Abstract
Across multiple sectors, including food, cosmetics and pharmaceutical industries, there is a need to predict the potential effects of xenobiotics. These effects are determined by the intrinsic ability of the substance, or its derivatives, to interact with the biological system, and its concentration–time profile at the target site. Physiologically-based kinetic (PBK) models can predict organ-level concentration–time profiles, however, the models are time and resource intensive to generate
Introduction
Humans, like other animals, are exposed daily to a multitude of chemicals of anthropogenic origin, including pharmaceuticals, food additives, pesticides, consumer goods and cosmetic ingredients. The safety assessment of chemicals is a legal requirement that is essential to ensure their safe use by workers and consumers, and to ensure the protection of domestic/farm animals and environmental species. However, for the majority of chemicals, there is a lack of available data for safety assessment — hence predictive models are essential. Predicting toxicity requires knowledge of both the intrinsic activity of the chemical (or its derivatives) and the extent to which the organism is exposed. Whilst external exposure, or dose, has traditionally been used in assessments, it is recognised that the dose at the target site (i.e. organ-level exposure) is the more relevant measure, being causally linked to observed toxicity. 1 This reasoning has long been applied in drug design, where the internal exposure level can be linked more reliably to the desirable, therapeutic effect. As discussed by Pistollato et al. 2 in terms of legislation, kinetic data are a specific requirement for plant protection and biocidal product safety assessment, and, whilst not formally required, the incorporation of such data is widely recommended in other regulations such as Classification, Labelling and Packaging (CLP) and the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH). Guidance documents from the European Chemicals Agency (ECHA) 3 and the Scientific Committee on Consumer Safety 4 recommend making use of all available data (including kinetic data) to support decision-making. Whilst general information regarding absorption, distribution, metabolism or excretion (ADME) may be useful, more accurate prediction requires organ-level concentration–time profiles. Physiologically-based kinetic (PBK) models (synonymous with physiologically-based pharmacokinetic, toxicokinetic or biokinetic (PBPK, PBTK or PBBK) models) are employed in numerous industries to provide such predictions.
In a PBK model, the body is represented as a series of compartments (e.g. individual organs) connected by blood flow. The models use knowledge of physiology and anatomy (such as organ volumes and cardiac output), in combination with chemical-specific information (such as solubility and partitioning behaviour) to predict the concentration–time profile of the chemical in tissues, cellular compartments or sub-compartments. Differential equations are used to describe the rate of change of concentration of the chemical in each compartment, as summarised in Figure 1. Detailed information on how to construct and validate PBK models, their applications in different sectors and tools available to support PBK modelling have been well-reported previously.1,5–9 Of particular note is the recent Organisation for Economic Co-operation and Development (OECD) The key characteristics of PBK models and the data captured in the PBK model data set.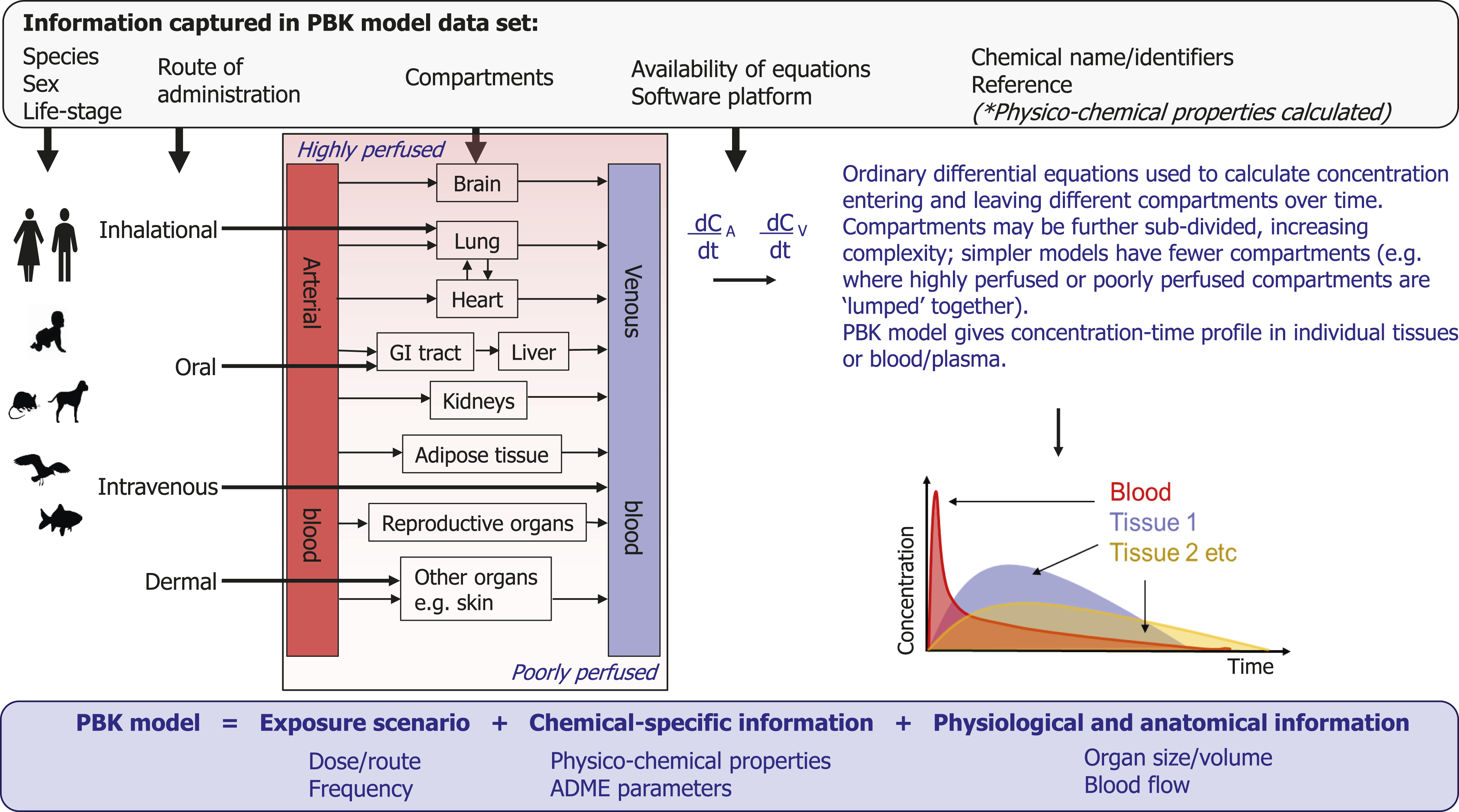
For environmental chemicals, the numerous applications of these models include: determining the dose at target tissues following external exposure; route-to-route extrapolation; dose extrapolation; inter-species and intra-species extrapolation (accounting for species, population or genetic variability through adaptation of physiological and anatomical parameters);
The ECHA reports that read-across is the most commonly used alternative method to reduce or replace animal testing in safety assessment.
