Abstract
Mice and humans branched from a common ancestor approximately 80 million years ago. Despite this, mice are routinely utilized as animal models of human disease and in drug development because they are inexpensive, easy to handle, and relatively straightforward to genetically manipulate. While this has led to breakthroughs in the understanding of genotype–phenotype relationships and in the identification of therapeutic targets, translation of beneficial responses to therapeutics from mice to humans has not always been successful. In a large part, these differences may be attributed to variations in the alignment of protein expression and signaling in the immune systems between mice and humans. Well-established inbred strains of “The Laboratory Mouse” vary in their immune response patterns as a result of genetic mutations and polymorphisms arising from intentional selection for research relevant traits, and even closely related substrains vary in their immune response patterns as a result of genetic mutations and polymorphisms arising from genetic drift. This article reviews some of the differences between the mouse and human immune system and between inbred mouse strains and shares examples of how these differences can impact the usefulness of mouse models of disease.
Keywords
Despite having diverged nearly 80 million years ago (Yang et al. 2004), mice are routinely used in scientific research as models of human diseases. The use of mouse models for immunological research has many advantages: (1) the mouse immune system is relatively well characterized, (2) the mouse genome has been sequenced for several commonly used mouse strains, (3) genetic manipulation is relatively straightforward, and (4) their size, life span, and fecundity allow for easy handling and study designs. While the use of mouse models for understanding diseases and for evaluating potential therapeutics has been enlightening, the translation of discoveries from mice to humans has not always been successful. In mouse models of immune diseases and therapies, these differences may be attributable in part to the variations in the protein expression and functional alignment of immune responses between the two species. Complicating the translation of mouse model findings to human diseases and therapies is in part the result of genetic polymorphisms or pseudogenes between the various commonly used inbred mouse strains. This review highlights some of the differences among inbred mouse strains and between the mouse and human immune system as well as shares examples of how these differences can impact the usefulness of mouse models of human diseases. This is not intended to be an exhaustive review of these differences but rather aims to offer illustrative examples that emphasize the importance of researching and understanding genetic variation before developing or using animal models of immune diseases or for immune therapies.
Mouse Model Enigma
It has been said that mice are “lousy models for clinical studies” (Davis 2008, 835). This sentiment seems to be increasingly common as drugs that are successful preclinically fail in clinical trials (Begley and Ellis 2012; Prinz, Schlange, and Asadullah 2011; Mak, Evaniew, and Ghert 2014). While it seems that success has been limited in translating drugs that “cure” mice into effective drugs for humans, many failures may, in part, be related to suboptimal study design (Begley and Ellis 2012). The converse may also be true: some drugs that failed in mouse models might have worked well in patients. For these models to be useful, there needs to be an in-depth understanding of mouse pathobiology and its relationship to the human disease to be modeled. Fundamentally, mouse models of human disease are artificial. For example, models of colitis in which disease is induced in mice by administration of dodecyl sodium sulfate (DSS) is essentially a model of mucosal injury and the immune response to that injury (DeVoss and Diehl 2014). This model does not specifically recapitulate the pathogenesis of human ulcerative colitis (UC) or Crohn’s disease (CD), but it continues to be used as a model for disease mitigating therapies because of its ease and reproducibility (Benight et al. 2012; Li et al. 2015). Although genetically engineered mice (GEM) have allowed phenotypic evaluation of individual genes and gene mutations as well as the development of mouse models of human diseases such as UC or CD, they do not exactly replicate the spectrum of human disease (DeVoss and Diehl 2014; Valatas, Vakas, and Kolios 2013). Indeed, simply replicating the underlying genetic predispositions to human disease in mice may not be adequate for disease expression, as they cannot recapitulate the myriad influences that drive the pathogeneses. While underlying genetic polymorphisms in the human population may predispose to disease, manifestation of disease typically also requires additional environmental, microbial, or physiological triggers that often cannot be easily captured in the animal model (Rosenblum, Remedios, and Abbas 2015). A detailed understanding of the disease to be modeled and the immune context of the different stages of that disease are essential to make valid comparisons from mouse to human. When modeling immune-mediated diseases, there are a number of factors that must be considered.
