Abstract
The past decade has seen an increase in the development and clinical use of biomarkers associated with histological features of liver disease. Here, we conduct a comparative histological and global proteomics analysis to identify coregulated modules of proteins in the progression of hepatic steatosis or fibrosis. We orally administered the reference chemicals bromobenzene (BB) or 4,4′-methylenedianiline (4,4′-MDA) to male Sprague-Dawley rats for either 1 single administration or 5 consecutive daily doses. Livers were preserved for histopathology and global proteomics assessment. Analysis of liver sections confirmed a dose- and time-dependent increase in frequency and severity of histopathological features indicative of lipid accumulation after BB or fibrosis after 4,4′-MDA. BB administration resulted in a dose-dependent increase in the frequency and severity of inflammation and vacuolation. 4,4′-MDA administration resulted in a dose-dependent increase in the frequency and severity of periportal collagen accumulation and inflammation. Pathway analysis identified a time-dependent enrichment of biological processes associated with steatogenic or fibrogenic initiating events, cellular functions, and toxicological states. Differentially expressed protein modules were consistent with the observed histology, placing physiologically linked protein networks into context of the disease process. This study demonstrates the potential for protein modules to provide mechanistic links between initiating events and histopathological outcomes.
The liver is a primary organ of xenobiotic metabolism and is particularly susceptible to toxicant-induced injury. Liver biopsy is the current clinical gold standard for definitive diagnosis of liver injury, but biopsies are time-consuming, invasive, and subject to sampling variability (Tan et al. 2014). Circulating biomarkers specific to organ injury can be an alternative for diagnosing liver damage, provided that these markers afford sufficient sensitivity and specificity. Measuring serum aspartate aminotransferase (AST) and alanine aminotransferase (ALT) has historically been used to assess liver damage, but these enzymes are not specific for liver tissue, severity of injury, or differential diagnosis of liver disease (Green et al. 2010; X. Yang et al. 2015; Bailey et al. 2012). The past decade has seen significant clinical advancements in combining multiple biomarkers with complex algorithms for assessing liver disease to improve clinical diagnosis (e.g., fibrotest/fibrosure, fibrometer, and hepascore; Chin et al. 2016). Use of these tests has been reported to reduce the need for liver biopsy by 50% to 80% (Chin et al. 2016). Panels of biomarkers may increase diagnostic specificity by permitting the simultaneous measurement of multiple contributors to pathogenesis. A diagnostic biomarker panel can provide a distinctive or unique signature specific to histological features of disease (AbdulHameed et al. 2014; Ippolito et al. 2016; Tawa et al. 2014).
With more than 80,000 registered chemicals in commerce and hundreds more introduced into the commercial sector each year, developing toxicology assays specific for each chemical is financially and logistically impractical (U.S. Environmental Protection Agency 2017). Understanding the molecular mechanisms underlying disease pathogenesis after toxicant exposure may facilitate the development of new, chemically agnostic diagnostic tools based on adverse health effects. In previous research in our laboratory, we used in silico approaches to identify common gene modules associated with liver toxicologic pathology, including histological fibrosis and steatosis (AbdulHameed et al. 2014; Tawa et al. 2014). These gene modules were associated with histopathological features of fibrosis and steatosis regardless of the toxicant inducing the response. Gene modules outperformed individual genes in predicting severity of histological damage (AbdulHameed et al. 2014; Tawa et al. 2014). Protein products of some of these genes were identified in the plasma as possible mechanistic biomarkers of fibrosis in both in silico and in vivo experiments (AbdulHameed et al. 2014; Ippolito et al. 2016; Tawa et al. 2014). Gene modules associated with chemically induced steatosis supported mechanistic links to the lipid accumulation profile (Tawa et al. 2014), but we did not test the steatosis predictions in vivo.
In this study, we chemically induced either fibrosis or steatosis by orally administering the reference chemicals bromobenzene (BB) or 4,4′-methylenedianiline (4,4′-MDA) to male Sprague-Dawley rats. The industrial chemicals BB and 4,4′-MDA are well-established hepatotoxicants (Heijne et al. 2005; Wong, Card, and Racz 2000; Chen et al. 2008; Dugas et al. 2004). BB exposure initially reduces glutathione, resulting in the formation of reactive oxygen species (ROS). Increased ROS leads to lipid peroxidation and the pathologies of steatosis and necrosis (Casini, Pompella, and Comporti 1985). The DrugMatrix database reports that daily doses of 785 mg/kg cause steatosis within 5 days in rats (Ganter et al. 2005).
4,4′-MDA is an ingredient in the industrial synthesis of 4,4′-methyldiphenyldiisocyanate, an aromatic diamine used in the production of polyurethane products (Dugas et al. 2004). Liver toxicity resulting from 4,4′-MDA exposure is well-documented. Occupational and accidental human 4,4′-MDA exposures have resulted in jaundice, toxic hepatitis, and cholestasis (Chen et al. 2008). Rats administered 25 mg/kg/week 4,4′-MDA subchronically developed portal vein hyperplasia, bile duct fibrosis, and necrosis of biliary epithelial cells (Dugas et al. 2004). The DrugMatrix database reports that the daily dose of 162 mg/kg caused fibrosis within 5 days of oral administration in Sprague-Dawley rats (Ganter et al. 2005).
The objective of this study was to determine the associations between toxicant-induced histological features and biomolecular changes in liver tissue, serum proteins, and serum chemistries in order to develop more specific and facile diagnostic assays. We placed these changes in the context of protein modules associated with liver injury and used pathway analysis to infer mechanistic associations between networks of proteins and histological changes in the liver. An improved mechanistic understanding of protein changes underlying well-known histopathological changes could potentially inform the identification of biomarkers for improved staging of liver diseases in the progression from steatosis to fibrosis and long-term damage.
Materials and Methods
Test Chemicals
BB (chemical abstract service [CAS] No. 108-86-1) and 4,4′-MDA (CAS No. 101-77-9) were obtained from Sigma Chemical (St. Louis, MO). Chemicals were diluted in corn oil and administered at a dose volume of 5 ml/kg. Doses were based on range-finding studies using published literature reference doses for steatosis (BB) and fibrosis (4,4′-MDA; Ganter et al. 2005, National Toxicology Program 2002, 2010).
Animals
Integrated Laboratory Systems (ILS, Research Triangle Park, NC) performed all in vivo rat experiments as previously described (Ippolito et al. 2016). All animal use procedures were approved by the ILS Institutional Animal Care and Use Committee. Research was conducted in compliance with the Animal Welfare Act, and other federal statutes and regulations relating to animals as stated in the Guide for Care and Use of Laboratory Animals (National Research Council [US] 2011). Male Sprague-Dawley rats (CD IGS [CRL: CD (SD)]), aged 8 weeks, were supplied by Charles River Laboratories (Raleigh, NC). Animals were housed 2 per cage in a temperature and humidity controlled environment with a 12-hr light/dark cycle. Reverse osmosis-treated tap water (City of Durham, NC) and Purina Rodent Diet No. 5002 (Ralston Purina Co., St. Louis, MO) were supplied ad libitum.
Chemical Exposures and Specimen Collection at Necropsy
Animals were randomized into vehicle control and dosing groups using Statistical Analysis System version 9.2 (SAS Institute, Cary, NC), controlling body weight across groups using analysis of variance. BB and 4,4′-MDA were administered in corn oil by oral gavage as a single dose or in 5 consecutive daily doses. Dosing volume was based upon daily body weight. Animals were euthanized 24 hr after the final dose administration. Livers were harvested at necropsy. One half of the left lobe of the liver was flash frozen in liquid nitrogen and stored at −80°C, and the other half was fixed in 10% formalin for histochemical staining. Plasma and serum were collected by cardiac puncture at necropsy. Using a butterfly needle, whole blood was collected into ethylenediaminetetraacetic acid tubes for plasma or tubes with no additive for serum. Plasma was isolated by centrifugation at 1,300 g for 15 min at 4°C and stored in aliquots at or below −70°C. Serum was isolated by allowing whole blood to clot at room temperature for 60 min, centrifuging at 1,300 g for 15 min at 4°C, and freezing aliquots at or below −70°C. Clinical chemistry analyses were performed by AniLytics, Inc. (Gaithersburg, MD), by standard procedures for the following analytes: alanine aminotransferase (ALT), lactate dehydrogenase (LDH), AST, sorbitol dehydrogenase (SDH), alkaline phosphatase (ALP), nonfasting glucose, and total cholesterol. Serum and plasma hemoglobin levels were measured by Nanodrop-1000 Spectrophotometer (v.3.8.1, ThermoFisher Scientific, Waltham, MA; Tolan et al. 2013).
Histopathology
The formalin-fixed liver specimens were embedded in paraffin. A 5-μm section of liver from each animal was stained with hematoxylin and eosin, Oil Red O for lipid accumulation, and Masson’s trichrome for collagen deposition. Livers from animals administered BB were also stained with periodic acid Schiff’s histochemical stain for glycogen accumulation. The tissues were evaluated by a pathologist certified by the American College of Veterinary Pathologists (author Molly Boyle). Tissues were scored for degree of pathology on a scale with the following distribution: none (score 0, 0% of tissue affected), minimal (score 1, >0–30% of tissue affected), mild (score 2, 30–60% of tissue affected), moderate (score 3, 60–80% of tissue affected), and marked (score 4, >80% of tissue affected).
Plasma Immunodepletion for iTRAQ (isobaric tag for relative and absolute quantification)
Plasma proteins were immunodepleted using the Agilent Multiple Affinity Removal System (MARS) for Human-14 column (Agilent Technologies, Inc., Richardson, TX). For the MARS depletion, approximately 90 μl of the sample was filtered through Agilent 0.22 μM spin filters (part number [PN] 5185-5990), and 60 μl of the filtered sample was diluted with 440 μl of Agilent Buffer A (Agilent, PN 5185-5987). Two hundred microliters of diluted sample was injected onto an Agilent HU-14 4.6 × 100 mm column (Agilent, PN 5188-5218) maintained at 23°C. An Agilent 1100 system was used to perform separation by high-performance liquid chromatography (HPLC), with sample well plate and fraction collector set to 8°C. A gradient was used for the separation (100% buffer A for the initial 15 min at 0.5 ml/min; 100% buffer B [Agilent PN 5185-5988] until 22 min at a flow rate of 1 ml/min; a gradient to 100% buffer A from 22.1 to 33 min at a flow rate of 1 ml/min). The 280 nm wavelength was monitored to ensure injection reproducibility. Starting at 2 min, the fraction collector collected fractions every 1 min, stopping at 18.9 min. Two injections were made for each sample; fractions 1 to 12 were pooled from both injections to create the flow-through fraction.
iTRAQ Analysis of Plasma and Liver Tissue
Agilent 5 kDa molecular weight cutoff filters were used to buffer exchange the depleted plasma samples. The pooled fractions were reduced to a volume less than 500 μl. Three mililiters of 10 mM tetraethylammonium bicarbonate was added, and the sample was centrifuged and again reduced to less than 500 μl. This procedure was repeated 2 more times. After the final centrifugation to 500 μl or less, the volume was adjusted to 500 μl. Protein concentration was determined by bicinchoninic acid assay (ThermoFisher Scientific, Rockford, IL [PN 23227]).
For iTRAQ labeling, 200 μg of the protein was dried by speed evaporator and processed according to the iTRAQ protocol using iTRAQ 8-plex reagents (Sciex; Redwood City, CA [PN 4390812]). Samples were dissolved in 40 μl of 50 mM tris-(2-carboxyethyl) phosphine dissolution buffer and reduced in 2% sodium dodecyl sulfate with 200 mM methyl methanesulfonate to block cysteines. Proteins were digested overnight with 10 μg trypsin and isotopically labeled according to manufacturer instructions. Isotopically labeled samples were combined and vacuum centrifuged for 15 min at 45°C to remove most of the isopropanol. Samples were diluted with strong cation exchange mobile phase (900 μl of 3% formic acid: 2 mM ammonium formate: 25% acetonitrile in water).
The samples were fractionated at a flow rate of 0.95 ml/min using a Polysulfoethyl A column (200 × 4.6 mm, 5 µm, 300 Å [PolyLC, Columbia, MD; PN 204SE0503]). The Agilent 1100 HPLC system described above was used to perform the separation with the column compartment set to 35°C. Sample injection volume was 1,600 μl with the following separation solvents: solvent A (3% formic acid in water), solvent B (500 mM ammonium formate and 3% formic acid in water), and solvent C (100% acetonitrile). The initial conditions were 73% A: 2% B: 25% C. A linear gradient was performed to 75% B: 25% C in 20 min, stopping at 35 min. Fractions were collected every 1 min from 3 min until 34.9 min for a total of 32 fractions. Samples were evaporated by vacuum centrifugation at 45°C. Fractions 9 to 20 were reconstituted for analysis with 85 μl of 0.1% formic acid in water and filtered through 0.22 μM Agilent spin filters for analysis.
Orbitrap Analysis
Peptides were separated by Thermo Proxeon easy nano-LC (Thermo Fisher Scientific) using Acclaim pepmap trapping columns (100 C18, 2 cm, 75 inner diameter µm, 3 µm 100 Å (Thermo Fisher Scientific, PN 164705) and an Acclaim pepmap analytical column (100 C18, 15 cm, 75 µm inner diameter, 3 µm particle size 100 Å [PN ES800]). The sample was added to 80 µl of 0.1% formic acid in water. The injection volume was 16 µl. A gradient of 0.1% formic acid in water (pump A) and 0.1% formic acid in acetonitrile (pump B) was initiated with 3% B, with 45% B at 90 min, 95% B at 100 min, 95% B at 105 min, and 1% B at 106 min. The analysis was terminated at 110 min.
