Abstract
Cerebral small vessel disease (cSVD) is a major contributor to stroke, dementia, and cognitive decline. Despite significant progress through large-scale genome-wide association studies (GWAS) for cSVD and stroke, the genetic architecture underlying these conditions remains poorly understood. This review highlights recent advancements in statistical tools and provides a comprehensive overview of current insights into the genetic underpinnings of cSVD and stroke. We focus on the relevance of non-additive effects, local heritability, and polygenicity in shaping these traits. While single nucleotide polymorphism (SNP)-based heritability estimates for stroke and cSVD traits remain lower than pedigree-based estimates, we explore challenges and opportunities in addressing this “missing heritability.” In particular, we emphasize the importance of investigating both common and rare variants, to better characterize the genetic basis of cSVD. Furthermore, we discuss the role of negative selection in shaping complex disease traits and the relevance of the “omnigenic” model in the context of cSVD traits. In summary, we aim to provide a more nuanced understanding of cSVD and stroke genetics, paving the way for future research into their molecular mechanisms.
Introduction
In his groundbreaking 1918 paper, R.A. Fisher significantly advanced our understanding of the genetic underpinnings of variability in human traits. Fisher proposed that the inheritance patterns of quantitative traits, such as human height, could be explained by the combined effects of multiple genetic units within the framework of Mendelian inheritance. 1 This foundational concept laid the groundwork for the field of quantitative genetics. By the early 2000s, technological advancements in genomics made it possible to systematically study the genetic units through single-nucleotide polymorphisms (SNPs), enabling the analysis of millions of SNPs and their associations with phenotypic traits and diseases. These developments culminated in the creation of genome-wide association studies (GWAS), a powerful statistical approach used to identify associations between individual SNPs and complex traits or disease phenotypes across large cohorts of individuals.
The NHGRI-EBI catalog, curated by the National Human Genome Research Institute and the European Bioinformatics Institute, now encompasses nearly 6,500 GWASs investigating SNPs that commonly segregate within populations and their associations with various complex traits and diseases. Notably, nearly half a million SNPs from these GWASs exhibit statistical associations with complex diseases, even after accounting for the stringent genome-wide significance threshold.2,3 These associations span the entirety of the autosomes, involving numerous genetic loci, and extend to some degree into the sex chromosomes, emphasizing the polygenic nature of complex disease traits. In the context of neurological and neurodegenerative diseases, the polygenicity observed in GWASs amounts to more than 5,000 genetic loci conferring disease susceptibility. The polygenicity of complex traits is best explained by the “infinitesimal” genetic architecture, which suggests that the effects conferred by individual SNPs – typically those with a minor allele frequency (MAF) exceeding 1% or common and low-frequency variants – are modest and distributed across the genome, with each SNP contributing only a minimal proportion to the overall variance in trait levels or disease risk. 4 Noticeably, GWAS findings for any given genetic locus that may harbor a causal variant often include not only the sentinel SNPs achieving genome-wide significance but also other tagging SNPs in linkage disequilibrium (LD) with similar allele frequencies within the population. 5
In the following sections, we explore the evolving genetic architecture of cSVD and stroke, drawing on published GWAS findings as well as our own additional analyses that leverage recent advances in statistical genetics. We describe the contributions of GWAS to understanding both cerebral small vessel disease (cSVD) and stroke, including efforts to characterize shared genetic susceptibility between the two conditions, and to assess causal relationships between cSVD, stroke and its risk factors. Particular emphasis is placed on quantitative magnetic resonance imaging (MRI) markers of cSVD, which represent the different components of cSVD pathology but also serve as statistically powerful intermediate traits (endophenotypes) for applying the different methods. We begin by examining the contribution of non-additive genetic effects – such as epistasis – to heritability and highlight genomic regions that are particularly susceptible to gene-gene interactions. We then discuss the concept of polygenicity, which is increasingly viewed as a measurable spectrum rather than a binary feature. This perspective has important implications for trait prioritization in genetic studies, particularly for identifying traits where a smaller number of larger-effect cSVD genes may play a key role in stroke pathophysiology. We also provide a perspective on the sample sizes that future GWASs on cSVD may require to achieve heritability saturation levels comparable to well-studied complex traits like body mass index (BMI) and height. Finally, we provide an outlook on how cell-type-specific insights and early-life markers can inform and guide future genetic discovery efforts.
Current state of GWASs for cerebral small vessel disease (cSVD) and stroke
Under the infinitesimal genetic architecture model, detecting statistically significant associations for common and low-frequency genetic variants typically requires large sample sizes, ideally in the range of several tens of thousands to millions of participants. 6 Advances in statistical methodologies and meta-analysis techniques have enabled large-scale, population-wide GWASs, significantly expanding the GWAS studies. For example, a recent GWAS on stroke analyzed the additive effects of common SNPs in approximately 1.6 million participants, including 0.1 million stroke cases, by regressing stroke status on each additional count of the minor allele. 7 Similar large-scale studies with sample sizes ranging from 17,467 to 52,708 have investigated the additive associations of common SNPs with continuous markers of covert pathological processes, such as MRI markers of cSVD – a major cause of stroke that affects the brain’s small penetrating vessels.8–12 These studies incorporated traditional MRI-derived markers of cSVD, such as white matter hyperintensities (WMH) and perivascular spaces (PVS), which are predominantly detectable in older individuals and reflect advanced tissue damage. Additionally, they included markers of white matter microstructure from diffusion MRI (dMRI) that are more sensitive to structural alterations, capturing early-life changes predisposing to cSVD. Among dMRI measures, diffusion tensor imaging (DTI) is the most widely used, offering metrics like fractional anisotropy (FA), mean diffusivity (MD) and peak width of skeletonized mean diffusivity (PSMD), a robust DTI-based marker for cSVD and related cognitive impairment in older populations that has also been linked to slower processing speed in young adults (mean age: 22 years). 13 Together, GWASs of MRI and dMRI markers of cSVD, alongside studies of stroke and its subtypes, have identified approximately 140 LD-independent genetic loci – highlighting not only the shared genetic architecture between cSVD and stroke but also pointing to potential causal pathways linking subclinical vascular injury to clinical cerebrovascular events.
Shared genetic susceptibility, provides a lifespan perspective
Positional overlap of genome-wide significant variants from GWASs highlights the shared genetic architecture between cSVD and stroke that extends to specific stroke subtypes (large artery stroke [LAS], and small vessel stroke [SVS]) as well as to markers of white matter microstructures (FA and MD) (Table 1). Notable shared loci include ICA1L/NBEAL1/CARF1 (chr2q33.2), SH3PXD2A/STN1 (chr10q24.33), COL4A2 (chr13q34), and LAMC1 (chr1q25.3), between WMH, FA, MD and stroke subtypes (LAS, and SVS). The shared genetic susceptibility in addition to suggesting common molecular mechanisms also offers insight into the biology of cSVD across the lifespan pointing to early disease processes.9,11 For example, risk variants for WMH burden – particularly at KLHL24 (chr3q27.1), VCAN (chr5q14.2), SH3PXD2A (chr10q24.33), and NMT1 (chr17q21.31) – identified in older adults are also associated with variation in DTI metrics in young adults, including PSMD. 11 Similarly, several risk loci for white matter PVS burden in older adults (OPA1 [chr3q29], SLC13A3 [chr20q13.12], and CENPF/KCNK2 [chr1q41]) show significant associations with PVS burden in individuals in their twenties. 9
Positional overlap of genomic regions between cSVD traits and stroke and its subtypes.
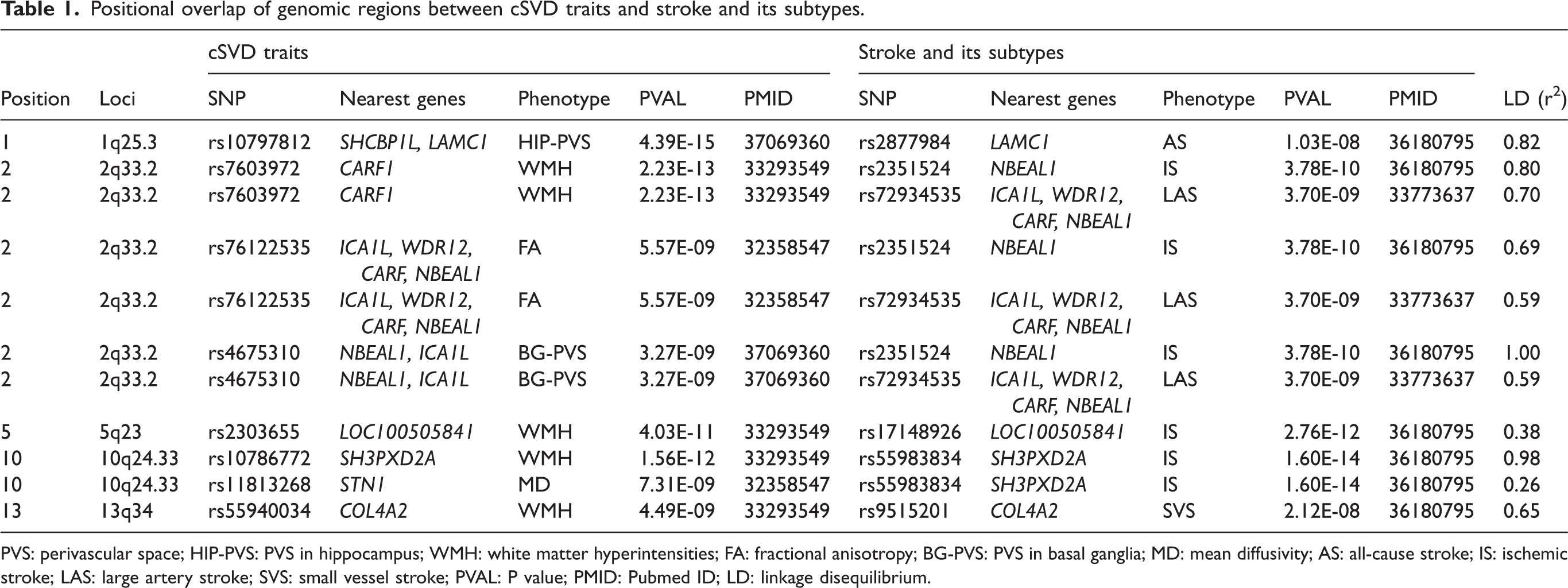
PVS: perivascular space; HIP-PVS: PVS in hippocampus; WMH: white matter hyperintensities; FA: fractional anisotropy; BG-PVS: PVS in basal ganglia; MD: mean diffusivity; AS: all-cause stroke; IS: ischemic stroke; LAS: large artery stroke; SVS: small vessel stroke; PVAL: P value; PMID: Pubmed ID; LD: linkage disequilibrium.
