Abstract
Recent trends in toxicological pathology include implementation of digital platforms that have gained rapid momentum in the field. Are we ready to fully implement this new modality? This opinion piece provides some practical perspectives on digital pathology such as its cost limitations, relative time requirements, and a few technical issues, some of which are encountered for specific lesions, that warrant caution. Although the potential for digital pathology assessment with whole slide images has made great strides, we are of the opinion that it is not yet ready for complete replacement of glass slides in toxicologic pathology safety assessments.
This is an opinion article submitted to the Toxicologic Pathology Forum. It represents the views of the authors. It does not constitute an official position of the Society of Toxicologic Pathology, British Society of Toxicological Pathology, or European Society of Toxicologic Pathology, and the views expressed might not reflect the best practices recommended by these societies. This article should not be construed to represent the policies, positions, or opinions of their respective organizations, employers, or regulatory agencies.
Like a growing number of other toxicologic pathology groups, we, at Greenfield Pathology Services, Inc., offer and use digital toxicological pathology capabilities. Digital imaging is used for live viewing of histology during pathology rounds and consultations. However, we remain skeptical regarding implementing digital pathology for a routine full tissue primary screening evaluation of a clinical trial-enabling nonclinical study. We are still finding that histopathological evaluation of glass slides is almost always the best choice for the sponsor and for the scientific integrity of the study. One primary factor is cost because digital slide reading requires an additional process (scanning and the associated infrastructure and support) and additional time for pathology evaluation to achieve a relatively equivalent level of critical evaluation compared with glass slides. As Dr LaPerle in her companion article addresses some of the advantages of digital evaluations, this current opinion piece serves as a counterpoint discussion on current limitations toward complete transition to digital pathology platforms as a community. 1
How many of us envision a future where our work involves portable laptops instead of requiring our use of bulky microscopes? The answer is obvious. That stated, even as commerce moves globally toward digital platforms across multiple realms, our work as toxicologic pathologists remains diagnostic with its objectives oriented toward patient safety and public health. As change is inevitable, it is imperative that our transitions to digital microscopy remain data-driven and tempered by critical assessments. Decisions driven by data from robust validation studies will position digital pathology appropriately within our tool set in toxicologic pathology. The following paragraphs highlight some key challenges encountered with digital pathology relative to microscopic pathology as applied to safety evaluations. Because nonclinical evaluations in toxicology studies remain the core of safety assessments in clinical trials, it is up to us as scientists and professionals to ensure that the decisions to adopt new techniques are data-driven and not based merely on ease of use at the bench. Given that patient safety is not a portable asset, our opinion is to tread cautiously as we transition into the new era of digital spaces while emphasizing the maintenance of key features available with traditional microscopic pathology.
There are three interrelated areas that are key considerations when selecting whole slide imaging (WSI) for histopathology, namely, expense, time requirements, and technical issues impacting scientific rigor.
First, digital pathology adds expenses to nonclinical study pathology evaluations which can be significant. The glass slides still need to be prepared from the wet tissue specimens—that cost is unchanged. Each additional preparation for the digitized slide is at an additional cost. Slide scanning is priced variably, from as much as $20/slide down to a proposed $3 to $4/slide. The $3 range has been projected to be the price point once it becomes routine with consistent and high-volume outputs. The $3 slide scanning cost assumes that all the infrastructure is there to implement it, where infrastructure includes purchase and maintenance of scanners, personnel to quality control (QC) slides, clean slides as needed, loading and unloading of slides from scanners, and personnel and/or software to QC the whole slide images once scanning is complete. If during the scan process it is determined that there is wax or debris on the coverslip, then the slide needs to be scanned again, which in turn translates to additional time and costs (Figure 1). Moreover, there is additional technical IT infrastructure to maintain (eg, servers, cloud storage/computing, laptops with sufficiently large hard drives to work with numerous WSIs, ergonomic mice, high-definition monitors) by laboratories as well as at the study pathologist end assessing WSIs. If GLP compliance is required, then validation is mandatory generating additional costs; validation includes data integrity and security as well as validating that the images on the monitor are true to the original glass. In fact, a whole new IT infrastructure would need to be developed around this new process step. On the other hand, while IT infrastructure has components that tend to have a limited lifespan, often measured in years, a microscope has a lifespan measured in decades.