An LTQ Orbitrap Velos (Thermo Fisher Scientific) was used to conduct parent ion scans using 60,000 resolution over the mass range of 400 to 1,800 m/z. The top 10 peptides were fragmented by high collision dissociation (settings: minimum signal threshold 500, resolution of 7,500, isolation width of 2 m/z, stepped collision energy at 40 and 50 V, default charge state of 2, and an activation time of 0.1 ms) with dynamic exclusion (repeat count = 1; repeat duration = 20 s; exclusion list ≤ 50 precursors). Exclusion was done relative to mass with an upper limit of 2.5 amu and a lower limit of 1.25 amu. A charge state of 1 was rejected. Because iTRAQ reagents interfered with the natural isotopic patterns of the peptides, the monoisotopic precursor selection was disabled. When fractions contained more than 100 identified peptides, mass exclusion lists containing the precursors for high confidence peptides were created, and fractions were rerun to improve coverage. Mass spectral data were processed using Proteome Discoverer v.2.1 (ThermoFisher Scientific). The peak lists were searched by the Sequest data analysis program against the RefSeq protein sequences in the rat database available from the National Center for Biotechnology Information (Retrieved August 3, 2015; fixed modifications: cysteine [metylthio] and any N-terminus or Lysine [iTRAQ 8-plex]; variable modifications: methionine [oxidation], serine/tyrosine/threonine [phosphorylation], tyrosine [iTRAQ 8-plex], and asparagine and glutamine [deamidation]). Proteome Discoverer processed the raw data with the following parameters: precursor ion mass tolerance = 10 ppm; fragment ion mass tolerance = 0.02 Da; enzyme = trypsin; maximum number of equal modifications per peptide = 3; maximum number of dynamic modifications per peptide = 4; maximum number of missed cleavages = 2. Only scans with a 1% false discover rate (FDR) based on posterior error probability as determined by the percolator node of Proteome Discoverer were used for protein identification. Exclusion lists were made from all peptides exceeding the 1% FDR threshold and the samples reinjected.
iTRAQ Data Analysis
Scaffold Q+ (version Scaffold 4.4.6, Proteome Software Inc., Portland, OR) was used to quantitate iTRAQ peptide and protein identifications. Thresholds for acceptable peptide identifications were >50.0% probability and acceptable protein identifications were >99.0% probability with ≥2 identified peptides by the Scaffold Local FDR algorithm with probabilities assigned by the Protein Prophet algorithm (Nesvizhskii et al. 2003). Proteins sharing significant peptide evidence were grouped into clusters, correcting channels as described in i-Tracker (Shadforth et al. 2005). Normalization was performed across all acquired peptide intensities within each assigned protein as described (Oberg et al. 2008). Proteins were considered differentially expressed if they were expressed at ±1.3-fold of control as described by Ahn et al. (2014) with p < .05 as determined by the permutation test with Benjamini–Hochberg p value correction.
Protein Set Enrichment Analysis and Pathway Analysis
Protein fold changes relative to controls and p values from Scaffold were imported into Ingenuity Pathway Analysis (IPA; v.9.0; Redwood City, CA) for analysis. Threshold parameters were set at ±1.3-fold change from control with a Benjamini–Hochberg adjusted p value < .05. Search terms relating to either lipid accumulation (steatosis) or collagen accumulation (fibrosis) were used to filter the data to determine the number of proteins associated with each pathology. p Values were calculated by Fisher’s exact test, right tailed. IPA software was used to generate comparison analysis heat maps to visualize trends and clusters of pathway scores among experimental observations. Heat maps were plotted using Fisher’s exact test p values for pathways and disease functions filtered for relevance to either lipid accumulation or fibrosis pathologies. Unfiltered data were used to conduct canonical pathways analyses in the IPA software to determine the number of proteins, which were significantly modulated relative to controls in each pathway. Statistical significance was determined using Fisher’s exact test, right tailed (p < .05).
Gene-protein Module Assessment and Adverse Outcome Pathway (AOP) Mapping by Transcriptomics
RNA was isolated from livers using a miRNeasy 96 kit (Qiagen, Valencia, CA). RNA samples were evaluated with a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA) using the Agilent RNA 6000 Nano Reagents and a multiwell NanoDrop 8000 Spectrophotometer (Thermo Fisher Scientific). Transcriptomics was performed using the AffyMetrix Rat Gene 2.1 ST 96 Array Plate and analyzed on the GeneTitan System (Affymetrix, Santa Clara, CA) as described (Ippolito et al. 2016). Microarray data have been deposited in the Gene Expression Omnibus database (accession number GSE70559). Significantly differentially expressed proteins were matched with corresponding messenger RNA (mRNA) transcripts and mapped to gene modules associated with fibrosis or steatosis identified in our previous study (Tawa et al. 2014). Functions ascribed to the differentially expressed protein products of genes within each module were assessed by manual data curation and IPA software. The “network interaction” feature of IPA software was used to determine proteins in gene/protein networks. Protein modules were mapped to molecular mechanisms associated with the steatosis (Mellor, Steinmetz, and Cronin 2015) or fibrosis pathology (Horvat et al. 2017).
Hierarchical Clustering
Normalized intensities of the differentially expressed proteins identified by Scaffold Q+ software were imported into Partek Genomics Suite (v.6.6, Partek, St. Louis, MO). Hierarchical clustering was performed by centroid clustering and shifting all columns to a mean of zero and scaling to a standard deviation of 1. Data were clustered using Euclidian distance measure of dissimilarity. Venn diagrams were created in the list manager to determine the number and identity of proteins differentially expressed in both plasma and tissue.
Results
To evaluate global protein changes at different times in disease progression, reference toxicants were administered in a single dose or once daily for 5 days at concentrations reported in the literature to induce steatosis or fibrosis. Protein profiles were linked by network analysis to gene modules of hepatotoxicity and published mechanisms of hepatotoxicity.
Steatogenesis
Histopathology and clinical chemistries after BB administration
BB administration resulted in a dose- and time-dependent increase in the frequency and severity of subacute inflammation and lipid accumulation relative to controls (Figure 1). Inflammation was characterized by disruption of hepatic architecture by random and periportal dispersions of mononuclear cells, neutrophils, multinucleated giant cells, and macrophages (Figure 1B). A single dose of BB at 0, 124, 229, 424, or 785 mg/kg resulted in minimal to mild inflammation in 0%, 57%, 88%, 86%, or 86% of animals (n = 7 per group), respectively. Mild to marked inflammation was observed in 0%, 0%, 20%, 60%, and 100% of animals (n = 5 per group) after 5 consecutive days of dosing at 0, 124, 229, 424, and 785 mg/kg, respectively (Online Supplemental Figure 1A). A single BB administration resulted in minimal to moderate periportal vacuolation in 0%, 43%, 100%, 86%, and 86% of animals at 0, 124, 229, 424, and 785 mg/kg, respectively. Specific morphological features of the cytoplasmic vacuolation were suggestive of intracytoplasmic lipid accumulation (compare Figure 1D). After 5 days, 0%, 80%, 80%, 80%, and 100% showed histological evidence of vacuolation (lipid accumulation) at 0, 124, 229, 424, and 785 mg/kg, respectively (Online Supplemental Figure 1B). Lipid accumulation was confirmed by Oil Red O histochemical staining (Figure 1E). There was no histological evidence of fibrosis at any dose level. There was no evidence for cytoplasmic accumulation of glycogen (e.g., periportal and bridging cellular swelling with accumulation of wispy cytoplasmic material suggestive of glycogen). Periodic acid Schiff’s histochemical stain was negative for glycogen accumulation.
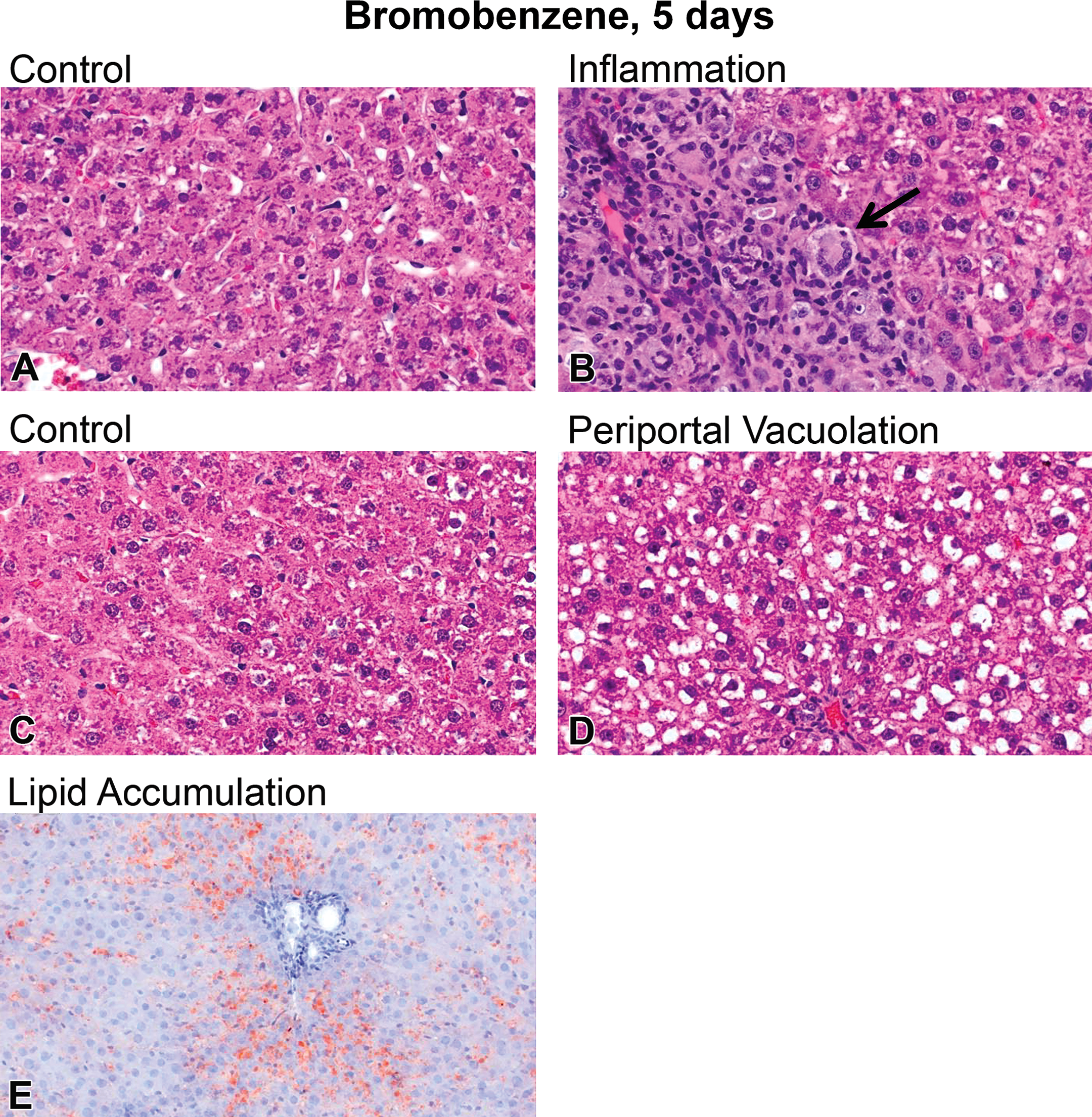
Histological evidence of liver inflammation and lipid accumulation after 5 days of bromobenzene (BB) administration in male Sprague-Dawley rats. Representative liver images from (A) control and (B) BB at 785 mg/kg. BB administration resulted in histological evidence of periportal inflammation (multinucleated giant cell denoted by black arrow). (C) control liver, (D) Histological evidence of vacuolation was observed after 785 mg/kg BB. (E) Oil Red O histochemical staining for lipid accumulation.
Changes in the levels of circulating liver enzyme were consistent with liver injury (Figure 2, Online Supplemental Table 2). The concentrations of cholesterol and the activity levels of the liver enzymes AST, alanine aminotransferase (ALT), ALP, LDH, and SDH were measured in serum after administration of BB at a single dose or after 5 consecutive days (Figures 2 and 3A). After a single dose, AST activity was significantly elevated at 229, 424, and 785 mg/kg (a 2.3-, 2.9-, and 1.7-fold increase, respectively, over control; Figure 2A). After exposure to 424 mg/kg BB, ALT was significantly elevated 2.0-fold over control, although exposure to 785 mg/kg resulted in ALT levels that were indistinguishable from controls (Figure 2B). There were no significant changes in ALP or cholesterol at any dose level (Figures 2C and 3A). Additionally, LDH levels were elevated at the 424 and 785 mg/kg dosing levels (Figure 2D). SDH levels were significantly elevated following 229, 424, and 785 mg/kg BB (a 11.2-, 13.3-, and 5.3-fold increase, respectively, over control; Figure 2E). Interindividual variability was high among BB-administered animals, with a trend toward increases with no statistically significant differences in liver enzymes after a single dose or 5 consecutive days. There was a trend toward elevated SDH after 5 days (p = .06 at 424 and 785 mg/kg; Figure 2E). At day 5, cholesterol levels were significantly elevated at 424 and 785 mg/kg (a 1.5- and 1.7-fold increase, respectively, over control; Figure 3A). No significant changes were noted following a single administration. Cholesterol levels were significantly elevated above control after BB administration (Figure 3A). Following a single exposure to 229, 424, or 785 mg/kg BB, nonfasting serum glucose was significantly and dose dependently reduced relative to vehicle control (1.4-, 1.5-, and 1.6-fold, respectively; Figure 3B). A similar dose-dependent effect was seen after 5 days, with nonfasting serum glucose was lower than vehicle control by 1.6- and 1.5-fold after 785 and 424 mg/kg BB, respectively (Figure 3B).
