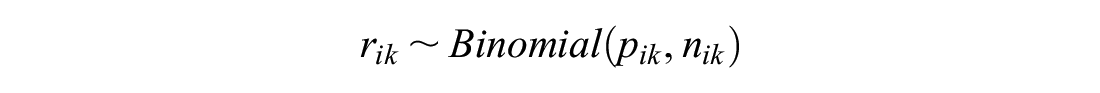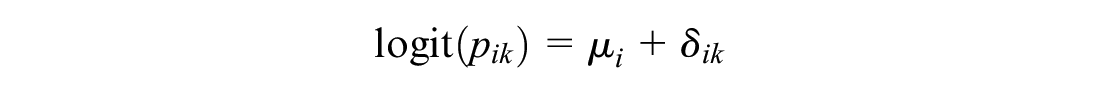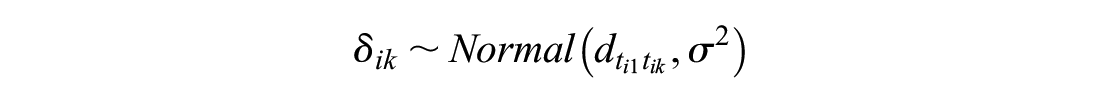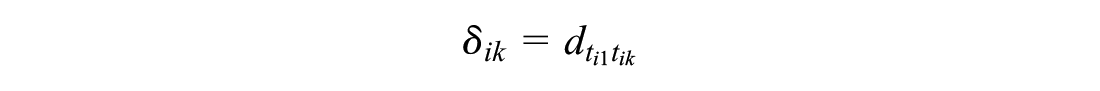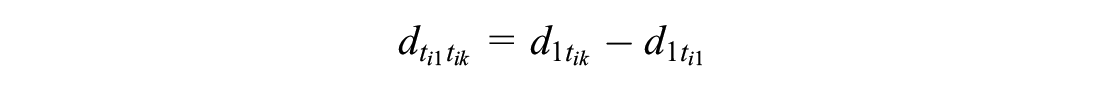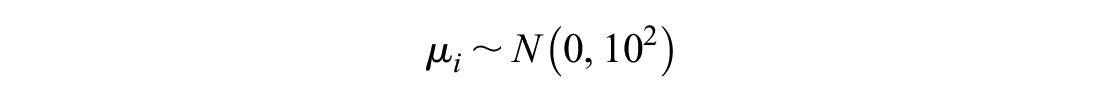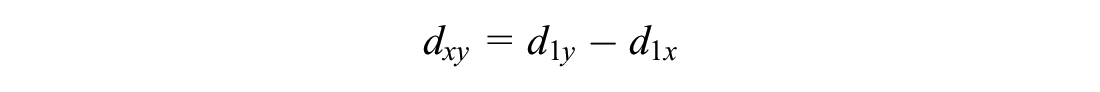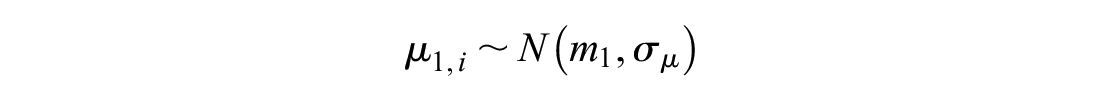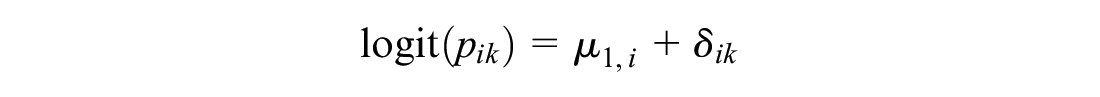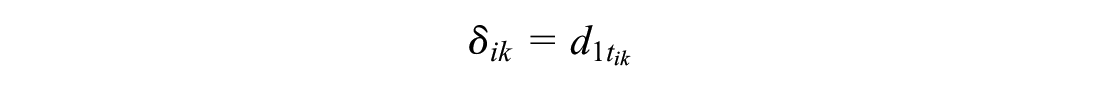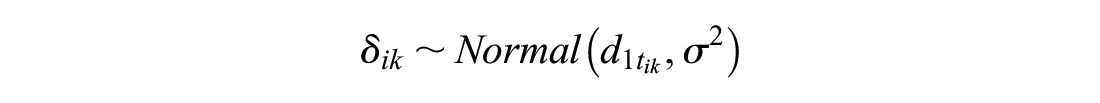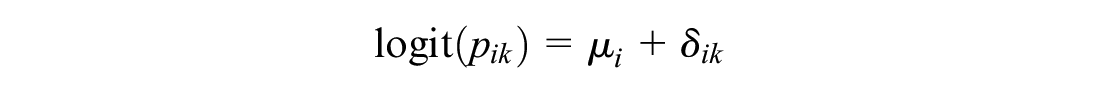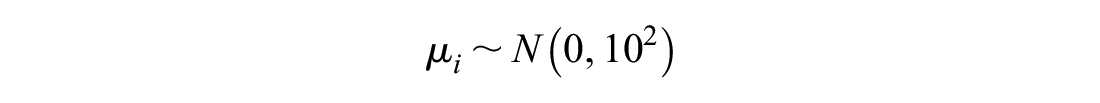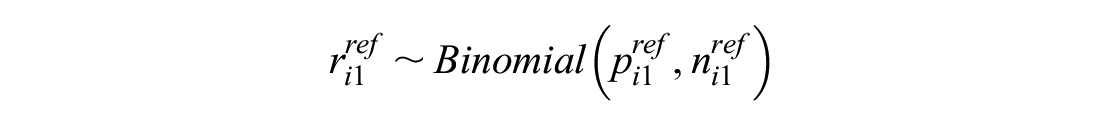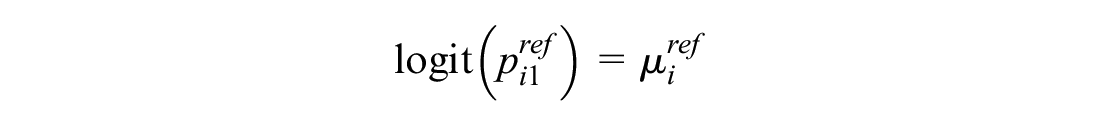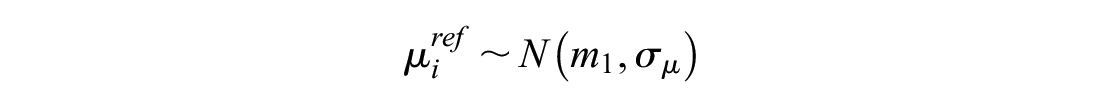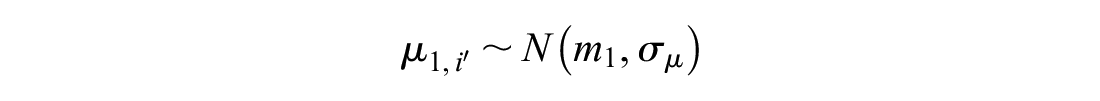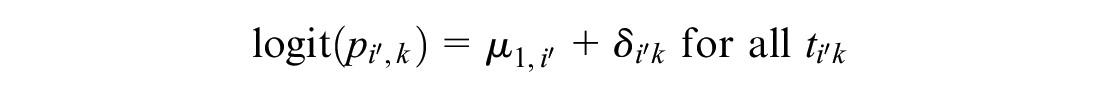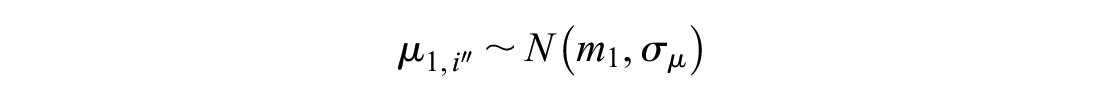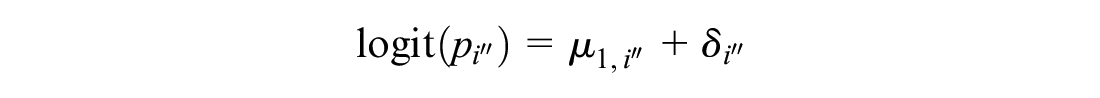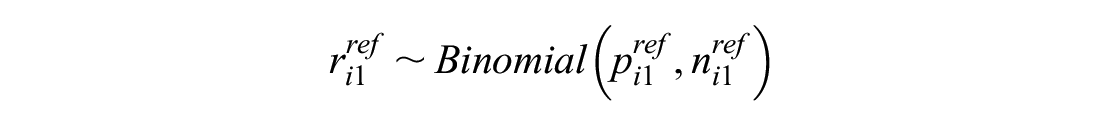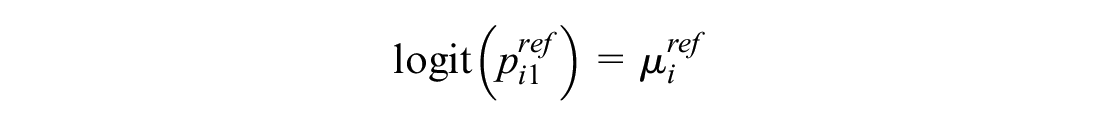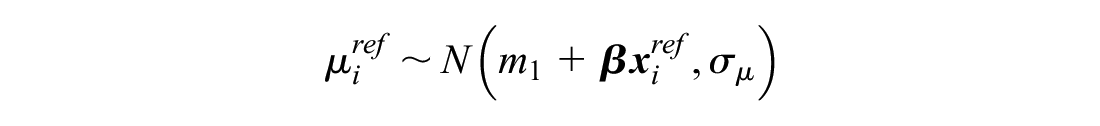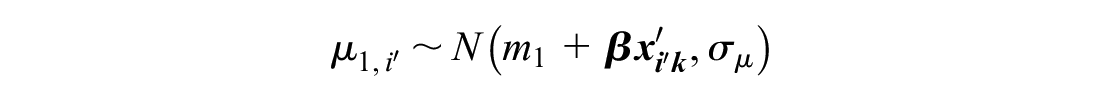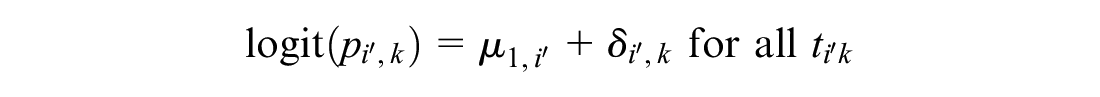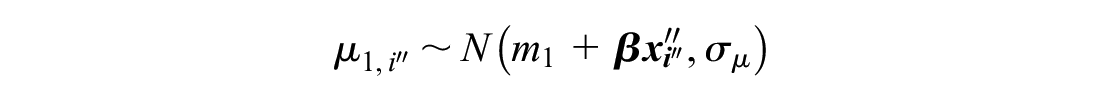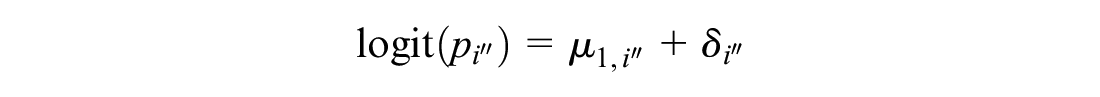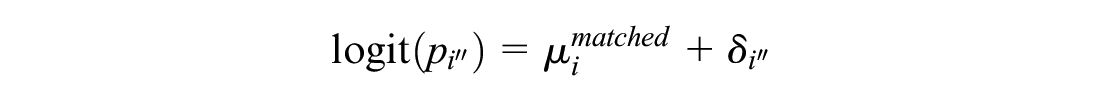Abstract
Background
Network meta-analysis (NMA) requires a connected network of randomized controlled trials (RCTs) and cannot include single-arm studies. Regulators or academics often have only aggregate data. Two aggregate data methods for analyzing disconnected networks are random effects on baseline and aggregate-level matching (ALM). ALM has been used only for single-arm studies, and both methods may bias effect estimates.
