Abstract
Objective
To investigate the relationship between the interleukin 10 (IL10) gene single nucleotide polymorphisms (SNP) −1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) and risk of type 2 diabetes mellitus in a Chinese population.
Methods
This case–control study recruited patients with type 2 diabetes mellitus and healthy control subjects. Genotyping of the −1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) SNPs was conducted and genotype frequencies were compared between the two groups.
Results
The study recruited 364 patients with type 2 diabetes mellitus and 677 healthy controls. Patients carrying the −1082 GG genotype had a significantly increased risk of type 2 diabetes mellitus (adjusted odds ratio [OR] 1.57, 95% confidence interval [CI] 1.03, 2.68), as did those patients carrying the −592 AA genotype (adjusted OR 1.63, 95% CI 1.06, 2.53). Subjects carrying both the −1082 GA + GG and −592 AC + AA genotypes had a significantly increased risk of type 2 diabetes mellitus (adjusted OR 2.03, 95% CI 1.24, 3.15).
Conclusions
The SNPs −1082G/A and −592 A/C increased the risk for type 2 diabetes mellitus, and could be potential targets for screening for the early detection of the risk of type 2 diabetes mellitus.
Introduction
Type 2 diabetes mellitus is a common chronic disease, and its complications have become a major cause of morbidity, mortality and disability in both developed and developing countries. 1 There were approximately 285 million people worldwide living with diabetes in 2010, 9.2 million of them in China. 1 It is estimated that the number of patients with type 2 diabetes mellitus worldwide will reach 300 million by 2025. 1 Although type 2 diabetes mellitus is induced by various environmental and genetic factors, its detailed aetiology is not yet fully understood. 2 Several studies have reported that multiple cytokines may be involved in the regulation of the immune response, which has in turn been shown to play a role in the pathogenesis of type 2 diabetes mellitus.3,4
Interleukins are a diverse constellation of small cell signalling protein molecules, or cytokines, that regulate the function of the immune system in humans.5,6 Interleukins are produced predominantly by T cells, monocytes, macrophages and endothelial cells. 7 Their functions include facilitating communication among immune cells, controlling genes, regulating transcription factors, and governing inflammation, differentiation, proliferation and the secretion of antibodies. 8 Genetic polymorphisms of interleukins play a critical role in their activity, and could alter cytokine function and dysregulate their expression. 9 Therefore, individual genetic differences may be closely related to the development of type 2 diabetes mellitus.
Interleukin 10 (IL-10) is a multifunctional regulatory cytokine involved in the inflammatory response that functions as a general inhibitor of the proliferative and cytokine response of both type 1 and type 2 helper T cells. 10 The gene encoding IL-10, IL10, is located on chromosome 1 (1q31–1q32).11,12 Three functional promoter single nucleotide polymorphisms (SNPs) in the IL10 locus at –1082G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) from the transcriptional start site have been confirmed, and there are indications that they influence IL10 gene transcription. 13 One meta-analysis found there was an association between the IL10 gene −1082G/A polymorphism and type 2 diabetes mellitus, but no association was found for −819 T/C or −592 A/C. 14 However, the results of various studies have been inconsistent.4,14 The aim of the present case–control study was to investigate the relationship between SNPs of the IL10 gene (−1082 G/A [rs1800896], −819 T/C [rs1800871] and −592 A/C [rs1800872]) and the risk of type 2 diabetes mellitus in a Chinese population.
Patients and methods
Study population
This case–control study recruited consecutive patients diagnosed with type 2 diabetes mellitus according to World Health Organization criteria who were screened at the Department of Endocrinology, First Affiliated Hospital of the People’s Liberation Army, Beijing, China between January 2010 and November 2012. 15 Patients with acute or chronic inflammatory disease, infections, cancer, or end-stage liver or kidney diseases were excluded from the study. Healthy control subjects were recruited from a health examination centre of the First Affiliated Hospital of the People’s Liberation Army during the same period, and were excluded if they had a history of acute or chronic inflammatory disease, infections, cancer, end-stage liver or kidney disease, coronary artery disease, or other metabolic disorders. A ‘super control group’ (age >75 years) was selected from the control group.
Baseline clinical and demographic data for all study participants were collected from the medical records. Body mass index (BMI) was calculated as weight (kg)/height2 (m2). Venous blood samples (5 ml) were obtained from all study participants after an 8-h fast. The blood samples were clotted at room temperature for 15 min and then serum was obtained by centrifugation at 2000
The Ethics Committee of the First Affiliated Hospital of the People’s Liberation Army reviewed and approved the study (no. 201001037). All participants provided written informed consent.