Multi-trait GWAS approaches offer valuable insights to positional overlaps by leveraging the genome-wide genetic correlations between related traits to identify risk loci with concordant effects and potential causal links. 14 This powerful strategy has been particularly effective for ischemic stroke (IS) subtypes, which often suffer from low statistical power due to limited sample sizes. Joint analyses of IS subtypes with coronary artery disease (CAD), atrial fibrillation and WMH, have uncovered several novel loci for LAS and cardioembolic stroke (CES). 7 For SVS, while the joint analysis with WMH confirmed several known loci (NBEAL1 [chr2q33.2], LOC100505841 [chr5q23], FOXF2 [chr6p25.3], HTRA1 [chr10q26.13], and COL4A2 [chr13q34]), no novel loci have been reported.7,15,16 This may be due to phenotypic heterogeneity in SVS case definitions, which can lead to misclassification, reduced effect size estimates, and lower statistical power.17,18 Encouragingly, studies using more refined phenotyping – such as radiologically confirmed SVS cases alongside standard clinical definitions – have begun to uncover novel loci, including VTA1-GPR126 [chr6q24.1-chr6q24.2], which is also associated with WMH. 15 Additional emerging strategies incorporating dMRI markers and deep learning-based segmentation that offers accurate, automated cSVD lesion characterization, 19 could further strengthen the genetic discovery addressing the low polygenicity of IS subtypes.
Since the majority of shared risk variants falls in the non-coding regions, they often provide limited clarity on the functional genes, requiring integration of GWAS data with other multi-omics to systematically characterize cSVD and stroke risk loci. Transcriptome wide association study (TWAS) framework – testing for the genes whose change in the expression levels explain the trait-association strength for a genetic variant – has identified genes such as NBEAL1 and ICA1L (chr2q33.2), where their higher expression levels are linked to lower white matter integrity (FA), larger WMH burden, and increased stroke risk (Table 2). Similar associations have been found for FAM117B (lower FA values) and SLK (larger WMH burden), with stroke risk.7,11,12 These findings are further supported by colocalization analyses showing that the same SNPs influence both gene expression (expression quantitative trait loci – eQTLs) and relevant brain traits. However, co-regulation of multiple nearby genes by the same eQTLs can complicate interpretation and limit the ability to pinpoint causal genes from TWAS alone. 20 In contrast, proteomics offers a more targeted approach by focusing on biologically discrete, stable entities rather than the more dynamic nature of gene expression. Recent proteome-wide association studies (PWAS) have identified SNPs functioning as cis-protein quantitative trait loci (pQTLs) influencing the protein levels of proximal genes.21–23 When applied within a Mendelian randomization (MR) framework – where cis-pQTLs are less likely to directly associate with the outcome (Figure 1), 24 MR studies showed that higher levels of TFPI, IL16RA, and CD40 in plasma are associated with reduced risk of SVS, with the effect of TFPI likely mediated through lower WMH burden.25,26 Similar protective associations have been observed for F11, KLKB1, and PROC, which are involved in coagulation and inflammatory pathways. Notably, many of these proteins are targets of existing drugs with established safety profiles, thereby underscoring the potential of GWAS-informed drug repurposing.7,27
TWAS identified shared genetic susceptibility between cSVD and stroke.
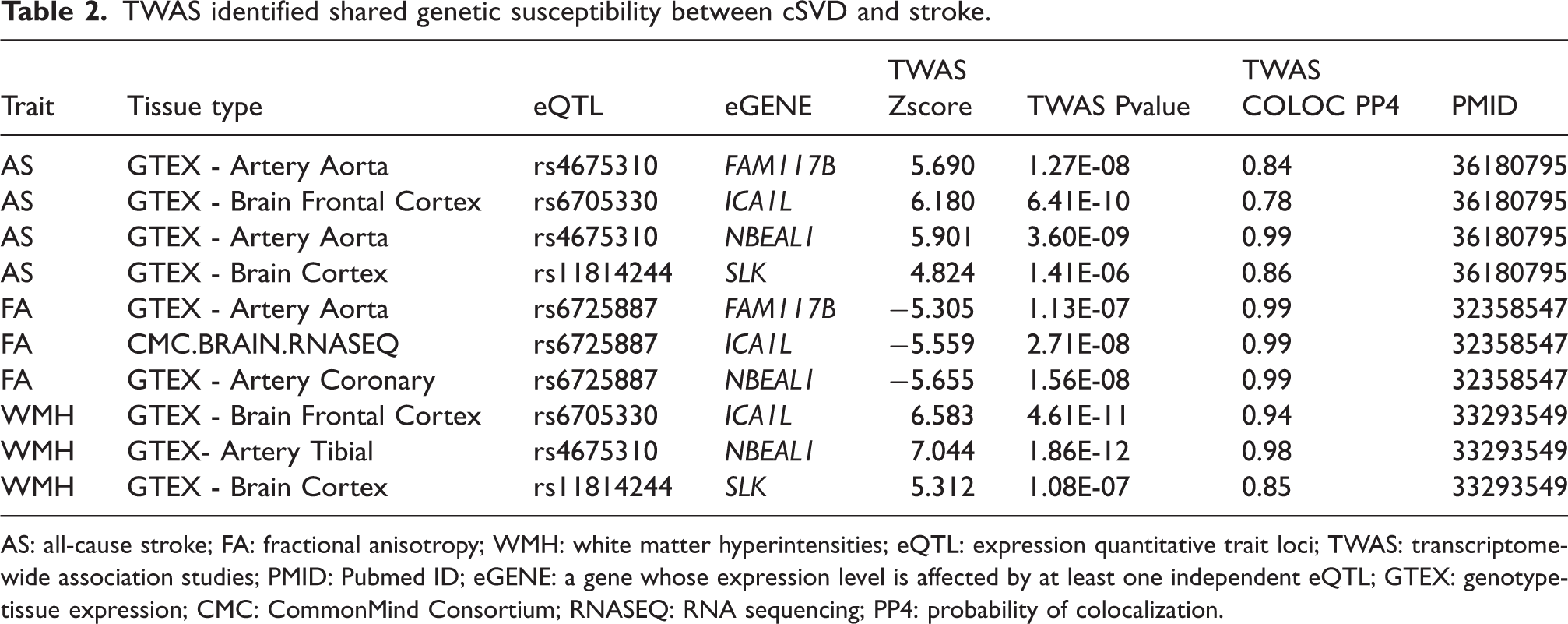
AS: all-cause stroke; FA: fractional anisotropy; WMH: white matter hyperintensities; eQTL: expression quantitative trait loci; TWAS: transcriptome-wide association studies; PMID: Pubmed ID; eGENE: a gene whose expression level is affected by at least one independent eQTL; GTEX: genotype-tissue expression; CMC: CommonMind Consortium; RNASEQ: RNA sequencing; PP4: probability of colocalization.
