Abstract
To address the challenges posed by large-scale development, validation, and adoption of artificial intelligence (AI) in pathology, we have constituted a consortium of academics, small enterprises, and pharmaceutical companies and proposed the BIGPICTURE project to the Innovative Medicines Initiative. Our vision is to become the catalyst in the digital transformation of pathology by creating the first European, ethically compliant, and quality-controlled whole slide imaging platform, in which both large-scale data and AI algorithms will exist. Our mission is to develop this platform in a sustainable and inclusive way, by connecting the community of pathologists, researchers, AI developers, patients, and industry parties based on creating value and reciprocity in use based on a community model as the mechanism for ensuring sustainability of the platform.
Developments in high-throughput slide scanning and data storage have revolutionized the field of pathology by enabling whole slide imaging (WSI) of histopathological specimens. Combined with the unprecedented possibilities of recent artificial intelligence (AI) techniques such as deep learning and hardware, we are now on the verge of accelerating “AI pathology” and spur the use thereof across the entire value chain—from drug discovery toward clinical diagnostics. In short, our goal is to bring to digital pathology the same acceleration that publicly sourced repositories of images such as ImageNet 1 have allowed for the development of AI for general-purpose computer vision. However, there are clearly several challenges that need to be overcome regarding large-scale development, validation, and adoption of AI in pathology.
To address the challenges related to the availability of large sets of digital slides for AI models, we have proposed the BIGPICTURE (http://www.bigpicture.eu/) project to the Innovative Medicines Initiative (IMI) Call 18. IMI is Europe’s largest public–private partnership initiative organized by a joint undertaking between the European Union and the European Federation of Pharmaceutical Industries and Associations (EFPIA) to support collaborative research and develop networks of industrial and academic experts in Europe. 2
As a consortium of academics, small enterprises, and pharmaceutical companies, our vision is to become the catalyst in digital transformation of pathology by creating the first European, ethically compliant, and quality-controlled WSI platform, in which both large-scale data and AI algorithms will exist. BIGPICTURE’s mission is to develop this platform in a sustainable and inclusive way by connecting the community of pathologists, researchers, AI developers, patients, and industry parties on the basis of value creation and reciprocity in use through a community model.
The project is guided by 4 principles. First, availability of data and algorithms to all legally and ethically entitled stakeholders, with ease of use and accessibility in mind. Second, functionality in terms of unique data processing and search tools based on state-of-the-art AI will be demonstrated through use cases focusing on the interests of various users within the community: quantitation of tumor-infiltrating lymphocytes, AI-assisted scoring of transplant kidney biopsies, and AI-assisted evaluation of nonclinical safety studies. Third, value will arise from bidirectional incentive streams between contributors and users of data and AI algorithms. In this community-based model, contributing digital slides or open-source implementation of algorithms will give access rights to other data and tools. Finally, sustainability beyond the 6 years of the funded action will be considered for every deliverable of the project with the aim to establish a platform that becomes the first European hub for hosting and using extensive WSI data sets and AI algorithms and tools that will be used by scientists, pathologists, and industrial parties. The components of the project and their relations are presented in Figure 1.