Genotyping
All study participants provided 5 ml of venous blood, with 0.5 mg/ml ethylenediaminetetra-acetic acid used as the anticoagulant. The blood samples were kept at –20℃ until use, Genomic DNA was extracted using the TIANamp Blood DNA Kit (Tiangen, Beijing, China) according to the manufacturer’s instructions. The genotyping of the IL10 gene SNPs −1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) was conducted in a 384-well plate format on the MassARRAY® Analyzer 4 system (Sequenom®, San Diego, CA, USA), which combines polymerase chain reaction (PCR) and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry technologies. The primers used for the IL10 gene SNPs −1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) for PCR amplification were designed using Sequenom® Assay Design version 3.1 software (Sequenom®), according to the manufacturer’s instructions. For –1082 G/A (rs1800896), the forward and reverse primer sequences were 5′-AGAAGTCCTGATGTCACTGC-3′ and 5′-AAGTCAGGATTCCATGGAG-3′, respectively. The forward and reverse primer sequences for −819 T/C (rs1800871) were 5′-ATGGTGTACAGTAGGGTGAG-3′ and 5′-TTTCCACCTCTTCAGCTGTC-3′, respectively. The forward and reverse primer sequences for −592 A/C (rs1800872) were 5′-AAGAGGTGGAAACATGTGCC-3′ and 5′-TACCCAAGACTTCTCCTTGC-3′, respectively. Each PCR reaction mix comprised 50 ng genomic DNA, 200 µM dNTP, 2.5 U Taq DNA polymerase (Promega, Madison, WI, USA) and 200 μM primers, in a total volume of 20 µl. The cycling programme involved preliminary denaturation at 94℃ for 2 min, followed by 35 cycles of denaturation at 94℃ for 30 s and annealing at 64℃ for 30 s, with a final extension at 72℃ for 3 min. The PCR products were verified by 1.0% agarose gel electrophoresis and visualized using ethidium bromide staining and ultraviolet light. The PCR products of rs1800896, rs1800871 and rs1800872 were 269 base pairs (bp), 238 bp and 300 bp, respectively. Reproducibility was verified by repeat analysis of a randomly chosen subgroup of 10% of the subjects.
Statistical analyses
All statistical analyses were performed using the SPSS® statistical package, version 11.0 (SPSS Inc., Chicago, IL, USA) for Windows®. Continuous and categorical variables were expressed as mean ± SD and n (%) of study participants, respectively. Categorical variables from patients and control subjects were compared using the χ2-test and continuous variables were compared using the Student’s t-test. The Hardy–Weinberg equilibriums between groups were compared using the χ2-test. Unconditional logistic regression was conducted to assess the effects of the IL10 gene SNPs –1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) on the risk of type 2 diabetes mellitus in co-dominant, dominant and recessive models, with results expressed as odds ratios (OR) and corresponding 95% confidence intervals (CI). All P-values were two sided and a P-value < 0.05 was considered statistically significant.
Results
Clinical and demographic characteristics of patients with type 2 diabetes mellitus and healthy control subjects who were included in a study investigating the association between single nucleotide polymorphisms of the interleukin 10 gene and the risk of type 2 diabetes mellitus.

Data presented as mean ± SD or n (%) of study participants.
Comparisons between patients and control subjects were made using the Student’s t-test for continuous variables and χ2-test for categorical variables.
BMI, body mass index; HbA1c, glycosylated haemoglobin; TC, total cholesterol; TG, triglycerides; HDL-C, high-density-lipoprotein cholesterol; LDL-C, low-density-lipoprotein cholesterol; NS, no statistically significant difference (P ≥ 0.05).
Genotype frequencies of three single nucleotide polymorphisms (SNP) of the interleukin 10 (IL10) gene in patients with type 2 diabetes mellitus and healthy control subjects.

Data presented as n (%) of study participants.
The Hardy–Weinberg equilibriums between groups were compared using the χ2-test. Unconditional logistic regression was conducted to assess the effects of the IL10 gene SNPs −1082 G/A (rs1800896), −819 T/C (rs1800871) and −592 A/C (rs1800872) on risk of type 2 diabetes mellitus, with results expressed as odds ratios (OR) and corresponding 95% confidence intervals (CI).
Adjusted for age, BMI, fasting glucose, fasting insulin, HbA1c, TC, TG, HDL-C and LDL-C.
BMI, body mass index; HbA1c, glycosylated haemoglobin; TC, total cholesterol; TG, triglycerides; HDL-C, high-density-lipoprotein cholesterol; LDL-C, low-density-lipoprotein cholesterol; NS, no statistically significant difference (P ≥ 0.05).
Interaction between two single nucleotide polymorphisms (SNP) of the interleukin 10 (IL10) gene in patients with type 2 diabetes mellitus and healthy control subjects.
