Abstract
Background
Type 2 diabetes is a chronic disease that has severe individual and societal consequences, which is forecast to worsen in the future. A new field of investigation is variations in circadian rhythm genes, in conjunction with diet and sleep variables, associations with, and effects on, type 2 diabetes development.
Objective
This systematic review aimed to analyse all current literature regarding circadian rhythm gene variations and type 2 diabetes, and explore their interplay with diet and sleep variables on type 2 diabetes outcomes. This review was registered with PROSPERO (CRD42021259682).
Methodology
Embase and Pubmed were searched on 6/8/2021/11/8/2021 for studies of all designs, including participants from both sexes, all ethnicities, ages, and geographic locations. Participants with risk alleles/genotypes were compared with the wildtype regarding type 2 diabetes outcomes. Studies risk of bias were scored according to the risk of bias in non-randomised studies – interventions/exposures criteria.
Results
In total, 31 studies were found (association n = 29/intervention n = 2) including >600,000 participants from various ethnicities, sexes, and ages. Variations in the melatonin receptor 1B, brain and muscle arnt-like 1 and period circadian regulator (PER) genes were consistently associated with type 2 diabetes outcomes.
Conclusions
Individuals with variations in melatonin receptor 1B, brain and muscle arnt-like 1 and PER may be at higher risk of type 2 diabetes. Further research is needed regarding other circadian rhythm genes. More longitudinal studies and randomised trials are required before clinical recommendations can be made.
Introduction
Type 2 diabetes (T2D) is a chronic disease that is forecasted to be associated with 1.59 million deaths per year by 2025 (Lin et al., 2020). The estimated cost of diabetes to the global economy was $1.31 trillion in 2015 and is predicted to rise to $2.5 trillion by 2030, equivalent to 2.2% of global gross domestic product (Bommer et al., 2018; Zhang and Gregg, 2017). Approximately 90% of diabetes cases are type 2 and currently 422 million people aged between 20 and 79 years have diabetes, forecast to rise to 629 million by 2045, which is about 6.3% of global population (Diabetes, no date; Khan et al., 2020).
The main symptom of T2D is hyperglycaemia caused by ineffective insulin secretion and/or action, characterised by eventual pancreatic β-cell failure (Olokobaet al., 2012). T2D is most frequently onset in adults aged 45 years and over, but its prevalence is increasing in younger populations (Lascar et al., 2018). The aetiology of T2D includes obesity, lack of physical activity (PA), age, family history, genetics, high consumption of sugar-sweetened beverages and red and processed meats, and low consumption of fruits and vegetables (Ali, 2013; Forouhi and Wareham, 2014).
Recently, circadian rhythm (CR) genes have been implicated in the development of T2D (Javeed and Matveyenko, 2018). CR genes are mostly expressed in the suprachiasmatic nucleus of the hypothalamus and in peripheral tissues including pancreatic β-cells. They influence physiological processes, including the sleep-wake cycle, metabolism and the immune system, by variable expression over general diurnal/nocturnal phases (Rijo-Ferreira and Takahashi, 2019). CR processes can interact with hormones including insulin and melatonin, and processes including gluconeogenesis, which may increase T2D risk (Dashti et al., 2020; Rijo-Ferreira and Takahashi, 2019). Single nucleotide polymorphisms (SNPs) in CR genes may therefore further modify this risk. Transcription in CR genes oscillates via an autoregulatory feedback loop, triggered by external cues including light exposure, PA, and diet (Cagampang and Bruce, 2012). CR gene expression is mediated by the retinoic acid-related orphan receptor alpha gene, which triggers a central positive feedback loop consisting of circadian locomotor output cycles kaput (CLOCK) and brain and muscle arnt-like 1 (BMAL1) heterodimer. The central positive feedback loop results in the expression of tissue-specific genes including melatonin receptor 1B (MTNR1B), as well as triggering a negative feedback loop consisting of period circadian regulator (PER) 1/2, cryptochrome circadian regulator (CRY) 1/2, and nuclear receptor subfamily 1 group D member (NR1D) 1/2. NR1D1/2 mediate transcription of REV-ERBα/β proteins which repress transcription of BMAL1, ultimately halting CR gene expression (Jakubowicz et al., 2017; Rijo-Ferreira and Takahashi, 2019). A greater understanding of the circadian cycle's relationship with T2D may lead to clinical strategies which limit T2D prevalence.
Previous research has revealed that CR disruptions, sometimes due to modifiable lifestyle zeitgebers diet and sleep, can further modify T2D risk (Dashti et al., 2015; Javeed and Matveyenko, 2018; Poggiogalle et al., 2018; Sinturel et al., 2020). Poor dietary regulation, including breakfast-skipping, consumption of a traditional ‘Western’ diet and night-time feeding can dysregulate secretion of CR-controlled hormones, including glucagon-like peptide 1 (GLP-1), which is key for glucose-dependent insulin release, therefore increasing T2D risk (Froy, 2010; Jakubowicz et al., 2017; Rijo-Ferreira and Takahashi, 2019). Sleeping patterns can also moderate CR gene expression via changes in light-dark cycle stimuli (Jakubowicz et al., 2017). Sleep disruptors such as excessive light exposure, shift work and enforced clinical laboratory settings can limit secretion of CR gene-mediated anti-diabetic hormones including GLP-1, and also limit insulin sensitivity and β-cell function, leading to metabolic dysregulation and increased T2D risk (Jakubowicz et al., 2017; Javeed and Matveyenko, 2018). A previous meta-analysis has been conducted of associations and interactions between CR gene variations, diet, sleep and T2D using studies from the cohorts for heart and aging research in genomic epidemiology consortium (Dashti et al., 2015). However, to the best of our knowledge, no systematic reviews encompassing all existing literature have taken place. Therefore, the primary aim of this systematic review was to determine whether variations in CR genes had an association with, or effect on T2D outcomes. And secondly, to determine whether diet and sleep moderate CR gene variations associations/effects on T2D outcomes.