14
In this approach, information from a data-rich (source) chemical is used to predict information for a data-poor (target) substance that is considered similar.
3
Kinetic information plays a key role in supporting read-across predictions3,15 and recent efforts have aimed to increase the accessibility of such data. Sayre and co-workers
16
published a database of time-series concentration data, extracted from an extensive search of the literature, and Pawar and co-workers
17
identified 38 databases containing a range of ADME-relevant data, as part of their overall review of resources to support read-across and
Over the past 30 years, the number of published PBK models and their applications has increased significantly.
18
In 2016, Lu and co-workers published a PBK Knowledgebase, comprising 307 chemicals for which PBK models were available from papers published between 1977 and 2014.
19
In their report, the authors described two case studies wherein PBK models from the Knowledgebase were used to inform the development of PBK models for ‘similar’ chemicals. In their study, chemical analogues were identified based on similarity of physico-chemical properties, although it is recognised that there is no consensus as to the best method to determine similarity.
20
Ellison and Wu
21
successfully demonstrated an analogous approach wherein a PBK model for a target chemical was evaluated by using information from source chemicals identified as structural or functional analogues. In order to assist researchers in identifying existing PBK models, a spreadsheet of those collated from the literature by the US Environmental Protection Agency (EPA), was made available
An enriched version of this PBK Knowledgebase was recently used as a proof-of-principle, to demonstrate that information from an existing PBK model could be used, in a read-across approach, to inform safety assessment. 23 In the analysis, methyleugenol was considered as a target chemical, with estragole and safrole being identified as suitable source chemicals (with respect to structural similarity). This approach was also successful, exemplifying how information from an existing PBK model could assist the development of a model for a similar chemical. Making best use of existing data and in particular the application of the read-across approach are recognised as important tools in reducing animal testing. 24 In order to facilitate the application of this approach, it is essential to identify chemicals for which PBK models are available. As ‘similarity’ is often considered in relation to structure or physico-chemical properties, it is also important to ascertain the nature of the chemicals for which models are available, comparing their characteristics to existing chemical data sets. Having information regarding the chemicals and the models in a readily accessible and updateable resource would be a significant asset for researchers, industry and regulators, with the potential to reduce the number of animals used in drug development and chemical safety assessment.
Several key features (which are represented in Figure 1) characterise an individual PBK model and include species, sex, life-stage, route of administration and the compartments required to accurately describe the time-course of the chemical. In some models, key organs (such as the liver, lungs, etc.) are incorporated individually as compartments; in others, these are further divided into constituent sub-compartments (for example, considering histopathological regions or explicitly including lymph or interstitial/vascular space) giving higher-level, more complex models. In other scenarios, organs are grouped together (referred to as ‘lumping’) to create simpler models, for example, all poorly perfused organs are considered as one compartment and all highly-perfused organs are considered as another. In addition to the physiological and anatomical information required, chemical-specific data are also a prerequisite. A substance may be identified using common names or chemical identifiers such as the Chemical Abstracts Service (CAS) Registry Number, a Simplified Molecular Input Line Entry System (SMILES) string or the International Chemical Identifier Key (InChiKey). Ideally, multiple identifiers should be incorporated in the data set to avoid ambiguity. Model development can be performed with a range of software, and the equations employed may be specified within the publication itself or as part of the supplementary information accompanying the article. Within this systematic review, key model characteristics, such as species, sex, life-stage, route of administration, compartments, availability of model equations and chemical identifiers, were captured within the PBK model data set, as summarised in Figure 1.
The second part of the analysis relates to the assessment of the chemical space coverage of the PBK model data set. There is no simple process by which a chemical can be designated as being a particular ‘type’ — for example, cosmetic ingredients may also be food additives, botanicals may have pharmaceutical properties, etc. Consequently, in order to assess the nature of the chemicals in the PBK model data set, key physico-chemical properties were generated and compared to those of chemicals appearing in other data sets. The data sets studied were: botanicals, pesticides, pharmaceuticals, food, cosmetic ingredients and REACH chemicals. The number of chemicals in the PBK model data set that also appeared in each of the other data sets was ascertained.
In summary, the aim of this systematic review was the curation of a data resource for existing PBK models. Relevant information for the models (species, sex, life-stage, substance identity, software used, etc.) was captured in a flexible spreadsheet format. The chemical space occupied by the PBK models (in terms of physico-chemical properties) was compared to that of other chemical types, by using six existing data sets. This resource has been created to assist the development and evaluation of PBK models based on existing data, thereby reducing the need to generate new data from animal studies.
Methods
Systematic review
This systematic review was prospectively registered on PROSPERO, the National Institute for Health Research’s international prospective registration system with the review question stipulated as: “For which substances are physiologically based kinetic (PBK) models available and which species, genders, life-stages and routes of administration have been investigated for these substances? This will include determining the chemical space coverage of the models and the availability of the associated model equations within the literature”. 25 The review complies with the PRISMA reporting standards; the PRISMA checklist is available as Supplementary Material S(i).
Briefly, following a scoping study of potentially useful databases and search terms, Scopus (https://www.scopus.com/), PubMed (https://pubmed.ncbi.nlm.nih.gov/) and Web of Science (https://www.webofknowledge.com) were selected as the most appropriate databases for identifying published papers on PBK models. The search of these databases was completed in October 2020. The search terms (“pbpk” OR “pbk” OR “pbbk” OR “pbtk” OR “pbpd” OR “pbpm” OR “physiologically based”) AND (“pharmacokinetic” OR “toxicokinetic” OR “biokinetic” OR “pharmacodynamics” OR “biopharmaceutical”) were used to search abstracts, titles and keywords of papers within each database, across all years available. The systematic review management tool Covidence was used for processing papers for the review (https://www.covidence.org/; accessed May 2021). A total of 14,803 papers were initially identified; however, following automated removal of duplicates in Covidence, 6771 remained. All abstracts were screened independently by two researchers with all conflicts being resolved by discussion. The inclusion criteria encompassed PBK models for all routes of administration for chemical, biological and carrier systems, including cases where normal physiology was altered or interactions between administered substances were investigated. Models that could not be associated with a specific substance (such as generic models applicable to large groups of compounds) were excluded. Where an abstract was associated with a paper that had subsequently been retracted, it was ensured that this model was excluded from the data set. Although standard practice in other systematic reviews, assessment of the quality of the reported models and risk of bias in reporting was considered unnecessary for this review. Our intention here was to document all available models, enabling interested researchers to rapidly identify potentially useful models to assist with future model development. The assessment of PBK model quality (aside from fundamental considerations relating to good modelling practice) needs to be considered in terms of fitness for a given purpose. 10 Therefore, it is context dependent and remains the decision of the model user. Following abstract screening, 3120 abstracts were retained for full text screening. PBK model data were extracted from 1649 of these papers, resulting in 7541 individual models being captured. Note that, if oral and intravenous dosing were used for both male and female subjects for the same chemical, this would be extracted as four individual models, hence there are many more models than individual chemicals. Reasons for the exclusion of papers during full text screening included: PBK model not being reported in the article; the article referenced a previously published model with no adaptations (information on the PBK model was extracted from the original publication); and full article not being available in English or not being reasonably accessible.