Methods
We modified random effects on baseline to separate RCTs connected to and disconnected from the reference and any single-arm studies, minimizing the introduction of bias. We term our modified method reference prediction. We similarly modified ALM and extended it to include RCTs disconnected from the reference. We tested these methods using constructed data and a simulation study.
Results
In simulations, bias for connected treatments for ALM ranged from −0.0158 to 0.051 and for reference prediction from −0.0107 to 0.083. These were low compared with the true mean effect of 0.5. Coverage ranged from 0.92 to 1.00. In disconnected treatments, bias of ALM ranged from −0.16 to 0.392 and of reference prediction from −0.102 to 0.40, whereas coverage of ALM ranged from 0.30 to 0.82 and of reference prediction from 0.64 to 0.94. Under fixed study effects for disconnected evidence, bias was similar, but coverage was 0.81 to 1.00 for reference prediction and 0.18 to 0.76 for ALM. Trends of similar bias but greater coverage for reference prediction with random study effects were repeated in constructed data.
Conclusions
Both methods with random study effects seem to minimize bias in treatment connected to the reference. They can estimate treatment effects for disconnected treatments but may be biased. Reference prediction has greater coverage and may be recommended overall.
Highlights
Two methods were modified for network meta-analysis on disconnected networks and for including single-arm observational or interventional studies in network meta-analysis using only aggregate data and for minimizing the bias of effect estimates for treatments only in trials connected to the reference.
Reference prediction was developed as a modification of random effects on baseline that keeps analyses of trials connected to the reference separately from those disconnected from the reference and from single-arm studies. The method was further modified to account for correlation in trials with more than 2 arms and, under random study effects, to estimate variance in heterogeneity separately in connected and disconnected evidence.
Aggregate-level matching was extended to include trials disconnected from the reference, rather than only single-arm studies. The method was further modified to separately estimate treatment effects and heterogeneity variance in the connected and disconnected evidence and to account for the correlation between arms in trials with more than 2 arms.
Performance was assessed using a constructed data example and simulation study.
The methods were found to have similar, and sometimes low, bias when estimating the relative effects for disconnected treatments, but reference prediction with random study effects had the greatest coverage.
The use of reference prediction with random study effects for disconnected networks is recommended if no individual patient data or alternative real-world evidence is available.
Keywords
Introduction
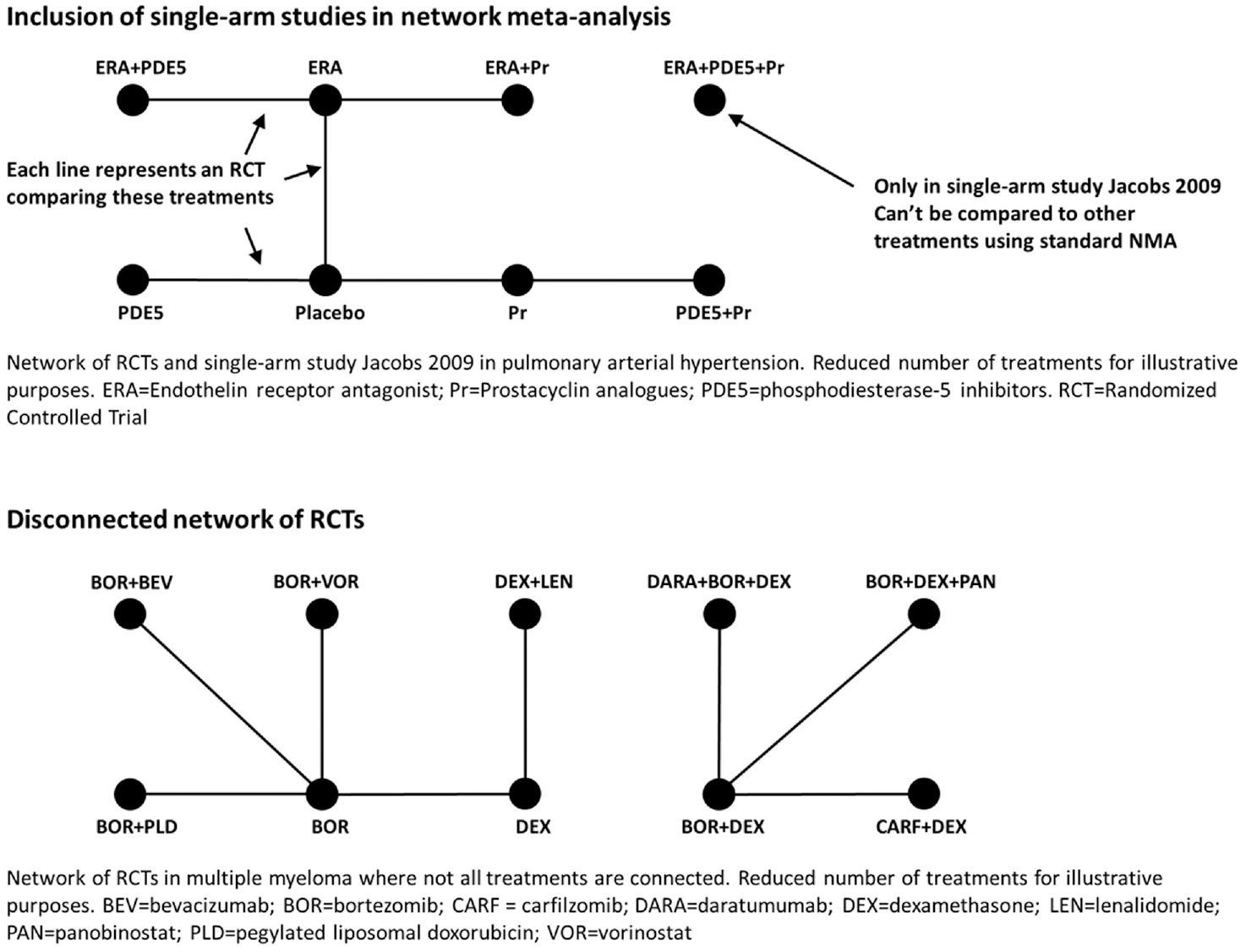
Network meta-analysis (NMA) is a method endorsed by the UK National Institute for Health and Care Excellence (NICE) and international scientific societies for estimating relative treatment effects between interventions based on randomized controlled trials (RCTs) in which each study compares a subset of the interventions of interest.1,2 NMA requires treatments to be connected by a network of RCTs to benefit from randomization and minimize the risk of biased relative to treatment effect estimates. 3 Disconnected networks are common in rare diseases, for which RCT recruitment is challenging.4–6 Disconnected networks can also arise if analyzing rarely reported outcomes or splitting treatments by doses or time points. Furthermore, traditional NMA is restricted to RCTs and cannot incorporate single-arm observational or interventional studies, which are again common in rare disease or in cases in which a control arm would be unethical or there has been a breakthrough in treatment efficacy and RCTs are not judged necessary.7–9 Single-arm studies and disconnected RCT networks are illustrated with examples from the literature in Figure 1.