Methodology
This systematic review was reported according to the preferred reporting items for systematic reviews and meta-analysis (PRISMA) guidelines (Page et al., 2021).
Search strategy
Search terms for Pubmed were formulated by HS, GV and LL, and were modified for Embase by CG. CG, LL and GV conducted searches of Embase (6/8/2021), and Pubmed (11/8/2021). English-language, human studies from any date prior to the search were included.
Criteria for study inclusion (PI/ECOS)
Population: Studies regarding CR genes and T2D and/or T2D-related metabolic traits were searched for in populations including both sexes, all ethnicities, ages, and geographic locations. Participants were recruited from various methods, including from existing cohorts, from the general population, or from hospitals and medical registers. Participants remained included if they were suffering from comorbidities including but not limited to cardiovascular disease, obesity and metabolic syndrome (MS), or if they were taking T2D medication. Studies were required to state the number of participants included. Pregnant females and participants suffering from other forms of diabetes were excluded.
Intervention/exposure: The intervention group were participants with T2D risk alleles and/or genotypes in CR genes. Accepted genotyping methods included DNA isolated from samples including blood and saliva, and genotyped via methods including TaqMan and Biobank Axiom arrays (AxiomTM Biobank Plus Genotyping Array, no date; Real-Time PCR Assays – UK, no date). Studies were required to report all CR genes and SNPs that were analysed. Secondary interventions included study-specific dietary and sleep variables (controlled and uncontrolled). Dietary variables included diet patterns (e.g., Mediterranean diet) and individual nutrients (e.g., fat consumption). Dietary patterns were measured via any method, including lab observations and self-reports such as 24-h food frequency questionnaires and food diaries. Sleep variables were measured by any method, including controlled sleeping hours under clinical conditions and actigraphy, as well as self-reported sleeping habits recorded via questionnaire.
Comparison: The primary comparison group were wildtype participants in the aforementioned CR genes. Secondary comparisons included diet and sleep variables (controlled and uncontrolled).
Outcomes: Primary outcomes were incidence of T2D and T2D-related metabolic traits, measured by methods including medical records, self-report, and metabolic tests (OGTT, fasting glucose, and HbA1c data). Studies could report T2D outcomes from a diverse number of measures, including relative risk, odds ratio, or comparison of metabolic outcomes. All outcomes related to T2D were collected. Studies were required to include statistical analysis at a significance value p < 0.05 prior to multiple comparisons. Studies were not required to have carried out corrections for multiple testing. Studies that included no T2D outcome data were excluded.
Study design
Study selection
All identified studies were manually screened by HS and LL, and duplicates were removed. Remaining studies were exported to Rayyan (Ouzzani et al., 2016), and were screened according to title and abstract by HS, LL and GV. After, remaining studies were full-text screened by HS, LL and GV. Full texts were accessed via St Mary's University library services. Remaining studies underwent reference screening by HS, LL and GV for an additional literature search. All conflicts were discussed and resolved by HS, LL and GV.
Data extraction
Data was extracted using a checklist formulated by HS, LL and GV, by two researchers per study. Data included author's details, study characteristics (location, setting etc.), methodology data, participant characteristics, interventions, comparisons, results, strengths and limitations, conclusions, and areas for future research. All conflicts were discussed and resolved by HS, LL and GV.
Quality assessment
Studies were scored according to criteria from the risk of bias (ROB) in non-randomised studies – interventions/exposures (ROBINS-I/E) tools (Risk of bias tools – ROBINS-E tool, no date; Sterne et al., 2016), by HS, LL and GV. Criteria included control of confounders, participant recruitment, classification of exposures/interventions, deviations from original protocol, missing data, measurement of outcomes, and selection of reported results. Studies were assigned a low/moderate/serious/critical ROB for each category, and overall, according to the ROBINS-I/E guidelines.
For ease of comparison, remaining studies were grouped by (a) genes and SNPs, (b) study design (e.g., intervention/association), and (c) study quality (ROB).
Registration
This systematic review was submitted to PROSPERO (‘PROSPERO International prospective register of systematic reviews’, 2021) for registration on 27/7/2021, and its protocol was accepted on 13/8/2021 (registration: CRD42021259682). Modifications were made to the registration regarding the use of Rayyan software on 23/9/2021.
Results
Search results
Figure 1: On 6/8/2021/11/8/2021, the aforementioned search terms applied to Embase and Pubmed returned 2,990 and 1,827 results, respectively (n = 4,817). Following title screening, 894 duplicates were excluded (n = 3,923). Following export to Rayyan, title and abstract screening excluded a further 3,983 studies (n = 30). After full-text screening, a further 10 studies were excluded (n = 20). Following citation searching of the remaining studies, 14 further studies were included (n = 34). Following full-text screening of studies identified via citation searching, 3 were excluded (n = 31).