Extraction of data from available physiologically-based kinetic models
Data were manually extracted from these 1649 papers by one reviewer, with information being acquired from text, tables, figures and supplementary information. The data were entered into a Microsoft Excel® spreadsheet that captured details of the chemical: parent and metabolites (specified where appropriate), species (with sub-category where relevant), sex, life-stage, route of administration, literature reference for the model (with DOI), compartments considered in the model, the software employed and the availability of PBK model equations within the article. Where possible, controlled vocabulary was used to ensure consistency of data extraction and to enable the resulting spreadsheet to be readily filtered and searched for specific types of models, that is, controlled vocabulary was used for species, sex, life-stage, route of administration, availability of equations and software used. The vocabulary was empirically derived, to enable the most efficient searching — the full rationale is given in the Supplementary Material S(ii). For example, life-stages can be reported in multiple ways — number of weeks, months, years of age, young adult, adult, neonate, young child, juvenile, etc. Therefore, for consistency, this information was allocated to the more generic categories of: pre-birth or pre-hatch; from birth or hatch up to adult; adult; pregnant; old age (if specified); as well as a generic category for health-compromised (excluding old age) individuals.
Chemical identifier information was obtained by manually inputting the chemical name (as given in the publication) into PubChem (https://pubchem.ncbi.nlm.nih.gov/; last accessed May 2021) and extracting the molecular weight, canonical SMILES, isomeric SMILES, InChiKey and CAS number. The CAS Registry Number from PubChem was used as input for the COSMOS database, version 2 (https://cosmosdb.eu; accessed April 2021). Where available, the CAS Registry Number and chemical name, as recorded in COSMOS, were extracted to confirm the identity of the chemical; the COSMOS ID was also extracted. This information was captured by one reviewer. An assessment of the reliability of screening and data extraction was undertaken and is reported below.
Assessment of the chemical space coverage of the physiologically-based kinetic model data set in relation to other chemical data sets
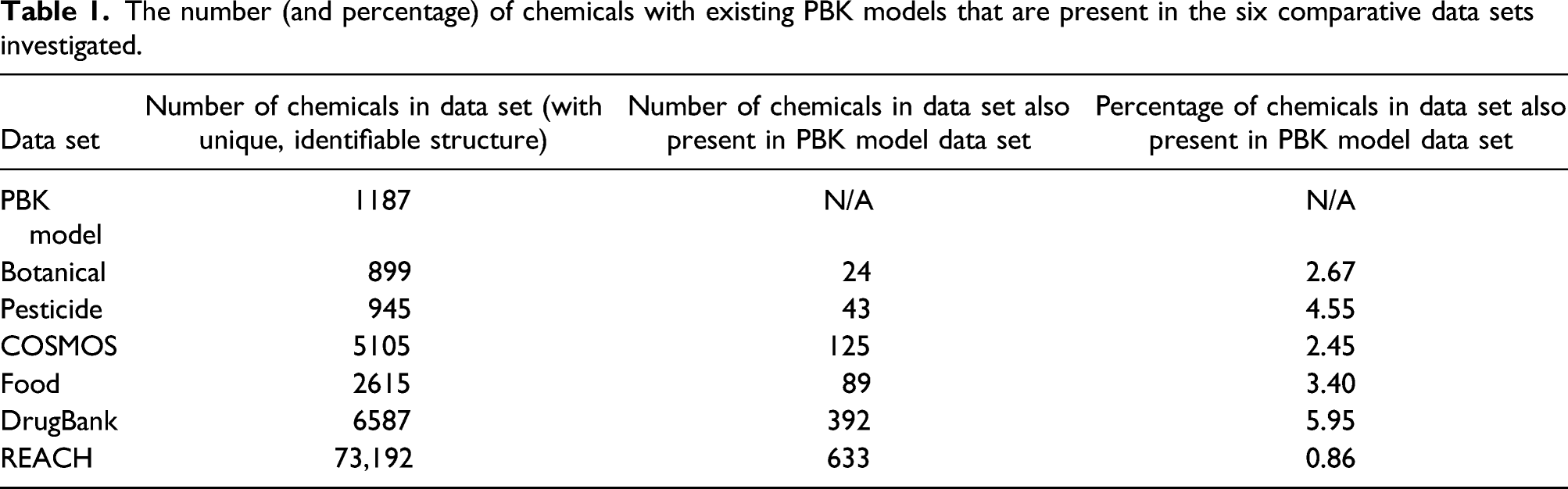
The number (and percentage) of chemicals with existing PBK models that are present in the six comparative data sets investigated.
Canonical SMILES for all chemicals in these data sets were generated by using OpenBabel (v.3.0.0; http://openbabel.org/wiki/Main_Page; accessed April 2021). From the PBK model data set, 1150 unique SMILES were identified with 1187 unique InChiKeys (note that chemical isomers may have the same SMILES string but different InChiKeys). In order to determine how many chemicals with PBK models were present in each of the other six data sets, the InChiKeys were compared.
The SMILES strings for all data sets were inputted into the RDKit (v. 2020.03.6; www.rdkit.org) Descriptor Node, accessed through KNIME software (v. 4.3.1; www.knime.com), in order to obtain the physico-chemical properties for all chemicals. The properties included molecular weight, number of hydrogen bond donors/acceptors, predicted logarithm of the octanol:water partition coefficient (SlogP) and the topological polar surface area (TPSA); the number of Lipinski rule violations were calculated from this information. Whilst it is possible to generate thousands of physico-chemical properties, here only a few readily calculable properties were selected, representing those most often used to broadly characterise chemicals in terms of size, polarity and partitioning behaviour. These simple properties were also used to determine Lipinski rule violations (frequently used to indicate potential oral absorption — a common route of administration for these models). The minimum, maximum, mean and median values, and the interquartile ranges of these properties, were calculated by using Minitab version 19.2 for all data sets. Histograms were also generated with Microsoft Excel to enable a visual comparison of the property ranges between the different data sets. The results of the statistical analysis are available as Supplementary Material S(iii).