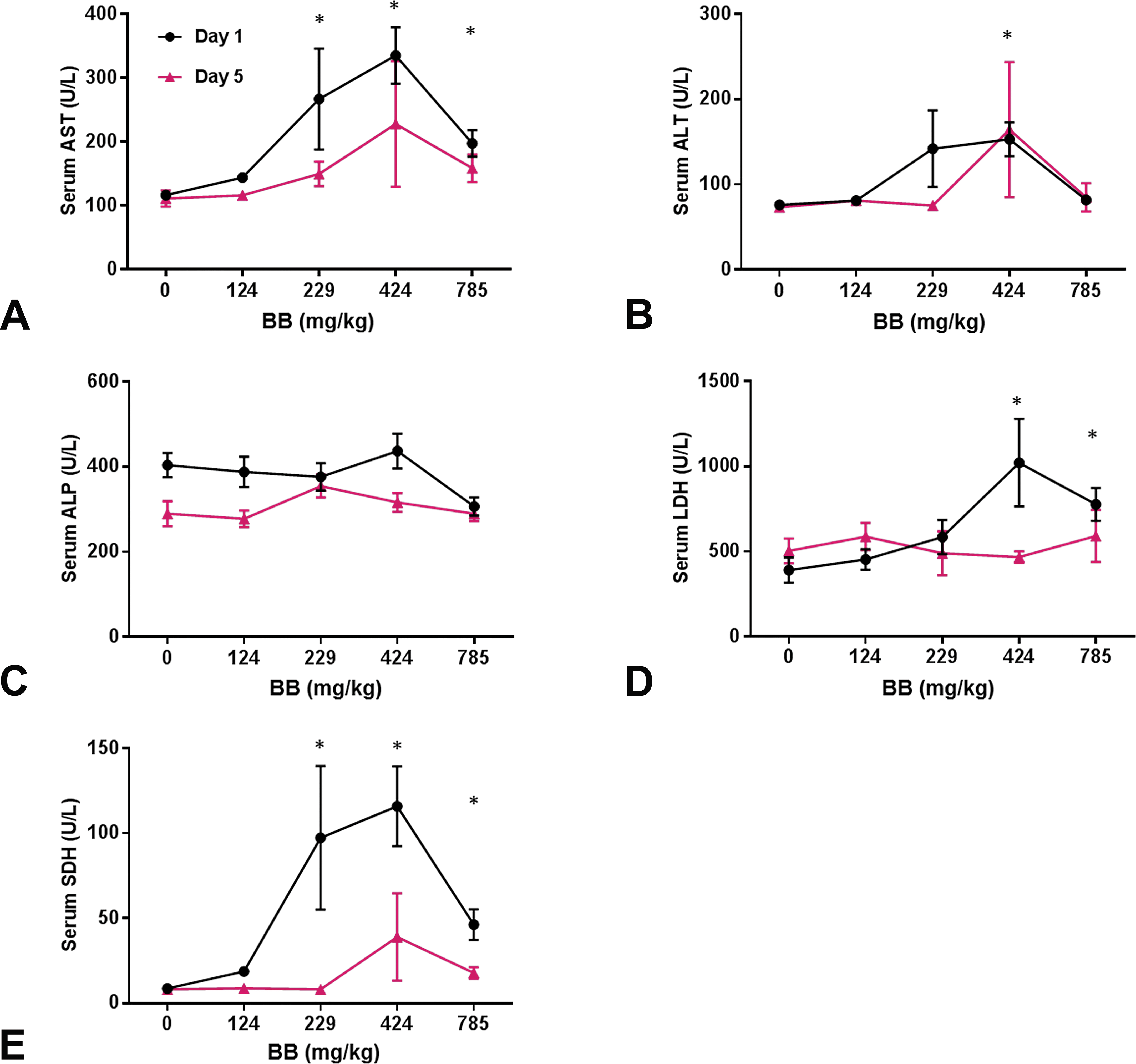
Modulation of serum clinical chemistries after bromobenzene (BB) administration. Serum (A) aspartate aminotransferase (AST), (B) ALT, (C) alkaline phosphatase (ALP), (D) lactate dehydrogenase (LDH), and (E) sorbitol dehydrogenase (SDH) after a single dose (black) or 5 consecutive days (red) of BB (doses in mg/kg). *p < .05 by analysis of variance with Bonferroni correction; error bars represent standard error of the mean.
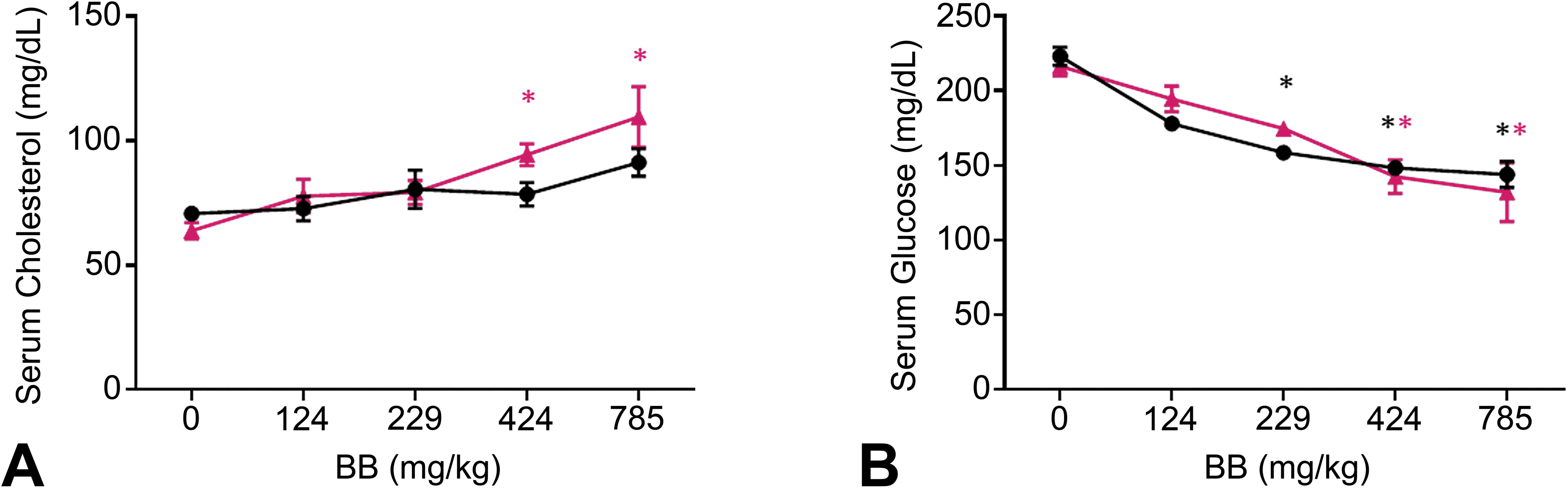
Modulation of serum cholesterol and glucose after bromobenzene (BB) administration. Serum (A) cholesterol and (B) glucose concentrations after a single dose (black) or 5 consecutive days (red) of BB (doses in mg/kg). *p < .05 by analysis of variance with Bonferroni correction; error bars represent standard error of the mean.
In 1 animal after a single BB administration at 424 mg/kg, there was a histopathological observation of hepatocellular necrosis. Moderate necrosis was observed in another animal after BB administration. Several clinical chemistry values (LDH, AST, and ALT) for this animal were greater than 2 standard deviations from the mean and were removed from the analysis (Online Supplemental Table 2). Additionally, the serum from this animal was orange colored and had a high hemoglobin content (54.4 ± 1.7 mg/dl), suggesting hemolysis. Collectively, the serum chemistries and histopathology suggested circulatory failure for this animal. This animal was excluded from further analysis.
Global analysis of differentially expressed proteins after BB administration
To correlate protein expression data with coregulated gene modules identified in our previous studies (Ippolito et al. 2016; Tawa et al. 2014), we identified differentially expressed proteins in liver tissue by conducting a global quantitative proteomics analysis using iTRAQ labeling reagents. For proteomics analysis, we compared proteins isolated from livers from rats exposed to vehicle and 424 mg/kg BB for 1 or 5 days and their vehicle-only controls (n = 4 rats per dose group with n = 4 controls per 8-plex iTRAQ experiment). This dose was the lowest BB dose with evidence of significant pathology and significant deviations from control in clinical chemistries (Figures 1 –3; Online Supplemental Tables 1 and 2).
In liver tissue, global proteomics analysis with iTRAQ labeling identified 81 (of 1,416 total identified) proteins differentially expressed in liver tissue after a single BB administration and 324 (of 1,644 total identified) proteins after 5 days of consecutive doses. We performed hierarchical biclustering analysis of the differentially expressed proteins. The BB animals clustered together away from vehicle-treated controls (Figure 4A). We used IPA to determine the cellular and toxicological functions enriched in the list of differentially expressed proteins (Figure 4B). Pathways associated with lipid accumulation, carbohydrate metabolism, and steatosis were significantly overrepresented in the list of proteins (p < .05). All were also significant at day 5 than after a single administration (Figure 4B). More proteins were associated with lipid metabolism after 5 days than after a single dose, with lower p values at day 5 than day 1 (Figure 4C) as expected. IPA was used to map differentially expressed liver tissue proteins to canonical pathways. Compared with a single dose, more proteins representing each canonical pathway were enriched after 5 days (Figure 4D). Many of the top enriched canonical pathways contain biomolecules that have been previously identified as mechanistically linked to the steatosis pathology, including liver X receptor (LXR), pregnane X receptor (PXR), and peroxisome proliferator-activated receptor (PPAR) dimerization with retinoid X receptor (RXR; Mellor, Steinmetz, and Cronin 2015; Figure 4D). Following a single administration of 424 mg/kg BB, the most enriched canonical pathways were oxidative stress responses (nuclear factor erythroid 2-related factor 2 [NRF2]) and RXR-associated nuclear receptor activation events as well as regulators of extracellular matrix (ECM) interactions (integrin-linked protein kinase [ILK] signaling; Figure 4D). After a single dose, 4 proteins were identified in each of the RXR-related pathways, 2 of which (APOA1 and FASN) were present in all 3 pathways. The RXR nuclear receptor activation is a molecular initiating event in steatosis development (Vinken 2013, 2015; Willett et al. 2014). After 5 days of BB administration, oxidative stress, and nuclear RXR-associated receptor signaling, canonical pathways remained significantly enriched (Figure 4D).
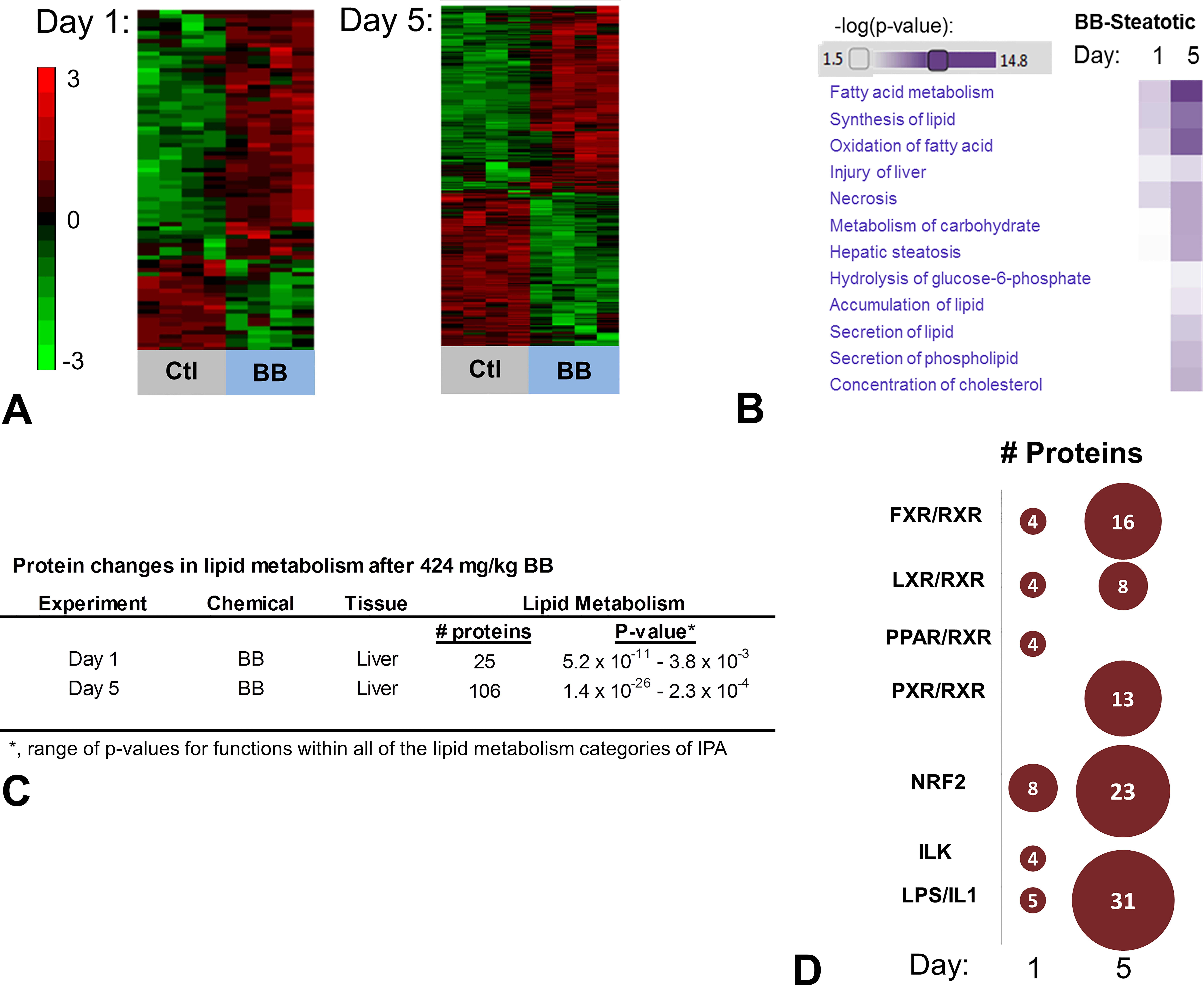
Alteration in protein expressions linked to cellular functions in liver tissue after a single administration of bromobenzene (BB) or 5 consecutive doses. (A) Hierarchical biclustering of proteins in rat liver tissue after 1 or 5 consecutive days of 424 mg/kg BB methylenedianiline administration. Each column represents a single animal. Differential abundances of iTRAQ-labeled proteins between control and exposed animals (FDR < .05) were imported into Partek Genomics Suite and normalized for clustering analysis (n = 4 animals per group; red, greater than mean; green, less than mean; see also Online Supplemental Figure 3 for clustering dendrograms). (B) Hierarchical clustering by p value identified cellular and disease functions differentially expressed at days 1 or 5 after 424 mg/kg BB. (C) Cellular and toxicity functions related to fatty acid metabolism are increased at 5 days relative to a single administration of BB. (D) Bubble plots indicate the top signaling pathways at 1 or 5 days. Size of the circles is relative to protein number.