There are many methods available to overcome disconnected RCT networks or to incorporate single-arm studies, and these have been described in a recent thorough review. 5 Real-world evidence, such as registry or insurance claims data, can be used to fill RCT evidence gaps and connect networks or to replace the RCT network entirely, as has been done in hip replacement surgery.10,11 Real-world evidence is not always available, is likely focused on only established treatments, and may be subject to implicit bias. 12 If individual patient data are available from all RCTs and single-arm studies, propensity matching or regression adjustment techniques could be used to conduct comparisons. 13 Even if individual patient data are available from only 1 RCT or 1 single-arm study, matching-adjusted indirect comparison, simulated treatment comparison, and multilevel network meta-regression could be used to conduct unanchored indirect comparisons on disconnected networks.14,15 However, individual patient data from commercial RCTs or single-arm studies are frequently not available to academics or to authors of national clinical guidelines. Furthermore, it should be stated that these methods are not as reliable as connected networks of RCTs are. 15
If only aggregate data are available, methods are still available to connect networks. Model-based NMA models dose-response relationships and can be used to connect disconnected networks if studies on multiple doses are available and if the dose-response relationship can be correctly specified.16,17 A class effect NMA can be used to connect networks if disconnected treatments have a similar mechanism of action or molecular structure to a connected treatment. 5 If interventions in a disconnected network are composed of common components, component NMA has been proposed to reconnect networks.18,19 Modeling the relation between treatment effects on different scales, multiple outcomes NMA can leverage connected networks on 1 outcome to connect disconnected networks, or strengthen sparse networks, on another outcome. 20 All of these rely on special circumstances and strong assumptions.
Random effects on baseline can be used to connect a disconnected network when only aggregate data are available. 4 Random effects on baseline assumes the response to control treatments are exchangeable across studies and thus removes the need for a connected network with the price of interfering with randomization in RCTs already connected to the intervention of interest.21–23 Another method termed aggregate-level matching (ALM) can include single-arm studies by matching them with an arm of a selected RCT that is connected to the intervention of interest, creating a new RCT. However, this method has not yet been extended to RCTs disconnected from the intervention of interest. 24
We first introduce the standard method of contrast-based NMA with independent baselines in the next section (“Contrast-Based Network Meta-analysis Model for a Connected Network with Independent Baseline”) before giving more detail on the issues with random effects in subsequent section (“Network Meta-analysis Model with Random Effects on Baseline and Its Problems”). In the “Modified Evidence Synthesis Models for Disconnected Networks with Aggregate-Level Data” section, we present our modified methods for the analysis of disconnected networks using only aggregate data and prevent bias in the RCTs already connected to the target comparator. We test these on an artificially disconnected network based on a real literature review in atrial fibrillation and in a simulation study in the sections “Assessment by Constructed Data” and “Assessment by Simulation Study,” respectively.25,26 We close by drawing conclusions and making methodology recommendations.
Existing Evidence Synthesis Models
Contrast-Based Network Meta-analysis Model for a Connected Network with Independent Baseline
We consider only NMA on binomial data with a logistic link function, but the methods we present can be applied to any generalized linear model for any type of data.
If each arm
The probability of event
The log odds of an event on the baseline treatment
The
The
The treatment denoted 1 is the so-called “reference” treatment, which is common across the network of evidence and relative to which the basic log odds ratios
The baseline effects
They are thus assumed to be independent of each other, so information on the relative treatment effects
This independent baseline model preserves randomization within trials. Comparison of any 2 treatments
However, this requires a connected network of RCT evidence between treatments x and
Under random effects in which
Network Meta-analysis Model with Random Effects on Baseline and Its Problems
The model described by Thom et al.
4
and Beliveau et al.
22
and generally called the “random effects on baseline” puts a random effect model on reference treatment
This models the log odds of the event on the reference treatment, even in trials that do not include the reference treatment. For any arm of a connected RCT, disconnected RCT, or single-arm study, the probability is modeled as
The treatment effects
or
This thus allows the inclusion of RCTs disconnected from the reference or single-arm studies. Unlike the independent baseline model, this approach assumes exchangeability of reference log odds
The downside to using random effects on baseline is that treatment effects
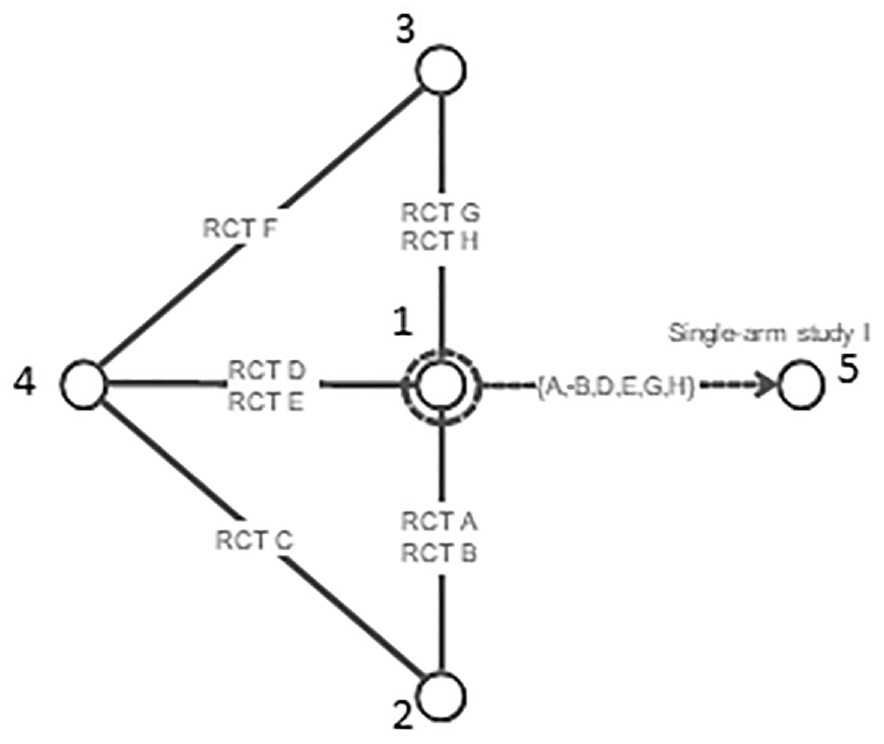
Using random effects on baseline to predict response on reference treatment A using studies that included A. However, the random effects on baseline model interferes with randomization in randomized controlled trials on treatments 2, 3, and 4.
Typically, for random effects models, the same heterogeneity variance
Modified Evidence Synthesis Models for Disconnected Networks with Aggregate-Level Data
We propose modifications and extensions of random effects on baseline and ALM to minimize bias when analyzing disconnected networks using only aggregate data.
Reference Prediction
Our method of reference prediction predicts outcomes on the reference treatment in RCTs disconnected from the reference and single-arm studies. This involves two modifications of random effects on baseline. The first is to analyze RCTs connected to the reference separately from those disconnected from the reference and from single-arm studies. We also modify the model under random study effects to correctly account for RCTs with more than 2 arms and to estimate the heterogeneity variance separately in RCTs connected to the reference and those disconnected from the reference. In the section titled “Reference Prediction with Covariates,” we further modify the method to include covariate effects and improve the prediction of the reference response.
To avoid interfering with randomization in RCTs connected to the reference, we propose splitting them into
However, we make a second use of the
We importantly note that there is no link in the model between
This meta-analysis model is then used to generate predictions of the log odds of response on reference treatment 1 in RCTs disconnected from the reference
The meta-analysis is therefore called a “reference prediction” model. As in random effects on baseline, the
As in the independent baseline model, fixed or random effects can be used for the treatment effect
Finally, the reparameterization of
Reference Prediction with Covariates
A further modification for reference prediction models is to include covariates. We assume we have some vector of length
The
Although the
For single-arm studies, we use
Note that we still assume independent
Modified ALM
Our modification to ALM extends the method to include RCTs disconnected from the reference. As for reference prediction, we also modify the method to keep the RCTs connected to the reference separate from those that are disconnected from the reference and from single-arm studies, thus minimizing the introduction of bias. Finally, we modify ALM on random study effects to correctly account for RCTs with more than 2 arms.
ALM, under contrast-based NMA with independent baselines, instead uses a
We model RCTs disconnected from the reference as
This ensures that the same
As the
Similarity could be measured by the Euclidean distance
ALM has been previously applied to single-arm studies.7 As illustrated in Figure 3, the approach to single-arm studies
The connected RCT should have baseline characteristics
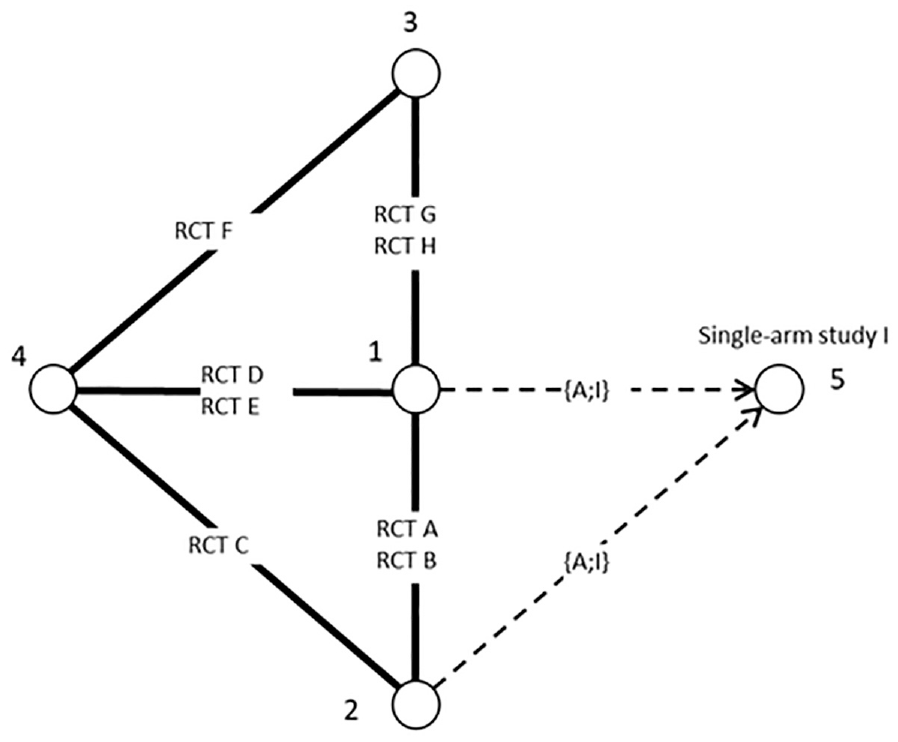
Using aggregate-level matching to connect a single-arm study to randomized controlled trials (RCTs) with similar design and patient characteristics. In this case, the baseline odds of RCT A is used to match to RCT I.