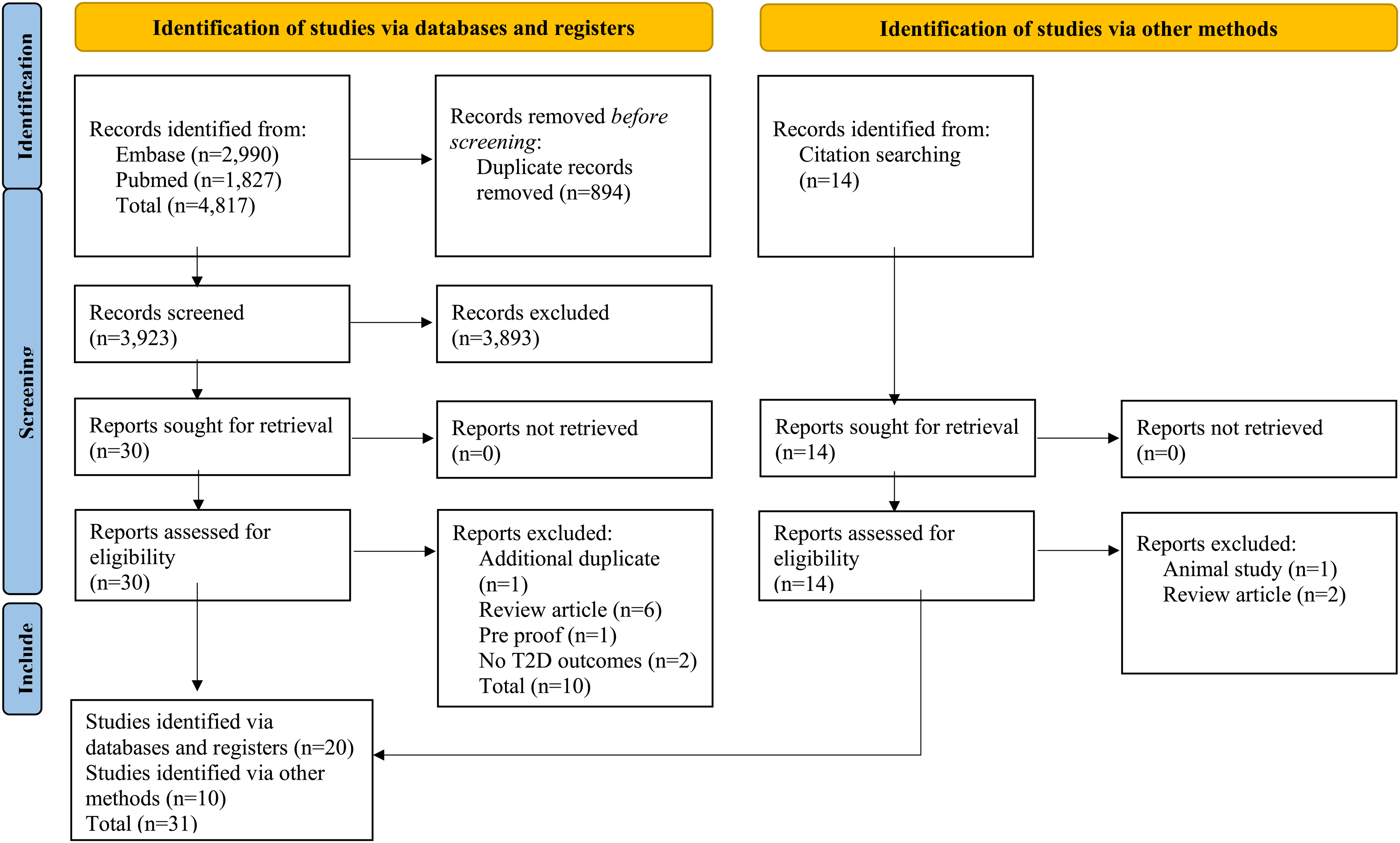
Search results flow diagram (Page et al., 2021).
Study characteristics
Table 1: 31 remaining studies (MTNR1B: n = 19/CLOCK: n = 8/BMAL: n = 4/PER n = 6/CRY n = 5/REV-ERBα/β n = 1) included 604,825 participants (median n = 1,675). Of studies that provided requisite participant data, the mean participant age was 46.1 years/51.5% male. Participants were from 18 different ethnicities. Study designs were case-controls (n = 13), cross-sectional studies (n = 13), GWAS (n = 2), an RCT (n = 1), and studies of mixed design (n = 2) (association n = 29, intervention n = 2). T2D outcome measures included metabolic data (n = 21) (OGTT results/fasting glucose/insulin resistance/β-cell function/HbA1c data) and T2D incidence (n = 10). Sleep and dietary variables were included in n = 5 and n = 4 studies, respectively.
Study characteristics including genes, participant variables, study design, genotyping method, and T2D outcome measure.

T2D: type 2 diabetes; MTNR1B: melatonin receptor 1B; CLOCK: circadian locomotor output cycles kaput; BMAL: brain and muscle arnt-like 1; PER: period circadian regulator; CRY: cryptochrome circadian regulator; OGTT: oral glucose tolerance test; IR: insulin resistance; SNP: single nucleotide polymorphism; RCT: randomised controlled trial; GWAS: genome-wide association study.
Number of studies.
Results calculated from studies with published participant characteristics.
OGTT results, fasting glucose (FBG), IR, β-cell function, HbA1c data.
Quality assessment
Table 2: A quality assessment undertaken by HS, GV and LL, shows 18 studies had a low; 6 had a moderate; 3 had a serious; and 4 had a critical ROB. Common reasons for bias were lack of HWE, lack of control of confounders, missing data regarding participant characteristics and recruitment data, and possible T1D participant inclusion.
Quality assessment of 31 studies using the ROBINS-I/E Guidelines (Sterne et al., 2016; Risk of bias tools - ROBINS-E tool, no date).