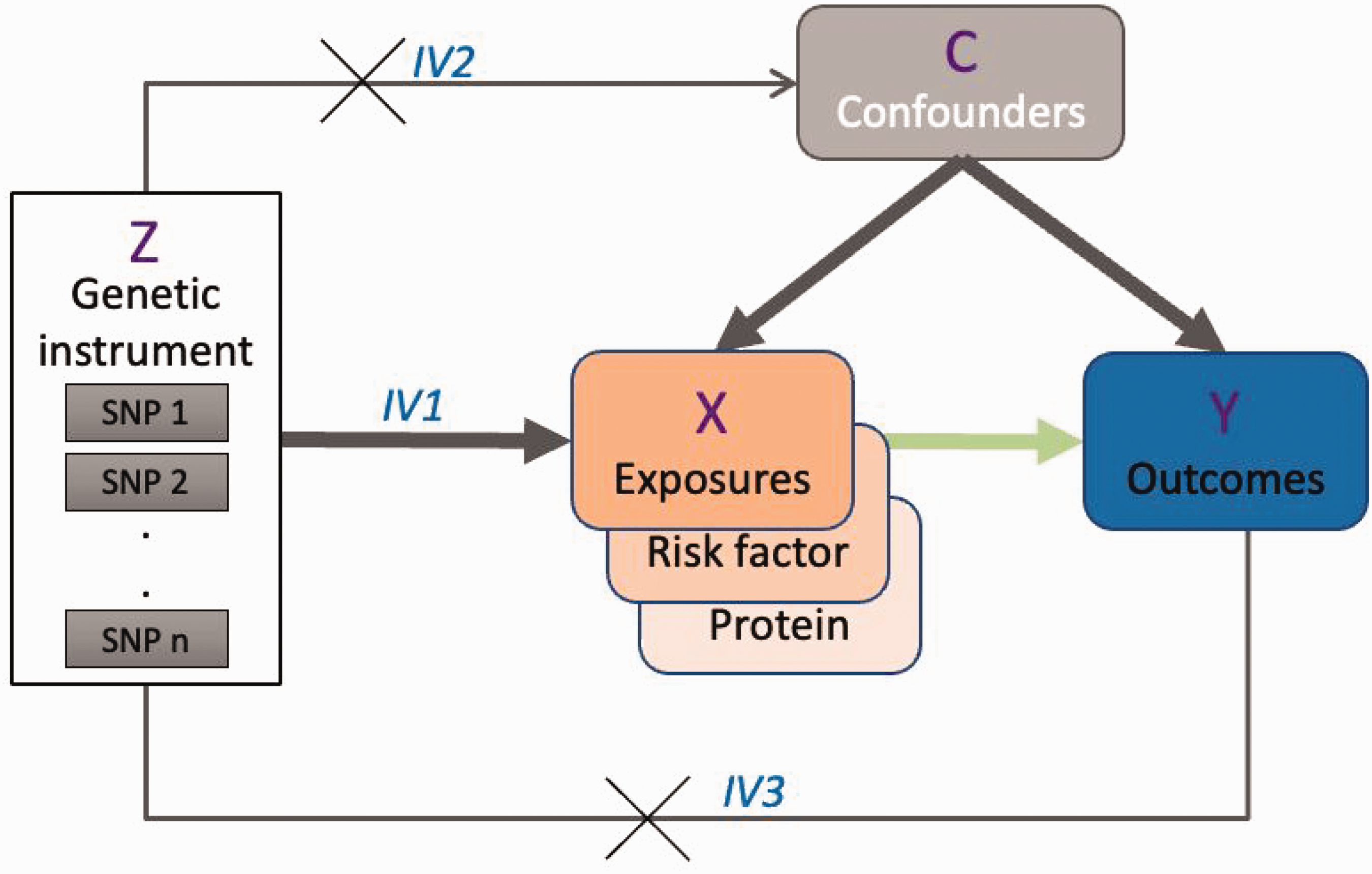
Directed acyclic graph on the causal association of a given exposure (risk factor or protein levels) with an outcome. Genetic instruments (Z) satisfying the key MR assumptions: IV1 – Z strongly predictive of the exposure (X); IV2 – absence of correlated pleiotropy of Z through confounders (C); IV3 – absence of Z directly associated with the outcome (Y).
MR applications in distinguishing causation from correlation
Individual genetic variants explain only a small portion of the risk for cSVD and stroke, limiting their usefulness in distinguishing causation from correlation. However, by combining multiple variants that collectively capture more variation in a risk factor, MR can help assess whether that factor causally influences an outcome through a specific biological pathway (Figure 1), while reducing confounding from shared influences. 28 A substantial body of literature has applied MR to examine the causal roles of various exposures in relation to stroke and cSVD. For instance, a MEDLINE search using terms like “Mendelian,” “randomization/randomisation,” and “stroke” or “cerebrovascular disease” yields 142 relevant observational studies. While a comprehensive review of this literature is beyond the scope of this discussion, several key findings on shared risk factors for stroke and cSVD warrant emphasis.
Notable findings include a putative causal link between genetically predicted higher WMH volume and increased risk of SVS and deep intracerebral hemorrhage. 11 A similar trend was observed for both ischemic and hemorrhagic stroke when incorporating risk variants associated with perivascular space (PVS) burden, particularly in the basal ganglia and hippocampus. 9 These findings strengthen prior epidemiological evidence by providing genetic support for a shared small vessel arteriopathy underlying both stroke subtypes.29,30 Moreover, as MR aggregates effects across multiple loci, the results point to the involvement of additional cSVD-related genes – such as NBEAL1, ICA1L, and PMF1 31 – alongside the well-established role of COL4A2 in hemorrhagic stroke. 32 Such observations linking intermediate markers and a range of related clinical endpoints also highlight potential implications for therapy, as antiplatelet agents are often prescribed empirically for individuals with extensive MRI markers of cSVD but no clinical stroke. The potential adverse effects of WMH on both stroke types highlight the need for randomized trials to assess the risk-benefit profile. Indeed, the Secondary Prevention of Small Subcortical Strokes (SPS3) trial showed a significant increase in hemorrhagic events and mortality with dual antiplatelet therapy in patients with symptomatic stroke. 33
At the risk factor level, genetic predisposition to elevated systolic and diastolic blood pressure (SBP & DBP) has shown consistent associations with increased IS risk, particularly for the LAS and SVS subtypes, while weaker associations were observed for CES. 34 This underscores the central role of hypertension in the pathogenesis of LAS and SVS, while atrial fibrillation appears more strongly associated with CES. 35 Genetically predicted blood pressure also showed concordant effects on cSVD markers such as WMH and PVS burden.9,11 Notably, larger WMH volumes have been observed even in individuals without clinically defined hypertension at the time of MRI. 11 Given findings from the SPRINT-MIND trial, which demonstrated that intensive blood pressure control reduces WMH progression and lowers dementia risk,36,37 such results support exploring similar interventions in high-risk individuals who do not yet meet the clinical threshold for hypertension. Additionally, increased pulse pressure – an indicator of arterial stiffness – has been independently associated with both IS and WMH burden,34,38 suggesting that age-related vascular changes contribute to cerebrovascular pathology through impaired cerebral blood flow, endothelial dysfunction, and small vessel remodeling. 39 Lipid traits also exhibit subtype-specific effects. Higher genetically predicted low-density lipoprotein (LDL) cholesterol levels are associated with greater LAS risk,40,41 whereas elevated high-density lipoprotein (HDL) levels are associated with reduced risk for SVS and WMH. 42 Further dissection of lipid subfractions implicates apolipoprotein B-containing lipoproteins as the principal mediators of LDL-related stroke risk. 43 Among metabolic and lifestyle factors, type 2 diabetes and smoking severity showed the strongest associations with increased SVS and WMH burden, 11 but not CES,44,45 indicating their more pronounced influence on microvascular brain injury than on macrovascular atherosclerosis.
Challenges and emerging approaches
Despite these significant advancements, GWAS, in principle, have captured only a limited proportion of phenotypic variance (heritability) for the cSVD traits and stroke. This limitation stems from their exclusive focus on common and low-frequency SNPs and their predominantly reductionist approach. Indeed, most studies to date have concentrated on genetic loci associated with additive effects, while non-additive effects, such as dominance, epistasis (gene-gene interactions), and gene-environment interactions, are rarely considered. Methodological challenges in genotype-phenotype association at the individual-cohort level, such as the complexity of modelling interaction variables, analyzing sex chromosomes, and sample size constraints, impede the ability to account for such factors that substantially influence individual genetic susceptibility. However, continued advancements in statistical methods leveraging association results from the meta-analysis of individual cohorts, enables a systematic data-driven learning on the genetic architecture of cSVD and stroke.
Heritability and the future of GWASs for cSVD and stroke
Under the assumption that genetic effects from common variants are uniformly distributed, GWAS summary statistics are increasingly used to estimate the proportion of phenotypic variance explained by common SNPs, often termed ‘SNP-based heritability’.46,47 A widely used method in this domain is LD-score regression (LDSC), a variance-component approach that regresses the variance of SNP-level effect sizes against an LD score.46,48 The LD score measures the genetic correlation between a given SNP and its neighboring tagging SNPs, which may harbor the causal variant. The slope derived from this regression provides an estimate of the trait’s SNP-based heritability (h2) across the genome. h2 values are expressed on a scale from 0 to 1, where values near 0 suggest that phenotypic variation is primarily due to environmental factors, while values near 1 indicate that genetic factors account for most of the variation. When applied to GWAS data for stroke and imaging-derived (MRI, DTI) measures of cSVD, LDSC typically produces SNP-based heritability estimates ranging from 0.2% to 0.6% and 6% to 20%, respectively (Table 3). These values are markedly lower than estimates derived from pedigree-based methods, which capture correlations between large genetic segments shared by identity by descent and disease risk. Incorporating genetic interactions and shared environmental influences, family/pedigree-based approaches have explained up to 40–80% of the phenotypic variance in stroke49–51 and as much as 80% in cSVD traits.52–55 However, large-scale pedigree data have shown that phenotypic correlations between spouses can be as strong as those observed among biological relatives. 56 Importantly, genetic components commonly used to adjust for population stratification (i.e. correlations in the allele frequency due to ancestry sharing) have also been found to correlate between spouses. 57 This indicates that assortative mating, where individuals with similar traits or social backgrounds are more likely to pair, introduces non-linear, shared environmental and lifestyle exposures that can confound or upward bias the heritability estimates. 58 Nevertheless, SNP-based heritability estimates for stroke remain substantially lower than those from pedigree-based models – a gap referred to as the missing heritability problem. This discrepancy likely reflects the highly polygenic nature of stroke and cSVD, where numerous common variants contribute small individual effects. It also underscores methodological limitations of SNP-based approaches, which often fail to capture non-additive genetic effects, rare genetic variations and gene-environment interactions. Bridging this gap is essential for reconciling heritability estimates across methods and gaining a more complete understanding of the genetic architecture underlying stroke and cSVD.
