and NA inhibitors (oseltamivir, zanamivir, peramivir, and laninamivir) are used clinically as antiviral drugs for influenza infection in humans.90–92 Owing to the S31N mutation in the M2 protein, emerging H7N9 viruses are reported to be resistant to adamantine antiviral drugs, 93 which are therefore not recommended for clinical therapy. Oseltamivir, generally used as treatment for influenza infection, reduces the viral load in the respiratory tract of patients. Drug resistance to NA inhibitors is unusual among influenza viruses. However, some patients with H7N9 AIV infection have poor clinical outcome. The arginine to lysine substitution at residue R292K and E119V mutation in NA protein are linked with resistance to NA inhibitors, including oseltamivir and peramivir.90,94–97 Ribavirin is a well-characterized, broad-spectrum nucleoside inhibitor, which is used for the inhibition of viral RNA synthesis and capping of mRNA. Ribavirin also has antiviral activity against NA inhibitor-resistant H7N9 virus infections. 98 Human infections with avian H7N9 influenza virus mainly originate from contact with infected poultry. An effective strategy to minimize the risk of human H7N9 infections would be to administer aerosolized ribavirin to the nasal mucosa. Regarding drug resistance of HPAI H7N9 isolates, A/Guangdong/17SF003/2016(H7N9) strain has been reported to have NA-292R residue, similar to A/Anhui/1/2013 (H7N9), and sensitivity to all three NA inhibitors. 75 Baloxavir marboxil is a novel inhibitor against influenza virus cap-dependent endonuclease, which was approved by the US Food and Drug Administration in October 2018 to treat influenza A and B viruses in Japan and the United States. 99 This new drug might serve as a potential treatment for H7N9 AIV infections in the near future. We summarize some characteristics and features of H7N9 virus-induced pathogenesis in Table 1.
Summary of the characteristics of H7N9 avian influenza A virus in humans

Vaccine development
To reduce the mortality and morbidity rates of influenza virus infection, HA- and NA-specific neutralizing antibodies can be efficiently generated via vaccination. Early detection of neutralizing antibodies and rapid availability of the vaccine are required to prevent H7N9 pandemics. In the development of H7N9 vaccines, the peptides of HA7 and NA9 molecules of A/Shanghai/2/2013 or A/Anhui/1/2013 are considered the most promising candidates. These peptide vaccines are currently undergoing evaluation in clinical trials.
New technologies for vaccine production, such as cell culture-based antigen vaccine, live attenuated influenza vaccine (LAIV), DNA vaccine, phylogenetically related seed strain vaccine, recombinant baculovirus expression vaccine, and adjuvant-containing H7-based vaccine may be serve to increase vaccination capabilities.100–108 These technologies would speed up the production of H7N9 vaccines; however, a long time is likely required to produce large quantities of available vaccine. Work is ongoing in developing vaccines based on the HA and NA genes of A/Anhui/1/2013-like H7N9 viruses reassorted with the high-yield master virus A/Puerto Rico/8/34 (PR8) to enhance H7N9 virus growth capability in eggs, with high total viral and HA protein yields and attenuated pathogenesis, which has already resulted in several vaccine candidates.29,109 LAIVs have been tested against a pandemic influenza virus.110–112