Unlike reference prediction, ALM requires only a single RCT connected to the reference to be identifiable, although more are recommended to ensure a sufficiently similar study is used for matching.
Assessment by Constructed Data
Methods of Constructed Data Example
Our constructed data example was created by artificially disconnecting a network drawn from a systematic literature review and contrast-based NMA on a connected network comparing treatments for the prevention of stroke, while minimizing the risk of clinically relevant bleeding, in atrial fibrillation.25,36 The advantage of artificially disconnecting a network is that we can compare the estimated of reference prediction and ALM to the results of the “true NMA” based on RCTs. Aligning with the methods presented in the sections “Existing Evidence Synthesis Models” and “Modified Evidence Synthesis Models for Disconnected Networks with Aggregate-Level Data,” we analyzed the binomial outcomes with a logistic link function. The NMA compared coumarin (international normalized ratio target range 2–3) with the oral anticoagulants apixaban (twice-daily 5 mg), dabigatran (twice-daily 150 mg), edoxaban (once-daily 60 mg), and rivaroxaban (once-daily 20 mg). This consists of 12 RCTs on 13 interventions reporting stroke and 17 RCTs on 24 interventions reporting clinically relevant bleeding.
We artificially disconnected the trials on dabigatran from those on other treatments in the evidence networks on stroke and clinically relevant bleeding. In the stroke network, there was 1 RCT on dabigatran, whereas in clinically relevant bleeding, there were 3 RCTs (details are given in the Appendix). In the first scenario, the coumarin arm was removed from the dabigatran trials, giving a disconnected network of 1 RCT on dabigatran 110 mg versus 150-mg doses in ischemic stroke and 3 RCTs on 9 different doses of dabigatran in clinically relevant bleeding. The second scenario consisting of splitting the RCTs on dabigatran into single-arm studies and excluding the coumarin control arms, giving 2 for ischemic stroke and 12 for clinically relevant bleeding. These disconnections are illustrated for ischemic stroke in Figure 4.
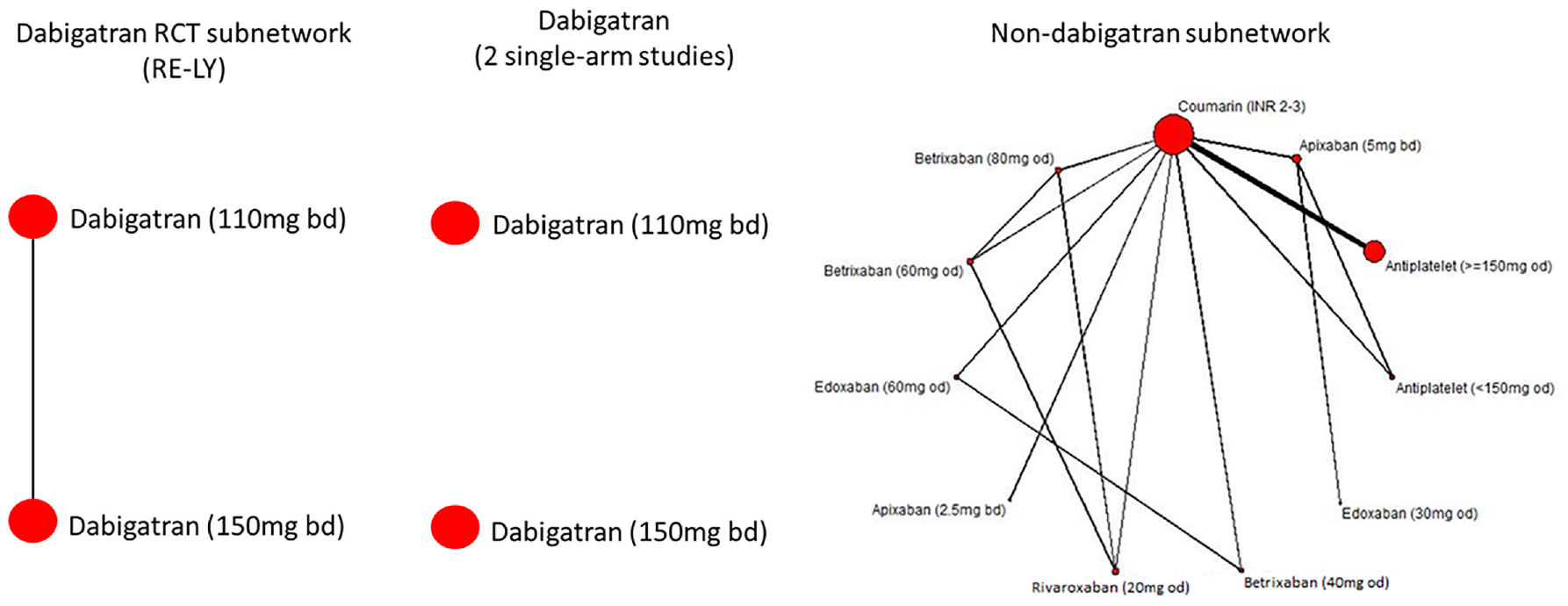
Disconnecting the ischemic stroke atrial fibrillation network. Twelve randomized controlled trials (RCTs) on 13 interventions are changed to either 1 RCT connected to the reference and 11 RCTs connected to the reference or to 2 single-arm studies and 11 RCTs to the reference.
We then applied the reference prediction and ALM to these disconnected networks in an attempt to reproduce the results of the original connected network, the true NMA. For reference prediction, we used meta-regression to explore which, if any, of mean age, proportion male, and CHA2DS2-VASc score should be included in the baseline response model. Covariates were centered at the across-study mean, and missing values for individual studies were set to the mean (i.e., zero in the regression due to centering). All 3 covariates, with no standardization, were used to calculate the Euclidean distance ALM, with missing values omitted from the summation.
Results of the Atrial Fibrillation Constructed Data Example
The random effects model with no covariates was selected as the regression model for reference prediction on ischemic stroke, because it had good total residual deviance (11.9 v. 11 data points), and its DIC of 67.14 was not substantially improved by including covariates (the DIC of the covariate models ranged from 66.96 to 67.93). All fixed effects had poor residual deviance (ranging from 68 to 79.8, which was substantially greater than the 11 data points) and worse DIC (ranging from 119 to 127.1) than the random effects models did. The same conclusions held for clinically relevant bleeding.
The ischemic stroke results for the methods with random study effects are presented in Figure 5 and those for the fixed study effects are shown in Figure 6. The tabulated results are provided in the Appendix.
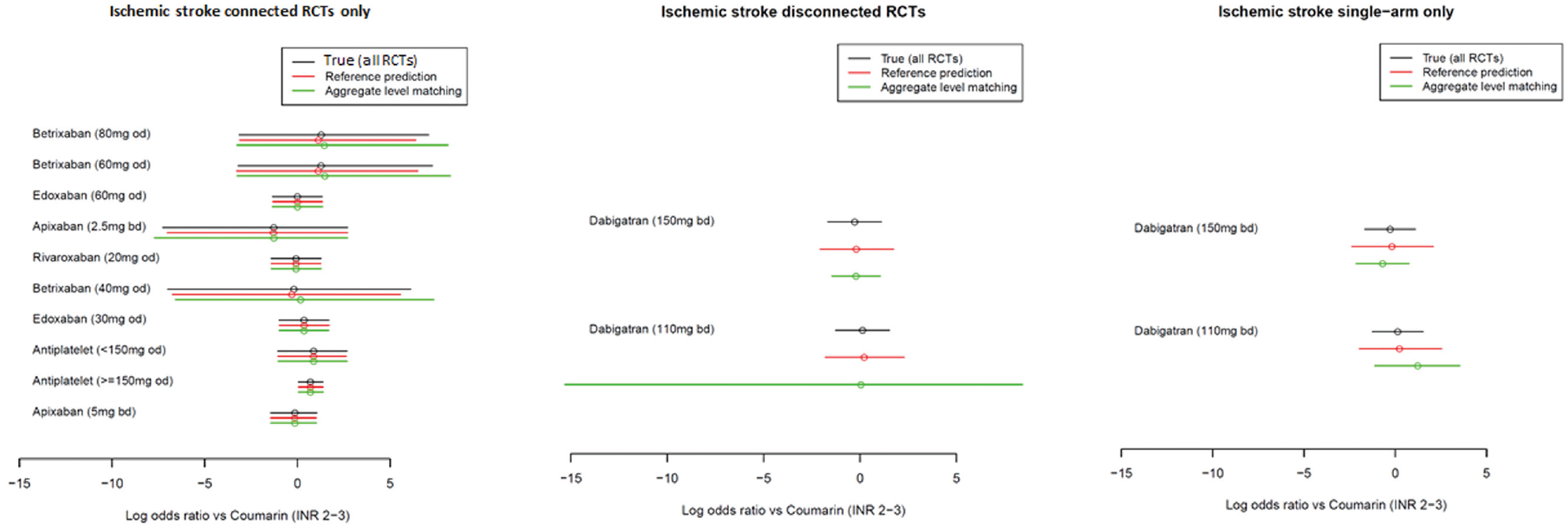
Comparison of the estimated log odds ratios using the true method (all randomized controlled trials), reference prediction, and aggregate-level matching under random study effects for the ischemic stroke outcome. Point estimates are means, and uncertainty intervals are 95% credible intervals.