Structural feature analysis
In order to determine the relative frequency of the occurrence of specific structural features in the chemicals comprising the PBK model data set, the SMILES strings were entered into the Chemotyper software (version 1.0; Molecular Networks, Erlangen, Germany; https://chemotyper.org/). A table which indicated the presence or absence of structural features identified by using ToxPrints (https://toxprint.org/) was created and exported into Microsoft Excel. In brief, ToxPrints represent a collection of 729 generic structural fragments covering (amongst others) commonly occurring functional groups, cyclic units and biomolecular substituents.
Results
Systematic review
Figure 2 summarises the number of papers considered at each stage of the review process and the final number of models extracted in the PBK model spreadsheet. Of the 6771 of papers initially identified, 3120 remained after abstract screening and data were extracted from 1649 of these. A summary of the papers considered at each stage of the review process, and the total number of models extracted.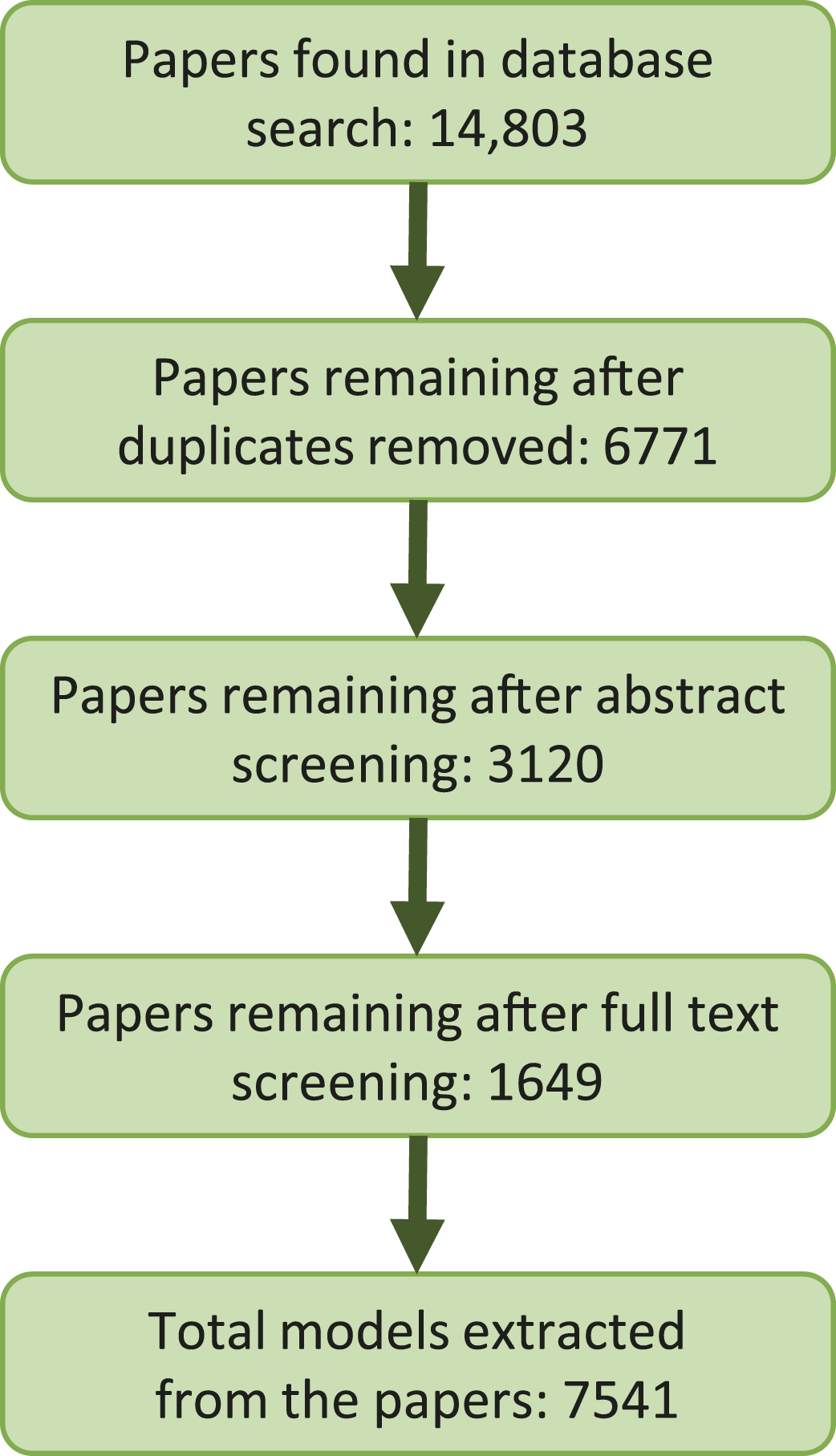
Extraction of data from available physiologically-based kinetic models
The number of models associated with different species.
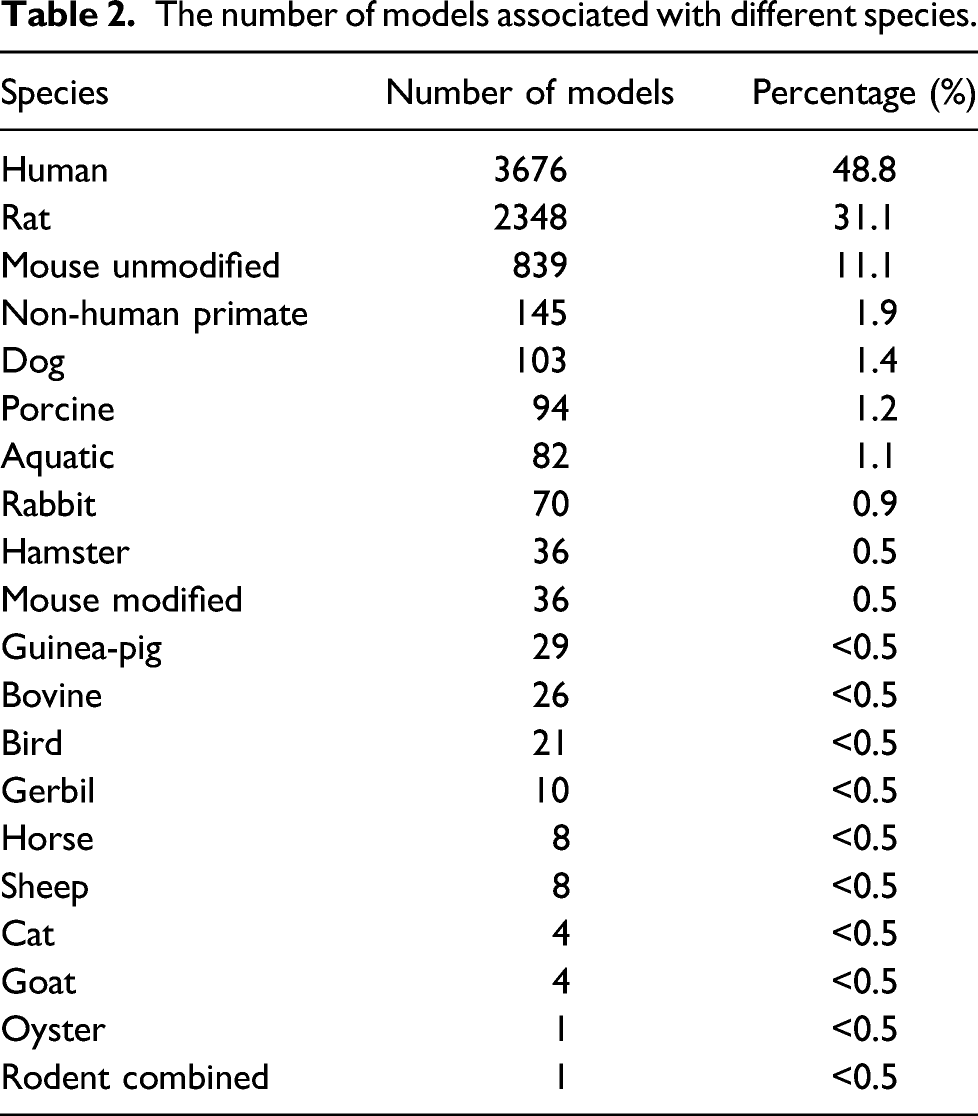
The number of models associated with different routes of administration.