Vaccination with a single dose of LAIV is known to be sufficient to stimulate an immune response against the homologous wild-type strain. Several human clinical trials are currently evaluating H7-based LAIVs.110,112–114 Previous outbreaks of A/mallard/Netherlands/12/2000 (H7N3) and A/Netherlands/219/2003 (H7N7) were owing to avian-derived H7 strains. These strains have been found to be phylogenetically related to the HA of H7N9 viruses. Several previous reports have shown that serum from individuals vaccinated with these vector-based or inactivated vaccines cross-reacted with H7N9.114–116 Baculovirus-insect cell expression vectors used in the rapid production of insect cell recombinant antigens is a practicable strategy in the development of HA-based vaccines. Other viral vectors (e.g., adenovirus and poxvirus) are also used to prepare vaccines. 31 The modified vaccinia virus Ankara and chimpanzee adenoviral ChAdOx1 vaccines are considered safe and immunogenic in humans for inducing long-term cross-reactive antibody against influenza A virus. These vaccines can be developed in a short time, to be used against emerging influenza strains. Moreover, an H7 and N9 molecule-containing vaccine, which is VLP-based from the A/Anhui/1/13 strain, has been reported to be producible by the baculovirus system. 107 A dose of this VLP vaccine administered twice together with an adjuvant significantly increases the neutralizing ability of HA- and NA-inhibiting antibody.32,107 The low-immunogenicity characteristics of H7-based vaccines have been reported in human 117 and animal models. 118 Currently, several clinical trials are investigating the effect of different adjuvants, including MF59 (Novartis), Matrix-M (Novavax), and AS03 (GlaxoSmithKline), to improve generation of protective antibodies.119–121 Regarding poultry vaccination against H7N9 AIVs, a series of inactivated vaccines has been used for the control and prevention of H7N9 outbreaks in poultry.100,122,123 For example, an H5/H7 bivalent inactivated influenza vaccine for chickens, ducks, and geese was introduced in September 2017;124,125 cases of H7N9 infections in human and poultry populations decreased after vaccination.126,127
Current diagnosis of H7N9 influenza virus infection
The first three human cases of infection with influenza H7N9 virus were reported in February and March 2013 in mainland China.42,80 Respiratory specimens from these patients were analyzed using real-time reverse transcription polymerase chain reaction (RT-PCR), viral culture, and sequencing analysis.128,129 Subsequently, cases were mostly confirmed using real time RT-PCR and in certain cases, viral isolation and serologic testing were also included. 42 The proper collection and handling of specimen contributes to the quality of test results. The most acceptable specimens for diagnosis or identification of influenza A (H7N9) virus are nasal wash, nasal aspirate, sputum, and nasopharyngeal swab specimens.36,130 Throat swabs are not recommended owing to the low viral load in these specimens; higher viral loads are found in sputum and tracheal samples. 131 The best time for sample collection is near symptom onset, within the first 3 days as viral shedding decreases after 4–5 days. However, shedding lasts up to 5 days in children.
Currently, several methods are used in the diagnosis of H7N9 AIV. Routinely, rapid influenza diagnostic tests (RIDTs) are used along with other confirmatory tests such as viral isolation, serologic tests, and RT-PCR. 130 RIDTs are widely used owing to their ease of use and quick results. RIDTs are based on antigen detection methods via the use of monoclonal antibodies, which attach to nucleoproteins on the targeted virus. 131 Compared with molecular methods, RIDTs have low sensitivity. 132 All RIDTs lack the capability of identifying or differentiating avian influenza subtypes. Because there are several different AIV subtypes, some more pathogenic than others, RIDTs that can differentiate between subtypes would greatly aid in screening, efficient diagnosis, surveillance, and control of HPAI.133,134 Such tests would also preclude the need for trained professionals to carry out testing. Although RIDTs only provide qualitative measures, the importance of obtaining a positive or negative result enhances efficient diagnosis, as positive samples can be sent for further analysis. 134