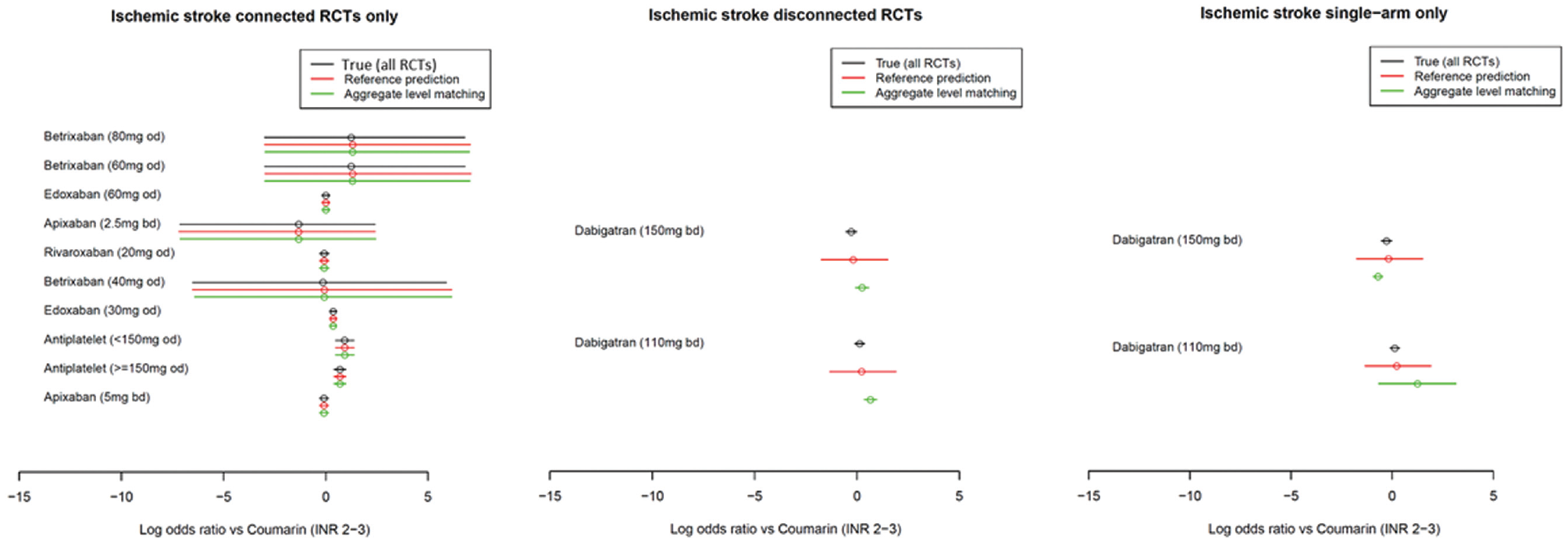
Comparison of the estimated log odds ratios using the true method (all randomized controlled trials), reference prediction, and aggregate-level matching under fixed study effects for the ischemic stroke outcome. Point estimates are means, and uncertainty intervals are 95% credible intervals.
Using both the fixed and random study effects, we observed that the reference prediction and ALM reproduced the results of the true NMA (i.e., independent baselines using all RCTs) in the connected portion of the network. For example, the log odds ratio for edoxaban (30 mg every day) was 0.36 (95% credible interval [CrI]: −0.94, 1.64) using true NMA, 0.36 (−0.92, 1.63) using reference prediction, and 0.36 (−0.93, 1.61) using ALM.
For the reference prediction on RCTs disconnected from the reference and single-arm studies, the point estimates were close to those of the true NMA. For example, if in an RCT disconnected from the reference, the log odds ratios for dabigatran (150 mg) were −0.28 (−1.55, 0.99) under true NMA, −0.19 (−1.98, 1.71) using reference prediction, and −0.21 (−1.36, 0.94) using ALM. The 95% CrIs were similar under the random study effects but much wider under the fixed study effects; however, in both cases, they overlapped with the point estimate of the true NMA. ALM had poor performance under fixed study effects with different point estimates and narrow 95% credible intervals, which did not always overlap with the true NMA estimate; for example, if in RCTs disconnected from the reference dabigatran (110 mg), the log odds ratios were estimated as 0.13 (−0.11, 0.37) using true NMA and 0.22 (−1.34, 1.91) using reference prediction, but 0.66 (0.35, 0.96) using ALM. Under random study effects for single-arm studies, the point estimates appeared to be more different from the true NMA than the reference prediction but similar on the RCTs from the reference. The 95% CrIs also depend greatly on the matched study with, sometimes very wide CrIs; for example, in RCTs disconnected from the reference, ALM estimated the log odds ratio for dabigatran (110 mg) to be −0.0065 (−19.59, 19.68) but the true NMA estimated 0.13 (−1.14, 1.39) and reference prediction 0.23 (−1.70, 2.24).
Random study effects models for clinically relevant bleeding were less reassuring (results are given in the Appendix). Both reference prediction and ALM had 95% CrIs that often did not overlap with the true NMA effect for the PETRO study, which reported clinically relevant bleeding but not ischemic stroke, so only in the former outcome network. This is likely because PETRO was a small study with only 70 patients on coumarin, whereas RE-LY, in both outcome networks, included 6022 patients taking coumarin. Both methods performed better for dabigatran doses studied in RE-LY. In the disconnected RCTs scenario, ALM had point estimates closer to the truth than the reference prediction. Conversely, ALM estimates were further from the truth, and with 95% CrIs not overlapping the truth, for the single-arm scenario. Reference prediction appears to be “safer” in general. Again, fixed study effects models performed poorly.
Assessment by Simulation Study
Methods of Simulation Study
We used a simple simulation study to assess the bias and coverage of treatment effect estimates of ALM and reference prediction in the scenarios of disconnected RCTs (i.e., where there are RCTs disconnected from the reference) and single-arm studies. Methods with lower bias and greater coverage are preferred.
We merged all covariates for matching and regression in ALM and reference prediction, respectively, as a single covariate. We explain below how this does not reduce the generality of the simulations. We assumed the covariates were not treatment effect modifiers. Effect modifiers are a problem for both connected and disconnected networks, and our methods do not purport to overcome the imbalance in effect modifiers.
The basic geometry of all of our simulations is illustrated in Figure 7. The performance of ALM and reference prediction depend on the quantity of the connected evidence and not on the disconnected evidence. We therefore varied only the number of RCTs connected to the reference and always included only 5 RCTs disconnected from the reference or 10 single-arm studies. The number of 2-arm and 3-arm RCTs to the reference was gradually increased, with scenarios described in Table 1. Simulations were conducted twice, once with the number of patients in trials fixed at 100 on each arm and again with 1000 patients. The patient number simulations were kept separate, and our primary case included 100 patients per arm.
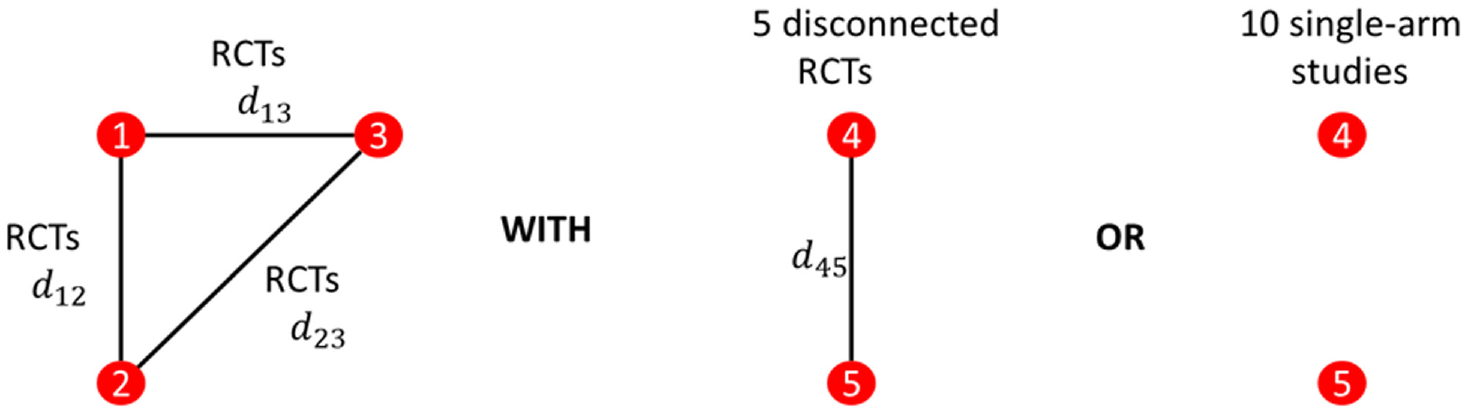
Network geometries considered in the simulation study. Either 5 randomized controlled trials (RCTs) from the reference or 10 single-are studies were considered. The number of RCTs to the reference was varied.