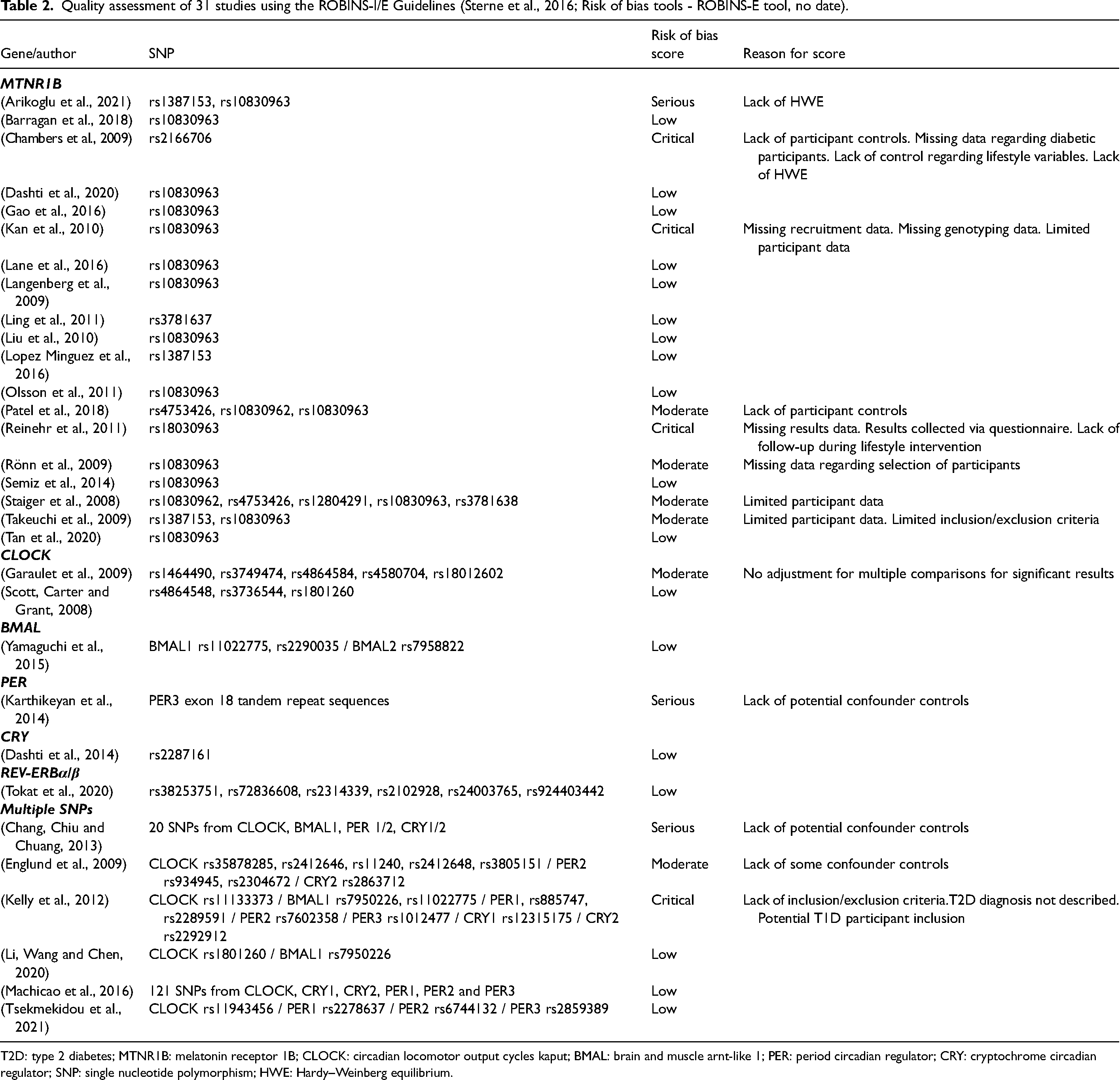
T2D: type 2 diabetes; MTNR1B: melatonin receptor 1B; CLOCK: circadian locomotor output cycles kaput; BMAL: brain and muscle arnt-like 1; PER: period circadian regulator; CRY: cryptochrome circadian regulator; SNP: single nucleotide polymorphism; HWE: Hardy–Weinberg equilibrium.
Findings
Associations/Effects of MTNR1B Gene Variations on T2D Outcomes: All 19 MTNR1B studies determined significant associations/effects of genetic variations on T2D outcomes (Tables 1, 2, and 3). SNPs that were associated with increased T2D outcomes were: rs1387153, 10830963, rs3781637, rs1083096, rs10830962, rs2166706, rs3847554. SNPs associated with reduced T2D outcomes were: rs780094 and rs3781638. However, one study only found significant results in participants aged <41 (Barragan et al., 2018) (Table 4). There were differences due to ethnicity, including a significantly greater association amongst Indian and Shanghai populations in comparison with Caucasian European and Beijing populations (Chambers et al., 2009; Liu et al., 2010). Also, rs10830963 association with reduced T2D was dependent on the rs780094 SNP in the glucokinase regulatory protein (Gao et al., 2016) (Table 3). Diet: A significant association between rs1387153, late meal timing and T2D was found (Lopez Minguez et al., 2016) (Table 5). Sleep: A significant association was found between rs10830963 and sleep variables including shift work, a ‘definitely morning’ chronotype, early rising, early sleep timing and T2D (Dashti et al., 2020; Lane et al., 2016). Other studies found no association between MTNR1B genetic variations, sleep variables and T2D (Liu et al., 2010; Olsson et al., 2011).
Case-control studies including genes and SNPs/authors/design/participant numbers/inclusion of diet and sleep variables/outcomes measured.