Viral isolation, considered the gold standard for influenza diagnosis, includes three different techniques: viral isolation using embryonated chicken eggs, viral culture, and shell viral culture. Viral isolation using embryonated eggs has some disadvantages, such as it is time consuming, has high biosafety level (BSL) requirements, and requires the use of eggs. 128 These disadvantages are drawbacks for efficient diagnosis; hence, viral isolation is more useful for subsequent viral analysis and, importantly, in the initial investigation of an outbreak. Molecular diagnostic tests such as real-time RT-PCR provide more rapid results for more efficient diagnosis. 128 Viral culture uses cell lines, which are infected, incubated, and monitored. Viral detection in cells can be done using different techniques, including specific antibody staining, hemadsorption, or immunofluorescence microscopy. In shell viral culture, cells are grown in 1 dram shell vials and stained with specific monoclonal antibodies for influenza detection. Serological tests include hemagglutination inhibition (HAI),135–137 virus neutralization (VN),136,138 complement fixation,139,140 single radial hemolysis tests (SHR),136,137,141 and enzyme-linked immunosorbent assay (ELISA).128,136,142–144 ELISA, one of the most commonly performed laboratory assays, uses mono- and polyclonal antibodies against nucleoproteins on the influenza virus. However, the sensitivity of ELISA is lower than that of molecular testing.128,132 HAI is a common assay based on HA-specific antibody that prevents influenza virus attachment to red blood cells, but HA has low sensitivity.128,136,138,145 VN assays require the use of infectious viruses, which are then neutralized with virus-specific antibodies. VN is more sensitive than HAI, but BSL2 or BSL3 laboratory conditions are required. The SHR assay measures complement-mediated hemolysis via antigen–antibody complex induction; this assay is more sensitive than HAI.128,132
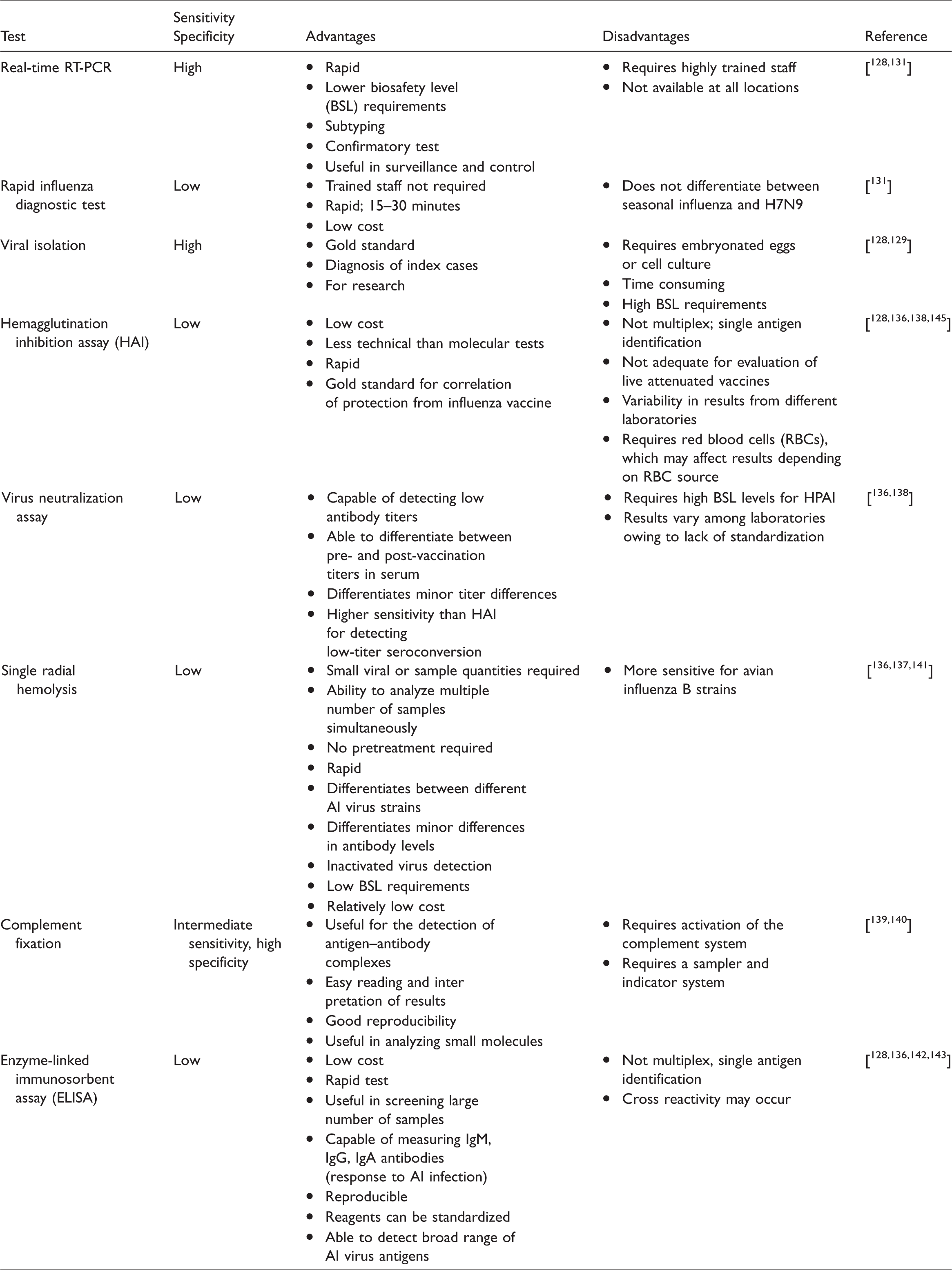
Molecular PCR-based diagnostic tests are commonly performed in the detection and subtyping of H7N9 AIVs as these are faster than viral isolation and more sensitive than serologic tests. PCR-based diagnostic tests require RNA extraction using reagents to inactivate the virus; hence, the BSL requirements are lower than for viral isolation techniques. The extracted RNA is transcribed with reverse transcriptase to complementary DNA (cDNA), then amplified using specific primers for each subtype. With advanced technology, primers are now able to detect avian influenza A even if the virus is genetically varied. 128 A recent advance in PCR-based detection methods for H7N9 AIV is the multiplex RT-PCR assay, which uses the GenomeLab Gene Expression Profiler (GeXP) analyzer, a multiplex gene expression analysis platform developed by Beckman Coulter.146,147 This assay simultaneously detects four NA serotypes of the H5 subtype. Both sensitivity and specificity are reportedly high; hence, this method can be useful in the detection, surveillance, and control of HPAI viruses. 148 Detection techniques such as immunochromatographic RIDTs and PCR-based techniques greatly contribute to rapid, simple, and efficient screening and confirmation of cases of HPAI H7N9 infection. PCR-based methods further aid in the surveillance and control of HPAI H7N9 infection as this method has the advantages of speed, sensitivity, and specificity. We summarize the main tests performed in the diagnosis or confirmation of H7N9 virus in Table 2.