Number of Randomized Control Trials (RCTs) in 3 Simulation Studies with 5, 15, and 50 RCTs Connected to the Reference a
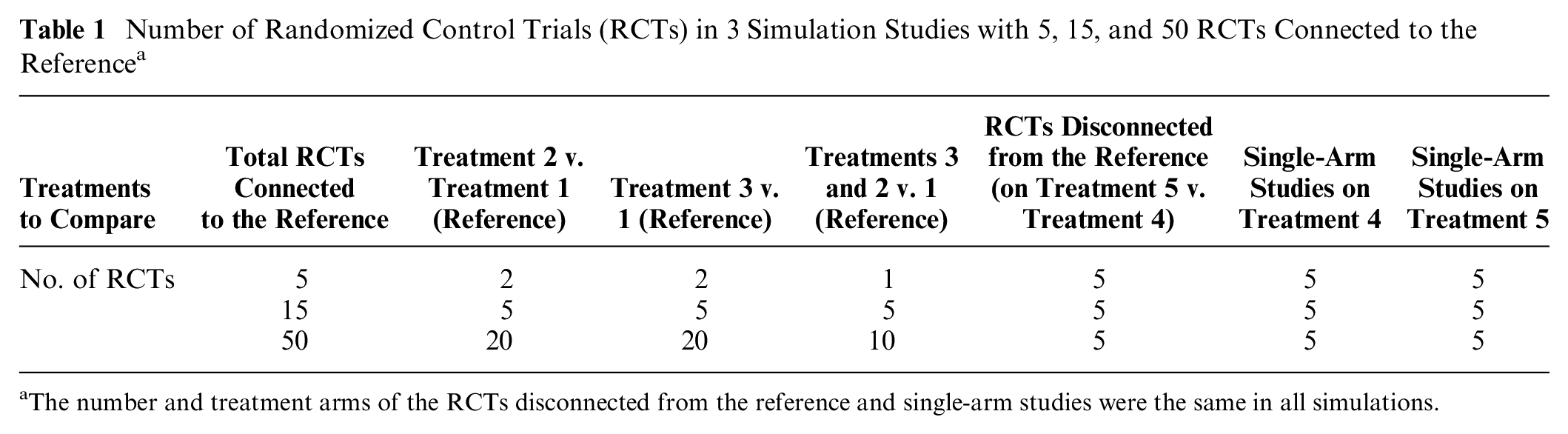
The number and treatment arms of the RCTs disconnected from the reference and single-arm studies were the same in all simulations.
Our simulated data followed the setup of the binomial-logistic NMA example that has been used throughout. The number of events on arm
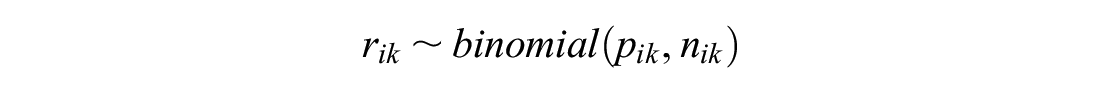
The probability of event
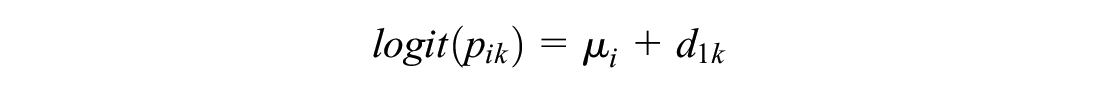
For numerical stability, we forced any arms with zero events (r = 0) to have at least 1 event (r = 1). The log odds ratios
The model we used to simulate the log odds for study
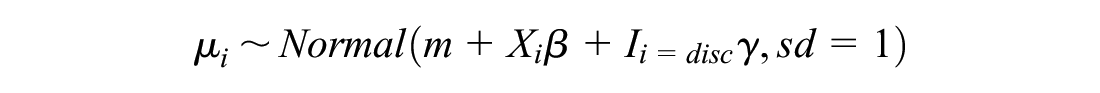
We simulated the across-study mean from
As noted above, we considered only a single prognostic factor
We applied contrast-based NMA, reference prediction, and ALM for all modeled scenarios and compared their treatment effect estimates to the “truth” in the (for reference prediction and ALM only) components connected to or disconnected from the reference. As we ran these for both fixed and random effects, and for 100 and 1000 patients per study arm, there were 96 scenarios in total:
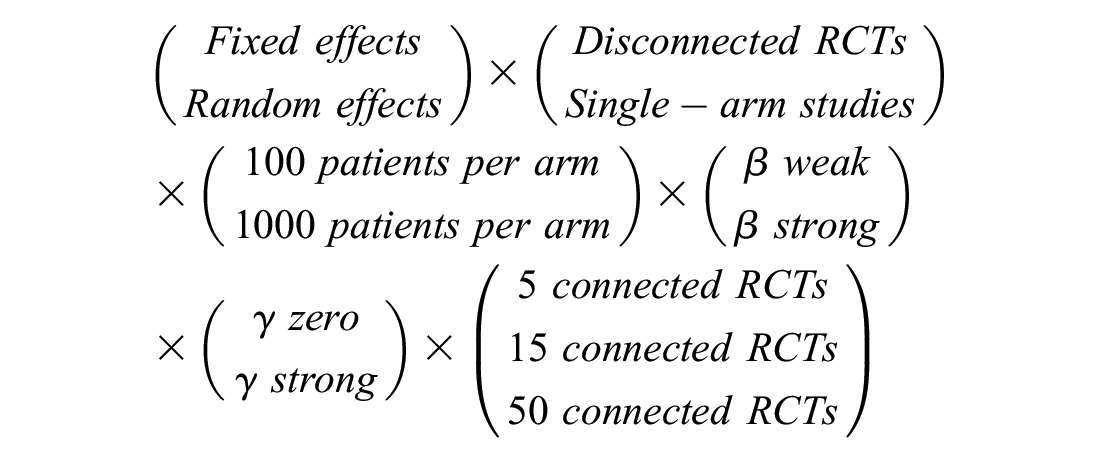

The 12 scenarios across assumptions on
List of Scenarios Used in the Simulation Study a
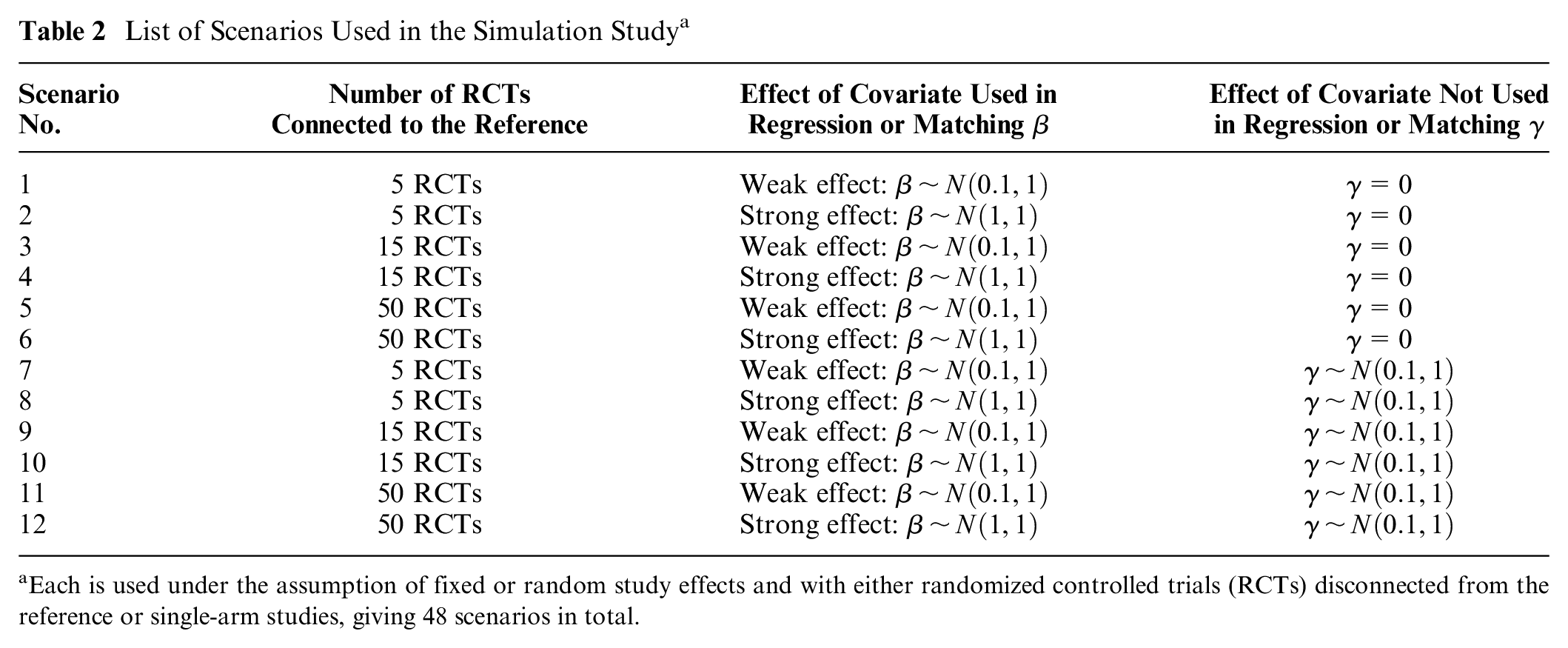
Each is used under the assumption of fixed or random study effects and with either randomized controlled trials (RCTs) disconnected from the reference or single-arm studies, giving 48 scenarios in total.