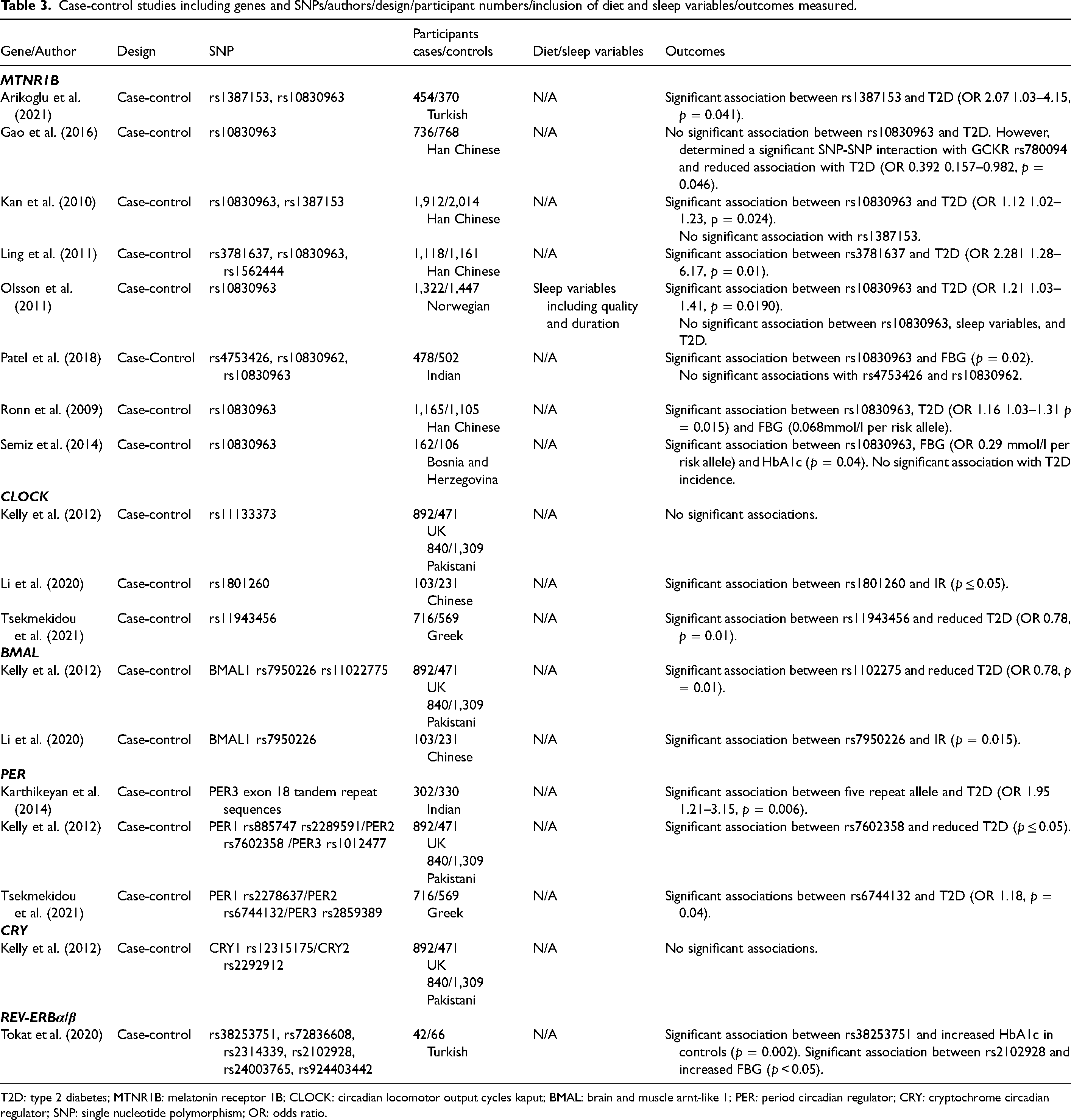
T2D: type 2 diabetes; MTNR1B: melatonin receptor 1B; CLOCK: circadian locomotor output cycles kaput; BMAL: brain and muscle arnt-like 1; PER: period circadian regulator; CRY: cryptochrome circadian regulator; SNP: single nucleotide polymorphism; OR: odds ratio.
Cross-Sectional studies including genes and SNPs/authors/design/participant numbers/inclusion of diet and sleep variables/outcomes measured.
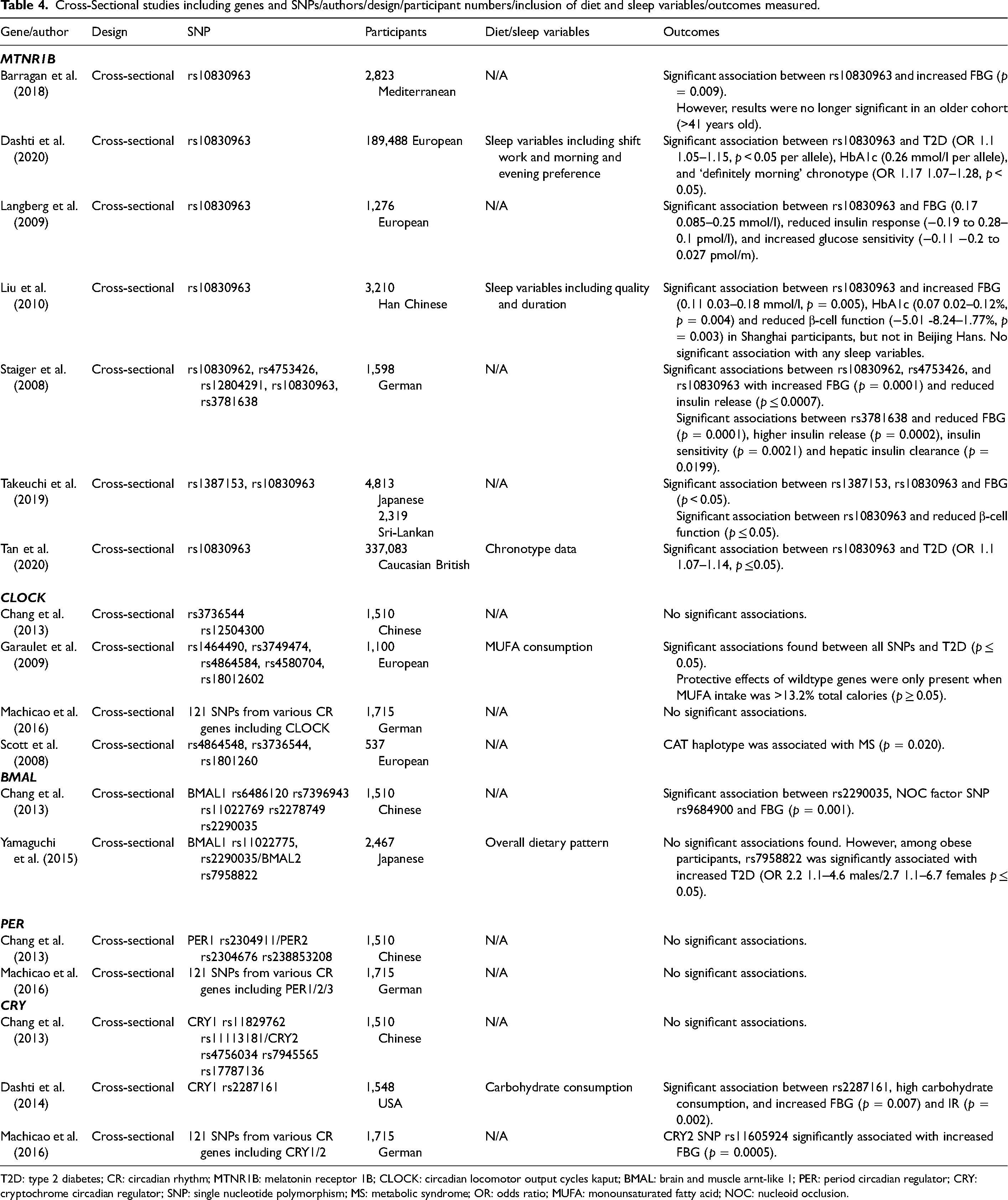
T2D: type 2 diabetes; CR: circadian rhythm; MTNR1B: melatonin receptor 1B; CLOCK: circadian locomotor output cycles kaput; BMAL: brain and muscle arnt-like 1; PER: period circadian regulator; CRY: cryptochrome circadian regulator; SNP: single nucleotide polymorphism; MS: metabolic syndrome; OR: odds ratio; MUFA: monounsaturated fatty acid; NOC: nucleoid occlusion.