Main tests for diagnosis and/or confirmation of H7N9 avian influenza A virus (AIV)
Discussion
Herein, we conducted a systemic literature review for the novel emerged H7N9 AIVs with respect to pathogenesis, vaccine development, and current diagnosis. Although current major H7N9 epidemics are in China, this reassorted virus has gradually evolved from LPAI to HAPI. Currently, HPAI H7N9 AIV has the capability of causing severe syndromes and inducing high mortality in both avians and humans. The WHO has also pointed out that among current AIVs, those that cause human infection are H5N1 and H7N9, thereby necessitating the continuous monitoring of genetic and antigenic change and variation in H7N9 AIVs. 26 Understanding the pathogenesis of the virus, developing prophylactic vaccines, and enhancing the sensitivity and specificity of H7N9 AIV diagnostic methods are urgently required.
As an atypical strain with mutations and genetic reassortment, emergence of the novel H7N9 AIVs in China raises concerns regarding new vaccine development and drug resistance. The influence of animal infections on human disease propagation places a large burden on health systems and animal trade controls to prevent the virus spreading via travelers and animal movement. Although AIVs usually do not infect people, rare cases of human infection with these viruses have been reported. 53 Regarding the transmission route, H7N9 AIVs have been found to cause human infection, mainly from poultry to humans. Infected birds can shed H7N9 AIVs in secretions like saliva, mucous, and feces, which can lead to human infection via mucosal surfaces of the eyes, nose, and mouth or through inhalation of droplets and dust containing the virus. Unprotected exposure to H7N9 AIVs from infected birds is currently considered the main risk factor and transmission route. The role of poultry farms, live animal markets, and retailers might be key in the spread of disease, as well as inadequate biosafety measures and poor handling of sick or dead animals. However, some laboratory-confirmed human cases lack evidence of a history of direct contact with animals, indicating the possibility of human-to-human transmission.53,149 To date, several studies have predicted and compared the route of poultry-to-human or human-to-human transmitted efficiency of H7N9 AIVs using the reproduction number (R);43,56,120,140,150 The R is a key indicator of transmission intensity estimated by fitting the number of poultry-to-human and from human-to-human infections into mathematical models. The data of these studies showed that the estimated R value for an H7N9 outbreak was below the epidemic threshold required for sustained human-to-human transmission, indicating that an H7N9 human-to-human pandemic is unlikely to occur and most cases of H7N9 AIV infection are owing to poultry-to-human transmission. Virlogeux et al. 149 recently used an ecological model to evaluate animal-to-human and human-to-human transmission of H7N9 during the first three epidemic waves in China; the authors indicated that there was a significant decrease in the incidence of H7N9 cases after live poultry market closures in several major cities of China. Accordingly, it can be concluded that although H7N9 human-to-human transmission might be possible, the major transmission route of H7N9 AIVs remains via poultry-to-human.