Heritability estimates based on the additive, non-additive effects & relationship between MAF and genetic effect size – negative selection (S).
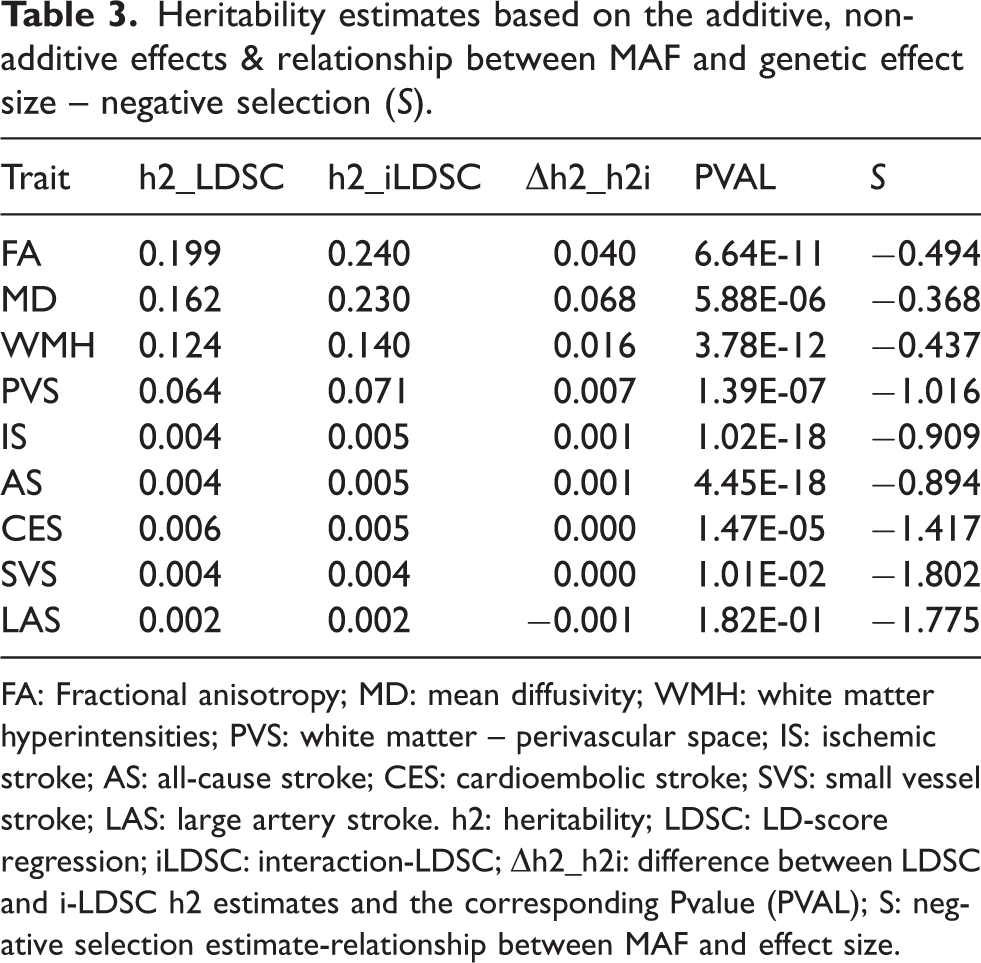
FA: Fractional anisotropy; MD: mean diffusivity; WMH: white matter hyperintensities; PVS: white matter – perivascular space; IS: ischemic stroke; AS: all-cause stroke; CES: cardioembolic stroke; SVS: small vessel stroke; LAS: large artery stroke. h2: heritability; LDSC: LD-score regression; iLDSC: interaction-LDSC; Δh2_h2i: difference between LDSC and i-LDSC h2 estimates and the corresponding Pvalue (PVAL); S: negative selection estimate-relationship between MAF and effect size.
Non-additive effects and its role in loci-level and genome-wide heritability estimation
While most variance-component methods focus on additive genetic effects, recent advances have made it possible to estimate non-additive contributions – such as epistasis – by leveraging SNP correlation structures. 59 One such method, interaction LD score regression (i-LDSC), extends the traditional LDSC framework by incorporating cis-SNP interaction LD scores, which capture local pairwise interactions between neighboring SNPs using population-matched reference genotype panels. By regressing standardized GWAS effect sizes on both standard and cis-SNP interaction LD scores, i-LDSC can estimate genome-wide heritability attributable to both additive and tagged non-additive effects. 60 Applying i-LDSC to European-only GWAS of stroke and cSVD imaging markers, we observed statistically significant contributions from epistatic components. Although the difference between the additive and the joint (additive + non-additive) heritability estimates remained modest (Δh2_h2i, Table 3), our findings align with evidence from the UK Biobank and other large population-based studies showing that genetic variance in complex traits, including white matter microstructure (e.g., FA), is predominantly additive, with only minor contributions from non-additive effects.61,62 However, such results should be interpreted with caution, as most methods assume a normally distributed effect of non-additive interactions across the genome – an assumption that may oversimplify the genetic architecture and prove invalid at specific loci.63,64 This can lead to underestimation of epistatic contributions to heritability. More targeted, locus-specific approaches are needed to capture the true impact of non-additive interactions. Such analyses could also reveal specific genomic regions where genes interact in network to influence key disease mechanisms. Supporting this, a genome-wide SNP-SNP interaction study of regional brain volumes (e.g., the temporal lobe) found concentrated epistatic effects in loci encoding transcription factors involved in inflammation, explaining nearly 6% of trait variance – substantial when compared to the 17.5% variance explained by additive effects alone.65,66
Extending the observation that epistatic effects may concentrate in specific genomic regions, we applied a locus-level heritability estimator – Heritability Estimator from Summary Statistics (HESS), 67 which, unlike many other methods, does not assume a predefined distribution of effect sizes. Instead, HESS estimates local heritability-h2 by projecting GWAS effect sizes onto the eigenvectors of the SNP correlation (LD) matrix for each locus, retaining only the most informative components to reduce noise. The squared contributions of these projections are then summed to yield the heritability estimate for each region. This data-driven approach allows HESS to learn the underlying effect size distribution directly from the observed LD structure.
In earlier work, we used HESS to identify genomic regions with disproportionately high contributions to WMH heritability. 11 In this review, we extend that analysis to additional cSVD traits and identify several loci with significant local heritability-h2. These loci may harbor genes that are more susceptible to interaction effects, whether through gene-gene (epistatic) mechanisms or through interactions with external environmental or lifestyle factors (Figure 2). This is consistent with prior findings suggesting that epistatic variance, though modest in total, tends to be concentrated in specific genomic “hotspots” where networks of interacting genes reside.61,68 Notably, several loci identified through HESS did not contain SNPs meeting genome-wide significance thresholds, suggesting the presence of potentially novel regions of interest – such as 6p22.1 and 11q11 for WMH, and 10q21.2 for MD (Table 4). A key strength of HESS is its ability to correct for the over-adjustment of effect sizes introduced by genomic control, which is commonly applied to account for population stratification but can inadvertently suppress true signal, especially in polygenic traits within homogeneous cohorts. 69 By reinflating association statistics, HESS recovers more accurate heritability estimates. When summing heritability across independent loci, HESS-derived genome-wide estimates (HESS-GW h2) often exceed those obtained from LDSC, suggesting that a greater portion of trait variance may already be captured through data-driven modeling of the underlying genetic effects (Table 4). Thus, although the genetic architecture of complex traits like cSVD is largely additive, partitioning heritability at the locus level may help identify regions that are particularly enriched for higher-order interactions. These regions represent promising candidates for future studies investigating how genetic susceptibility may be modified by environmental or lifestyle exposures.