Results of the Simulation Study
Estimated bias and coverage probabilities across all methods and all scenarios for random study effects models in which RCTs have 100 patients are illustrated in Figure 8 and where RCTs have 1000 patients in Figure 9. Bias is interpreted in comparison with the mean treatment effect of 0.5 (i.e., the mean of log odds ratio
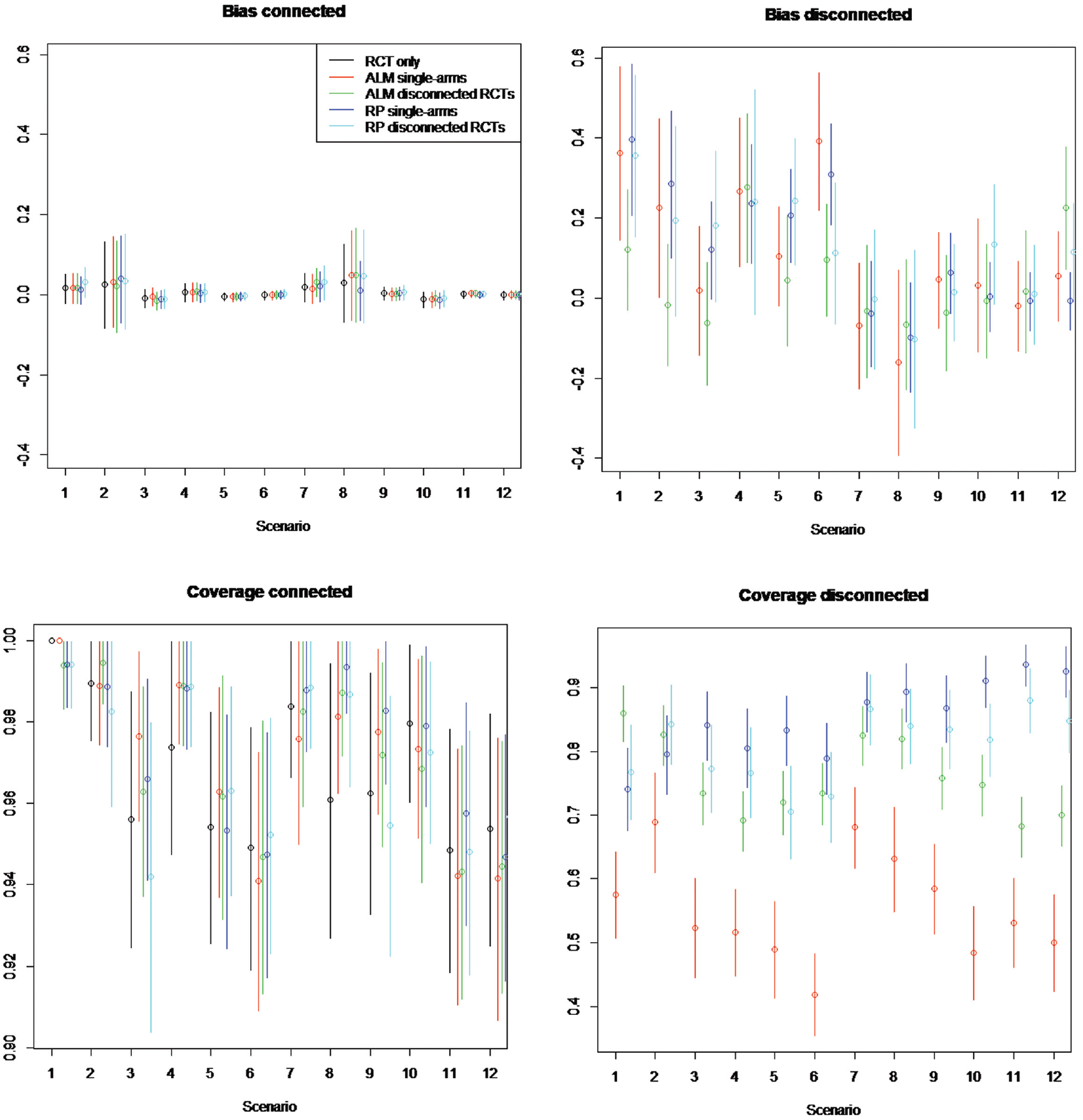
Simulation study estimated bias and coverage of each method in the connected and disconnected evidence scenarios. Random study effects of 100 patients per arm. Scenarios: scenario 1 = 5 randomized controlled trials (RCTs), β weak, γ = 0; scenario 2 = 5 RCTs, β strong, γ = 0; scenario 3 = 15 RCTs, β weak, γ = 0; scenario 4 = 15 RCTs, β strong, γ = 0; scenario 5 = 50 RCTs, β weak, γ = 0; scenario 6 = 50 RCTs, β strong, γ = 0; scenario 7 = 5 RCTs, β weak, γ≠ 0; scenario 8 = 5 RCTs, β strong, γ≠ 0; scenario 9 = 15 RCTs, β weak, γ≠ 0; scenario 10 = 15 RCTs, β strong, γ≠ 0; scenario 11 = 50 RCTs, β weak, γ≠ 0; scenario 12 = 50 RCTs, β strong, γ≠ 0. The β represents the impact of covariates included in aggregate-level matching (ALM) and reference prediction (RP) regression. γ represents the impact of the covariates not included in the ALM or RP regression. Scenarios are an analysis on RCTs only using single-arm studies (single) and RCTs from the reference (disconnected).
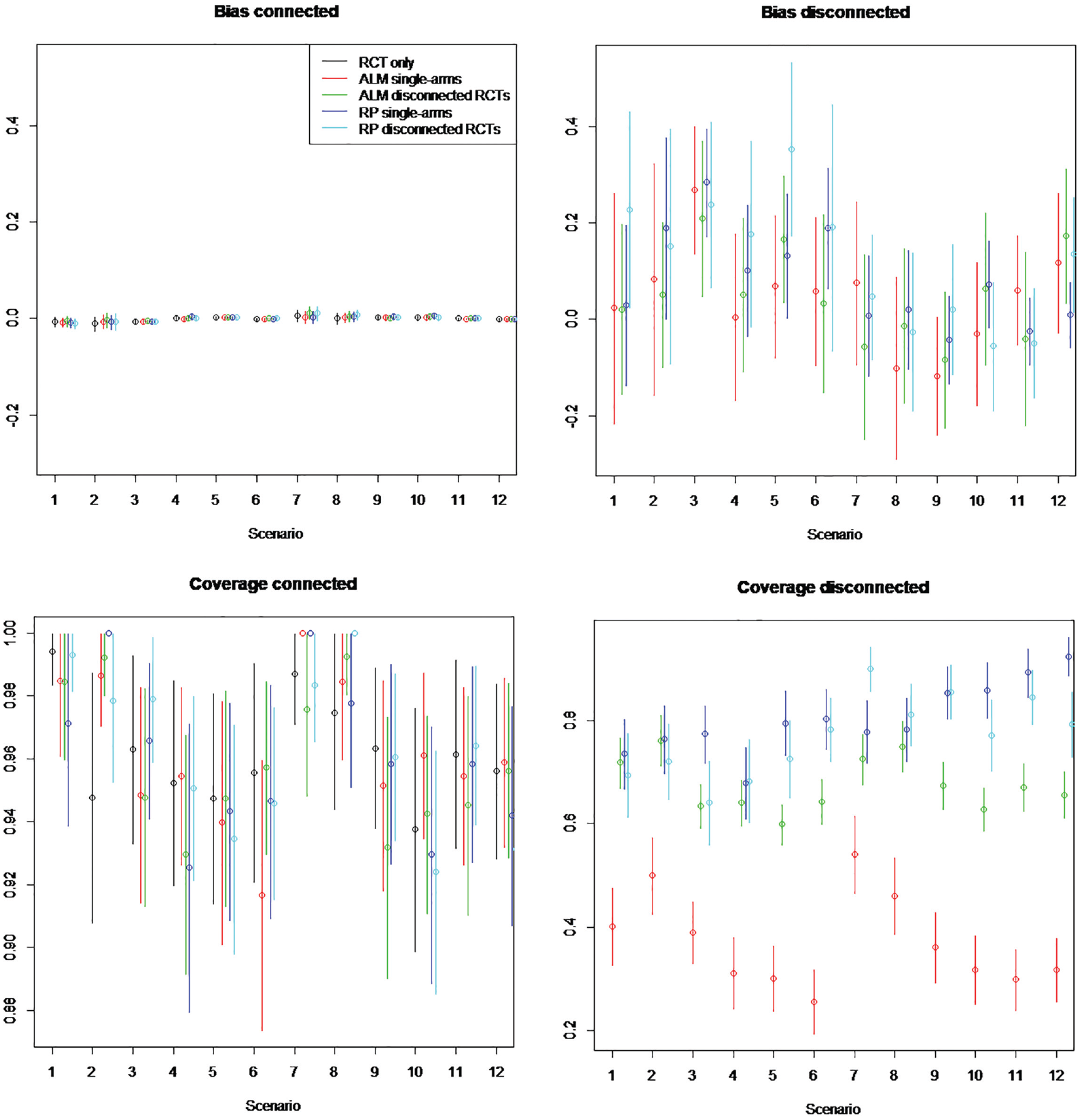
Simulation study estimated bias and coverage of each method in the connected and disconnected evidence scenarios. random study effects of 1000 patients per arm. Scenarios: scenario 1 = 5 randomized controlled trial (RCTs), β weak, γ = 0; scenario 2 = 5 RCTs, β strong, γ = 0; scenario 3 = 15 RCTs, β weak, γ = 0; scenario 4 = 15 RCTs, β strong, γ = 0; scenario 5 = 50 RCTs, β weak, γ = 0; scenario 6 = 50 RCTs, β strong, γ = 0; scenario 7 = 5 RCTs, β weak, γ≠ 0; scenario 8 = 5 RCTs, β strong, γ≠ 0; scenario 9 = 15 RCTs, β weak, γ≠ 0; scenario 10 = 15 RCTs, β strong, γ≠ 0; scenario 11 = 50 RCTs, β weak, γ≠ 0; scenario 12 = 50 RCTs, β strong, γ≠ 0. β represents the impact of covariates included in aggregate-level matching (ALM) and reference prediction (RP) regression. γ represents the impact of the covariates not included in ALM or RP regression. Scenarios are an analysis on RCTs only, using single-arm studies (single), and RCTs from the reference (disconnected).