BB-induced protein changes in the context of the protein modules
In our previous work (Ippolito et al. 2016; Tawa et al. 2014), we used an iterative signature algorithm to identify gene modules from Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways by hierarchical clustering associated with steatosis using data from the DrugMatrix gene expression database. We compared the genes associated with the protein modules from the current study with those derived from the in silico DrugMatrix study and determined which gene modules were enriched by performing a Fisher’s exact test (Figure 5A). We identified 3 enriched modules after a single administration of BB and 16 enriched modules after 5 days of consecutive doses. All 3 of the enriched modules at day 1 were also enriched at day 5 (Figure 5A). Most of the modules identified in this analysis for day 5 had previously been linked to either a steatosis or fibrosis phenotype (Figure 5B).
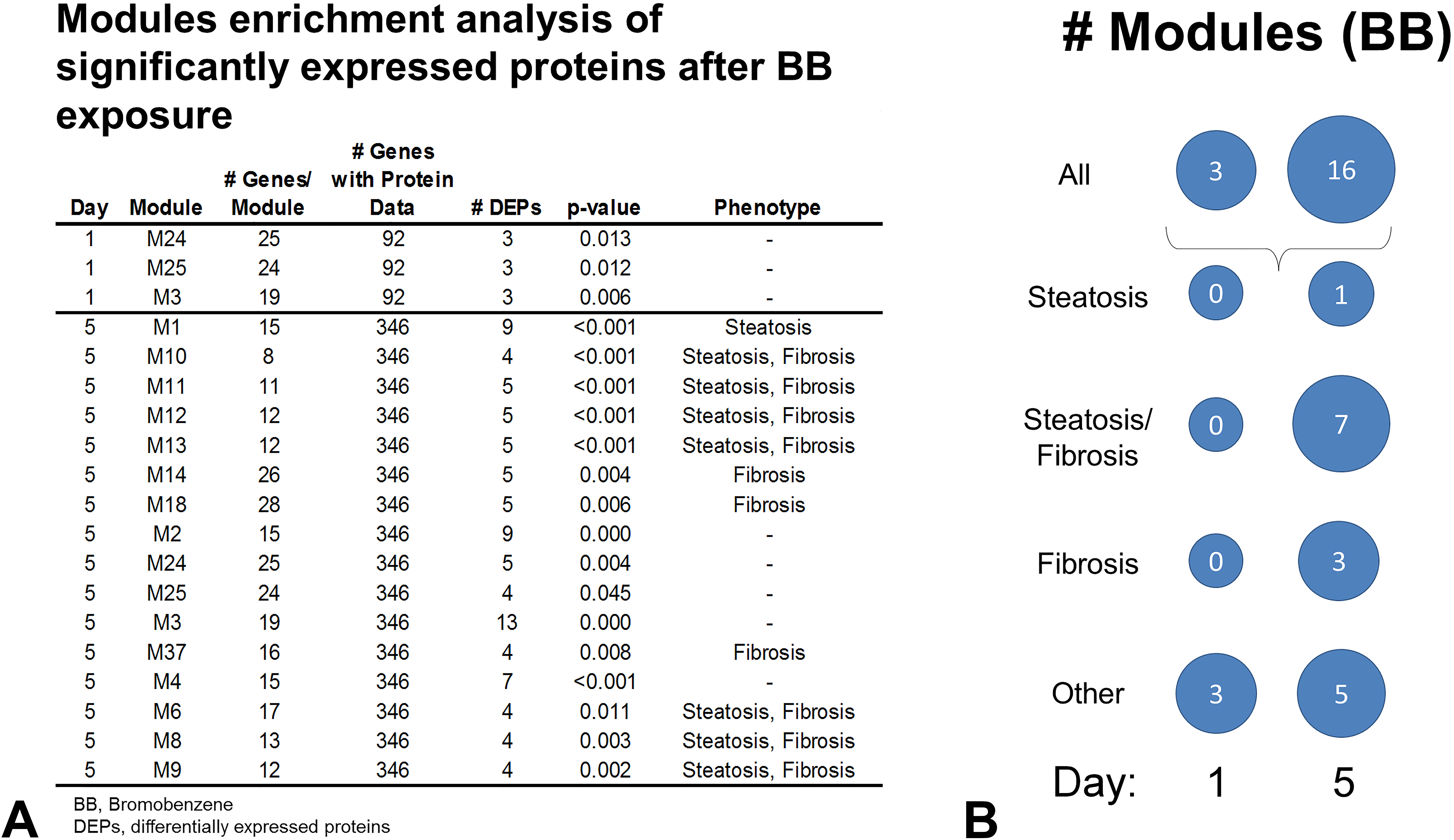
Differentially expressed protein modules in liver tissue after a single administration of bromobenzene or 5 consecutive doses. (A) Protein products of differentially expressed gene modules from Tawa et al. (2014) were identified in iTRAQ global proteomics data. (B) Modules at 1 or 5 days were mapped to steatogenic phenotypes.
Global proteomics analysis of plasma proteins after BB: Comparison with liver tissue
We performed global proteomics analysis to identify differentially expressed plasma proteins after a single administration of BB or 5 consecutive daily doses (Figure 6A). There were 37 (of 172) and 34 (of 188) differentially expressed proteins at 1 and 5 days, respectively. Most of the proteins identified were acute phase inflammatory proteins previously associated with the steatosis and/or fibrosis phenotype, including apolipoproteins, haptoglobin, and fibrinogen chains (Figure 6A, orange, bold text; see also Online Supplemental Table 4 for protein names). Proteins with known metabolic associations with lipid and carbohydrate functions that were enriched after 424 mg/kg administration included UGP2, SORD, APOA1, and APOB. UGP2 showed one of the most significant increases relative to controls after 424 mg/kg BB administered for either a single dose or 5 consecutive days (Figure 6A; 1.9 ± 0.4 and 1.5 ± 0.8 log2 fold change relative to control, respectively; p < .05 Benjamini–Hochberg FDR (Reiner, Yekutieli, and Benjamini 2003). UGP2 is a member of modules 3 and 4, which are both significantly enriched after 1 and 5 days of BB administration (see Figure 5). In plasma, UGP2 increased relative to controls after 424 mg/kg BB administered for either a single dose or 5 consecutive days (1.9 ± 0.4 and 1.5 ± 0.8 log2 fold change relative to control, respectively; p < .05 Benjamini–Hochberg FDR (Reiner, Yekutieli, and Benjamini 2003). In liver tissue, UDP-glucose-6-dehydrogenase (UGDH) levels increased relative to controls after 424 mg/kg BB administered for either a single dose or 5 consecutive doses (0.5 ± 0.2 and 1.3 ± 0.1 log2 fold change relative to control, respectively; p < .05 Benjamini–Hochberg FDR; Reiner, Yekutieli, and Benjamini 2003; see Online Supplemental Figure 4).
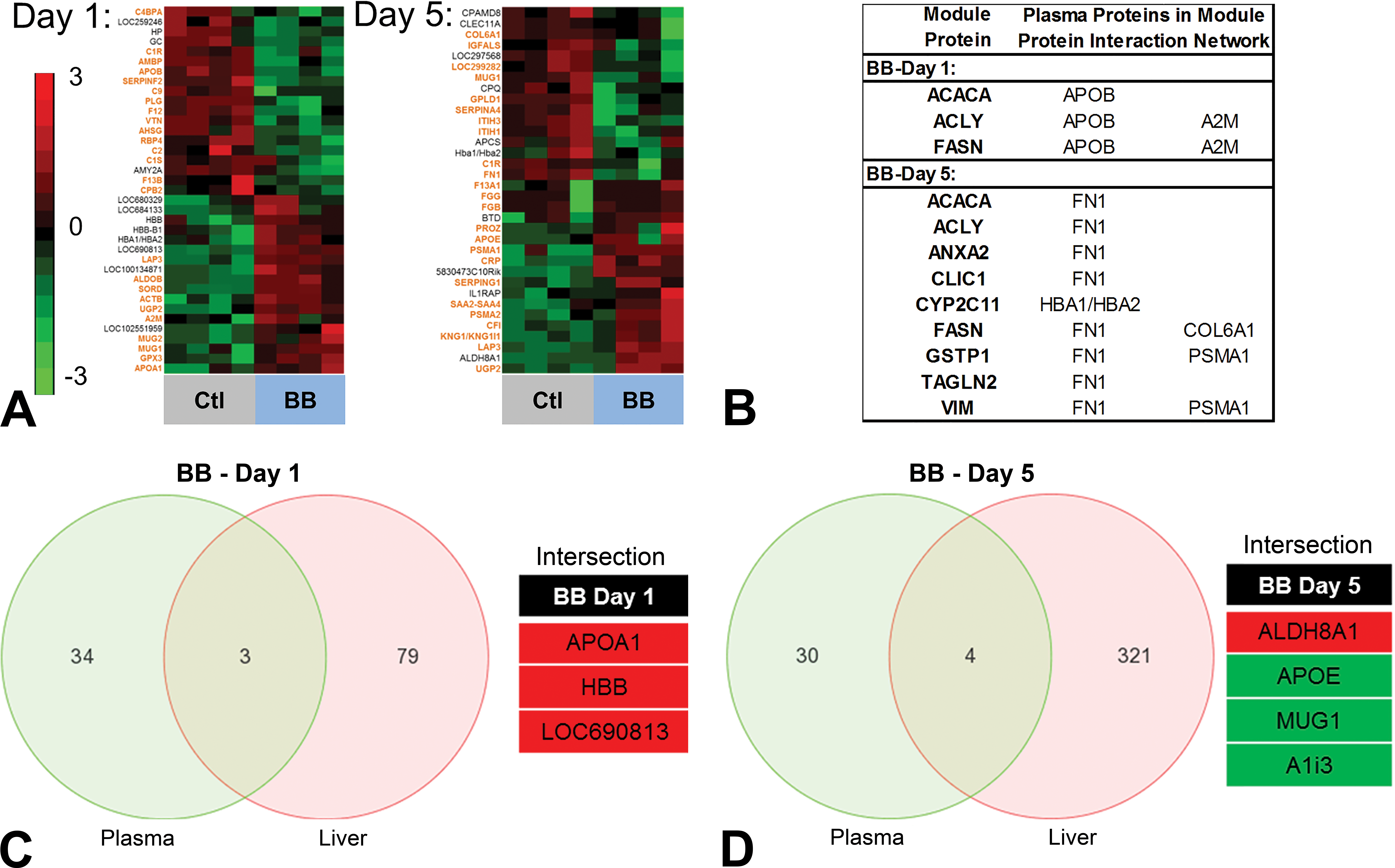
Comparison of differentially expressed plasma and liver proteins from rats administered bromobenzene (BB). (A) Plasma proteins from animals exposed to 4,4′-methylenedianiline for a single administration or 5 consecutive doses were quantified by iTRAQ reagent labeling and mass spectrometry analysis. Each column represents a single animal. Relative abundances (FDR < .05) were imported into Partek Genomics Suite and normalized for hierarchical biclustering analysis (n = 4 animals per group; red, greater than mean; green, less than mean; see also Online Supplemental Figure 3 for clustering dendrograms). Each column represents a single animal. Bold orange text indicates proteins implicated in steatosis and/or fibrosis mechanisms (see also Online Supplemental Table 4 for protein names). (B) Identification of plasma proteins in module gene/protein interaction networks. Ingenuity pathway analysis was used to identify all interaction networks for each module protein (see Figure 5), and the union of interaction proteins was used to identify plasma proteins in the interaction network of proteins expressed in liver tissue. Significantly differentiated proteins were identified in liver tissue and plasma from rats administered BB for a single dose (C) or 5 consecutive days (D). Venn diagrams were generated with Partek Genomics Suite. The intersection proteins are identified in the tables to the right of each diagram. Significance was determined by Permutation test with Benjamini–Hochberg corrected p value as well as a 1.3-fold change cutoff. Red indicates greater than mean, and green indicates less than mean.
To determine whether any of the differentially regulated proteins in liver tissue could also be detected in plasma, we compared the global proteomics results in both plasma and liver after BB administration for a single dose or 5 consecutive doses (Figure 6A and B). After a single dose or 5 consecutive doses of BB, 19 of 37 (51%) and 12 of 34 (35%) differentially expressed proteins in the plasma were acute phase response proteins, respectively. Proteins associated with carbohydrate metabolism were differentially abundant in both liver tissue and plasma. After a single administration of BB, 3 proteins were differentially abundant in both plasma and liver tissue: APOA1, HBB, and LOC690813 (Figure 5C, intersection table and Online Supplemental Table 5). After 5 days, 4 different proteins were differentially abundant in both plasma and tissue: ALDH8A1, APOE, A1I3, and MUG1 (Figure 5D, intersection table and Online Supplemental Table 5).
To further characterize the dose- and time dependencies of the BB effects, we determined the fold changes of the mRNA transcripts (as measured by Affymetrix gene array in our previous study; Ippolito et al. 2016) for the proteins differentially expressed in both plasma and liver tissue (Online Supplemental Table 6). For BB, gene transcripts for 5 of the 6 proteins were available for analysis; a gene probe was not present on the microarray for the transcript encoding LOC690813. A single administration of BB resulted in a dose-dependent increase in ApoA1 and a nondose dependent elevation in Aldh8a1 and Mug1 (Online Supplemental Table 6, upper left quadrant). After 5 days, all 5 transcripts were dose dependently elevated relative to controls (Online Supplemental Table 6, upper right quadrant).