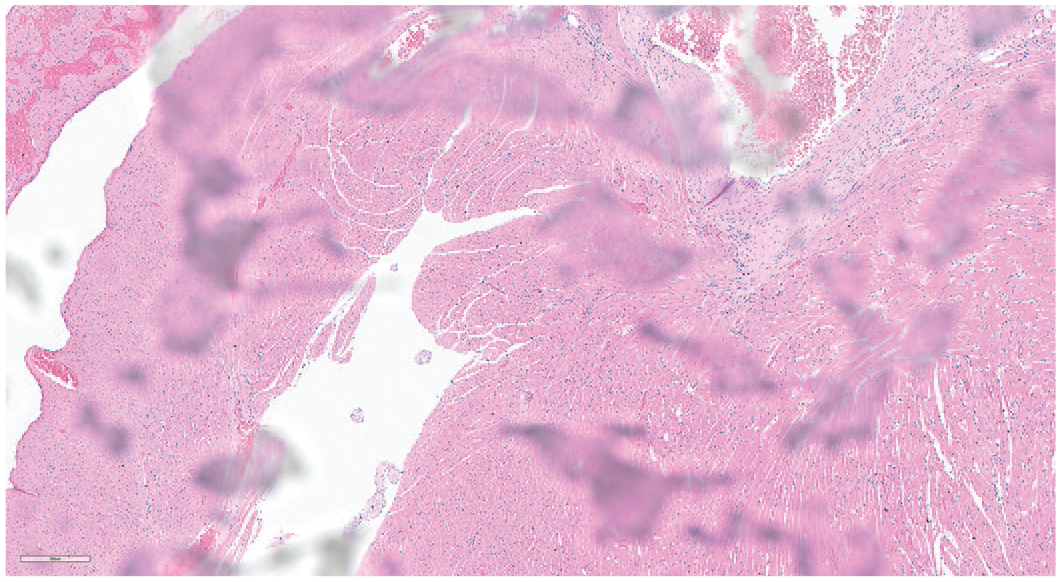
Portion of a whole slide image received for interpretation captured from a glass slide with wax on the coverslip.
Second, there are several considerations in assessing the difference in total study time requirements between glass slides and digital pathology evaluations across the timeline for pathology evaluation. Scanning slides adds to the timeline. For a fair assessment, the comparison should be based on the date that the slides are ready, and shipment initiated to the study pathologist (for glass slides) versus initiation of slide delivery to the individual responsible for scanning (for whole slide images). For both scenarios, the endpoint remains delivery of the draft pathology report. We had one experience where whole slide images were delivered a full week late because the scanners were overbooked. We have also experienced instances where slides had to be return shipped because the laboratory failed to scan them for the pathology peer reviewer prior to sending them to the study pathologist for a glass primary read. In such instances, scanning can become a huge time sink.
Ironically, after scanning, delivery of the images to the pathologist is not necessarily instantaneous. There are two routes. One is to ship an encrypted hard drive, while the second is to download the images from a secure portal. The time required for shipping a hard drive is generally equivalent to shipping the slides. Downloading from a secure portal can be variable and largely dependent on bandwidth. Even with smaller studies, this can take more than just an overnight session, which can be viewed as negating the time savings of delivering whole slide images versus shipping glass slides. With glass slides, there is of course a need to have proper chain of custody. For whole slide images, this is built into the digital platform. With glass slides, this may require some effort for things such as slide inventory, depending on the sponsor-specific requirements. However, glass slide inventory documentation is often a small part, ranging from a few minutes up to two hours for a large study relative to time involved for digital scanning plus download.