Other study designs including genes and SNPs/authors/design/participant numbers/inclusion of diet and sleep variables/outcomes measured.
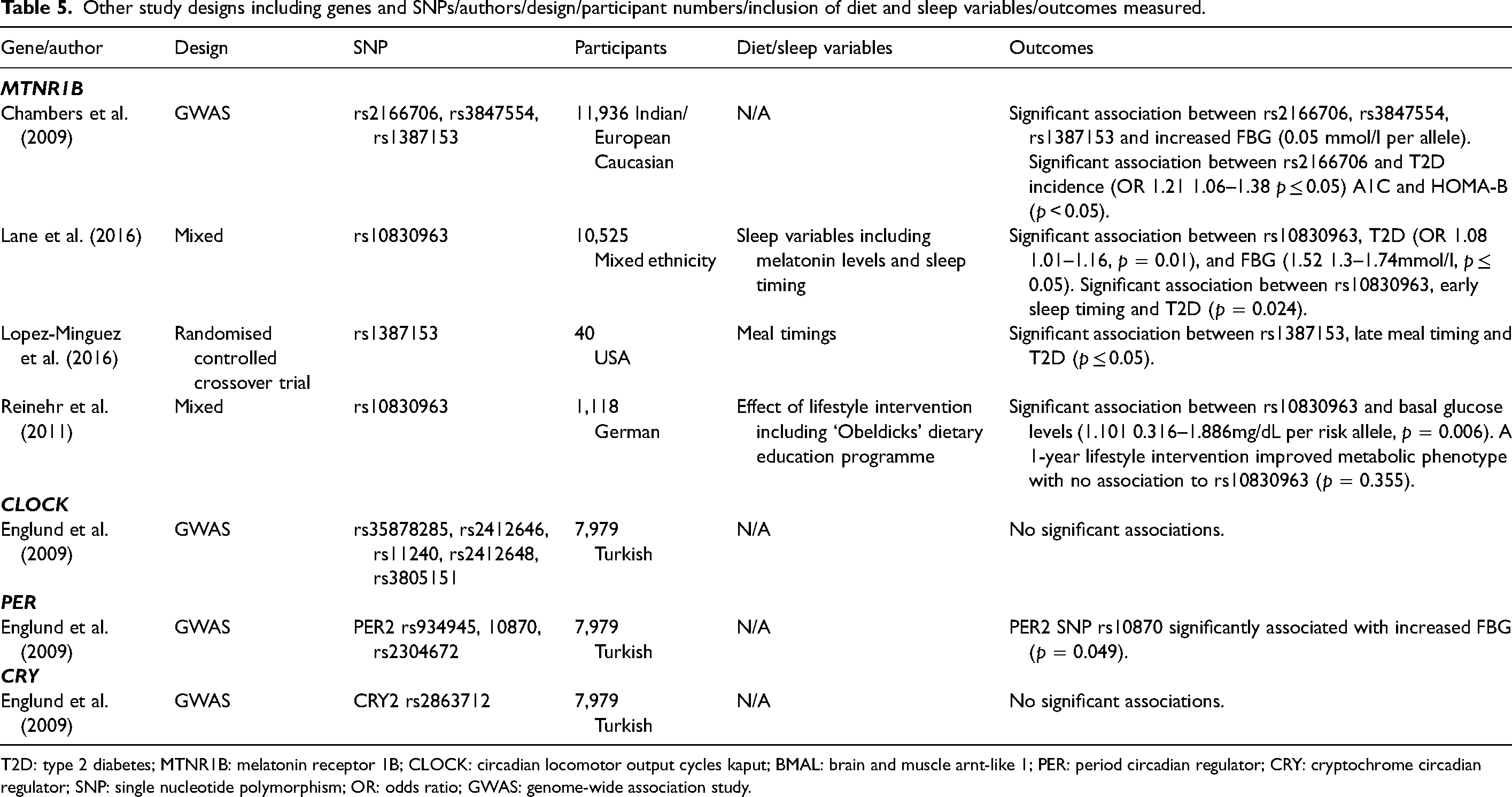
T2D: type 2 diabetes; MTNR1B: melatonin receptor 1B; CLOCK: circadian locomotor output cycles kaput; BMAL: brain and muscle arnt-like 1; PER: period circadian regulator; CRY: cryptochrome circadian regulator; SNP: single nucleotide polymorphism; OR: odds ratio; GWAS: genome-wide association study.