Fibrogenesis
Histopathology and clinical chemistries after 4,4′-MDA administration
4,4′-MDA administration resulted in a dose- and time-dependent increase in bile duct hyperplasia, inflammation, and periportal fibrosis (Figure 7). Bile duct hyperplasia was characterized by increased numbers of small bile ducts emerging in portal areas relative to controls (Figure 7B). Biliary epithelium formed normal ducts and was well differentiated. Bile duct hyperplasia was frequently associated with subacute inflammation, fibrosis, and oval cell hyperplasia. After a single dose of 4,4′-MDA, minimal to moderate bile duct hyperplasia was observed in 0%, 0%, 60%, and 80% of animals (n = 5 per group) administered 4,4′-MDA at 0, 22, 60, and 162 mg/kg, respectively. Minimal and moderate to severe bile duct hyperplasia was observed in 0%, 20%, 100%, and 100% of animals (n = 5 per group) after 5 days of 4,4′-MDA administration at 0, 22, 60, and 162 mg/kg, respectively (Online Supplemental Figure 2A). Inflammation was observed after 4,4′-MDA administration, as characterized by the disruption of hepatic architecture by predominantly periportal aggregates of neutrophils, mononuclear cells, and macrophages (compare Figure 7D). After a single dose of 4,4′-MDA, minimal to mild subacute inflammation was observed in 0%, 0%, 60%, and 80% of animals dosed at 0, 22, 60, and 162 mg/kg, respectively. After 5 days, 0%, 0%, 100%, and 100% of the livers showed evidence of mild to moderate subacute inflammation after dosing with 4,4′-MDA at 0, 22, 60, and 162 mg/kg, respectively (Online Figure 2B). Periportal fibrosis was not observed in any of the dose groups after a single dose but was evident after 5 consecutive days of dosing, as determined by Masson’s trichrome histochemical staining. Periportal fibrosis was characterized by increased thickness of the periportal collagenous matrices (Figure 7F). By 5 consecutive days of 4,4′-MDA administration, minimal to mild fibrosis was evident in 0%, 0%, 100%, and 100% of rats dosed with 0, 22, 60, and 162 mg/kg, respectively (Online Supplemental Figure 2C). Minimal to moderate cytoplasmic alteration, characterized by increased cytoplasmic eosinophilia and granularity periportal to the bridging areas, was observed in 0%, 0%, 20%, and 20% of animals following a single administration of 0, 22, 60, and 162 mg/kg 4,4′-MDA, respectively. After 5 consecutive doses of 0, 22, 60, and 162 mg/kg 4,4′-MDA, minimal to moderate alteration was observed in 0%, 0%, 100%, 100% of the livers, respectively (Online Supplemental Table 8).
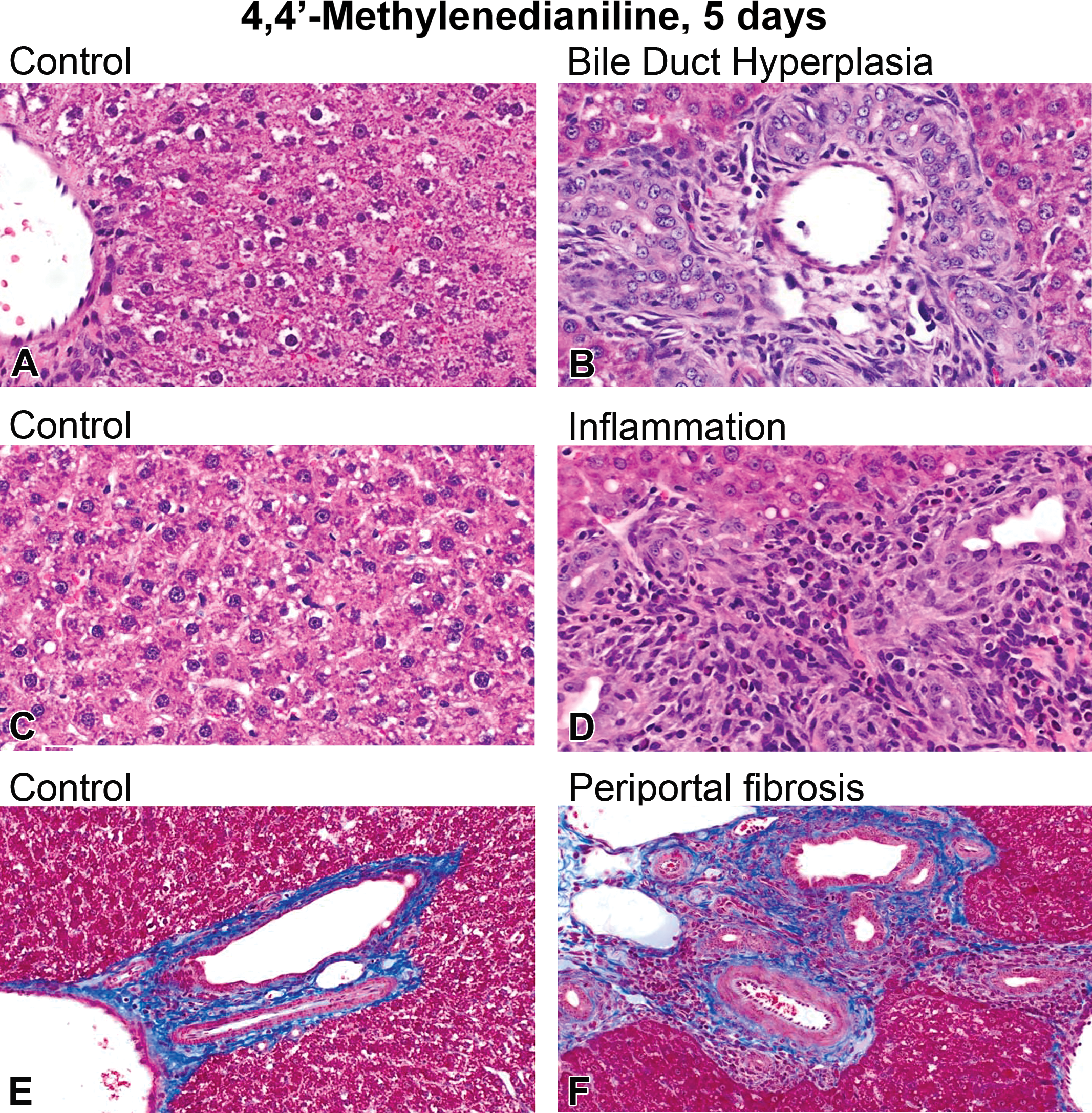
Histological evidence of hepatic bile duct hyperplasia, inflammation, and periportal fibrosis after 5 days of 4,4′-methylenedianiline (4,4′-MDA) administration in male Sprague-Dawley rats. Representative images from livers of (A) control and (B) 4,4′-MDA at 162 mg/kg. Histological evidence of bile duct hyperplasia was apparent after 4,4′-MDA administration. Representative images from (C) controls and (D) 4,4′-MDA at 162 mg/kg showed histological evidence of inflammation after 4,4′-MDA. Representative images from (E) control and (F) 4,4′-MDA at 162 mg/kg showed histological evidence of periportal fibrosis by Masson’s trichrome histochemical staining.
Changes in liver enzymes suggested the development of liver injury. On day 1, 4,4′-MDA administration resulted in a trend toward elevated liver enzymes and cholesterol, which was statistically insignificant at all doses due to interindividual variability among animals (Figure 8; Online Supplemental Table 1). By day 5, 4,4′-MDA administration resulted in a dose-dependent, statistically significant elevation over control in AST, ALT, ALP, and SDH (Figure 8A–E). For 59.7 mg/kg 4,4′-MDA, the average AST level increased 3.2-fold over control (Figure 8A), and the mean ALT level increased 3.9-fold (Figure 8B). At 162 mg/kg, mean AST, ALT, and ALP increased over control by 4.0-, 3.1-, and 1.5-fold, respectively (Figure 8A–C). Cholesterol levels were significantly elevated above control after 162 mg/kg 4,4′-MDA (2.3-fold over control) and were higher in 4,4′-MDA-treated animals than in BB-administered animals (Figure 9A). Rats administered 4,4′-MDA showed no evidence of a change in serum glucose after a single dose but did show a significant reduction (1.2-fold) in serum glucose after 5 days administration at 162 mg/kg (Figure 9B).
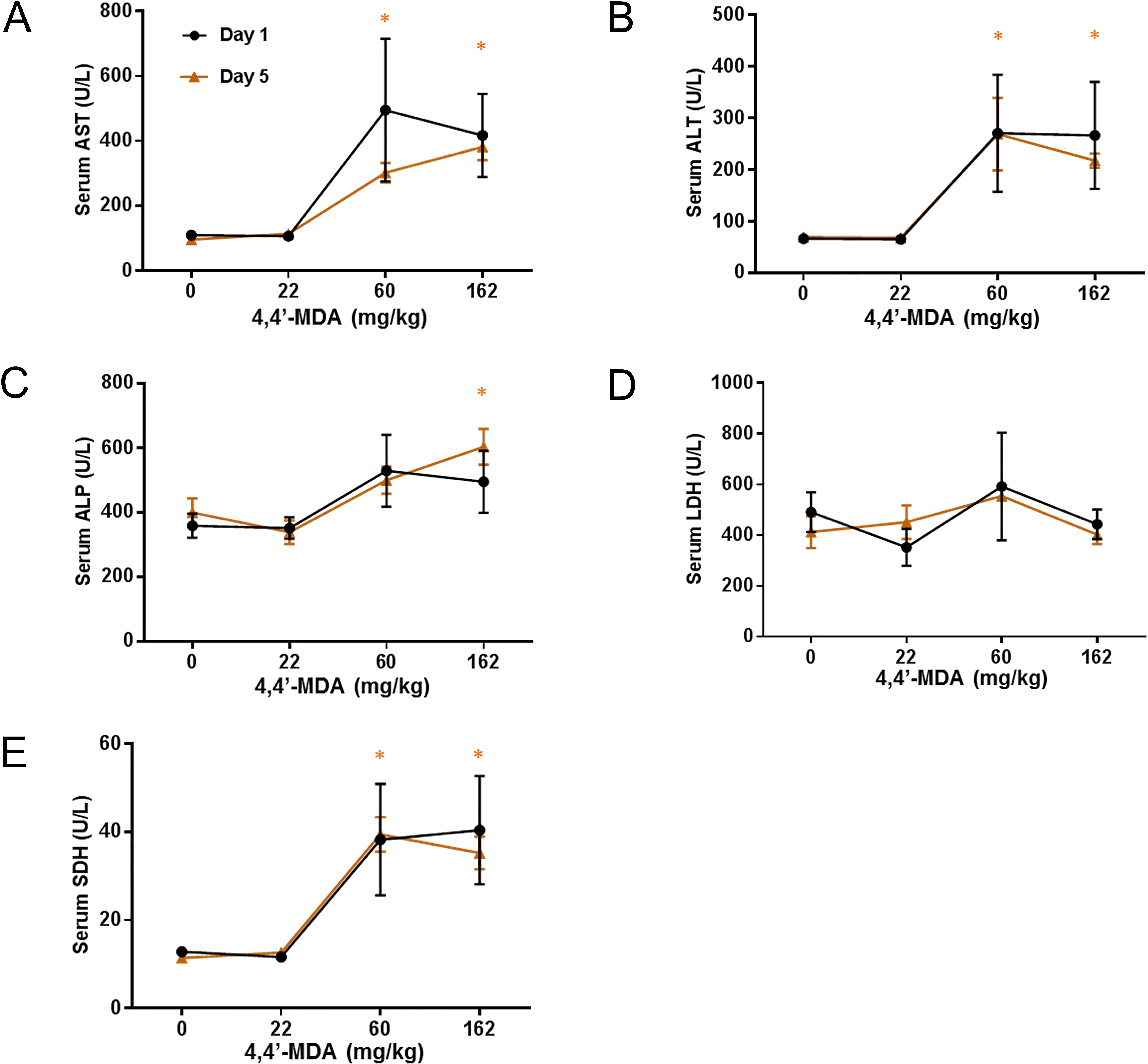
Modulation of serum clinical chemistries after 5 days of 4,4′-methylenedianiline (4,4′-MDA) administration. Serum (A) aspartate aminotransferase (AST), (B) ALT, (C) alkaline phosphatase (ALP), (D) lactate dehydrogenase (LDH), and (E) sorbitol dehydrogenase (SDH) concentrations after a single dose or 5 consecutive doses of 4,4′-MDA (doses in mg/kg). *p < .05 by analysis of variance with Bonferroni correction; error bars represent standard error of the mean.
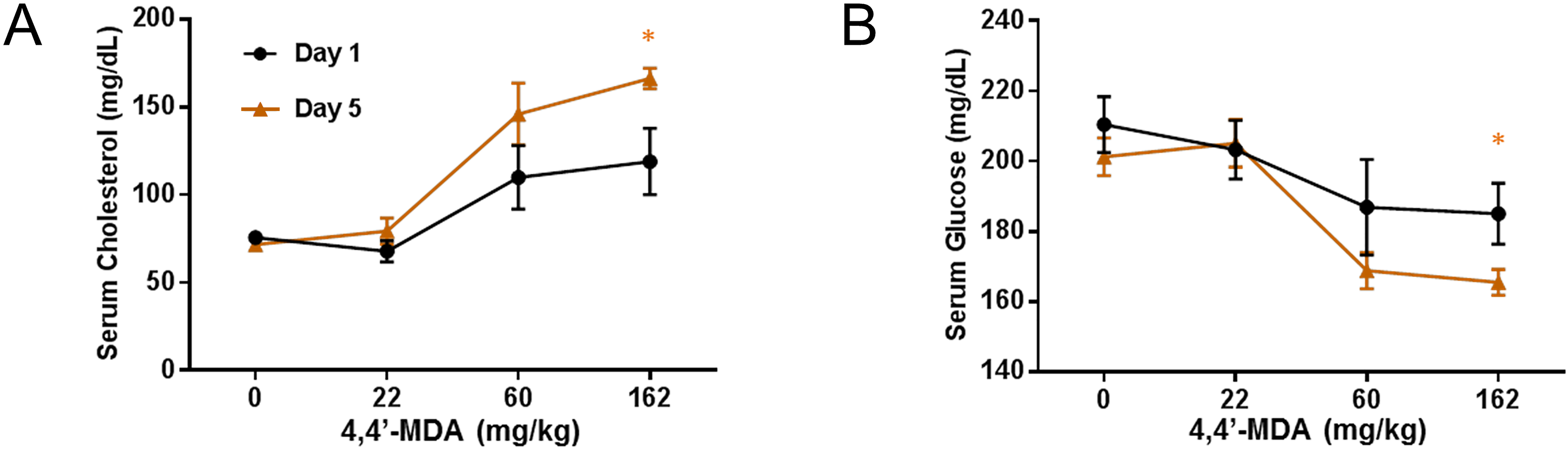
Modulation of serum cholesterol or glucose after 5 days of 4,4′-methylenedianiline (4,4′-MDA) administration. Serum (A) cholesterol or (B) glucose concentrations after a single dose or 5 consecutive doses of 4,4′-MDA (doses in mg/kg). *p < .05 by analysis of variance with Bonferroni correction; error bars represent standard error of the mean.