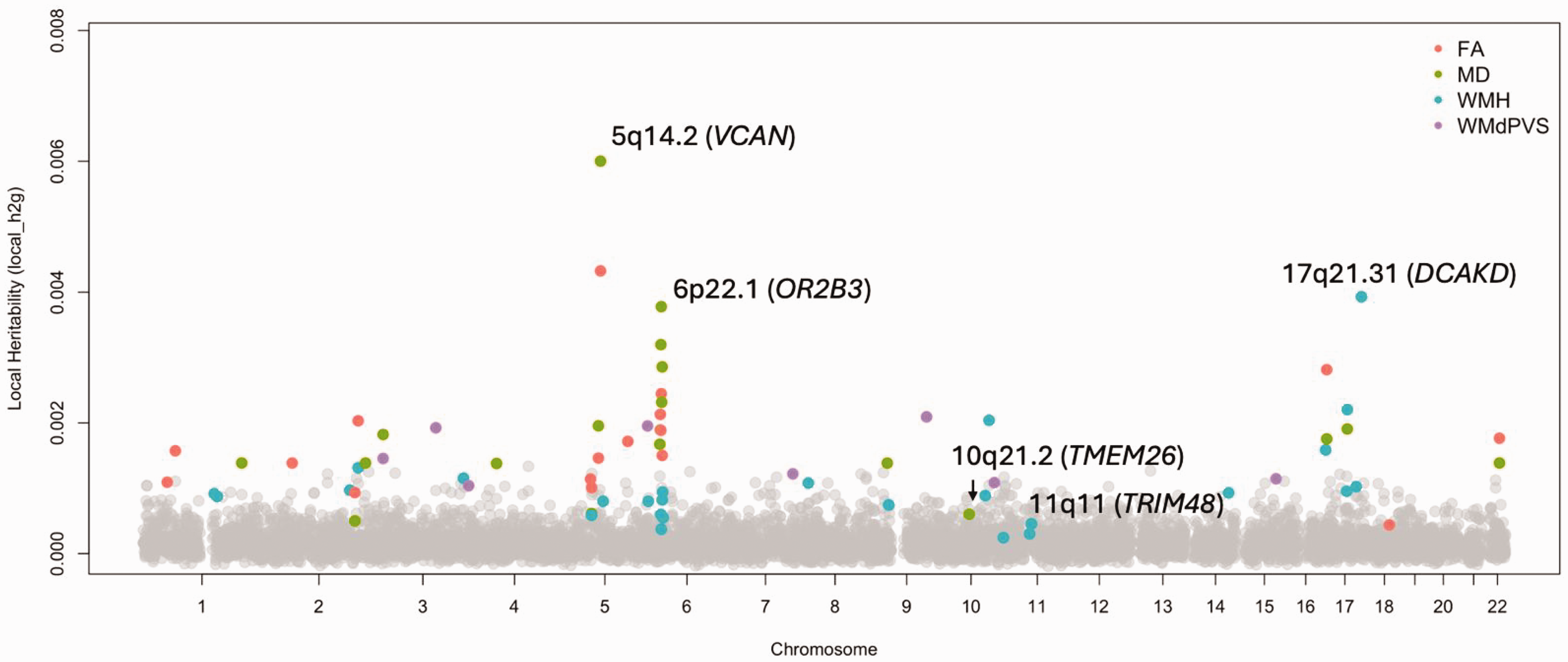
Loci-level heritability of cSVD traits. Colored dots represent non-zero h2 estimates (P < 0.05). Gene names correspond to loci with genome-wide significant (P < 2.94E-05) loci-level h2. FA: Fractional anisotropy, MD: Mean diffusivity; WMH: white matter hyperintensities; WMPVS: white matter – perivascular space.
Loci with significant regional-level heritability estimates across cSVD traits.
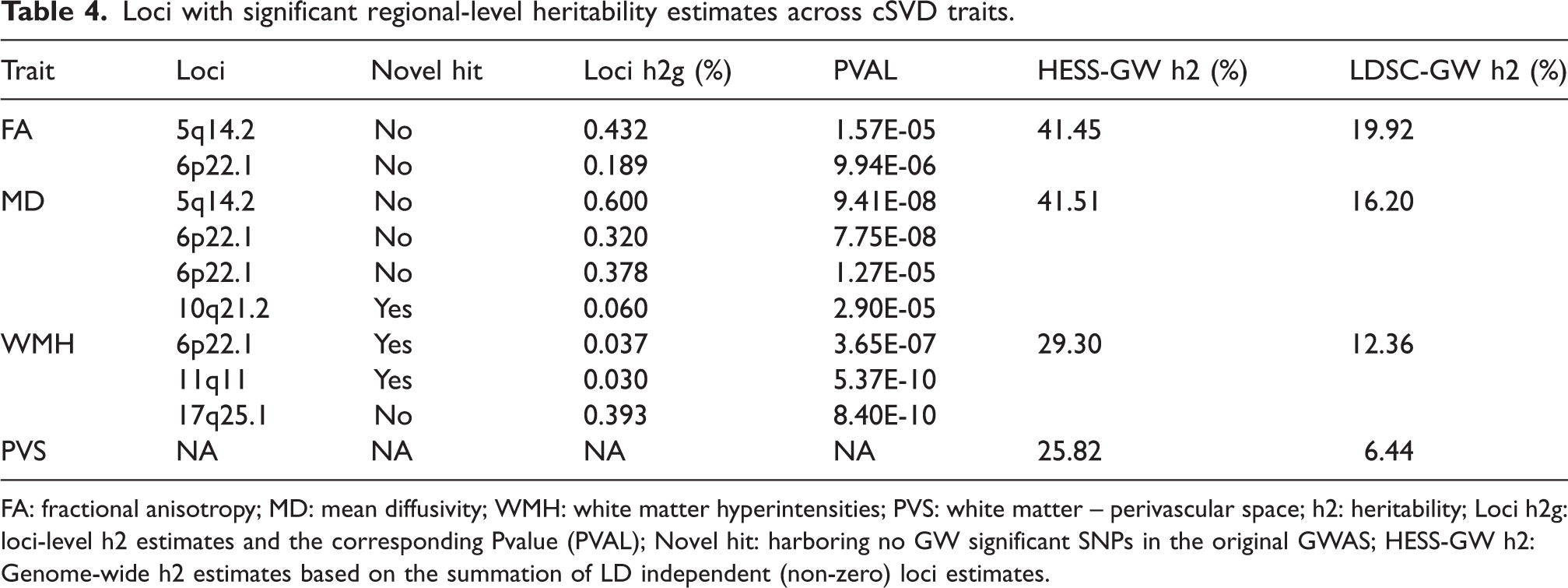
FA: fractional anisotropy; MD: mean diffusivity; WMH: white matter hyperintensities; PVS: white matter – perivascular space; h2: heritability; Loci h2g: loci-level h2 estimates and the corresponding Pvalue (PVAL); Novel hit: harboring no GW significant SNPs in the original GWAS; HESS-GW h2: Genome-wide h2 estimates based on the summation of LD independent (non-zero) loci estimates.
Addressing the challenge of missing heritability
Variance-component methods, by design, estimate heritability using only the variance (second moment) of the genetic effect size distribution. Recent developments have extended this framework to incorporate higher moments, such as the fourth moment, which capture the shape and spread of genetic effects.70,71 Fourier Mixture Regression (FMR) is one such method, that decomposes the effect size distribution into components ranging from the smallest to largest empirical variances and regresses these against modified LD scores tailored to each component. By including sample size as a predictor, FMR also estimates the sample size and number of genome-wide significant loci required to capture a target proportion (e.g., 50–90%) of SNP-based heritability for a given trait. Incorporating higher moments in this way has provided a more refined understanding of genetic architecture and the sample sizes needed to saturate SNP-based heritability in several complex diseases and traits. 72
In this study, we systematically applied FMR to continuous cSVD traits only - WMH, PVS, and MD, since stroke and its subtypes exhibit low genome-wide SNP-based heritability, limiting FMR’s applicability. It is worth noting that continuous traits generally yield narrower standard errors and more precise FMR estimates than binary outcomes like stroke, further justifying this focus. For the cSVD traits analyzed – where current SNP-based heritability estimates range from 11% to 32% – FMR suggests that a sample size of approximately 8 million would be needed for PVS to reach 50% SNP-based heritability (Figure 3). In contrast, WMH and MD would require far smaller samples – about 190,000 and 120,000, respectively. Notably, 50% of SNP-based heritability for WMH and MD could be explained by approximately 130 and 160 genome-wide significant loci, whereas PVS would require over 4,000, highlighting its more polygenic nature. Polygenic scores constructed from these subsets of loci captured most of the trait variance for WMH (r2 = 0.895) and MD (r2 = 0.902), suggesting which traits may benefit most from expanded GWAS efforts. Finally, consistent with findings in other complex traits, 72 achieving 90% SNP-based heritability would require sample sizes nearly ten times larger than those needed to reach 50%.