Regarding virus-induced pathogenesis and pathology, H7N9 AIV infection causes mild to severe syndromes to humans owing to either LPAI or HPAI strains, as compared with infections in poultry, which are often asymptomatic when infection is with LPAI H7N9 strains. The major element for distinguishing HPAI and LPAI is determination of the amino acid profiles at the HA cleavage site. Increased basic amino acids have been proven to enhance the capability of maturation and replication of AIVs. The mechanism through which LPAI H7N9 viruses induce different pathogenesis in birds and humans is still not fully understood, suggesting that to determine the virulence of influenza A virus, it is critical to check the occurrence of variations or mutations in other viral proteins that are involved in virus replication and dissemination. Reports have indicated that H7N9 Asian lineage and H5N1 HPAI Asian lineage strains are mainly responsible for most cases of human illness worldwide to date, and these also result in the most serious illnesses and highest mortality. Clinical observation reports indicate that severe symptoms including viral pneumonia, hypercytokinemia, and even ARDS in cases of death owing to H7N9 AIV infection, which mainly occur in older people, as compared with H5N1 infection in which severe symptoms generally occur in younger patients.9,81,151,152
In addition, comparative cohort studies have indicated that the sex ratio is much higher in urban than rural areas in infections with both H7N9 and H5N1 viruses.9,153,154 The reason might be correlated with sex-based differences in exposure to AIVs rather than to differences in immunity.9,153 For instance, higher numbers of cases of H7N9 infection have been reported among men in Shanghai city, where men have a greater tendency than women to have more frequent retail exposure to live poultry. Other possible reasons for this difference have been reported, including worse prognosis for infected men than infected women, and differences in health-seeking behaviors. 153 However, some reports indicate that differences in immunity between men and women remain a consideration. 155 Recently, Chen et al. 156 reported that an increasing trend of H7N9 human infections in rural areas of Zhejiang Province in China during 2013–2017 was the result of rural live poultry markets that remained open despite the epidemic; this suggests that closing live poultry markets is an effective way to control H7N9 human infections.
In this study, we report that human patients with H7N9 AIV infection develop severe illness, dysregulation of the cytokine and chemokine response, late viral clearance, and impaired immunity. 157 Induction of pro-inflammatory cytokines by H7N9 is the predominant factor leading to hypercytokinemia, an important risk factor triggering multiple organ damage, thereby increasing the mortality rate in human cases of infection with H7N9 AIV.81,152 Reports have indicated that peripheral blood mononuclear cells have greater susceptibility to H7N9 than H5N1 virus, and H7N9 could more efficiently infect B and T lymphocytes than H5N1 (p<0.01). 158 Another study indicated that hypercytokinemia is associated with interferon-induced transmembrane protein-3 dysfunction, which could be useful in the prediction of H7N9-induced mortality. 159 In addition, H7N9 AIVs show greater tropism for the respiratory epithelium as well as nasopharynx-associated lymphoid tissues than H5N1 viruses. 152 Similar results have been found in H7N9 virus-challenged mice and ferret models.152,160 Since the end of 2017, a new H7N9 AIV (A/Guangdong/17SF006/2017) has been identified as an HPAI strain in China. This HPAI H7N9 virus was generated from its LPAI counterpart22,46,74 and has high affinity for both human-type α2-6-linked sialic acid and avian-type α2-3-linked sialic acid receptors.65,66 These characteristics demonstrate the capability of H7N9 viruses for adaptation and binding, thus opening the door for the virus to develop drug resistance, thereby presenting a challenge for drug and vaccine development as the clinical symptoms of these mutants differ from the original H7N9 virus, including respiratory distress syndrome, septic shock, acute renal dysfunction, and multiple organ dysfunction.3,78,79 These symptoms may be related to the presence of IP-10 and IL-6 in patients with severe H7N9-induced disease, as these induce an acute inflammatory response and extensive tissue destruction. 86
Regarding drug use and design, targeting NLRP3 inflammosome formation may be effective in reducing the severity and mortality of H7N9 AIV infection, as humans lack immunity against H7 subtype influenza viruses. Some patients with H7N9 infection have shown poor outcomes with therapies such as M2 ion channel or NA blockers. 94 In addition, some reports have indicated that H7N9 viruses acquire drug resistance against NA inhibitors via substitutions, which is correlated with amino acid substitutions; R292K confers the highest increase in oseltamivir half-maximal inhibitory concentration (IC50), E119D confers the highest IC50 of zanamivir, and H274Y confers highly reduced inhibition by oseltamivir. 161 The need for new anti-influenza viral drugs is urgent. Testing some alternatives in H7N9 treatment is advisable, such as intranasal administration of ribavirin and baloxavir marboxil, as these have demonstrated effectiveness with other influenza viruses.98,99
In prevention and prophylaxis of H7N9 AIVs, it is necessary to develop vaccines that can easily induce neutralizing antibody and generate CD8+ cytotoxic T cells for virus neutralization and clearance, respectively. LAIV, H7 and N9 molecule-containing vaccine, and recombinant VLP vaccines based on A/Anhui/1/13 strain have shown effective stimulation of the specific immune response and no cross-reaction with other virus strains.100,162 Clinical trials and animal studies have revealed that much remains to be accomplished in the development of new vaccines, and their efficacy relies on targeting different phylogenetically related genotypic strains of H7N9 AIV. 108 Human contact with infected poultry and global travel is unlikely to diminish, and the pathogenesis of H7N9 AIVs will likely persist. Therefore, strategies should be focused on understanding more about the impact of new therapies and prophylaxis to control circulating viruses.