Fibrosis signatures identified by global liver tissue proteomic analysis after 4,4′-MDA administration
To determine the time course of protein expression after fibrogenic 4,4′-MDA administration, we conducted an unbiased, global proteomics analysis of liver tissue (Figure 10). As with the BB analyses above, we selected the lowest dose of 4,4′-MDA with evidence of pathology (histology and clinical chemistry) for the global analyses (i.e., the 162 mg/kg 4,4′-MDA dose; n = 4 rats per dose group with n = 4 controls per 8-plex iTRAQ experiment).
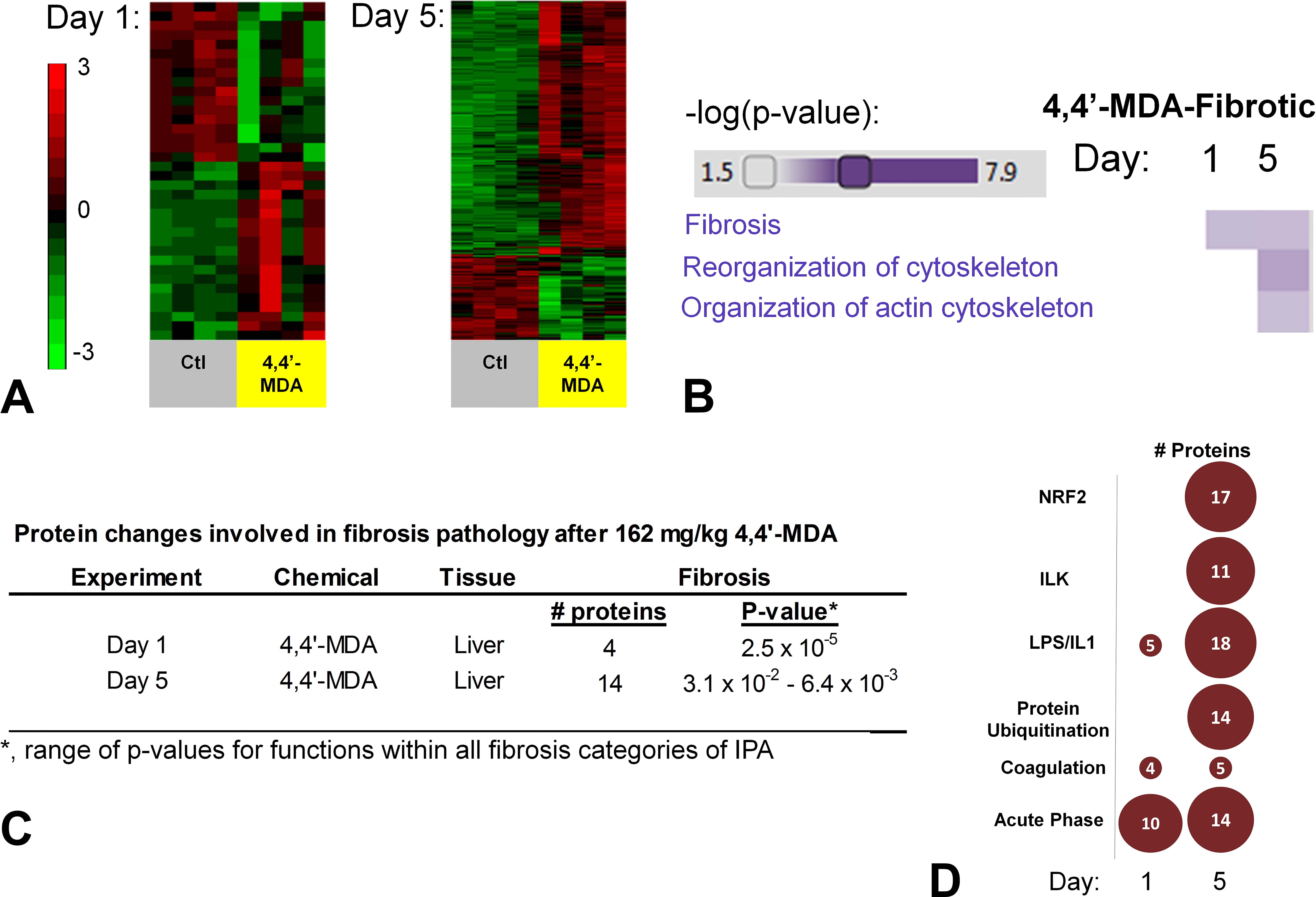
Increased expression of proteins in liver tissue associated with fibrogenic end points after 5 days of consecutive treatment with 4,4′-methylenedianiline (4,4′-MDA). (A) Hierarchical clustering of differentially expressed proteins (FDR < .05) in rat liver tissue after 1 or 5 administrations of 162 mg/kg 4,4′-MDA. Each column represents a single animal. Relative abundances of iTRAQ-labeled proteins (FDR < .05) were imported into Partek Genomics Suite and normalized for clustering analysis (n = 4 animals per group; red, greater than mean; green, less than mean; see also Online Supplemental Figure 5 for clustering dendrograms). (B) Enriched cellular functions in liver tissue after a single administration or 5 consecutive doses of 4,4′-MDA. (C) Number of differentially expressed proteins in the canonical fibrosis pathway in Ingenuity Pathway Analysis (IPA). (D) Top canonical pathways identified in IPA after MDA after 1 or 5 days.
In liver tissue, 36 (of 1,156 total identified) differentially expressed proteins were identified after a single administration (Figure 10A and B) and 235 (of 1,267) differentially expressed proteins were identified after 5 consecutive doses of 4,4′-MDA administration (Figure 10C). As with BB administration, hierarchical clustering identified a distinct patterns of protein expression in livers from rats administered 4,4′-MDA relative to their control counterparts (Figure 10A and B). Using IPA, we conducted functional analysis and identified significant enrichment of pathways and cellular functions associated with ECM reorganization and fibrosis phenotypes (Figure 10B and C). More proteins were significantly associated with the fibrosis pathology (as defined in [Ippolito et al. 2016; Tawa et al. 2014]) after 5 days than 1 day of 4,4′-MDA administration (Figure 10C). After a single administration of 4,4′-MDA, canonical pathways associated with inflammatory signaling were enriched (acute phase response signaling, LPS/IL1 (Lipopolysaccharide/interleukin 1) signaling, and the coagulation system; Figure 10D). After 5 days, canonical pathway analysis suggested predominantly the enrichment of oxidative stress, acute phase, and protein degradation pathways (Figure 10D). The number of proteins representing each canonical pathway increased after 5 days relative to a single administration.
Protein modules associated with fibrosis pathology after 4,4′-MDA administration
To determine whether the proteins identified by global proteomics mapped to expression modules identified in our previous research (Tawa et al. 2014), we mapped the differentially expressed proteins to their mRNAs in the liver injury modules (Figure 11A). After a single administration of 4,4′-MDA, there were 4 enriched modules; all 4 were associated with the fibrosis and/or steatosis phenotypes. The number of modules with significant liver expression proteins increased to 12 after 5 consecutive days of 4,4′-MDA administration (Figure 11A). Of these, 7 of the 12 were associated with the steatosis and/or fibrosis phenotypes (Figure 11B).
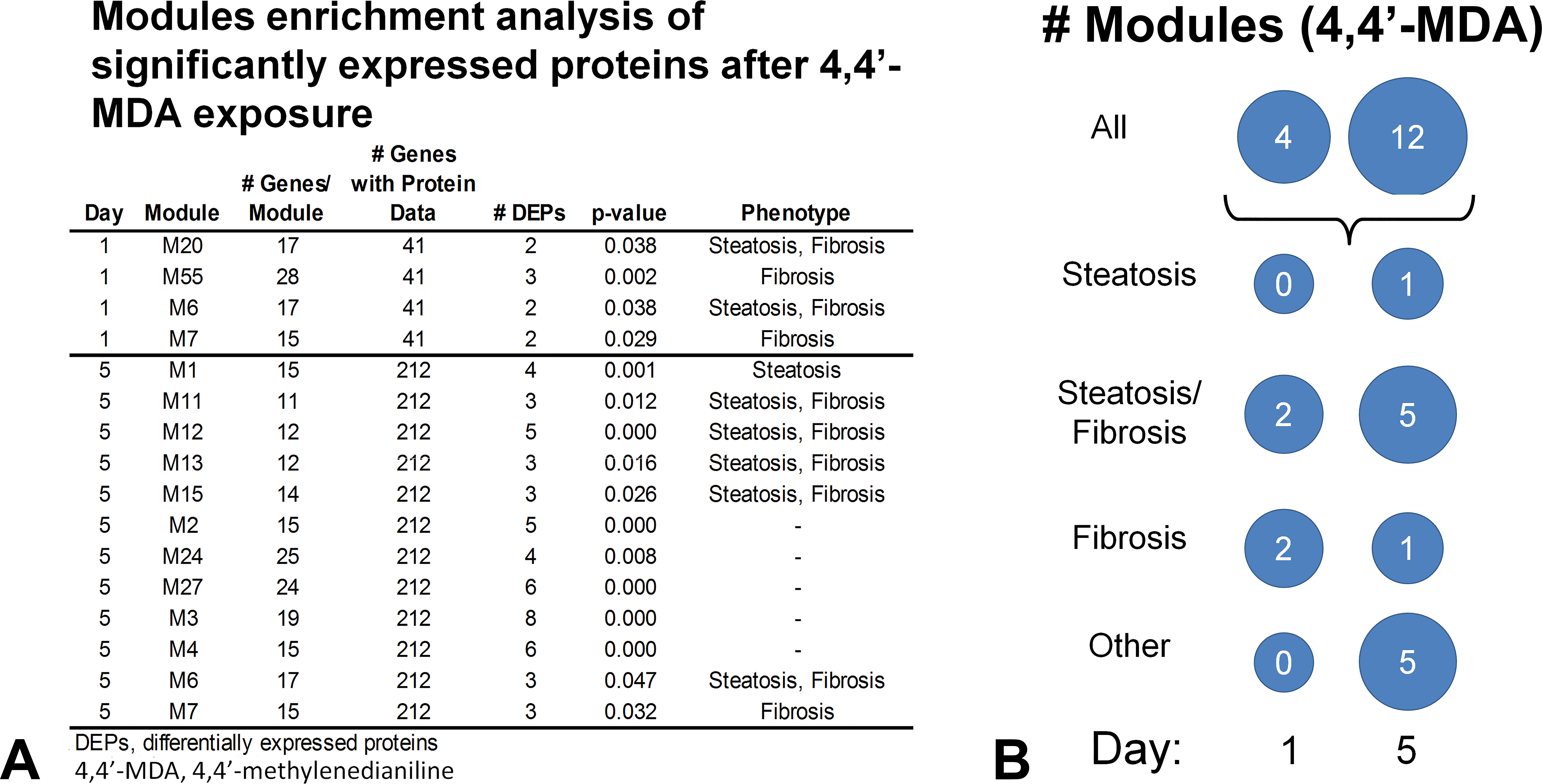
Differentially expressed protein modules in liver tissue after a single administration of 4,4′-methylenedianiline (4,4′-MDA; 162 mg/kg) or 5 consecutive doses. (A) Protein products of differentially expressed gene modules from Tawa et al. (2014) were identified in iTRAQ global proteomics data. (B) Modules at 1 or 5 days were mapped to fibrogenic phenotypes.
Differentially expressed plasma proteins after 4,4′-MDA: Comparison with tissue protein expression
A single administration or 5 consecutive doses of 4,4′-MDA resulted in 30 (of 137) and 66 (of 120) differentially expressed proteins in plasma, respectively (Figure 12A and B). Many of these proteins were associated with either the steatosis or fibrosis phenotypes (see bold orange text in Figure 12A; see also Online Supplemental Table 4 for protein names). A single administration of 4,4′-MDA resulted in differential regulation of 7 proteins in both plasma and tissue: FGA, FGB, FGG, HBB, HP, ITIH4, and KNG1/KNG1|1 (Figure 12C, intersection table, and Online Supplemental Table 5).