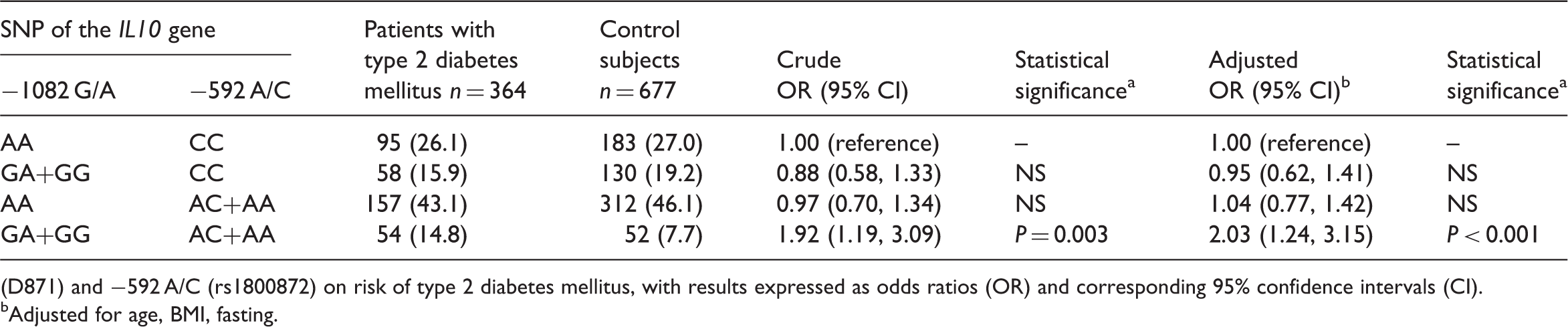
(D871) and −592 A/C (rs1800872) on risk of type 2 diabetes mellitus, with results expressed as odds ratios (OR) and corresponding 95% confidence intervals (CI).
Adjusted for age, BMI, fasting.
Discussion
Interleukin 10 is an important immunoregulatory cytokine that is produced by activated T cells, monocytes, B cells and thymocytes. 10 It plays an important role in stimulating and suppressing the immune response, and functions as an immune response modulator.16–18 Previous studies have reported several polymorphic sites in the IL10 gene promoter region upstream of the transcription start site, including three polymorphisms at positions −1082 A/G, −819 C/T and −592 A/C. 13 Some previous studies reported that these IL10 gene promoter polymorphisms were associated with the risk of type 2 diabetes mellitus.19–21 This present case–control study investigated the association between genetic polymorphisms of the IL10 gene promoter region and the risk of type 2 diabetes mellitus. The main findings of the present study were that the −1082G/A and −592 A/C variants carried a significantly increased risk of type 2 diabetes mellitus, while the −1082 GA + GG and −592 AC + AA genotypes when carried together had a synergistic effect on the risk of type 2 diabetes mellitus. However, the −819 T/C gene polymorphism did not appear to be associated with the risk of type 2 diabetes mellitus.
Previous studies have found an association between IL10 genetic variants and the risk of type 2 diabetes mellitus.10,19–22 These current findings are consistent with several studies.23,24 One study conducted in India indicated that the IL10 gene −1082 A/G genotype significantly increased the risk of type 2 diabetes mellitus. 23 Another study from Iran found that the IL10 gene polymorphism −592 A/C was associated with immune diseases, including type 2 diabetes mellitus with and without nephropathy. 24 A recent large meta-analysis pooled ten case–control studies and concluded that the IL10 −1082 A/G polymorphism was strongly associated with the risk of type 2 diabetes mellitus, but no association was found between the −819 C/T or −592 A/C polymorphisms and the risk of type 2 diabetes mellitus. 4 However, results have been inconsistent. For example, several studies reported no significant association between the −1082 G/A and −592 A/C variants and the risk of type 2 diabetes mellitus.4,25 Since these studies were conducted in different populations, it is difficult to make direct comparisons between them. Discrepancies may be due to differences in variant frequencies among various ethnic groups, and IL10 gene polymorphisms may play different roles in the development of type 2 diabetes mellitus depending on the population.
This present study had a number of limitations. First, it was conducted in a single hospital and the participants may not have been representative of other areas within China. Secondly, the aetiology of type 2 diabetes mellitus involves multiple genes and environmental factors, so other genetic and environmental factors should be considered in further studies. Secondly, the genotype distributions of −1082 G/A and −819 T/C in both the case and control subjects were not consistent with Hardy–Weinberg equilibriums, which suggests the sample of this study might not represent the Chinese population. The main reason might be the relative small sample size. Therefore, further studies with large sample sizes are needed to confirm our results.
In conclusion, the findings of this present study suggest that the IL10 −1082 G/A and −592 A/C gene polymorphisms increase the risk for type 2 diabetes mellitus, while carrying the combination of the −1082 GA + GG and −592 AC + AA genotypes had a synergistic effect on the risk of type 2 diabetes mellitus in a Chinese population. However, the −819 T/C polymorphism was not associated with the risk of type 2 diabetes mellitus. These results suggest that these polymorphisms could be potential targets for screening for the early detection of the risk of type 2 diabetes mellitus.
Footnotes
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