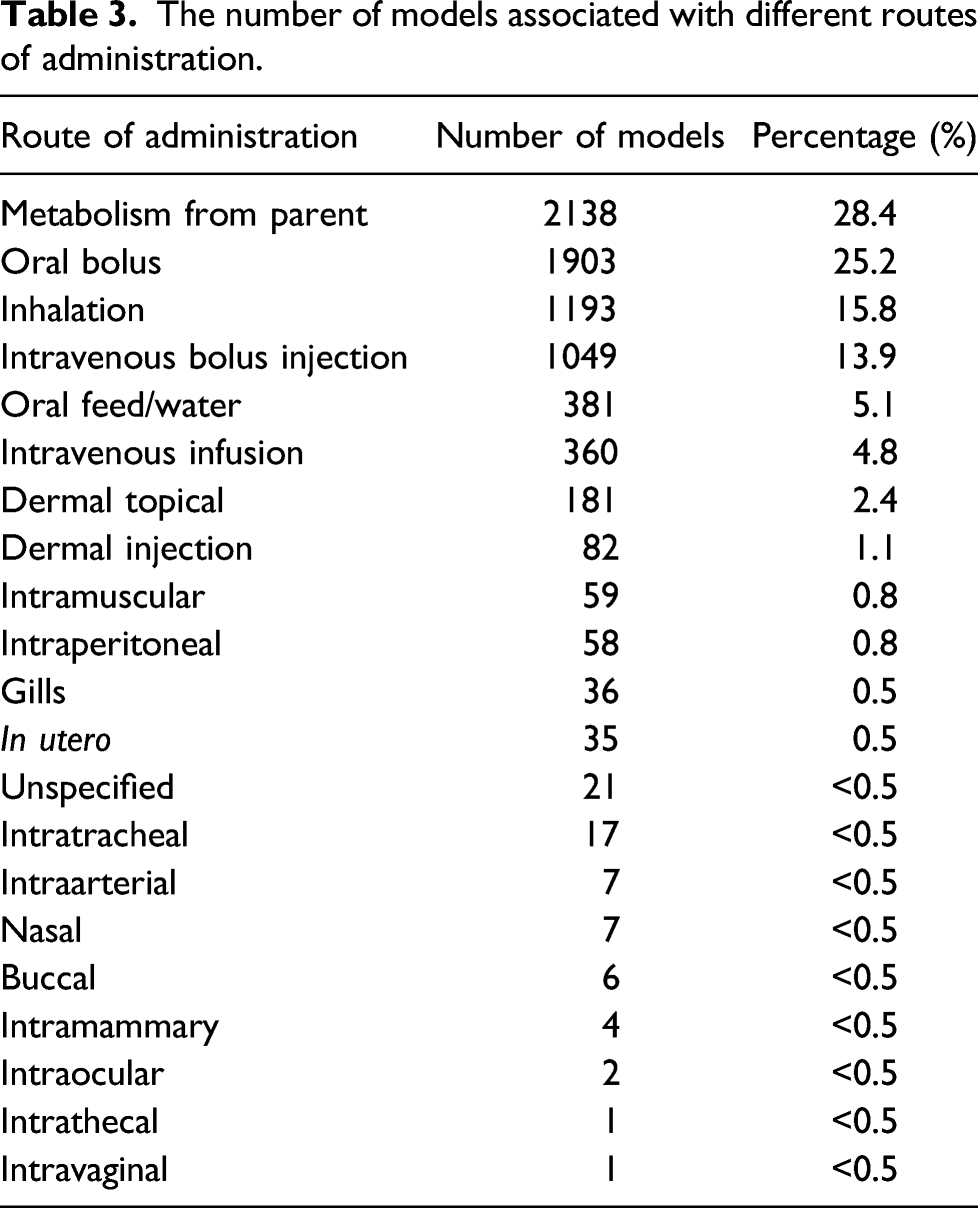
Assessment of the chemical space coverage of the physiologically-based kinetic model data set in relation to other chemical data sets
The results for the comparison of InChiKeys for chemicals in the PBK model data set to those for the six comparative data sets are shown in Table 1.
Statistical analysis of the physico-chemical properties of the chemicals in the PBK model data set.