While no mouse model is perfect, failure of models to predict therapeutic targets or human responses to therapies lies in part with flaws in research design and implementation. When designing studies in mice, careful evaluation of the mouse strain and substrain genetics must be considered as well as the comparability of the immune expression and immune response patterns between mice and humans. Beyond genetics, however, studies must be designed with attention to the use of adequate animal numbers and appropriate controls and with an understanding of how the environment, diet, housing, husbandry, and microbiome could impact the outcome of the studies (Kilkenny et al. 2010; Justice and Dhillon 2016; Reardon 2016). In addition, the use of different mouse models or immunologically divergent mouse strains to address a research question may be mechanistically informative and improve the predictive value of these studies (Begley and Ellis 2012).
Genetically Engineered Mouse Models—The Importance of Genetic History
Knowledge of the genetic history (the source and specific substrains used and the backcrossing history) of GEM can be important in interpreting the data and understanding its relevance to human disease (Barthold 2002). Although many researchers obtain their GEM from vendors, it is also quite common for them to obtain these mice from collaborators. When GEM are obtained from collaborators, they may have little insight into the genetic history of those animals, which is often compounded by the lack of inclusion of substrain information in the original publications on the GEM (e.g., Embryonic Stem [ES] cell origin, backcrossing strain source, etc.). Mice generated by ES cell technology are often backcrossed with the various C57BL/6 substrains. Mice are considered fully backcrossed (“congenic”) after 10 or more generations. However, even fully backcrossed mutant mice retain about 0.2% of the ES donor cell DNA flanking the inserted genetic material (up to 3.3 cM on each side) as well as some distributed randomly throughout the other regions of the DNA (Schalkwyk et al. 2007; Eisener-Dorman, Lawrence, and Bolivar 2010). This flanking DNA (congenic footprint) and randomly distributed DNA may contain hundreds of genes, some of which may be modifier genes for the target of interest or pertain to phenotypic characteristics unrelated to the gene of interest (Schalkwyk et al. 2007). It is important to understand that despite repeated backcrossing, unexpected biological consequences may arise from donor gene retention. Methods such as the clustered regularly interspaced short palindromic repeats (CRISPR) that can generate mutations on selected genetic background(s) can mitigate this concern, and offer opportunities to leverage differences between strains, by generating mutations in different strains. It is essential that pathologist and basic researchers work together not only interpreting the data from these models but also designing the studies. The importance of understanding the genetic history of the GEM and their concurrent controls cannot be understated (Barthold 2002).
Strains and Substrains
A common misconception about the inbred mouse strains is that they are genetically identical, regardless of their source. In fact, there may be notable diversity between the same strain obtained from different sources as a result of genetic drift. In some cases, these differences may impact immune system responses. While there are many inbred strains of mice commonly used in translational research, examples of strain and substrain variation will be detailed here for the two most commonly used in the development of genetically modified mice: C57BL/6 and 129. However, when using other strains of mice, the same source/substrain issues must be considered (Casellas 2011).