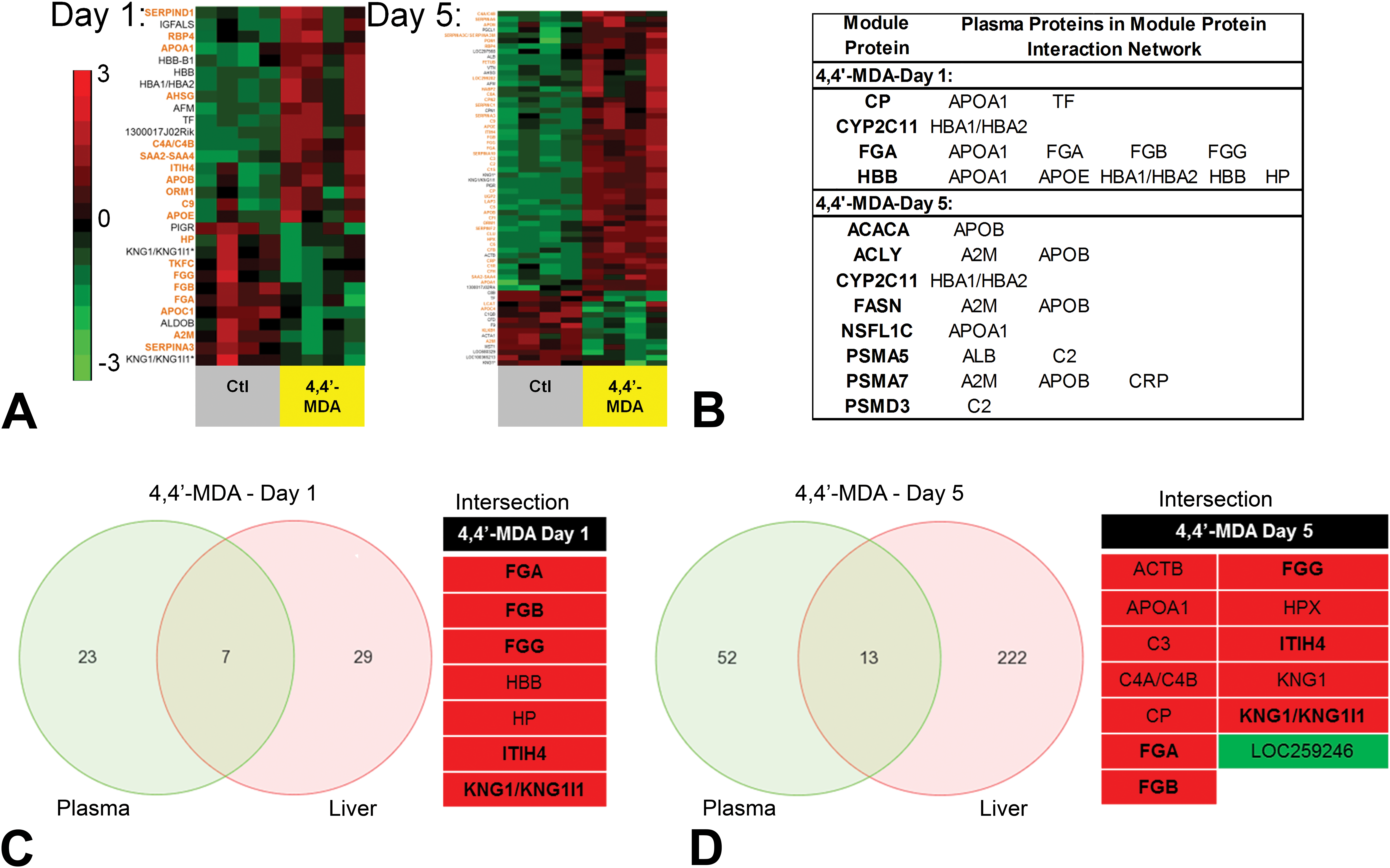
Comparing differentially expressed plasma and liver proteins from rats administered 4,4′-methylenedianiline (4,4′-MDA). (A) Plasma proteins from animals exposed to 4,4′-MDA for a single administration or 5 consecutive doses were quantified by iTRAQ reagent labeling and mass spectrometry analysis. Relative abundances (FDR < .05) were imported into Partek Genomics Suite and normalized for clustering analysis (n = 4 animals per group; red, greater than the mean; green, less than the mean; see also Online Supplemental Figure 5 for clustering dendrograms). Bold orange text indicates proteins associated with steatosis or fibrosis (see also Online Supplemental Table 4). (B) Identification of plasma proteins in module gene/protein interaction networks. Ingenuity pathway analysis was used to identify all interaction networks for each module protein (see Figure 11), and the union of interaction proteins was used to identify plasma proteins in the interaction network of proteins expressed in liver tissue. Proteins with significantly different abundances between control and exposed animals (FDR < .05) were identified in liver tissue and plasma from rats administered 4,4′-MDA for a single dose (C) or 5 consecutive days (D). Data were normalized for clustering as in Figure 10. Red indicates greater than mean, and green indicates lower than mean. Venn diagrams were generated with Partek Genomics Suite. The intersection proteins are identified in the tables to the right of each diagram. Significance was determined by Permutation test with Benjamini–Hochberg corrected p value as well as a 1.3-fold change cutoff.
Following a single dose of 162 mg/kg 4,4′-MDA, 1 animal clustered with the control group instead of the treatment group (see the third column of the 4,4′-MDA treatment group in Figure 12A). Unlike the other treated animals, this animal did not have significant changes in serum liver enzymes, although this animal did have histological evidence of liver inflammation and bile duct hyperplasia (Online Supplemental Table 3). Further, this animal was the only one in the group to have (a) moderate cytoplasmic alteration and (b) moderate mitotic alteration characterized by increased hepatocyte mitotic figures (Online Supplemental Table 8). The serum AST and ALT levels for this animal were 150 U/L and 62 U/L, respectively (i.e., 2.7- and 4.3-fold lower than the average of the 162 mg/kg 4,4′-MDA dose group). However, given the histological features of bile duct hyperplasia and inflammation, this animal was not removed from the analysis.
After 5 consecutive doses, 13 proteins were differentially regulated in both plasma and tissue: ACTB, APOA1, C3, C4A/C4B, CP, FGA, FGB, FGG, HPX, ITIH4, KNG1, KNG1/KNG1|1, and LOC259246 (Figure 12D, intersection table, and Online Supplemental Table 5). FGA, FGB, FGG, ITH4, and KNG1/KNG1|1 were differentially regulated in both plasma and tissue after a single dose of 4,4′-MDA or 5 consecutive administrations (compare bold text in Figure 12C and D). A comparative analysis of the gene transcript precursors for the differentially expressed proteins at all doses is reported in Online Supplemental Table 6. After a single administration of 4,4′-MDA, none of the gene transcripts encoding FGA, FGB, FGG, HBB, HP, ITIH4, or KNG1/KNG1|1 were differentially expressed at any dose (Online Supplemental Table 6, lower left quadrant). After 5 days of consecutive administration, all gene transcripts with the exception of the fibrinogen chains were differentially expressed. Hbb and Itih4 were downregulated, and the remaining genes were upregulated (Online Supplemental Table 6, lower right quadrant).
Discussion
In this study, we identified differential protein expression modules in the context of mechanisms disease pathogenesis in steatosis and fibrosis. We administered the reference chemicals BB and 4,4′-MDA to male Sprague-Dawley rats and performed global proteomic analysis. We anchored differentially expressed protein modules to histological features characteristic of lipid accumulation and fibrosis. The results of the study identify unique protein modules expressed in liver after BB or 4,4′-MDA administration and identify plasma proteins in gene/protein interaction networks with these disease-associated liver proteins. Many of these proteins are associated with disease pathogenesis and associated histopathology for steatosis and fibrosis (Willett et al. 2014).
Liver Enzymes and Clinical Chemistries
As predicted, liver enzymes began to increase after 5 days of treatment but not after a single administration of BB or 4,4′-MDA. We did not observe a consistent dose response with the clinical chemistries (i.e., liver enzymes ALT and AST) in the BB-administered animals. At the highest dose, enzyme levels were lower than the dose selected for further protein analysis. This dose also caused significant necrosis affecting much of the liver. While the reason for this inconsistency is unclear, this dose caused overt clinical symptoms of wasting and toxicity, suggesting generalized systemic toxicity in addition to liver damage.
In addition to the traditional transaminases, serum SDH was increased after 5 days of 4,4′-MDA administration. A single administration of either chemical also resulted in increased SDH concentrations. SDH was also upregulated in plasma by after a single dose of BB. Serum SDH elevation is liver-specific, reacts similarly to transaminases in response to liver injury, and has been used in conjunction with these biomarkers to assess liver damage (Burcham and Harman 1988; Singh et al. 2015). However, SDH is disease agnostic and tends to rapidly increase before returning to normal, which can be difficult to capture from a clinical chemistry panel (Weisner et al. 1965). Intracellularly, SDH facilitates the process of de novo fatty acid synthesis by using glucose to synthesize fructose, rapidly depleting ATP in the process (Luo et al. 2016; Marchesini, Petta, and Dalle Grave 2015; Abdelmalek et al. 2010). Further research is necessary to elucidate the mechanism by which SDH enters the blood.
In BB-treated livers, UGDH was a significantly upregulated glucose utilization protein relative to controls (Online Supplemental Figure 4). UGDH mediates the production of UDP glucuronic acid from UDP glucose. Prior to UGDH activity, UGP2 must convert glucose-1-phosphate into UDP glucose. This is a key step in glucose metabolism because UGP2 is the only enzyme capable of catalyzing this reaction. In summary, reduced serum glucose, along with increased plasma SDH, UGP2, and liver UGDH, supports the histological evidence of increased carbohydrate utilization with de novo fatty acid synthesis in the liver after BB intoxication.
UGP2 was not elevated in the liver tissue, suggesting an alternative source for UGP2 in plasma. UGP2 has been identified in the secretome or exosome of peripheral blood mononuclear cells (PBMCs), red blood cells, and neutrophils (Mathivanan Lab 2017). When stressed, PBMCs produce exosomes that can promote wound healing (Beer et al. 2015). The UGP2 could have entered the blood as part of the response to damage. Further studies are needed to confirm these hypotheses.
Histopathological and Toxicoproteomic Effects of Reference Chemicals
In our earlier work, we used in silico analysis to identify gene modules anchored to histological end points derived from the large set of gene expression and histological data in the DrugMatrix database (Tawa et al. 2014). The results of the present study provide confirmatory evidence at the protein level for the gene expression modules we previously identified in transcript data. Our study shows that many of the genes in the gene toxicity modules match differentially expressed protein products. Global proteomics analysis detected protein products of only about 4% of the transcripts detected by Affymetrix array analysis, yet, surprisingly, many of the differentially expressed proteins mapped to 1 or more gene toxicity module, with between 11% and 50% protein coverage of the total number of genes in a given module. Moreover, we identified liver tissue proteins in gene and protein interaction networks with differentially expressed plasma proteins. The number and identify of proteins in these interaction networks change in association with histological evidence of disease progression and severity. These results support a module-based approach to defining mechanistic biomarkers of disease progression. Interpreting the network associations between tissue and plasma proteins should be approached with caution. Differential expression in plasma could be associated with either leakage from the liver or another tissue or a physiological related or unrelated mechanism for liver-specific pathologies (e.g., a systemic acute phase reaction; Mellor, Steinmetz, and Cronin 2016).
Steatosis
Toxicants and drugs, including amiodarone, methotrexate, irinotecan, and glucocorticoids, are well recognized to cause steatosis (Rabinowich and Shibolet 2015). Although BB is best known as a metabolically activated necrosis-inducing toxicant, exposure has resulted in vacuolation with lipid accumulation indicative of steatosis (Heijne et al. 2005; Wong, Card, and Racz 2000; Dahl et al. 1990; Yoshioka et al. 2017). In our study, BB administration resulted in histological features associated with nonalcoholic fatty liver disease (NAFLD), characterized by mild inflammation with vacuolation and positive staining for Oil Red O (i.e., presumptive lipid accumulation).
We also observed histological features of steatohepatitis, including inflammation, hepatocyte necrosis, and hypertrophy. Our plasma and tissue protein profiles suggested early indications of fibrotic injury in the absence of histological evidence of collagen accumulation. Although fibrosis is not typically characteristic of NAFLD (Brunt and Tiniakos 2010), nonalcoholic steatohepatitis (NASH) progresses to fibrosis and ultimately liver cirrhosis. The primary distinction between simple steatosis and NASH is the presence of hepatocellular injury (Brunt and Tiniakos 2010; Sanyal 2011). We observed histological evidence of inflammation and lipid accumulation within 24 hr of BB administration in our study. Concomitantly, we observed protein signatures suggesting changes in glucose utilization, lipid metabolism, and carbohydrate metabolism. Steatosis-associated modules 3, 24, and 25 (enriched at days 1 and 5) contained proteins that mapped to molecular mechanisms proceeding from nuclear receptor activation or oxidative stress in the steatosis pathogenesis (Figure 13A). Modules 1, 11, and 12 (enriched at day 5) mapped to oxidative stress–related molecular mechanisms in the steatosis pathogenesis (Figure 13A). A complete list of proteins associated with each module is given in Online Supplemental Table 3. Many of the changes observed were associated with different phases of the steatosis pathogenesis. For example, SDH, UGP2, and serum glucose could be related to inflammation, glucose utilization, and lipid and/or carbohydrate metabolism.
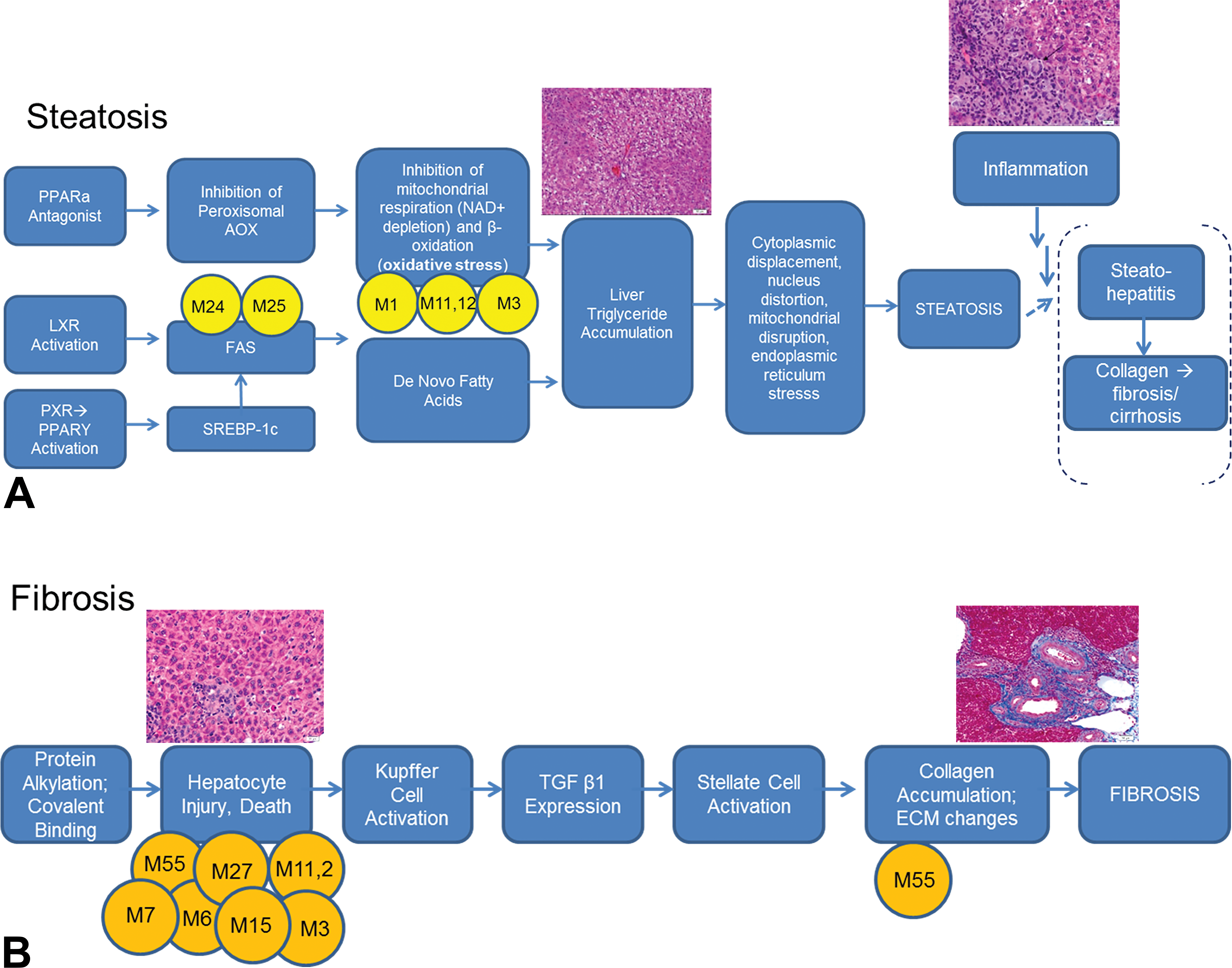
A modules approach to steatosis and fibrosis pathogenesis. (A) Modules were associated with histopathologies and relevant mechanisms of toxicity in steatosis pathogenesis based on analysis of gene function within each module. Over time, prolonged exposure to steatogenic compounds can lead to steatohepatitis in the presence of inflammation and ultimately to fibrosis and cirrhosis (dotted brackets). See Online Supplemental Table 3 for a full list of genes within each module (M = module; BB = bromobenzene; PPAR = peroxisome proliferator activator receptor; LXR = liver X receptor; PXR = pregnane X receptor; SREBP = sterol regulatory element-binding protein; AOX = fatty acyl-CoA). (B) Modules were associated with histopathologies and the fibrosis adverse outcome pathway based on analysis of gene function within each module. See Online Supplemental Table 3 for a full list of genes within each module (M = module; 4,4′-MDA = 4,4′-methylenedianiline; TGF = transforming growth factor; ECM = extracellular matrix).