The additional pathologist time is an important consideration. In our experience, we find that it can take up to four times longer to evaluate WSIs compared with the time required to evaluate the equivalent in glass slides. That may partially be because we are efficient with the microscope and sorting slides, but so far have not done many digital evaluations. There are claims of significant time savings on the part of the pathologist by going to review of whole slide images. There has been a lot of discussion about the increased efficiency of review of slides with WSI being in the range of 42%. 2 Although widely cited at meetings and in conversations, the actual details of methodology for how this conclusion has been reached have not yet been published or transparent. It is unclear whether the number was generated using real-world experiences in identifying and diagnosing lesions, or whether it is based on an observer with a stopwatch observing whole slide images user. It is also not clear how many studies or what types of studies and/or tissues were used to generate the data. In informal discussions, we have not heard of anyone actually allocating less time for the slide evaluation due to such purported savings. Our experience is that whole slide images results in more time required, even with slide sorting time accounted for when specific target organs need to be evaluated. The increased time required for assessment of WSIs translates into increased expense, reduced productivity, and a greater number of pathologists to accomplish the same amount of work.
Another consideration impacting time is a pathologist’s efficiency in handling slides versus changing digital images. For us, to take one slide off the microscope stage and replace with it a second slide is as quick as reaching over to press the return key on the laptop. When dealing with digital, sometimes it is that quick; sometimes not, depending on the capabilities of the given software available to the given pathologist.
Once a slide is positioned on the microscope, the optical image of the microscope has a larger field of view compared with a digital image for a given magnification. This increases viewing efficiency (Figure 2) with a microscope. A microscope provides a circular field of view. Viewed on a computer monitor, that field of view is rectangular. If we assume that the corners of the chip recording the image meet the circumference of that circle, and that the chip replicates an approximate 16:9 ratio rectangle at the same magnification, then pathologist can microscopically view almost twice the amount of tissue in a single field on a microscope compared with that viewed on a computer monitor with a WSI at the same magnification.

Schematic of the field of view in observing via a microscope and computer monitor (not drawn to scale). Assuming that the chip is 16:9 ratio and assuming that the corners of the chip are within and meet the circumference of the microscope field of view, a pathologist examining glass microscope slides will see almost twice as much tissue in the field of view at the same magnification.
Finally, there are technical reasons why WSIs may present some challenges. If whole slide images are not properly quality controlled, there can be artifacts that affect interpretation (Figure 1). When a pathologist receives the scanned image represented in Figure 1, there is nothing that the pathologist can do other than request that the slide be cleaned, rescanned, and resent. If instead, the pathologist has the glass slide in hand, it is a simple matter to wipe off the excess wax (or fingerprints blurring the image) and evaluate the slide without delay. The diligence that labs have in quality control of WSIs varies widely. A pathologist working commonly with a laboratory that does this well may be surprised to receive these low-quality whole slide images when working with a different laboratory.
Another unique issue is that different tissues often adhere to the glass slide variably such that a specific tissue is not always uniformly in the same plane of focus throughout the glass slide. This means that portions of the tissue are out of focus and cannot reliably be examined as a WSI. The common rejoinder from digital pathology vendors and proponents is to employ z-stacking (a digital image-processing method that combines multiple images taken at different focal distances to provide a composite image with a greater depth of field). However, that requires either doing z-stacked scans of all slides, or of specific tissues that commonly have this issue, or on specific glass slides. If specific slides, that requires a priori knowledge that the slide needs to be scanned with z-stacking capabilities. Although z-stacking may be one solution, it is important to realize that most of the major slide imaging viewing platforms do not yet support z-stacked images, so it remains unclear whether this is even a viable solution in the current environment as we surge forward into adopting digital pathology. Hopefully, the scanners as well as the various imaging platforms will implement this technology—but the point is that we are not there yet!