C57BL/6 Strain and Substrains
C57BL/6 is one of the most common mouse strains used in biomedical research and probably the most common strain involved in contemporary GEM. Although commonly referred to simply as C57BL/6 in communications and (peer-reviewed) publications, the substrains are not identical and research-relevant differences among substrains have been identified and reported (Mekada et al. 2009; Kiselycznyk and Holmes 2011; Simon et al. 2013b). Data derived from different substrains are routinely compared both between and within laboratories. A history of the commercially available C57BL/6 substrains can be found at http://www.envigo.com/resources/data-sheets/envigo-68-c57bl6-enhanced-technical-data-sheet_screen.pdf and https://www.jax.org/strain/000664. C57BL/6 substrains can be identified by the letter(s), referring to the laboratory codes, following the “6.” For example, C57BL/6J indicates the source as the Jackson Labs (the shortened version, B6, specifically refers to the C57BL/6J substrain). C57BL/6N substrains are derived from the National Institutes of Health (NIH); these mice were obtained by the NIH from the Jackson Labs in 1951. These 2 substrains were shared with other vendors and retained the C57BL/6J or C57BL/6N nomenclature but should have subsequent additional laboratory code(s) that indicate the supplier, such as Hsd (Harlan, now Envigo), Tac (Taconic), Crl (Charles River Labs), and so on. Thus, a C57BL/6 at Crl derived from the NIH substrain is C57BL/6NCrl. These all represent C57BL/6 substrains which have genetically drifted from their parent strain; the N substrains are more similar to each other than to the J substrain (Simon et al. 2013a; Zurita et al. 2011), and the differences should be reviewed before selecting a source for any biomedical research. These variations are often overlooked by researchers who may purchase mice from different vendors depending on contracts, cost, or inattention. An important example of the potential impact of substrain variation in immune responses was reported recently in C57BL/6NHsd mice and was confirmed by the supplier in some but not all of the lines named C57BL/6NHsd (http://www.envigo.com/assets/docs/c57-customer-communication-2-final-9jun16.pdf). In these mice, a genetic mutation was identified in the
Conclusions generated from GEM models may also be complicated by the use of purchased control comparators rather than wild-type littermates. For primarily fiscal reasons, it is common for researchers working with GEM to maintain only the null mice for use in their studies. Instead of using wild-type littermates, they purchase wild-type C57BL/6 (or other strain) from a supplier. This practice introduces several confounding factors into their research: (1) genetic drift may have introduced meaningful changes in the GEM genome as these research institution-maintained colonies tend to be quite small, (2) retained ES cell DNA, if from a different strain may impact study findings, (3) the control substrain may be from a different source (and thus may have unique polymorphisms) than that used for backcrossing, and (4) the environmental exposures (diet, caging, bedding, microbiome, etc.) between the 2 groups will be different, sometimes dramatically so (Stappenbeck and Virgin 2016; Reardon 2016). Additionally, some researchers keep their own C57BL/6 colonies to use as wild-type controls for their studies, which undoubtedly have notable genetic drift as a result of small colony size and suboptimal breeding programs. Thus, phenotypic conclusions on immune effects between in-house GEMs and vendor-sourced wild-type mice may have a significant impact on translational studies.
The 129 Substrains
Similar issues regarding substrains and suppliers of C57BL/6 mice exist also for the “129”mice. Genetic variation among 129 substrains is notably greater than that for C57BL/6, and current 129 nomenclature recognizes 16 substrains representing 4 major groups, which is reviewed in Simpson et al. (Simpson et al. 1997; Threadgill et al. 1997). The substrains of origin of most of the available 129-derived ES cell lines can be searched via the International Mouse Strain Resource (IMSR; http://www.findmice.org/index.jsp). Until recently, 129 strains of mice have been the primary source for ES cells used to generate mutant mice (e.g., “knock-out” or “knock-in” mice). As previously mentioned, the “congenic footprint” of the ES cell DNA may confound study data. Researchers should be aware of the 129 substrains and/or ES cell substrains (see https://www.jax.org/news-and-insights/2001/february/129x1-svj-genetically-contaminated-what-does-that-really-mean) being used in their investigations, as this may impact the validity of the data interpretation (Cantin and Mann 1999).