We saw no histological evidence of fibrosis at any time point in BB-treated animals, but over time, we observed a shift to upregulation in fibrosis-related protein signatures. On day 1, our study identified upregulation in protein patterns corresponding to early molecular mechanisms in steatosis pathogenesis, including lipid accumulation, oxidative stress, and nuclear receptor activation (Chin et al. 2016; Dugas et al. 2004; Ganter et al. 2005). The proteins FASN, acetyl-CoA carboxylase (ACACA), and ATP citrate lyase (ACLY) are upregulated in liver tissue at 1 and 5 days. FASN expression is one of the key events leading to the steatosis phenotype. ACACA) and ACLY () are both involved in fatty acid synthesis and have been implicated in steatosis (Meakin et al. 2014; Nishikawa et al. 2012; J. S. Yang et al. 2010). Early in disease progression, liver levels of FASN, ACACA, ACLY, and UGDH tissue are in gene/protein interaction networks with plasma proteins APOB, A2M, and UGP2. APOB secretion from the liver is positively correlated with lipid efflux (Angrish et al. 2016). A2M (α-2-macroglobulin) is a protease inhibitor synthesized by hepatocytes. The protein is an acute phase response protein and is secreted into the blood as part of generalized injury, stress, and inflammatory responses (Naveau et al. 1994; Shukla et al. 2015). After 5 days, FASN, ACACA, ACLY, and UGDH were still upregulated, but the proteins in gene/protein interaction networks changed in the plasma. The new interaction partners included UGP2, fibronectin 1 (FN1), and collagen 6A1 (COL6A1). FN1 and COL6A1 are associated with ECM reorganization and collagen formation in fibrosis (Alsafadi et al. 2017; Lettmann et al. 2014). Further, by 5 days, 4 proteins in a fibrosis-related expression module were differentially expressed in the liver tissue: annexin 2, vimentin, transgelin 2, and chloride intracellular channel 1. Differential expression of all 4 has been reported in fibrosis-related pathologies (Ippolito et al. 2016; Zhong et al. 2015). FN1 and PSMA1 (proteasome subunit 1) are differentially expressed plasma proteins in the gene/protein interaction network. Proteasomes (e.g., PSMA1) and proteasomal functions have been associated with alcoholic steatosis and liver injury (Wang et al. 2016). The upregulation of a fibrosis-related module may suggest the initial progression toward steatohepatitis and ultimately fibrosis (see Figure 13A). A longer time course is needed to verify this hypothesis.
Fibrosis
Chemicals such as carbon tetrachloride, thioacetamide, and dimethylnitrosamine are well recognized to cause fibrosis in the centrilobular region (Amin and Mahmoud-Ghoneim 2011; Liedtke et al. 2013; Wallace et al. 2015). Although not considered an archetypal fibrosis-inducing chemical, 4,4′-MDA has been reported to cause periportal fibrosis (Ishihara, Katsutani, and Aoki 2006; Dugas et al. 2004). In accordance with literature predictions, our histopathology results suggest that the toxicant 4,4′-MDA induced fibrosis by a mechanism independent of lipid accumulation and steatosis precursors (Ganter et al. 2005). Fibrosis may result from protein alkylation after chemical intoxication (Vinken 2013). In our study, 4,4′-MDA exposure may perturb energy metabolism or ECM reorganization through the production of hyaluronic acid (Fuhring et al. 2015; see Online Supplemental Figure 4). Further, there were fewer proteins involved in lipid metabolism affected by 4,4′-MDA than BB (see Online Supplemental Table 7). Carbohydrate utilization pathways were not perturbed by 4,4′-MDA until later in disease progression (i.e., after 5 days of administration; see Online Supplemental Figure 4).
In our study, many of the protein modules mapped to histopathological indicators of fibrosis (e.g., ECM formation and hepatocellular injury and death). These mechanisms lead to the pathological features of fibrosis. Of the 4 animals, 3 administered a single dose of 4,4′-MDA showed comparable protein changes in plasma and liver, which associated with histological features of fibrosis. A larger cohort is needed to determine whether the cytoplasmic alteration and/or mitotic alteration in 1 animal’s liver contributed to the differences in protein profiles between this animal and all others in this dose group. Interestingly, a single administration of 4,4′-MDA led to modulation of fibrosis-related modules in liver tissue and plasma proteins, including the α, β, and γ chains of the fibrinogens (FGA, FGB, and FGG), as well as the inflammatory proteins ceruloplasmin (CP), transferrin, and apolipoprotein A1 (APOA1; Ippolito et al. 2016). Many of the differentially regulated plasma proteins after 5 days of 4,4′-MDA administration associated with collagen accumulation, ECM reorganization, and inflammation leading to fibrosis (e.g., SERPINA3, C3, HPX, KNG1; Ding et al. 2011; Ippolito et al. 2016). Seven of the modules contained proteins previously reported in association with hepatocyte injury (Figure 13B). One of the modules (M55) contained ECM proteins and mapped to tissue-specific key events associated with collagen accumulation (Figure 13B). CP mapped to module 55, and FGA mapped to modules 55 and 20 (see Online Supplemental Table 3). Taken together, these results suggest that the changes in plasma proteins may be in interaction networks with liver fibrosis modules, possibly associated with acute phase responses or inflammatory responses following injury or death of hepatocytes. However, as indicated above for BB, interpreting changes in the plasma in the context of interaction networks needs to be approached with caution. Although many of the proteins are associated with acute phase response and ECM remodeling, some may be related to responses unrelated to liver pathology (e.g., dehydration or inappetence; Arias 1982). Plasma proteins identified using the iTRAQ labeling were high-abundance proteins associated with the acute phase response. More sensitive assays are needed to detect low-abundance proteins that are more specifically associated with liver disease progression of individual toxicants.
Mechanistic Framework
Systems toxicology seeks to unite molecular events with histological observations at the individual molecule and pathway levels to better predict disease progression and diagnosis. Both fibrosis and steatosis pathologies have been well described in mechanistic contexts (Willett et al. 2014). At the molecular level, toxicants may induce steatosis by perturbing cellular pathways at the level of specific nuclear receptors and proteins associated with lipid homeostasis disruption, leading to increased accumulation of triglycerides within hepatocytes. Dysregulation of lipogenesis, mitochondrial β-oxidation, and fatty acid transport can ultimately lead to steatosis (Zhou et al. 2006; Mellor, Steinmetz, and Cronin 2015; Kawano and Cohen 2013). Xenobiotics can activate nuclear receptors (e.g., LXR, RXR, PPAR), which serve as molecular initiating events for steatosis by perturbing normal homeostasis of fatty acid metabolism (Willett et al. 2014). LXR can heterodimerize with RXR and bind to the promoter region of genes involved in fatty acid metabolism (e.g., fatty acid synthase, apolipoprotein E, lipoprotein lipase, fatty acid uptake transporter, and genes encoding regulatory proteins such as carbohydrate response element binding protein and sterol response element binding protein). Regulatory proteins can activate enzymes orchestrating glycolysis and lipogenesis (e.g., ACACA and fatty acid synthase) as well as triglyceride accumulation (steroyl-CoA carboxylase). Caution should be taken when interpreting the IPA-based associations between proteins identified in our study and the LXR, PXR, PPAR, and RXR response, as many of these proteins may be associated with the acute phase response and/or generalized liver injury without specificity to the steatosis pathology.
Inhibition of mitochondrial function may be an important factor in BB-induced hepatotoxicity by disrupting β-oxidation with subsequent increases in fatty acids and triglyceride formation. This mechanism could be integral to activation of SREBPF1 (Sterol Regulatory Element Binding Transcription Factor 1) in the steatosis pathogenesis (Wong, Card, and Racz 2000). In our analysis of both tissue and plasma, we identified proteins directly downstream of these molecular initiating events, suggesting a mechanistic link between protein expression, cellular function, toxicological pathways, and histological features of disease. Many of the proteins found to be upregulated after BB administration were also associated with fibrotic end points, consistent with the literature indicating that unchecked steatosis can lead to steatohepatitis, fibrosis, and/or cirrhosis (Willett et al. 2014). It should be noted, however, that further research is needed to differentiate proteins involved in general liver toxicity and those specific to a given pathological response (Gruys et al. 2005). The differentially expressed proteins for BB affecting lipid accumulation and carbohydrates and the 4,4′-MDA-induced changes in proteins associated with the ECM, acute phase response, coagulation, inflammatory signaling, and degradation pathways may be nonspecifically associated with generalized liver injury. With further research, some of these biomolecular changes could be placed within the context of AOPs, which are conceptual constructs that map existing knowledge regarding links between molecular initiating events and adverse outcomes (Ankley et al. 2010).
Biomarker Discovery
In our study, we report differential expression of both liver and plasma proteins associated with early or later stage disease (day 1 vs. day 5). It should be noted, however, that it is not surprising that we observe little overlap at the protein level. Proteins secreted in response to tissue damage are likely to be present in low abundance in serum. However, the proteins that do overlap in plasma and serum could serve as a potential panel of novel biomarker candidates for liver injury in an accessible biofluid (e.g., plasma or saliva; Pinzani and Rombouts 2004).
Liver biopsies remain the gold standard for staging disease progression in order to inform therapeutic regimen. Circulating biomarkers specific for mechanistic features of liver disease could serve as surrogate histological indicators, obviating or reducing clinical indication for liver biopsy. Ongoing research seeks to identify biomarkers distinctly correlate with histological features.
Numerous serological tests for liver fibrosis have been developed in recent years, but diagnosing and staging fatty liver diseases have proven more challenging because of the multifactorial nature of steatogenesis (Munteanu et al. 2016). As antifibrotic drugs enter clinical trials, it becomes critical to identify and stage liver fibrosis before irreversible damage occurs (Munteanu et al. 2016). Specific biomarkers are necessary for effective assessment of hepatotoxicity. An ideal mechanistic biomarker would have a detectable increase in concentration before significant liver damage occurs. Tests such as the fibrotest/fibrosure, fibrometer, and hepascore include biomarker combinations to assess inflammation, collagen accumulation, and steatosis to better predict disease progression noninvasively (Chin et al. 2016; Munteanu et al. 2016). A better understanding of differentially expressed protein modules in liver tissue and the plasma protein in their interaction networks can facilitate the development of mechanistic biomarkers for more specific and sensitive tests to evaluate disease progression.
Conclusions
This study used global proteomic analysis to anchor differentially expressed protein modules to histological features of liver steatosis and fibrosis induced by the toxicants BB and 4,4′-MDA, respectively. Linking histology with molecular changes in the context of a mechanistic framework like the AOP improves the understanding of molecular mechanisms of hepatic disease, allowing for the discovery and clinical use of mechanistic biomarkers of hepatotoxicity.
Footnotes
Authors’ Note
Opinions, interpretations, conclusions, and recommendations are those of the author(s) and are not necessarily endorsed by the U.S. Army. Citations of commercial organizations or trade names in this report do not constitute an official Department of the Army endorsement or approval of the products or services of these organizations. This research complied with the Animal Welfare Act and implementing Animal Welfare Regulations, the Public Health Service Policy on Humane Care and Use of Laboratory Animals, and adhered to the principles noted in The Guide for the Care and Use of Laboratory Animals (National Research Council [US] 2011)
Acknowledgments
We thank LTC Vazquez and Dr. Valerie DiVito for program management oversight. We thank Integrated Laboratory Systems, Inc., for conducting the in vivo animal exposure experiments and for interpretation, particularly Dr. Cheryl Hobbs, and Mr. Michael Streicker. We thank Dr. Erica Carroll for consultation.
Authors’ Contribution
Authors contributed to conception or design (BM, MA, MP, WD, CB, JK, MB, SW. JL, DI); data acquisition, analysis, or interpretation (BM, MA, MP, WD, CB, JK, MB, SW, JL, DI); drafting the manuscript (BM, JK, SW, DI); and critically revising the manuscript (BM, MA, MP, WD, CB, JK, MB, SW, JL, DI). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported in part by an appointment to the Postgraduate Research Participation Program at the U.S. Army Center for Environmental Health Research administered by Oak Ridge Institute for Science and Education (ORISE) through an interagency agreement between the US Department of Energy (DOE) and United States Army Medical Research and Materiel Command (USAMRMC).
Supplemental Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