Some subtle lesions can be difficult to see. Such lesions often require manual fine focus adjustment available as a knob on the traditional light microscope. The ability to focus up and down in the plane of section generally increases the efficiency and accuracy of the pathologist in identifying such lesions (see Figure 3). Manual focus adjustment is unavailable on digital platforms. In fact, WSIs are generally limited to a single plane of focus. Although, z-stacking can, in theory, address this limitation, it is simply not routinely available at this time on currently marketed digital platforms. Therefore, the probability of capturing subtle lesions is further compounded on digital platforms relative to traditional light microscopic evaluation. This is especially relevant with tissues where heterogeneity is the norm, such as with the brain and/or spinal cord sections where neither neurons nor lesions look the same across the sectional landscape. For screening evaluations of heterogeneous tissues (where artifacts are not uncommon), additional tools generally aid in the identification of potential morphological changes (such as epifluorescence and polarization as discussed below). The lack of such tools on digital pathology platforms, compounded with the inability to adjust fine focus, diminishes the technical rigor employed to screen morphological changes. Unlike diagnostic pathology with clues from signalment and history, toxicologic pathology evaluations are screening toxicology studies designed to identify effects of test compounds with unknown potential for toxicity. Not having a priori knowledge of susceptible areas within the heterogeneous landscape for any given test compound or class increases the probability that it will be missed. Most importantly, it is unclear whether data from validation studies demonstrating the sensitivity [Type I (false positives) vs Type II (false negatives) errors] for digital pathology evaluations as compared to traditional microscopic evaluation with glass slides are available. Until such data become transparent, implementing digital pathology alone (ie, without concurrent and/or limited access to glass slides), especially in some specialty areas such as neuropathology, poses risk and negative implications for test compound safety screening assessments. If morphologic changes are visible, but less readily apparent on WSIs, then it would suggest that the pathologist needs to spend more time on each WSI (especially if glass slides are needed for confirmation). Such situations decrease overall efficiency contrary to the 42% increase in efficiency claim.
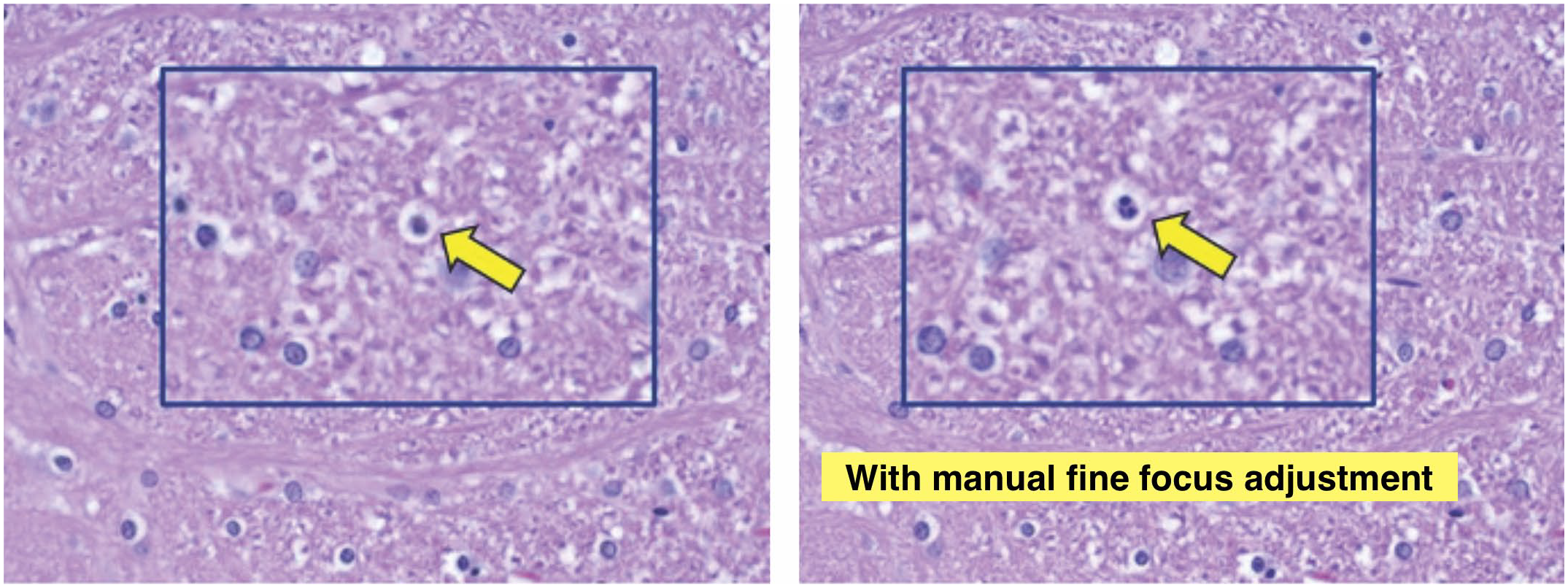
These images depict visualization of an apoptotic focus associated with nerve fiber degeneration of thinly myelinated sensory nerve fibers within a cross-section of the spinal trigeminal tract. The image on the left is depicted in a single plane of focus with the inset depicting the area of interest. The image on the right depicts the same area following manual focus adjustment on microscope. The associated inset depicts the same area of interest as image on the left. Note the clear individual apoptotic bodies in the image on the right following manual focus adjustment, thereby enabling confident diagnosis. Coronal section, Brain Level 6, Rat, original objective 40X.