Examples of Immune Variation Impacting Mouse Models
While common inbred mice are considered “immune competent,” many have specific immune deficits (Sellers et al. 2012; Doran et al. 2016; Mostafavi et al. 2014). A few examples of these differences will be discussed, but a more comprehensive (although not exhaustive) list of some of the better described differences is found in Table 1. Additional excellent resources (beyond the included references) on strain genetic variation/polymorphisms include the IMSR website http://www.findmice.org/ and the Mouse Genome Informatics website, http://www.informatics.jax.org/. In a number of cases, mouse strain-specific immunological peculiarities have been exploited to generate mouse models of disease. Susceptibility of various strains to infectious or immune disease induction is certainly useful: these models tend to be consistent and reproducible in certain strains. This susceptibility is typically due to immune deficits or polymorphisms that may or may not be associated with disease susceptibility in humans. For example, one of the most common strains of mice used for induction of experimental autoimmune encephalitis, a model of multiple sclerosis, is the SJL/J strain. These mice have a wide assortment of immune abnormalities, including hyperresponsive IL-12 (Zwiers et al. 2011) and hyporesponsive IL-4 (Yoshimoto et al. 1995) and abnormalities in T cell receptor (TCR)β-V8 receptor function as well as factors associated with SJL/J substrain variation (Behlke et al. 1986; Behlke and Loh 1986, Summers deLuca et al. 2010). In addition, strains may have different biases in T helper (TH)-type responses. For example, C57BL/6 mice have been demonstrated to have a TH1-type bias to pathogens, whereas strains such as BALB/c, A/J, and DBA/2 mice have TH2-biased responses. This does have an impact on how mice respond to infectious diseases, induction of autoimmune disease, and clearance of materials, such as nanoparticles used in immunotherapies (Watanabe et al. 2004; Jones et al. 2013).