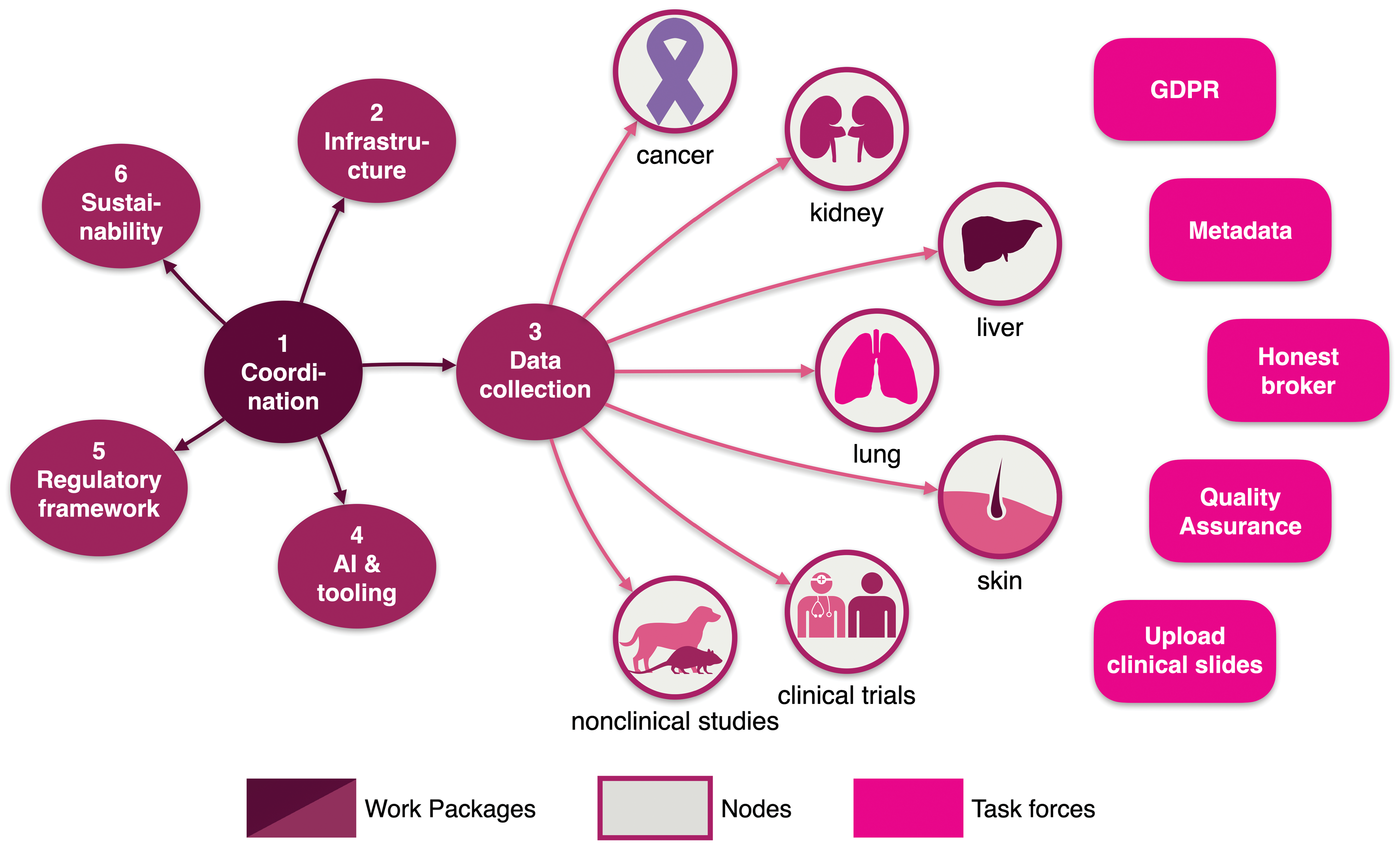
Modular structure of the project through WPs: WP1 focuses on efficiently managing a large public–private partnership with more than 40 partners and several third parties. WP2 will develop a cutting-edge, GDPR compliant-by-design infrastructure consisting of a central repository. WP3 will coordinate the collection of WSI by data collection nodes throughout Europe. WP4 will build the tools for accessing, annotating and mining digital slides. WP5 will accompany the evolution of the regulatory framework and standardization, including quality controls and quality assurance of WSI data sets. WP 6 will develop community-based business models that ensure BIGPICTURE keeps its value for the community in the future. The task forces will concentrate on specific transversal topics that play a role in several WPs. GDPR indicates General Data Protection Regulation; WP, work package; WSI, whole slide imaging.
To execute this project, we have established a consortium led by clinical and veterinary pathologists, centered around Europe’s leading researchers in digital and computational pathology, partnering with experts in research infrastructure and a strong network of pathology departments, professional societies, patient advocates, small and medium-sized enterprises (SME’s), and regulatory bodies. The project is articulated around 4 key objectives.
The first goal is to develop a sustainable, secure, and General Data Protection Regulation (GDPR) compliant repository in a federated and scalable design at 2 sites, capable of hosting over 4.5 petabytes of data. This will be based on the existing federated European Genome-phenome Archive (https://www.ebi.ac.uk/ega/home) technology established through the ELIXIR (https://elixir-europe.org) research infrastructure to manage the exchange of confidential information between contributors and users. 3
Second, our ambition is to collect and standardize more than 3 million high-quality, well-annotated clinical and nonclinical DICOM-formatted WSI with associated metadata. Of this data set, 2 million slides will be taken from animal models and toxicology studies and will include complete sets of tissues from several species, with a broad coverage of elementary lesions. Also, more than 1 million WSI will be collected from clinical pathology practice and clinical trials. This collection will create unprecedented opportunities for translational studies. In this perspective, a particular effort will be made to harmonize nomenclatures.
Third, to enhance the use of the repository, we will develop and deploy open-source and cross-platform tools to view, search, and navigate through slide data collections. Also included will be content-based image retrieval tools, so data sets may be searched not only by catalogue but also by image characteristics. 4 In addition, task-agnostic generic AI tools based on out-of-distribution data analysis and deep transfer learning will complement or even overcome the unending task-specific “collect data—train model—test model” cycle. Such tools will operate as synergistic building blocks, opening new avenues to develop algorithms that are more widely applicable. This approach will be a key enabler in several fields. Among others, it has the potential to boost efficiency of toxicity studies in animal experiments and in clinical trials, accelerating nonclinical research and thereby drug development. Several ways to interact with the WSI or the AI models available on the platform are presented in Figure 2.