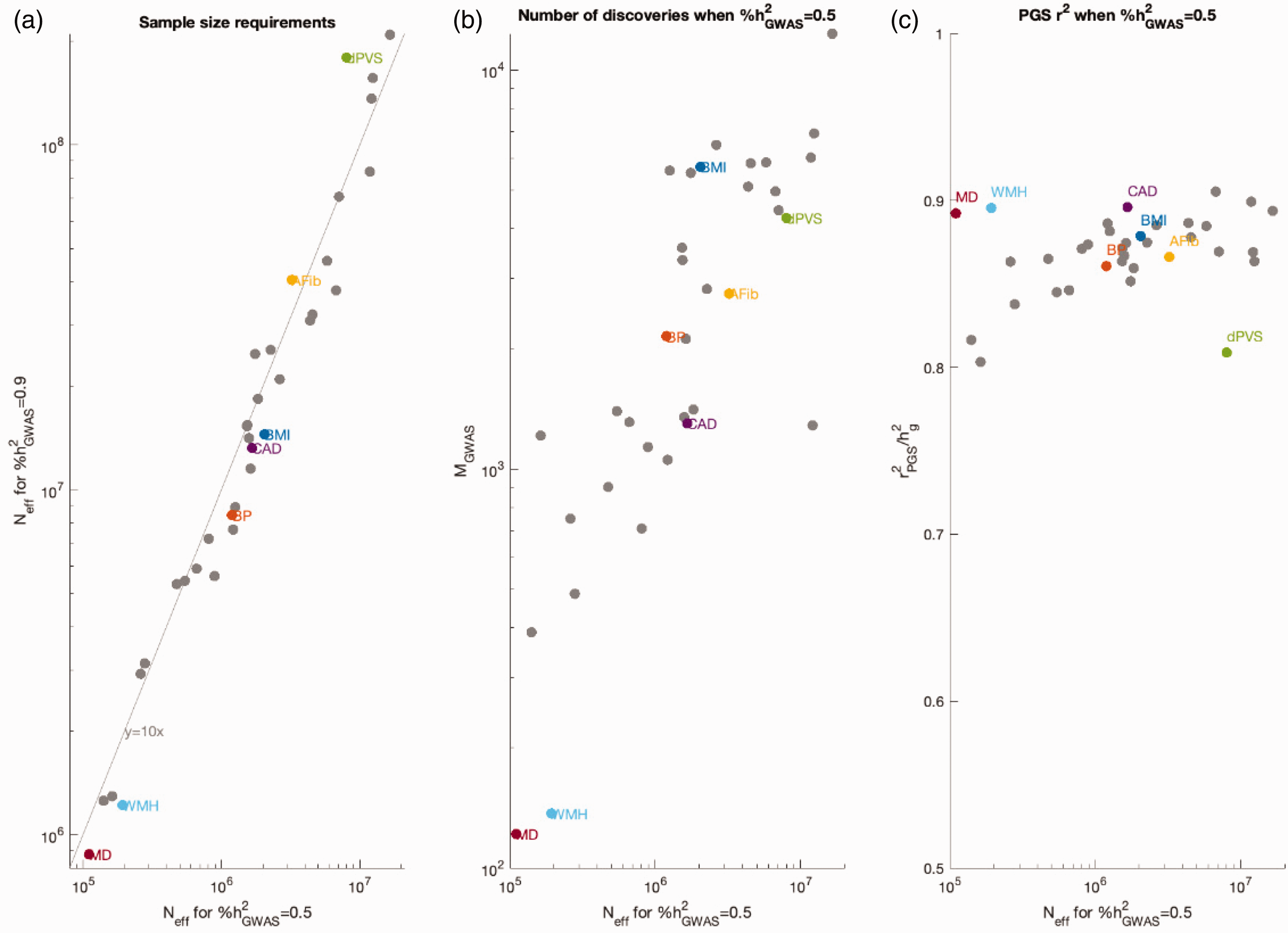
Sample size requirements and predictions for future GWAS, including cSVD traits. (a) Effective (Neff) sample size required for genome-wide significant SNPs to explain 50% or 90% of SNP heritability. (b) Predicted number of independent SNPs (MGWAS) at %h2GWAS = 0.5. (c) Theoretical maximum PGS accuracy, defined as PGS r2, at %h2GWAS = 0.5. MD: mean diffusivity; WMH: white matter hyperintensities; PVS: white matter –perivascular space; AFib: atrial fibrillation; CAD: coronary artery disease; BP: blood pressure; BMI: body mass index. (Reproduced with permission from Luke J. O’Connor, Connor L.J et al., 2021, for non cSVD traits).
Insights from rare-variant association studies on heritability and polygenicity
Unlike GWAS, which primarily captures common and low-frequency variants (MAF >1%), rare variants (MAF <1%) profiled through whole-exome or whole-genome sequencing (WES/WGS) efforts, aids in the identification of variants with potentially larger trait-effect sizes. Efforts such as the Trans-Omics for Precision Medicine (TOPMed) program have enabled harmonized WGS and WES analysis by pooling biological and clinical data across multiple cohorts. For instance, a recent WGS study of stroke (6,833 cases and 27,116 controls) identified novel loci, including 7q22, 13q33, AUTS2, RAP1GAP2, and TEX13C. 73 Rare variant analyses of cSVD traits have also begun to yield additional insights. WES-based studies of WMH burden identified novel rare variants in genes such as ALCAM and GBA3. 74 Notably, the association of ALCAM with WMH is also supported by suggestive associations in common variant GWAS, 11 suggesting convergence between rare and common variant architecture. ALCAM encodes an endothelial adhesion molecule involved in immune trafficking and inflammation, and circulating protein levels of ALCAM have been previously linked to stroke outcomes, 75 highlighting its potential as a shared biomarker for both risk and prognosis. This convergence underscores the utility of integrating rare and common variant data to refine fine-mapping efforts. In many cases, the phenotypic effects of rare variants may be partially tagged by common variants in the same gene or pathway. 76 This supports a model in which trait-associated genes act along a mechanistic continuum - from subtle regulatory effects that are observed at the GWAS level to rare, high-impact coding or structural variants that influence gene function and downstream biology. A particularly effective strategy to detect rare, functional variants has been the use of dichotomized intermediate phenotypes.77,78 Recently by combining GWAS and WES study of individuals with extreme cSVD burden (defined by WMH distribution and the presence or absence of lacunes) identified NOTCH3 – a gene earlier implicated in cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL) – a monogenic form of cSVD.10,79 This finding points to a phenotypic continuum between monogenic and sporadic forms of cSVD, highlighting shared mechanisms across the disease spectrum. 80
Evidence from WGS studies also underscores the critical role of rare variants in recovering the missing heritability of complex traits. Heritability estimates derived from WGS association statistics reveal that additive, or narrow-sense, heritability is predominantly concentrated in low-LD regions, with rare protein-altering variants contributing significantly. 81 This preferential enrichment of heritability in rare variants and low-LD regions highlights a fundamental limitation of current GWAS methods. Widely used reference panels, such as the 1000 Genomes and Haplotype Reference Consortium, lack sufficient resolution to accurately capture these regions. While advances such as deep-learning neural networks and larger reference panels (e.g., TOPMed consortium, >100 K samples) have enhanced imputation accuracy for some frequency bins (10−³ to 10−4), accuracy remains suboptimal in frequency ranges where most heritability is concentrated for complex disease traits.82,83 Consequently, WGS approaches are indispensable for saturating additive genetic variance and identifying actionable causal variants with potentially large effect sizes. Second, the inverse relationship between allele frequency and the phenotypic variance explained highlights the important role of negative selection, wherein evolutionary forces continually eliminate variants with large phenotypic effects, preventing them from becoming common.84–87 Interestingly, quantifying negative selection through the relationship between allele frequency and effect sizes reveals considerable variability across complex disease traits.85,88 For example, Zeng et al., 85 demonstrated that stroke is subjected to stronger negative selection compared to closely related vascular risk factors (e.g. blood pressure traits, type 2 diabetes [T2D]). Using the same method, as in Zeng et al., 85 “SbayesS” we estimated the selection signatures (S) across stroke and its subtypes and cSVD traits (Table 3). Briefly, “SBayesS” is a Bayesian framework that models standardized effect sizes from GWAS as a function of SNP correlation structure (LD matrix), the expected distribution of genetic effect sizes, and allele frequency. It estimates the selection parameter S, which quantifies the relationship between a variant’s minor allele frequency and the variance of its effect on the trait. An S-value less than zero indicates evidence of purifying (negative) selection, where variants with larger effects tend to be rarer in the population due to selective pressure against deleterious alleles. In addition to confirming the stronger negative selection observed for stroke, our additional analysis revealed that this selection pressure extends to its subtypes (SVS, LAS, and CES). In contrast, the cSVD traits (FA, MD, and WMH) exhibited relatively weaker selection pressures. Negative selection serves as a key determinant in flattening the heritability of complex traits by limiting the prevalence of large-effect variants. In support, the heritability values for stroke with substantial negative selection pressure are markedly lower compared to cSVD traits. This “flattening” could also drive the extreme polygenicity observed in complex traits by distributing the genetic effects across multiple loci as small effects. The varied selection pressures suggest possible differences in the polygenic architectures of complex traits, emphasizing the importance of systematic studies of polygenicity. Such investigations can provide valuable biological insights, even if the heritability for a given trait remains modest.
Genetic architecture of cSVD traits and stroke
Polygenicity is conventionally interpreted as a state distinct from monogenic disease forms, classifying a given trait as either polygenic or not (possibly monogenic). However, recent studies reveal that closely related traits can exhibit substantial variability in polygenicity despite having similar heritability. 89 Understanding this variability could help guide future genetic association studies to prioritize traits that are less polygenic and possibly driven by risk loci with large effects, better identifying the underlying mechanisms. Indeed, common variants within genes associated with monogenic forms of disease conditions are also linked to sporadic forms, demonstrating a genetic continuum between monogenic and polygenic traits. For example, disease traits, such as schizophrenia, serve as model examples of high polygenicity,90,91 characterized by the distribution of small effect sizes across numerous common variants in the genome. Conversely, neurodevelopmental disorders driven by a selective set of genes with large phenotypic effects,91,92 represent the opposite end of the spectrum. Despite these distinctions, studies reveal that common variants within genes implicated in neurodevelopmental disorders of monogenic basis can cumulatively influence the penetrance of rare variants, thereby increasing the overall risk of schizophrenia. 93 Similarly, cSVD-related phenotypes provide a compelling case study of such continuum. Genes such as HTRA1 and COL4A1 harbor both common and rare variants associated with sporadic, multifactorial cSVD, as well as rare causal mutations driving monogenic forms of the condition.16,80,94 Moreover, indirect evidence from the integration of GWAS with single-cell sequencing approaches across the lifespan suggests an important role of developmental factors for cSVD as well.9,11,95
In this context, we systematically quantified the polygenicity of cSVD, stroke, and related traits using a mathematical framework proposed by Connor et al., 2019. 89 This method referred to as stratified LD fourth moments regression (S-LD4M) regresses the GWAS association strength (or chi-square statistic) on to the underlying effect size distributions reflecting how evenly the explained heritability is distributed across genomic loci, and estimating the effective number of independently associated loci (Me) that serves as a proxy measure for polygenicity. The estimated Me values were then compared against pre-computed values for other complex traits and diseases. 89 Given that the polygenicity is likely to vary across the different biologically important functional categories,96,97 we additionally partitioned the polygenicity-Me across different genomic annotations, including coding, regulatory, and conserved regions, alongside baseline genomic estimates.