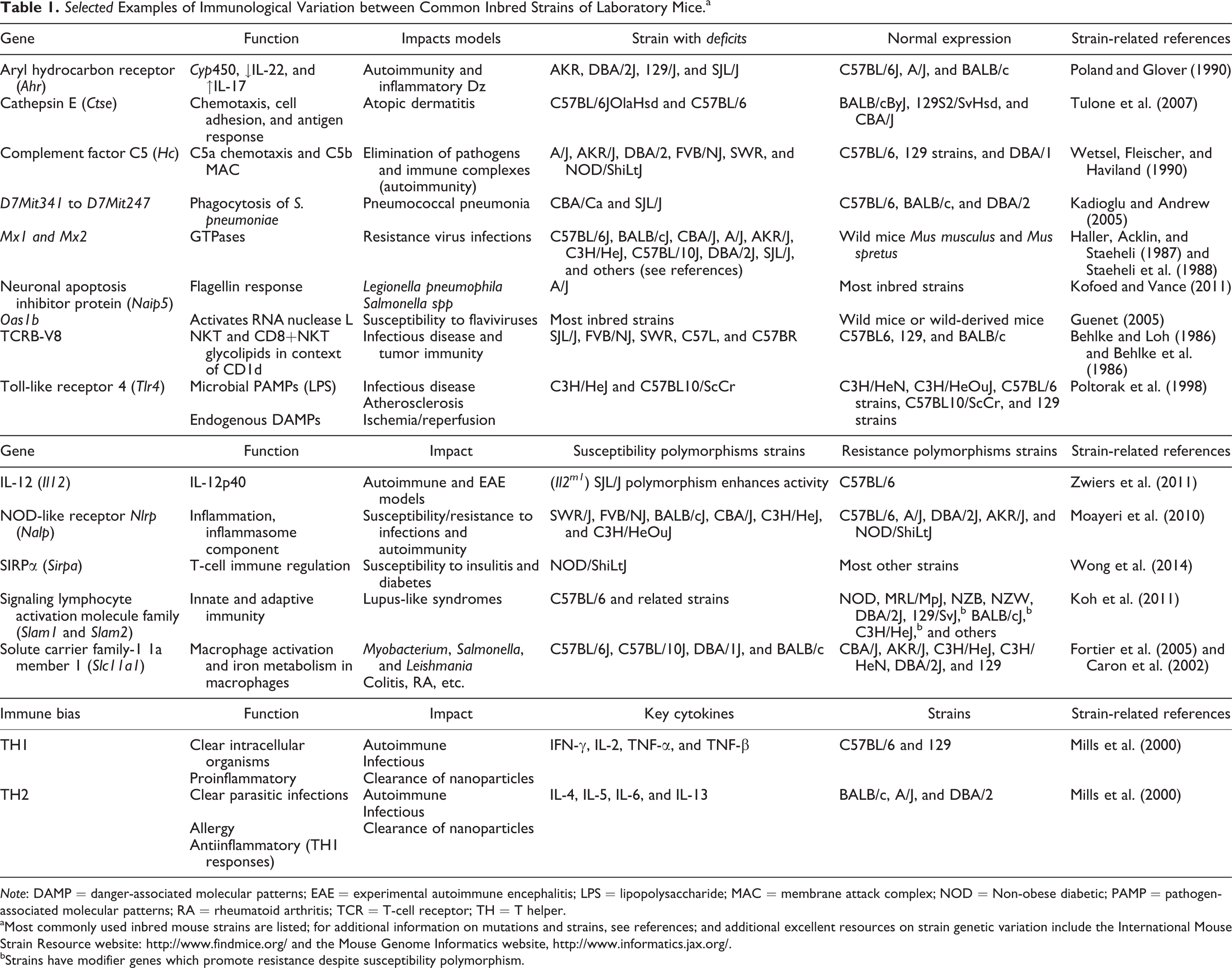
aMost commonly used inbred mouse strains are listed; for additional information on mutations and strains, see references; and additional excellent resources on strain genetic variation include the International Mouse Strain Resource website: http://www.findmice.org/ and the Mouse Genome Informatics website, http://www.informatics.jax.org/.
bStrains have modifier genes which promote resistance despite susceptibility polymorphism.
The C3H/He strains offer another example of an immune deficit that has been exploited to understand disease pathogenesis. The C3H/HeJ substrain has no functional toll-like receptor 4 (TLR4), while C3H/HeN (and other C3H strains) has fully functional TLR4 (Poltorak et al. 1998). TLR4 is a key regulator in innate immune responses to microorganisms and cell damage and is an important mediator of immune responses to lipopolysaccharide (Kawai and Akira 2010; Alexander and Rietschel 2001). This difference has allowed for studies evaluating the importance of TLR4 in the pathogenesis of diseases such as myocardial ischemia/reperfusion and in acute bacterial infections (Zhao et al. 2009; Moreno et al. 2013). Increased susceptibility to intracellular organisms such as
Another example of strains that have a similar name but with immune differences is the DBA/1 and DBA/2. Interestingly, the DBA/1 mouse is notably more susceptible to induction of autoimmune diseases such as collagen-induced arthritis and glomerulonephritis, and this may be due in part to the fact that DBA/2 but not DBA/1 strains have a loss of function mutation in the
A final example of how immune-relevant polymorphisms can impact data interpretation from mouse models is in susceptibility to
Mouse and Human Immune Comparability
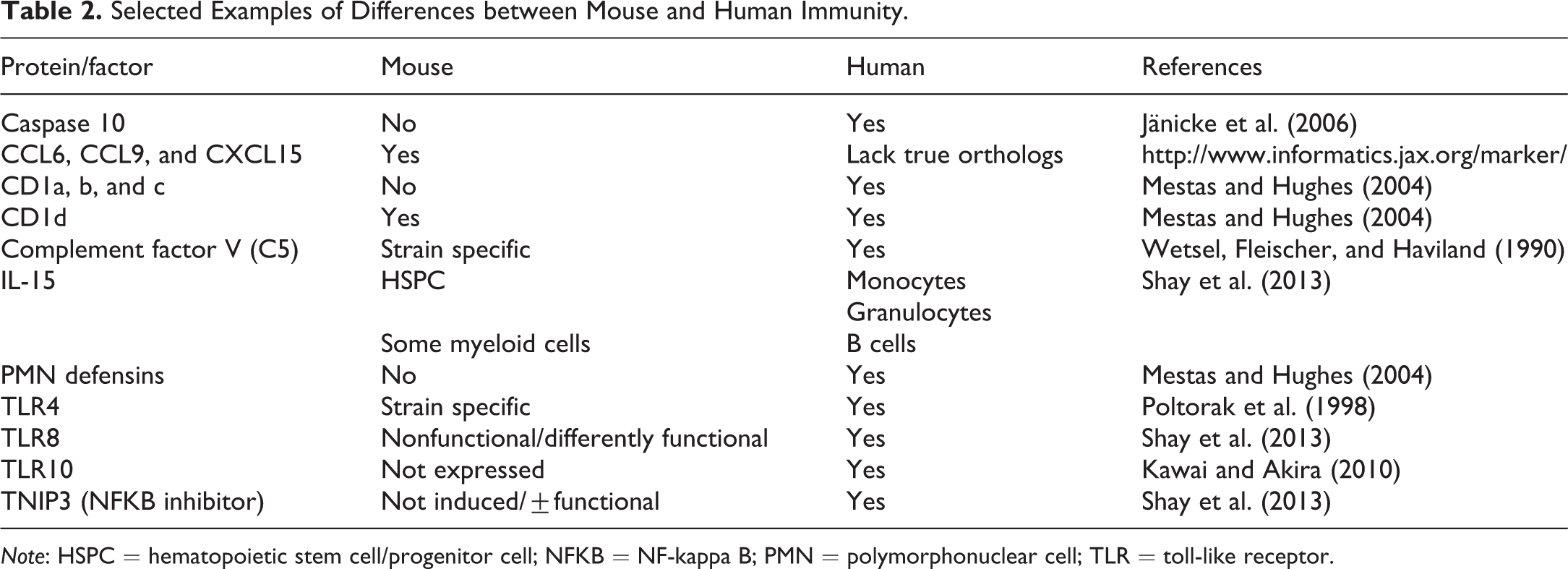