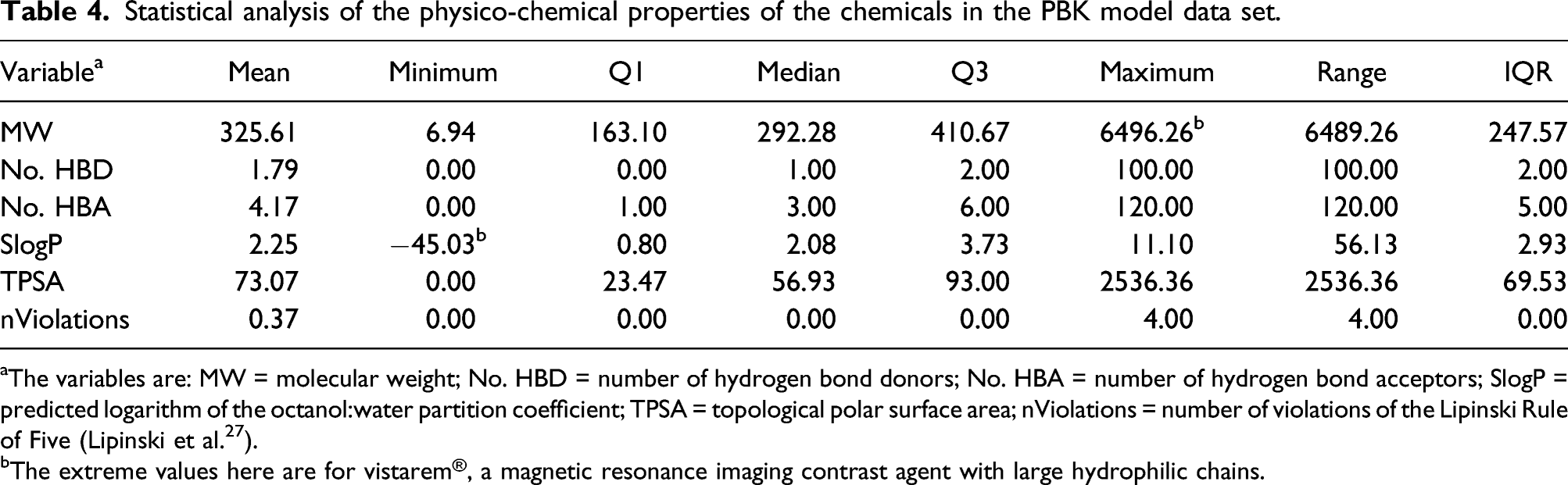
aThe variables are: MW = molecular weight; No. HBD = number of hydrogen bond donors; No. HBA = number of hydrogen bond acceptors; SlogP = predicted logarithm of the octanol:water partition coefficient; TPSA = topological polar surface area; nViolations = number of violations of the Lipinski Rule of Five (Lipinski et al. 27 ).
bThe extreme values here are for vistarem®, a magnetic resonance imaging contrast agent with large hydrophilic chains.
The Supplementary Material S(iii) shows the results of a similar statistical analysis of the physico-chemical properties for the six data sets to which the PBK model data set was compared.
Figures 3(a–f) shows the comparison of these key physico-chemical properties across the seven data sets. A comparison of the ranges of physico-chemical properties across the seven data sets investigated.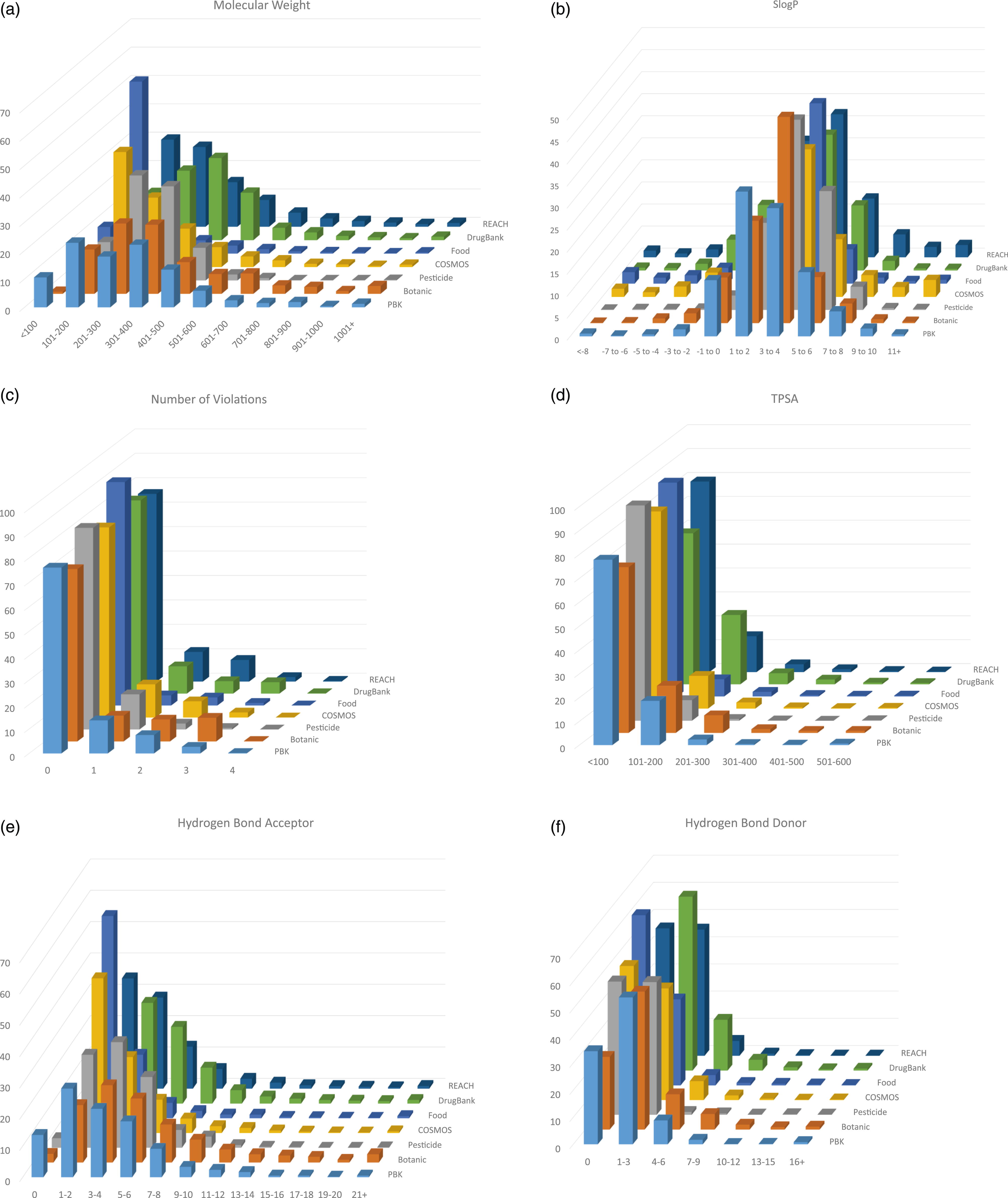
Structural feature analysis
The percentage of chemicals within the PBK model data set that contain the specified chemotypes identified by using ToxPrints.