Polygenicity-driven trait prioritization for future GWASs
At the baseline model, as expected, traits exhibiting an infinitesimal genetic architecture, characterized by small effects distributed across the genome, displayed higher Me values, indicative of greater polygenicity. In contrast, traits driven by large genetic effects concentrated within specific genomic regions demonstrated lower polygenicity, reflected by lower Me values (Figure 4). Lifestyle and vascular risk factors (e.g., smoking, SBP, and BMI) displayed high polygenicity comparable to height, which serves as a prototypical example of the infinitesimal genetic model. Stroke (all-cause stroke) also displayed a high level of polygenicity, consistent with its complex genetic architecture involving numerous small-effect loci. However, imaging traits capturing early-life changes predisposing to cSVD (FA and MD) and traits tied to specific pathophysiological mechanisms or molecular pathways, such as inflammatory processes, exhibited comparatively lower polygenicity. These traits tend to be influenced by a narrower set of genes, often including variants with larger, disease-specific effects – such as APOE in Alzheimer’s disease, DNASE1 in autoimmune disorders, and TCF7L2 in type 2 diabetes. Interestingly, MD, which is closely associated with microglial activity and implicated in the formation of inflammatory demyelinated lesions, 98 exhibited a similar level of low polygenicity. This finding is consistent with observations of weaker negative selection for imaging endophenotypes (Table 3), implying a genetic burden shifted toward fewer loci with larger effects.84,85 Weaker negative selection reduces the efficiency of purging deleterious mutations with large phenotypic effects, thereby allowing these loci to disproportionately influence the trait and ultimately decreasing its polygenicity. Supporting this, a comprehensive study comparing multi-organ endophenotypes (including 2,016 brain-derived phenotypes) with disease endpoints, using biobank-scale datasets such as UK Biobank and FinnGen, revealed that endophenotypes typically exhibit lower polygenicity and weaker negative selection effects compared to clinical endpoints. 99 This distinction highlights the unique genetic architecture of endophenotypes, which often capture intermediate phenotypic expressions that are closer to the causal mechanisms of disease. Overall, suggesting that intermediate phenotypes are more directly aligned with disease etiology within the causal pathway. 100 Focusing future GWAS efforts on such traits could yield deeper insights into the etiology of cSVD and stroke. Indeed, genetic risk score approaches jointly studying the genetic architecture of imaging endophenotypes of cSVD and vascular risk factors, and disease endpoints consistently show that such endophenotypes provide a more precise measure of organ damage resulting from risk factors.101,102
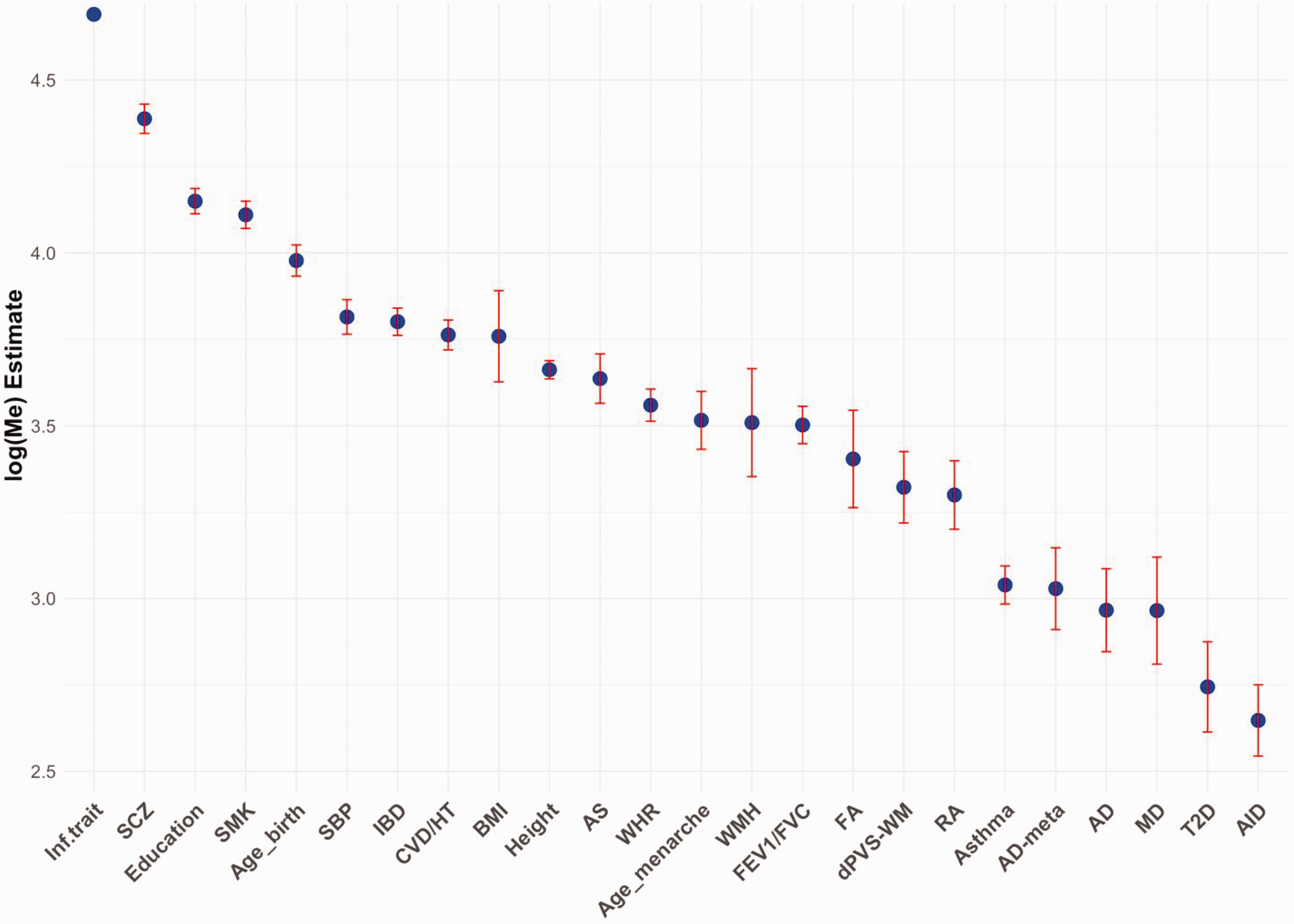
Comparison of polygenicity (Me) estimates for stroke and cSVD traits with other complex-disease traits. Inf. Trait: Model trait with infinitesimal architecture. SCZ: schizophrenia; SMK: smoking; SBP: systolic blood pressure; IBD: inflammatory bowel disease; CVD: cardiovascular disease; HT: hypertension; BMI: body mass index; AS: all-cause stroke; WHR: waist-hip ratio; WMH: white matter hyperintensities; FA: fractional anisotropy; PVS-WM: white matter – perivascular spaces; RA: rheumatoid arthritis; AD-meta: Alzheimer’s disease including parental history; MD: mean diffusivity; T2D: type 2 diabetes; AID: auto-immune disorder.
Second, for each trait, polygenicity estimates were partitioned across different genomic functional categories, and polygenicity enrichment was calculated. Polygenicity enrichment is defined as the proportion of polygenicity attributed to a specific annotation category relative to the baseline model. As per the infinitesimal genetic architecture, consistent enrichment values (>0.50) were observed across multiple genomic categories (Supplementary figure 1) for all the investigated traits. Regions involved in the regulation of gene expression and function exhibited a relatively higher concentration of polygenicity. These included gene-regulatory regions, such as promoters, enhancers, and transcription factor binding sites, as well as highly conserved genomic segments identified through Genomic Evolutionary Rate Profiling (GERP). Highly conserved regions are known to harbor genes integral to core cellular and molecular processes and are under strong purifying selection, which prevents the accumulation of deleterious mutations. 103 The observed high polygenicity enrichment in these regions suggests that selection processes constrain large-effect genetic variation, resulting in only small genetic effects from these core gene regions and flattening their overall contribution to trait values, effectively mitigating potential perturbations. Stroke and all cSVD traits showed polygenicity enrichment in GERP-conserved regions and regulatory marks, such as histone modifications, though FA and MD displayed broader patterns of enrichment across enhancers and intronic regions. For stroke, enrichment was most pronounced in coding and conserved regions, while WMH exhibited relatively lower enrichment in histone-related annotations. These findings highlight the functional specificity of polygenicity across traits, reflecting distinct biological underpinnings.