Associations/effects of CLOCK gene variations on T2D outcomes: Four of eight studies determined variations in CLOCK are associated with T2D outcomes (Tables 3, 4, and 5). SNPs that were associated with increased T2D outcomes were rs1801260, rs1464490, rs3749474, rs4864584, rs4580704, rs18012602 and rs3736544. However, rs11943456 was associated with reduced T2D outcomes. Diet: In one cross-sectional study, a protective association of wild-type CLOCK genes was only established when participants consumed >13.2% of calories from monounsaturated fatty acids (MUFAs) (Garaulet et al., 2009) (Table 4).
Associations/Effects of BMAL gene variations on T2D outcomes: Four BMAL studies found significant associations with T2D outcomes (Tables 3 and 4). SNPs that were associated with increased T2D outcomes were: rs7950226, rs2290035 and rs7958822. Conversely, rs1102275 was associated with reduced T2D outcomes. However, rs7958822 was only significantly associated with T2D in obese participants (Yamaguchi et al., 2015) (Table 4), and rs2290035 was only associated with FBG in the presence of the polymorphism rs9684900 in the NOC gene (Chang et al. 2013) (Table 4). Diet: No significant associations were found between overall dietary habits, BMAL variations and T2D outcomes (Yamaguchi et al., 2015) (Table 4).
Associations/effects of PER gene variations on T2D outcomes: Four of six PER studies found significant associations with T2D outcomes (Tables 3, 4 and 5). SNPs that were associated with T2D were a PER3 exon 18 tandem repeat sequence, rs6744132, and rs10870. However, rs7602358 was associated with reduced T2D.
Associations/effects of CRY gene variations on T2D outcomes: Two of five CRY studies found significant associations with T2D outcomes (Tables 3, 4 and 5). SNPs that were associated with increased T2D were rs2287161 and rs11605924. Diet: One study determined that high carbohydrate consumption (as determined by greater than median consumption in two cohorts: >/<41.65% Mediterranean/>/<49.14% USA) and the CRY1 SNP rs2287161 were significantly associated with higher levels of IR and FBG compared to the low carbohydrate consumption group (Dashti et al., 2014) (Table 4).
Associations/effects of REV-ERBα/β gene variations on T2D outcomes: The only REV-ERBα/β study determined rs38253751 was significantly associated with increased HbA1c amongst the controls, and there was a significant association between rs2102928 and increased FBG (Tokat et al., 2020) (Table 3).
Discussion
The current systematic review aimed to determine whether variations in CR genes had an association with, or effect on T2D outcomes. The evidence suggests variations in MTNR1B, BMAL and PER are associated with T2D, whilst more evidence is needed for other CR genes, including CLOCK, CRY and REV-ERBα/β. The systematic review found no consistent associations or effects of CR gene variations in combination with diet and sleep on T2D outcomes.
Variations in MTNR1B were consistently associated with increased T2D (Tables 3, 4, and 5). Gain-of-function MTNR1B variations can lead to increased melatonin release, a hormone that regulates the sleep-wake cycle, but which has been associated with increased IR, decreased glucose-stimulated insulin response, and increased T2D risk (Dashti et al., 2020). The SNPs rs10830963, rs10830962 and rs1387153 were consistently associated with T2D outcomes, whereas rs4753426, rs3781637, rs3781638, rs2166706 and rs3847554 had mixed results or a limited number of studies. More replication is needed for consistent results regarding MTNR1B variations interaction with diet and sleep factors, including shift work, late chronotypes, and early rising. Future MTNR1B studies should investigate location and ethnicity differences, including between Caucasian and Indian, and Shanghai and Beijing populations. Differences between these populations may be due to genetics, lifestyle variables that were uncontrolled during baseline comparisons, or seasonality and light exposure, particularly in the Caucasian population from Northern Europe. The SNPs rs780094 and rs3781638 require further investigation, as they were associated with reduced T2D, which was contrary to the majority of the evidence.
Variations in CLOCK genes had inconsistent links to T2D outcomes (Tables 3, 4, and 5). However, rs1801260 was associated with T2D outcomes in multiple studies (Garaulet et al., 2009; Li et al. 2020; Scott et al. 2008). Other SNPs associated with T2D outcomes that require more replications are rs1464490, rs3749474, rs4864584, rs4580704, rs18012602 and rs3736544. Variations in CLOCK downregulate sirtuin 1 and limit adipocytokine expression which may lead to T2D. However, the SNP rs11943456 requires further investigation, as it was associated with reduced T2D. An area for future research is if sleep and CLOCK variations can modify T2D outcomes, and the effects of MUFA, as one study found there may be a protective effect of high levels of consumption (Garaulet et al., 2009) (Table 4).
Variations in BMAL were consistently associated with T2D outcomes (Tables 3 and 4). However, replication studies are needed regarding rs7950226, rs2290035 and rs7958822 – all SNPs with an association – as none were repeated across multiple studies. Also, rs1102275 requires further study, as it was the only SNP associated with reduced T2D (Kelly et al., 2012) (Table 3). BMAL variations have been associated with lower β-cell function and decreased pancreatic islet development (Chang et al. 2013; Kelly et al., 2012). Also, similar to the CLOCK gene, BMAL also regulates sirtuin 1 expression, which has been associated with decreased β-cell function and IR (Li et al. 2020), which may increase T2D outcomes. More studies of variations in BMAL and T2D should be an area of future research, as currently, only four have taken place. Further studies involving diet and sleep variables are also required, as currently, only one study with a dietary variable has taken place, with no significant findings (Yamaguchi et al., 2015) (Table 4). Current BMAL studies were also of low quality, due to a lack of control of confounders, lack of clear inclusion and exclusion criteria, poor T2D diagnosis methodology, and possible inclusion of T1D participants (Table 2). Therefore, current evidence regarding BMAL may have low reliability, and improved quality studies should be a focus of future research.