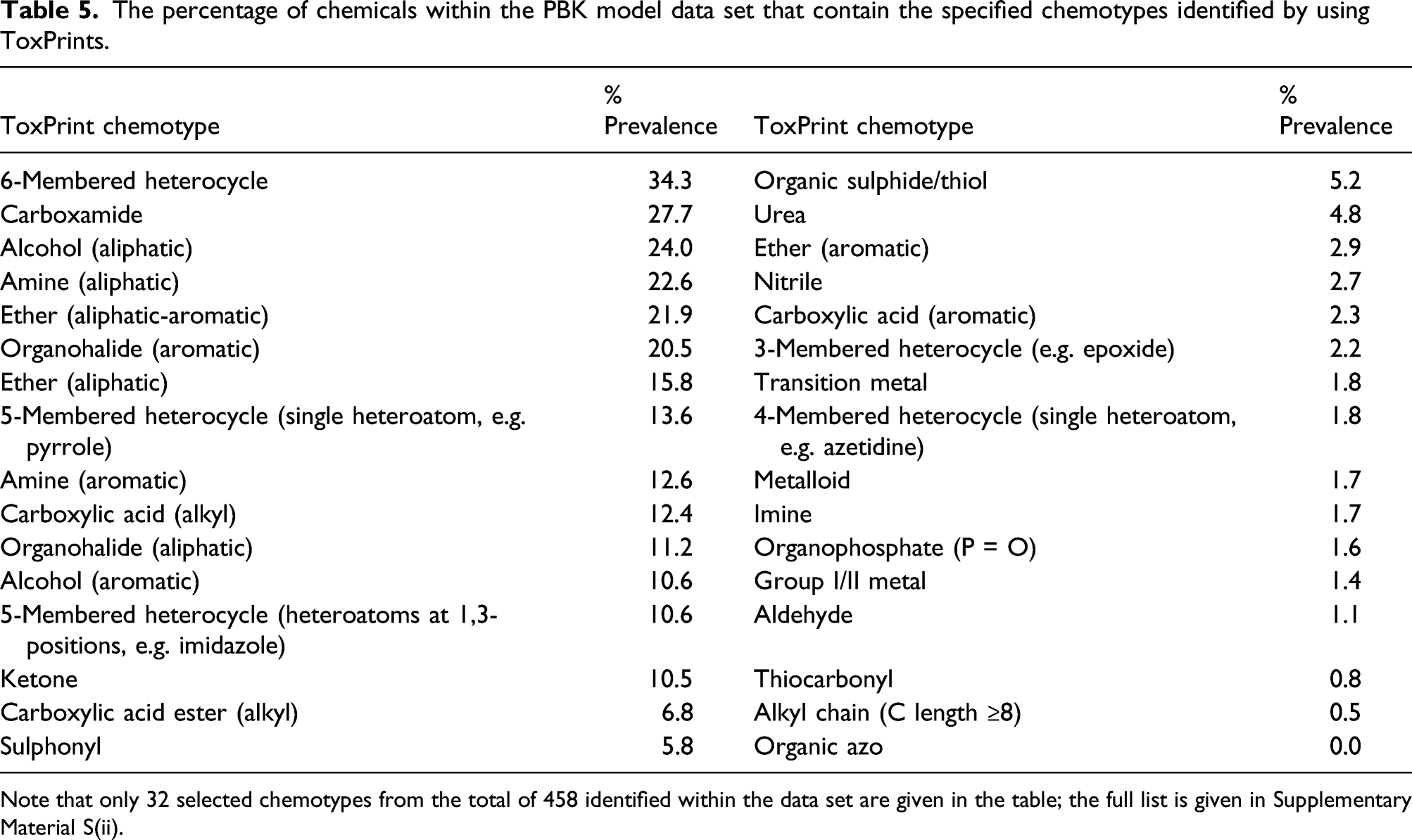
Note that only 32 selected chemotypes from the total of 458 identified within the data set are given in the table; the full list is given in Supplementary Material S(ii).
Assessment of screening and data extraction reliability
After screening all 6771 abstracts in duplicate, 3120 were taken forward to full text screening; of these, 1362 papers were rejected at this stage. In addition, 109 papers could not be readily obtained or were not in English. Therefore, data were extracted from 1649 papers, resulting in 7541 models. The resultant spreadsheet comprises over 150,000 individual entries, as for each model, the species, sex, life-stage, route of administration, availability of equations, compartments, references and chemical identifiers were captured. It is expected that errors will arise when assessing the suitability of papers for inclusion and performing extensive manual processing, hence a quality assessment exercise was undertaken. As part of this process, a representative sample from each of three stages of the screening and data extraction process was assessed by a second investigator: — 5% of the papers that had been excluded at the full text screening phase were reviewed; — 5% of PBK model data extracted from the papers (chemical information, species data (primary and secondary categories), sex, life-stage, route of administration, reference (DOI and PubMed ID if available), compartments investigated, availability of equations and simulation software were checked; and — 5% of the chemical identifier information from PubChem and COSMOS (chemical name, CAS Registry Number, molecular weight, canonical SMILES, isomeric SMILES, InChiKey and COSMOS ID) was obtained again from these sources and compared to the values in the spreadsheet.
The greatest source of ‘error’ was determined to be the exclusion of papers that were considered as potentially relevant by a second investigator, that is, 6% of excluded papers. In terms of the systematic review, this is not considered a highly significant problem. PBK models are continually being published, hence there can never be a finalised set of models. It is the intention to make this resource available in its current form, as a tool to assist researchers in finding relevant PBK models, and to update the resource in the future capturing models previously not identified or erroneously excluded.
For PBK model data, manually extracted from the papers, an error was detected in the information captured for 3.5% of the substances. This does not equate to 3.5% of the total information being incorrect, as this may indicate an error in only one (or possibly more) of the 13 columns that relate to the PBK model information.
An error was detected in the data for 2.4% of the chemicals in relation to the identifier information. As above, this does not equate to 2.4% of the total information being incorrect, but that for 2.4% of chemicals an error was detected in one (or more) of the seven columns associated with chemical identifier information.
The authors welcome any feedback from users regarding errors, omissions of existing models or updates for new models (note that models require a minimum level of information and novelty to be included); please email the corresponding author.
Discussion
In this systematic review, information concerning over 7500 PBK models were extracted from 1649 papers. The models encompassed 18 species (including rat, human, mouse, cow and guinea-pig) at various life-stages (e.g. juvenile, adult, pregnant and health-compromised) across 21 administration routes (e.g. oral, inhalation and
Although this is the largest collation of PBK models that the authors are aware of, it is not a complete list. As identified in the quality assessment exercise, some of the historic models were omitted. In addition, as this is such a dynamic area of research, the generation of a finite list of all models would not be possible. The publication of new models has shown a rapidly increasing trajectory in recent years. 26 However, the current data set serves as a basis for the continuing curation of existing models, which will provide an increasingly rich source of information for modellers in the future.