Relevance of the “omnigenic” model – insights from GWASs
While negative selection minimizes the phenotypic effects of core disease genes, the shared genetic susceptibility studies, as shown in Table 3, reveal that many of these core genes and pathways are strongly implicated in both cSVD endophenotypes and stroke.9,11,16 Furthermore, the core genes consistently show high colocalization probabilities for shared causal variants between cSVD (WMH) and stroke subtypes (SVS). 11 Notable signals include HTRA1 (implicated in CARASIL), COL4A1/2 (linked to microangiopathy), and FOXF2 (associated with Axenfeld–Rieger syndrome) – all of which are central to monogenic forms of cSVD. 11 TWAS which assess the mediating effects of gene expression on genotype-phenotype associations, further validate these insights.9,11,12,104 Collectively, these analyses help define core genes as those directly involved in disease biology – often implicated in monogenic forms and supported by strong functional evidence. Moreover, such downstream analyses also identify a broader set of peripheral genes that vastly outnumber core genes. While many peripheral genes lack strong statistical associations or known pathophysiological roles, they frequently show significant tissue-specific expression, especially in brain-derived tissues. 11 Conceptually, core genes are central to disease mechanisms, while peripheral genes contribute more diffusely through broader regulatory networks. Single-cell genomics and transcriptomics, integrated with GWAS data, show that both core and peripheral genes linked to cSVD and stroke are enriched in endothelial cells, pericytes, and oligodendrocytes – key cell types in cerebrovascular health.7,9,11,105 Partitioning heritability further reveals that SNPs associated with cSVD are enriched in regions of active chromatin and in genes expressed in these same cell types.12,106 Given the dense interconnectivity of cellular gene-regulatory networks, 107 perturbations in peripheral genes expressed in disease-relevant cell types can indirectly influence core genes like COL4A1/2 and FOXF2, amplifying their effects on disease risk.
Taken together, these observations lend support to the “omnigenic model,” which posits that small, non-zero effects from a densely interconnected network of peripheral genes in disease-relevant cell types propagate their influence toward a core set of genes, ultimately driving pathogenesis. 108 In this context, the cumulative contribution of numerous peripheral genes may explain a large share of heritability, while even subtle disruptions in a few core genes may be sufficient to drive disease onset and progression.
Perspectives to enhance discoveries on cSVD and stroke genomics
The shared genetic susceptibility observed across cSVD traits likely reflects common underlying pathophysiological mechanisms and offers valuable biological insight. However, part of this overlap may also be driven by limitations in segmentation algorithms that inadequately distinguish between imaging-derived cSVD endophenotypes. Traditional methods for quantifying cSVD lesions often rely on arbitrarily defined categories based on anatomical brain regions or vascular territories,109,110 which inadequately capture the heterogeneity and fail to provide a biophysically meaningful description of the underlying tissue. Recently, data-driven approaches summarizing lesion frequency across defined local regions have underscored the importance of spatial distribution and sub-endophenotyping in uncovering the mechanisms underlying different cSVD traits. 111 Clustering and deep-learning methods, which group data based on inherent feature similarities, eliminate the need for predefined regions, allowing for unbiased pattern discovery and identifying significant spatial patterns overlooked by conventional approaches.112,113 By preserving localized voxel-level variations and interactions, these methods generate clusters that are more homogeneous and explain a greater proportion of the phenotypic variation. Advanced imaging methods targeting brain microstructure have also emerged as promising tools for studying cSVD.114,115 Among these, novel biophysical models such as neurite orientation dispersion and density imaging (NODDI), also derived from multi-shell acquisitions, generate tissue-based markers of white matter microstructure. 115 A recent study on the genomic determinants of NODDI markers in young adults identified 21 genome-wide significant loci for various tissue-based markers of white matter microstructure, several of which were validated in follow-up analyses across different age groups and were associated with cognitive performance in young adults. 116 Interestingly, genetic risk variants for WMH identified in older age were significantly associated with lower neurite density index in young adults in their twenties, specifically in brain regions known to harbor the highest frequency of WMH later in life. 116 These findings underscore the potential of advanced imaging markers to bridge the gap between molecular mechanisms and clinical outcomes across the lifespan.
On clinical endpoints, consortium-wide initiatives like GIGASTROKE have highlighted the translational potential of genetic epidemiology findings, both for drug discovery and for risk prediction and stratification. Large cross-ancestry meta-analyses of GWAS or whole exome association studies, combined with cutting-edge bioinformatic and statistical techniques, have brought insights into promising drug repositioning opportunities for stroke, including SVS,7,95 and putative drug targets for experimental follow-up.10,94,117 Polygenic risk scores that integrate genetic liability to stroke and multiple vascular risk factors have shown promise in identifying individuals at high risk for ischemic stroke across diverse populations. 7 Expanding these methods to cSVD will require global efforts that integrate detailed neuroimaging with multi-omics data across ancestrally diverse cohorts. 118 Equally important are collaborative biobank networks that link genetic data to electronic health records and clinic-based imaging datasets. 119 These resources will be essential for increasing sample sizes, improving ancestry representation, and refining risk models.
Finally, to fully realize the potential of these approaches, interdisciplinary collaboration will be critical – bringing together clinicians, molecular biologists, statisticians, bioinformaticians, and computational scientists. Such integration is key to bridging methodological advances with clinical relevance and to developing tailored tools that reflect the unique genetic architecture of cSVD and stroke. Ultimately, these efforts could accelerate progress in prevention, diagnosis, and treatment of cerebrovascular diseases.
Summary
Our comprehensive study leveraging genetic association summary statistics for stroke, its subtypes, and imaging-derived endophenotypes of cSVD offers crucial insights into the potential of future GWASs for these disease-traits. By thoroughly examining the variance explained by common genetic variations, we demonstrate that cSVD endophenotypes, in general, are driven by lower polygenic variation concentrated in genomic regions associated with core disease biology, compared to the more heterogeneous stroke phenotype, which has a greater polygenic contribution. Downstream analyses like TWAS, consistently highlight the enrichment of disease risk loci in pathways critical to the maintenance of the cellular microenvironment, especially the cerebrovascular matrisome, implicating a network of genes involved in cell-type-specific functions. Furthermore, imaging markers quantifying microstructural changes, which have particularly low polygenicity, exhibit shared genetic susceptibility at both individual loci levels and in polygenic risk score settings with traditional MRI markers of cSVD. Given that alterations in white matter microstructure often precede the occurrence of visible cSVD lesions13,120–122, such markers present valuable opportunities for studying the molecular mechanisms leading to cSVD across the lifespan. To advance our understanding of the pathophysiological role of cSVD markers, future research must address several key limitations and opportunities, particularly in enhancing the scope and inclusivity of genetic studies. For example, current GWASs for MRI-based cSVD traits (and DTI-based measures) rely overwhelmingly on populations of European ancestry (>95%), highlighting the need for larger studies that include more diverse populations to improve the generalizability of findings. In addition, large-scale next-generation sequencing studies are essential for uncovering rare and structural variants, with promising advancements emerging from long-read sequencing technologies. As such studies could enable exploring the role of somatic mutations, which are increasingly recognized as significant contributors to both neurological phenotypes and their risk factors. Collectively, these approaches will not only expand our understanding of the genetic basis of cSVD and stroke but could also uncover novel molecular mechanisms.
Supplemental Material
sj-pdf-1-jcb-10.1177_0271678X251362977 - Supplemental material for Unravelling the genetic architecture of cerebral small vessel disease in the context of stroke
Supplemental material, sj-pdf-1-jcb-10.1177_0271678X251362977 for Unravelling the genetic architecture of cerebral small vessel disease in the context of stroke by Sathyaseelan Chakkarai, Quentin Le Grand, Lucas Wang Shaoxuan, Stephanie Debette and Muralidharan Sargurupremraj in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Data and code availability
All methods used in this study are publicly available. Analyses were conducted using default parameters as described in the original method documentation and vignettes. All GWAS summary statistics analyzed were publicly accessible, and statistical tools were implemented using open-source software packages available through R CRAN and the following URLs:
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: M.S. acknowledges the support of following grant mechanisms: R01NS017950-39, P30AG066546, RF1AG063507, Alzheimer’s Association Research Grant – 23AARG-1030407, and William and Ella Owens Medical Research Foundation. S.D. acknowledges support from the French National Research Agency and France 2030 (ANR-18-RHUS-0002, RHU-SHIVA; ANR-23-IAHU-0001, IHU-VBHI), Prix Burrus-FRM and NRJ-neurosciences, EU Horizon 2020 (grant No 754517). S.D. and Q.L.G acknowledge support from the Fondation Recherche Alzheimer. L.W.S was supported by NIH T32GM145432 and P30AG066546.
Acknowledgements
We thank the MD/PhD program at the University of Texas Health Science Center, led by Dr. José E. Cavazos, for their support. We also acknowledge the Computational Genomics Summer Institute (CGSI, GM135043) at the University of California, Los Angeles, and its faculty for the insightful discussions on various statistical genetic methods.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ contributions
S.C., Q.L.G., S.D., M.S., Designed and conceived the study, S.C., and M.S., conducted the analyses based on GWAS summary statistics, S.C., Q.L.G., L.W.S., S.D., M.S., wrote and edited the manuscript, S.D., and M.S., jointly supervised the work.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