Estimated Bias
On the component of the network connected to the reference, the bias of ALM ranges from −0.0158 to 0.051, for reference prediction from −0.0107 to 0.083, and for NMA based only on RCTs from −0.011 to 0.030. This suggests the bias is lower for NMA only but is similar between ALM and reference prediction and in any case low (usually less than 10%) compared with the true treatment effect.
In RCTs disconnected from the reference and in single-arm studies, the bias is much higher, ranging from −0.1 to −0.4 when disconnected studies are not systematically different (
Estimated Coverage
In the component of the network connected to the reference, the coverage of all methods was similar but greatest for NMA using only RCTs. The coverage ranged for ALM from 0.92 to 1.00, for reference prediction from 0.92 to 1.00, and for NMA using only RCTs from 0.95 to 1.00.
In RCTs disconnected from the reference and in single-arm studies, reference prediction has a coverage ranging from 0.64 (1000 patients, disconnected RCTs, 15 connected RCTs,
ALM appeared to perform worse for single-arm studies than for RCTs disconnected from the reference, with coverage on single-arm studies ranging from 0.26 to 0.69 and on disconnected RCTs from 0.60 to 0.86. Coverage of all methods appeared to decrease with increasing numbers of patients, going from a range of 0.42 to 0.95 on 100 patients to a range of 0.26 to 0.92 on 1000 patients. This may occur when there are systematic differences in prognostic factors between RCTs connected to the reference and single-arm studies or RCTs disconnected from the reference treatment (i.e.,
Fixed Effects Results
Fixed study effects results are presented in the Appendix. Bias was generally similar to the random study effects case in both the components connected to and disconnected from the reference for all methods and scenarios. The coverage probabilities were lower for reference prediction on disconnected treatments and ranged from 0.81 to 1.00. Coverage probabilities were so low, ranging from 0.18 to 0.76 for disconnected treatments across all scenarios, for ALM that the method could not be recommended with fixed study effects. It is worth recalling from above that coverage was worst for ALM under random effects (from 0.30 to 0.82).
Discussion
We have presented 2 modified methods for including RCTs disconnected from the reference, or any intervention of interest, or single-arm studies in contrast-based NMA when only aggregate data are available. Reference prediction is a modification of random effects on baseline with RCTs connected to the reference and RCTs disconnected from the reference kept separate, different heterogeneity variances used for random study effects models, and a corrected adjustment for multiarm studies. We modified ALM to allow the inclusion of RCTs from the reference, used different heterogeneity variances for RCTs connected to and disconnected from the reference, and corrected the adjustment for multiarm studies. These methods give similar point estimates and 95% CrIs to the independent baseline contrast-based NMA, which suggests that we have minimized the introduction of bias. Fixed study effects models performed poorly, with point estimates far from the truth and 95% CrIs that did not include the truth; this was matched by the simulation study in which fixed study effects had high bias and low coverage. The low coverage was a result of fixed effects being less “lenient” to poorly matching studies or to different populations than random effects. In the constructed data example, the performance of ALM was found to greatly depend on the matched study, illustrating the importance of choosing an appropriate study. The performance of reference prediction depended greatly on the similarity of the connected and disconnected evidence. This was confirmed by the simulation study in which coverage of both methods was worse when there were systematic differences in prognostic factors between connected and disconnected studies. This illustrates the importance of assessing whether these methods are appropriate in each specific circumstance. In both the constructed data and simulation study, reference prediction was found to have lower bias and greater coverage than ALM and could be viewed as the “safer” option for inference on disconnected networks.
There are many possible extensions to our methodology. Aggregate- or individual-level real-world evidence in the form of registries or cohort studies could be used to build the reference prediction models or control arm data for ALM. 11 This could allow predictions to be tailored to specific populations, rather than relying only on what has been studied in an RCT. Both methods could be used to incorporate single-arm studies on treatments already in the RCTs connected to the reference; hierarchical models could be used to keep effects separate or informative priors used to minimize any bias from single-arm studies on RCT data.7,38 The high uncertainty in the effect estimates of random study effects reference prediction or ALM suggests that these may be better suited to uncertainty quantification for input in value-of-information analyses using economic models. 39 This could provide an upper bound on the value of studying disconnected treatments in new head-to-head RCTs.
Equally, there are many limitations to our methods and assessments. Reference prediction, despite having good coverage and low bias, can provide highly variable and almost noninformative treatment effect estimates. Our constructed data situation was based on an example in atrial fibrillation and is unlikely to be generalizable to other clinical areas; that single-arm studies and RCTs from the reference were based on RCTs connected to the reference also means they are likely to be more similar in prognostic factors than a real scenario. Our constructed data example and simulation study have shown only that there are no problems in specific situations, and there may be cases in which both reference prediction and ALM lead to serious bias. We also restricted our analysis to binomial outcomes. Extension to count or continuous outcomes would be straightforward as similar generalized linear models for NMA on single time points exist. 1 However, time-to-event outcomes NMA such as fractional polynomials NMA, common in oncology, would require more significant modification. 40
We have presented methods that aim to get as close as possible to the results of a fully connected contrast-based NMA with independent baselines. However, we would term these as “last resort” methodologies for when networks are disconnected or if the RCT evidence is so limited as to be uninformative. If individual patient data are available from some or all trials, methods such as multilevel network meta-regression, propensity score reweighting, matching-adjusted indirect comparison, or simulated treatment comparison are recommended instead. If high-quality nonrandomized comparative evidence is available, this should instead be used through hierarchical models or informative priors. If only aggregate data from RCTs are available, and multiple outcomes, class effects, or dose-response modeling are not appropriate, reference prediction and ALM could be considered for health care decision making, as long as the limitations are recognized.
Supplemental Material
sj-docx-1-mdm-10.1177_0272989X221097081 – Supplemental material for Network Meta-analysis on Disconnected Evidence Networks When Only Aggregate Data Are Available: Modified Methods to Include Disconnected Trials and Single-Arm Studies while Minimizing Bias
Supplemental material, sj-docx-1-mdm-10.1177_0272989X221097081 for Network Meta-analysis on Disconnected Evidence Networks When Only Aggregate Data Are Available: Modified Methods to Include Disconnected Trials and Single-Arm Studies while Minimizing Bias by Howard Thom, Joy Leahy and Jeroen P. Jansen in Medical Decision Making
Supplemental Material
sj-docx-2-mdm-10.1177_0272989X221097081 – Supplemental material for Network Meta-analysis on Disconnected Evidence Networks When Only Aggregate Data Are Available: Modified Methods to Include Disconnected Trials and Single-Arm Studies while Minimizing Bias
Supplemental material, sj-docx-2-mdm-10.1177_0272989X221097081 for Network Meta-analysis on Disconnected Evidence Networks When Only Aggregate Data Are Available: Modified Methods to Include Disconnected Trials and Single-Arm Studies while Minimizing Bias by Howard Thom, Joy Leahy and Jeroen P. Jansen in Medical Decision Making
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institute for Health Research (NIHR) Bristol Biomedical Research Centre (BRC) and by the UK Medical Research Council (MRC) grant MR/S036709/1. The funding agreement ensured the authors’ independence in designing the study, interpreting the data, writing, and publishing the report.
Authors’ Note
Elements of this work have been presented by 1 or more of the authors: 2020, NMA on disconnected evidence networks: What can be done? Cochrane Training; 2019, NMA on disconnected evidence networks: What can be done? Liverpool University invited seminar; 2017, Thom H, Leahy J, Jansen J. Reference prediction to connect evidence networks. Methods in Meta-analysis meeting, London. Oral presentation; 2017, Thom H, Leahy J, Jansen J. Disconnected or Limited Evidence in Network Meta-Analysis: What can be done? ISPOR Glasgow. Workshop, Oral presentation.
Supplemental Material
Code and Data Availability
R and OpenBUGS Code and data to run ALM and reference prediction on a simple data set are provided in the Appendix. This is also available in a project, which will be easier to load into RStudio, at the repository https://github.com/Bogdasayen/disconnectednma-simple-example. Full code and data for the constructed data example are available at https://github.com/Bogdasayen/disconnectednma-constructed-data. Full code and data for the simulation study are available at ![]() .
.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.