Trends in model availability: Coverage of chemical types
Chemicals can be used for a variety of purposes, and often it is not feasible to allocate a chemical to a unique group (e.g. there is a significant cross-over between chemicals used as food additives and as cosmetic ingredients, hence the same chemicals may appear in more than one of the different data sets). It is therefore difficult to determine the chemical ‘types’ for which there are the most PBK models; however, some trends are discernible from the analysis undertaken. Unsurprisingly, given that PBK modelling evolved in drug development, the greatest proportion of models correspond to chemicals in the DrugBank data set. Pesticides are generally well studied and data-rich; therefore, the second most common type of chemical with PBK models are the pesticides. For food additives and cosmetic ingredients, where there are often chemicals in common, similar proportions of chemicals have PBK models. Due to the size and generality of the REACH data set, it would be anticipated that relatively few chemicals would have existing PBK models. The results confirm the paucity of PBK models available in relation to different areas of chemical space, and underline the importance of using existing PBK models to help fill data gaps.
Trends in model availability: Coverage of physico-chemical properties
Figure 3(a) shows the distribution of molecular weight across the seven data sets. As expected, the majority of the chemicals fall within the range of 100–600 Da, but notable differences exist between the data sets. There are a relatively high number of chemicals in the PBK data set with low molecular weight — these will include the volatile chemicals for which respiratory uptake has been extensively studied. Food additives and cosmetic ingredients (which have chemicals in common) show a relatively high proportion of low molecular weight chemicals. Chemicals that are designed to be biologically active, such as drugs and pesticides, tend to be developed in accordance with guidelines relating to preferred physico-chemical properties. For example, the Lipinski Rule of Five stipulates that drugs with poor oral absorption are associated with chemicals with: molecular weight > 500 Da, log P > 5, and > 10 hydrogen bond acceptors or > 5 hydrogen bond donors. 27 Other research has also suggested that a topological polar surface area (TPSA) higher than 140 Å2 is also unfavourable for oral absorption. Consequently, certain chemical types are designed to fall within narrower property ranges and such trends are evident in the property ranges mentioned in this article. A correlation between molecular weight and log P is often observed amongst groups of chemicals (although there are many exceptions to this); here a similar pattern to the range of values is generally observed for log P and molecular weight. For both properties, for the majority of chemicals the values fall within a narrow range; however, there are also extreme values for a few chemicals. Pesticide and botanical data sets have a greater percentage of chemicals in the log P ranges 3–4 (43% and 47%, respectively), whereas the PBK data set only has 28% of chemicals in this range.
Pesticide and botanical data sets comprise fewer molecules capable of carrying a charge (associated with increased hydrophilicity) — hence, on average, they have higher log P values. This is significant, as partitioning behaviour (often estimated by using log P) is a key element in building PBK models. Whilst the extreme values for log P, calculated by the software used here, may be unrealistic (and therefore unsuitable for model building), when used for comparison purposes they are still useful for demonstrating the trends in the data. The range in values for all of the physico-chemical properties of pesticides, is narrower than for the other chemical types, indicating the more restrictive chemical properties required for these chemicals.
Botanicals generally show a wider range of values for each of the physico-chemical properties (in particular, more chemicals show properties at the upper extremes of the ranges). A significant number of compounds within this set are large and complex. Whilst in other data sets, molecules tend to be designed for a specific purpose, and those out‐ with the given property ranges are filtered out, the same exclusions would not be applicable to this data set. The diversity of structural features present in the chemicals with existing PBK models is demonstrated by the ToxPrints analysis, with 458 chemotypes identified as being present within this data set. The diversity of these chemicals, in terms of their molecular complexity, is demonstrated by the number of chemotypes identified — individual chemicals were shown to contain between one and 69 chemotypes within their structure.
Conclusion
Understanding the kinetic behaviour of a chemical within the body, particularly its concentration–time profile at a target site, is essential to accurately determine its potential effect. For the majority of chemicals, there is a lack of data concerning toxicity and kinetics. However, generating such information
Continuing effort is required to curate existing PBK models. The ability to extract relevant data from models and reproduce those published in the literature would be facilitated by researchers embracing the use of systematic methods to record PBK models. Consistent formats for publishing PBK models have been proposed previously,28,29 with the template proposed in the recent OECD guidance 10 drawing on previous recommendations, most significantly the reporting format proposed by Tan and co-workers. 30 The use of consistent reporting formats is strongly encouraged, as this assists other researchers in re-using or re-purposing existing models.
Future effort is required to curate available models, identify appropriate similarity metrics to assist in the identification of PBK models for analogues to chemicals of interest, and, ideally, make such resources widely available, for example, by incorporation into free webtools. Such endeavours will help to leverage the wealth of information available in existing PBK models.
Supplemental Material
sj-pdf-1-atl-10.1177_02611929211060264 – Supplemental Material for A Systematic Review of Published Physiologically based Kinetic Models and an Assessment of their Chemical Space Coverage
Supplemental Material, sj-pdf-1-atl-10.1177_02611929211060264 for A Systematic Review of Published Physiologically based Kinetic Models and an Assessment of their Chemical Space Coverage by Courtney V Thompson, James W Firman, Michael R Goldsmith, Christopher M Grulke, Yu-Mei Tan, Alicia Paini, Peter E Penson, Risa R Sayre, Steven Webb and Judith C Madden in Alternatives to Laboratory Animals
Supplemental Material
sj-pdf-2-atl-10.1177_02611929211060264 – Supplemental Material for A Systematic Review of Published Physiologically based Kinetic Models and an Assessment of their Chemical Space Coverage
Supplemental Material, sj-pdf-2-atl-10.1177_02611929211060264 for A Systematic Review of Published Physiologically based Kinetic Models and an Assessment of their Chemical Space Coverage by Courtney V Thompson, James W Firman, Michael R Goldsmith, Christopher M Grulke, Yu-Mei Tan, Alicia Paini, Peter E Penson, Risa R Sayre, Steven Webb and Judith C Madden in Alternatives to Laboratory Animals
Supplemental Material
sj-xlsx-3-atl-10.1177_02611929211060264 – Supplemental Material for A Systematic Review of Published Physiologically-based Kinetic Models and an Assessment of their Chemical Space Coverage
Supplemental Material, sj-xlsx-3-atl-10.1177_02611929211060264 for A Systematic Review of Published Physiologically based Kinetic Models and an Assessment of their Chemical Space Coverage by Courtney V Thompson, James W Firman, Michael R Goldsmith, Christopher M Grulke, Yu-Mei Tan, Alicia Paini, Peter E Penson, Risa R Sayre, Steven Webb and Judith C Madden in Alternatives to Laboratory Animals
Footnotes
Acknowledgments
JCM and CVT gratefully acknowledge the funding and scientific advice of the European Partnership for Alternative Approaches to animal testing (EPAA) with respect to the funding of a studentship for CVT.
Author’s Note
The views expressed in this article are those of the authors and do not necessarily represent the views or the policies of the US Environmental Protection Agency.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the European Partnership for Alternative Approaches to animal testing (EPAA).
Supplementary Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