In neuropathology-focused studies, the use of tools such as epifluorescence to identify degenerating neurons can be enormously helpful. This same technique can often also be useful for assessment of the retina and retinal pigment epithelium. Being limited only to brightfield scans means that this technique cannot be leveraged, and that the identification of affected neurons becomes much more painstaking. Digital scans of slides labeled with fluorescent markers are not yet a practical consideration. Similarly, screening for crystals/collagen/foreign body matter (medical device pathology) is very efficient using a polarizer on a microscope which is also not currently possible with digital images. And then there are the situations where a magnification higher than the scanned magnification is desired—which is not a possibility with a digital image compared to the occasional use of a 60X or 100X objective. In situations where only the WSIs are available (such as when limited by international shipment of nonhuman primate tissues), and the flexibility of an optical microscope is desired or even necessary (such as the conduct of an independent peer review), this would mean an additional time delay for the glass slide(s) to be delivered.
Beyond the limitations of the scanner, there may be limitations of the computer monitor. Depending on the settings, features such as basophilia may be visible on one monitor but not on a colleague’s monitor. It is important to have the appropriate equipment and calibration of monitors to eliminate this error. Monitors also have differences in resolution. That can impact ease of identification of findings on whole slide images. Many pathologists we have talked with indicate that they routinely do their reviews at a higher magnification with a whole slide image than they normally would with glass, since they have anxieties about not identifying all findings. Reviewing at a higher magnification means a longer time to cover the same tissue area. It may be that this changes with time and experience.
Because majority of the work in toxicologic pathology involves screening studies, generally without a priori knowledge of what might be revealed during evaluation of the tissue section, having the most robust methods for evaluating slides is important to minimize the risk of missing subtle yet important microscopic findings. As in toxicology, subtle does not reflect insignificant. Our experience is that toxicology screening is still best done with evaluation of the glass slides that allows use of additional tools to focus, capture autofluorescence, and employ polarization as warranted.
Digital pathology has developed a level of sophistication that can indeed be usefully employed in certain niches and special situations; it is especially useful for the evaluation of the presence/absence of carefully pre-selected and validated morphological changes. Many of us are becoming familiar with the technology and are embracing the ease-of-use. However, before it becomes routine for our screening toxicology studies which are determining endpoints for the first-in-human dose enabling clinical studies, sponsors must have a valid reason to choose this option. For example, we may expect added value in the form of analytical packages from current machine learning software advances. As compelling new opportunities arise, we will be in a position to effectively deploy them, but we should do so responsibly without compromising existing features. So, our answer to the question “Are we there yet?” for digital pathology in our core human dose enabling studies is “no, not yet,” with an emphasis on the “yet” and guarded optimism that we will eventually get there.
Footnotes
Acknowledgements
The authors would like to thank Dr Dennis Hoover for helpful discussions in the preparation of this opinion article and Ms Beth Mahler for her assistance with the figures.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