The majority of PER studies found associations with T2D outcomes (Tables 3, 4, and 5). However, the variations associated with T2D outcomes – a PER3 exon 18 tandem repeat sequence, rs644123 and rs10870 have not been replicated in multiple studies. Also, rs7602358 requires further investigation, as it was associated with reduced T2D (Kelly et al., 2012) (Table 3). Gain-of-function PER variants may increase risk of T2D via repressing the CLOCK/BMAL heterodimer, which has been associated with reduced pancreatic β-cell function, islet development and insulin release (Englund et al., 2009). Another area for future study is if PER variations interaction with diet and sleep variables modify T2D outcomes, as currently, none have taken place.
The majority of CRY studies found no association with T2D outcomes (Tables 3, 4, and 5). However, when rs2287161 was paired with high carbohydrate consumption, it was associated with increased FBG and IR (Dashti et al., 2014) (Table 4). Also, rs11605924 was associated with increased FBG (Machicao et al., 2016) (Table 4). The CRY gene can limit gluconeogenesis via downregulating cyclic adenosine monophosphate (cAMP)/cAMP response element binding (CREB) protein signalling, and repressing glucocorticoid receptor and nuclear forkhead box protein O1 (Rijo-Ferreira and Takahashi, 2019). Reduced expression of CRY due to genetic variations therefore upregulates gluconeogenesis, increasing T2D risk (Rijo-Ferreira and Takahashi, 2019). Future CRY studies should further investigate its associations with high carbohydrate consumption, as this could lead to actionable changes to reduce T2D risk. Also, future studies could investigate CRY variations interactions with sleep variables to modify T2D outcomes, as currently none have taken place.
The singular REV-ERBα/β study found significant associations between rs38253751, rs2102928 and T2D outcomes (Table 3). Gain-of-function REV-ERBα/β variations increase phosphoenolpyruvate carboxykinase and glucose-6-phosphatase expression, therefore upregulating hepatic gluconeogenesis (Jakubowicz et al., 2017; Tokat et al., 2020). REV-ERBα/β variations may also increase hepatosteatosis, which may increase T2D risk (Jakubowicz et al., 2017; Tokat et al., 2020). However, only one study regarding REV-ERBα/β variations and T2D has taken place; therefore, more are needed to increase the quality of evidence.
Literature evaluation
A limitation of current research is a lack of longitudinal studies. All but two included data measurements from only one-time point (Lopez Minguez et al., 2016; Reinehr et al., 2011). T2D is a chronic disease that develops over long periods, therefore, longitudinal designs that include variables such as diet and sleep are essential to produce robust results (Forouhi and Wareham, 2014). In addition, a lack of intervention trials (n = 2) (Lopez Minguez et al., 2016; Reinehr et al., 2011) mean causal links between CR genes, lifestyle factors and T2D are yet to be established. The majority of included literature was case-control or cross-sectional studies (n = 26). A strength of case-control and cross-sectional studies is many variables, including diet, sleep, and participant characteristics can be studied simultaneously. Also, they can generate hypothesis for future intervention studies. However, case-control and cross-sectional studies are liable to suffer recall bias and cannot establish cause and effect. Another limitation was that data regarding diet and sleep were frequently recorded via questionnaire. Questionnaire data is susceptible to bias and may limit reliability of results (Resnicow et al., 2000). Intervention studies with more strict controls over diet and sleep are therefore required.
Another limitation of current research was thirteen studies only found differences in metabolic traits, rather than T2D incidence. Therefore, the associations/effects of CR gene variations on T2D risk cannot be established. For example, one study determined rs10830963 was associated with higher FBG and HbA1c, but not increased T2D risk (Semiz et al., 2014). This suggests the overall effect size of variations in CR genes and T2D outcomes may be negligible, reducing clinical and practical relevance. Another limitation was a number of studies included participants taking T2D medication. This may have confounded studies that measured T2D status via metabolic phenotypes. However, excluding all participants taking T2D medication would have been too limiting for the current systematic review. A strength of current literature was similar male (51.5%) female (48.5%) inclusion, as results can be generalised to both sexes.
An interesting finding was the BMAL and PER SNPs (rs1102275/rs7602358) associated with reduced T2D were in the same study, which had a critical ROB due to a lack of inclusion/exclusion criteria, lack of description of T2D diagnosis methods, and possible T1D participant inclusion (Kelly et al., 2012) (Tables 1 and 3). Therefore, these SNPs require further investigation to confirm their interaction with T2D.
Another area for future research is increased replication in East and South-East Asian, and Caucasian European populations, as current literature, indicates populations in these locations may have significantly different outcomes to variations in CR genes (Chambers et al., 2009; Liu et al., 2010).
Conclusion
T2D is a chronic disease that places a huge burden on individual's lives and health services globally, which is forecast to worsen in coming years. Recently, the effects and associations between genetic variations in CR genes and T2D have been investigated. This novel systematic review aimed to assess the current literature to determine CR genes associations and/or effects on T2D, as well as the modifying effects of diet and sleep. The results of this systematic review suggest consistent associations of variations in MTNR1B, BMAL and PER with T2D outcomes. For confirmatory results, and before practical and clinical recommendations can be made, further longitudinal and intervention studies and further diet and sleep studies need to be performed.
Supplemental Material
sj-docx-1-nah-10.1177_02601060231179777 - Supplemental material for A systematic review of variations in circadian rhythm genes and type 2 diabetes
Supplemental material, sj-docx-1-nah-10.1177_02601060231179777 for A systematic review of variations in circadian rhythm genes and type 2 diabetes by Harry Stevens, Giulia Verdone, Leonie Lang, Catherine Graham, Leta Pilic and Yiannis Mavrommatis in Nutrition and Health
Footnotes
Availability of data and materials
Supplementary data is published on the Figshare repository platform.
Consent for publication
All authors have expressed consent for publication.
Declaration of conflicting interests
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: LP is the founder of Optimyse Nutrition LTD, a personalised nutrition company offering genetic testing to clients.
Ethical approval
All reported studies including human participants, human data, or human tissue include a statement on ethical approval and consent. This systematic review was registered with PROSPERO on 13/8/2021 (registration: CRD42021259682).
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