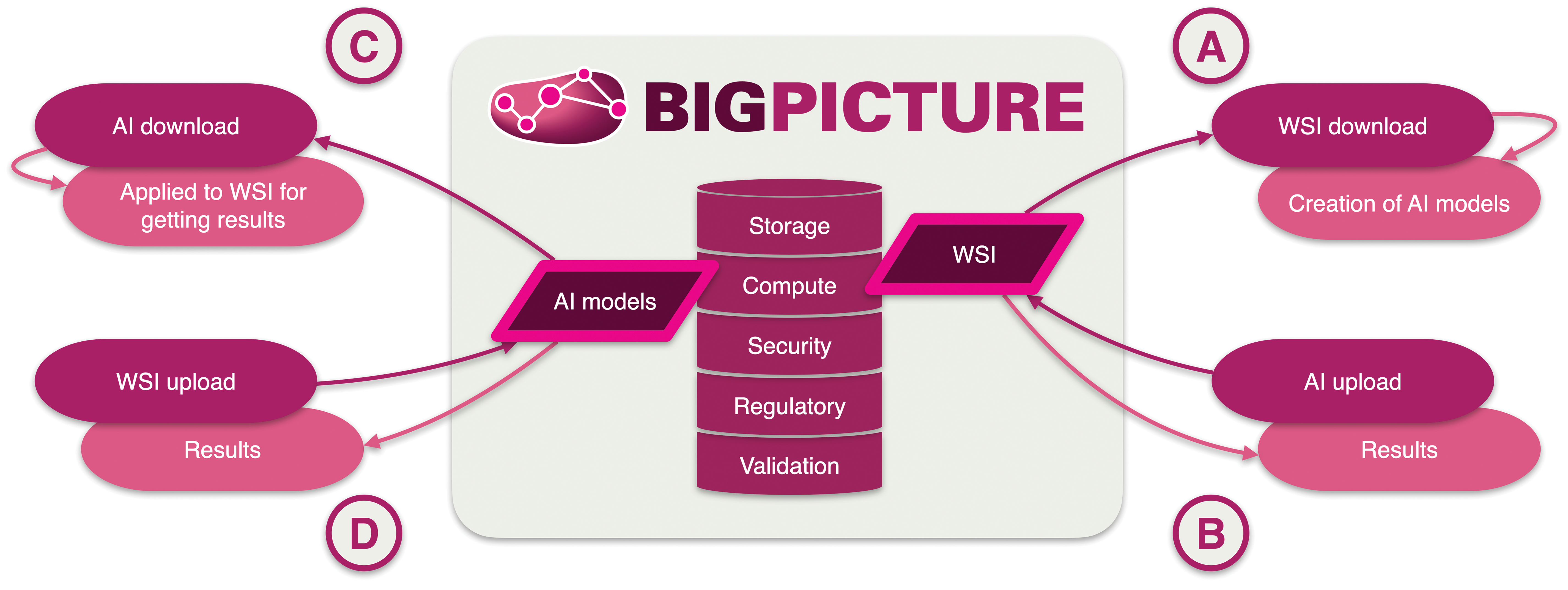
Mode of access to the data (WSI, AI models): A, Direct access to WSI data—the slide data can be accessed from the platform after approval, for example for use for the development of DL models. B, Indirect access to WSI data—the slide data never leaves the platform but is available for the validation of deep learning models, without the user ever seeing the WSI data themselves. C, Direct access to AI-models—AI tools are readily available for downloading by users and can then be used to develop new models or for analysis of datasets or clinical cases not located in the platform. D, Indirect access to AI-models—any digital slides data uploaded to the platform can be analyzed by available AI-models. The user does not get access to the AI-model itself but only receives the results from the analysis. This is well suited for the validation of data sets, clinical reference, and quality assessment purposes. AI indicates artificial intelligence; WSI, whole slide imaging.
Fourth, we aim also at coordinating the work of this consortium in order to advance the regulatory, legal, and ethical framework pertaining to the use of WSI for nonclinical safety testing and clinical use, in full compliance with GDPR. To achieve this, we will leverage ongoing dialogues with regulatory authorities FDA/EMA (eg, the working group of DPA-FDA, and the ESTP position paper featured in this issue) and by interacting with standardization organizations and the European Society for Pathology to lay down a comprehensive roadmap for the approval of advanced AI tools and digital slides for use in clinical and nonclinical assessments.
We are confident that the BIGPICTURE consortium will become a catalyst of research for AI Pathology in Europe and beyond and contribute to the higher goal of improving patients’ lives through enabling better understanding of diagnostic criteria, disease mechanisms, and therapeutics.
Footnotes
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article. Pierre Moulin is employee of Novartis AG and holds stocks and restricted stock units from Novartis AG. Erio Barale-Thomas is employee of Janssen Pharmaceuticals and holds stocks and restricted stock units from Janssen Pharmaceuticals; Member of DICOM WG26 and 33; Scientific advisor pro bono for Deciphex. Jeroen van der Laak is a member of the advisory boards of Philips, The Netherlands and ContextVision, Sweden, and received research funding from Philips, The Netherlands and Sectra, Sweden in the last five years; and acknowledges The Knut and Alice Wallenberg foundation for generous support.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement No 945358. This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA.