Diagnostic tools or platforms with high sensitivity and specificity for detection of viral antigens or antibodies of H7N9 AIVs have been widely developed. Although viral culture is conventionally viewed as the gold standard in virological assays, real-time RT-PCR is currently suggested as an alternative gold standard as the former method requires specific BSL laboratory conditions and experienced staff. However, RIDTs are quick and easy to perform, so these tests have become essential at sites where real-time RT-PCR is unavailable. Current RIDTs have the capability of detecting strains of both influenza A and B. However, they have low sensitivity with new strains and are unable to differentiate between subtypes; therefore, the rate of false negatives is higher. 131 RIDTs provide rapid detection of the presence or absence of viral analytes in patient samples; a major advantage of RIDTs is that results can be obtained within 15–30 minutes. 131 As mentioned, there are several different subtypes of H7N9 AIVs circulating in China, some more pathogenic than others. With the advantage of speedy results, RIDTs that can differentiate between subtypes would provide a breakthrough for rapid screening and diagnosis, especially of HPAI H7N9 and AIVs such as H5N1. 134 RIDTs can be performed at points-of-care for first-line influenza virus screening. 134 This would help in patient diagnosis as well as the detection of asymptomatic cases or mild symptom cases, thus preventing further spread. 163 In addition, several advanced or modified assays and platforms for detecting H7N9 AIVs are under investigation, such as modified surface plasmon resonance, 39 enzyme-induced metallization, 164 reverse transcription-loop-mediated isothermal amplification assay, 165 and chip-based biosensors. 166
This study has some limitations. We selected articles written in English only; in addition, we excluded small cohort studies and clinical trials and focused mainly on study results in human cases of AIV infection. These selection criteria might have resulted in the loss of certain important information and experimental results published in Chinese or other non-English languages, as well as some important observations from small cohorts and animal studies.
Conclusion
Currently, H7N9 influenza virus is disseminated among aquatic birds and poultry, increasing their adaption and virulence in humans. Although this virus primarily exists in mainland China, increasingly more imported cases have been reported in other countries. Clinical H7N9 AIV isolates have been proven to have resistant to adamantine and oseltamivir, which are commonly used in influenza therapy. This highlights that new antiviral drugs, continuous genotype identification, and new vaccines against H7N9 influenza viruses are urgently needed. In addition, the development of accurate and rapid testing methods is necessary for the surveillance and control of H7N9 virus infections.
Footnotes
Author’s contribution
WHW prepared the manuscript. EME, MRCI, and CYL helped to prepare the draft. EME and MRCI assisted in language editing. WA and AT revised the draft. SFW conceived the study and revised the draft.
Acknowledgements
The authors wish to thank the staff from Kaohsiung Medical University Hospital and Taiwan CDC for their assistance in providing essential and meaningful information of current H7N9 influenza virus.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by grants from the Center for Infectious Disease and Cancer Research, Kaohsiung Medical University (KMUTP105E00 and KMUTP 105E04), a grant from Higher Education Sprout Project - Ministry of Education, Taiwan (TB0357), and grants from the Ministry of Science and Technology, R.O.C. (108-2918-I-037-001).