While much of the understanding on immune cell differentiation and function derives from mice, molecular comparisons between immune cell gene expression and function on a large scale have begun only recently. Several well-characterized differences between mouse and human immunity have been reported and include an assortment of both innate and adaptive immune system differences. In the innate immune system, for example, mice do not express TLR10 (Kawai and Akira 2010), have nonfunctional or differently functional TLR8 (Shay et al. 2013), and do not seem to produce functional TNIP3, an inhibitor of NF-κB (Shay et al. 2013). In addition, there are many cytokines and chemokines in humans for which no known ortholog exists in mice and vice versa (Shay et al. 2013). In some cases, surrogate markers can be used (Zlotnik and Yoshie 2012). More examples of these differences are present Table 2.
Selected Examples of Differences between Mouse and Human Immunity.
Natural killer (NK) cells and the gamma–delta (γδ) T cells are present in both mice and humans and bridge the innate and adaptive immune system (Tyler et al. 2015; Schuster et al. 2016); however, there are notable differences between the cell-type-specific receptors between the 2 species (Table 3). While mouse and human NK cells function similarly, their receptors are highly divergent and reflect a coevolutionary process (Mestas and Hughes 2004; Makrigiannis and Anderson 2000; Rahim and Makrigiannis 2015). NK cells of both species function in an MHC I fashion and express the activating receptor NKG2D and also have activating and inhibiting receptors. However, the activating and inhibitory receptors have diverged significantly. The mouse retains the more ancient Ly49 C-type lectin family receptors for this purpose, and primates use the more recently evolved killer cell immunoglobulin-like receptor family (KIR; Uhrberg 2005). While humans do have one of the
Comparison of Mouse and Human Natural Killer Cells.a
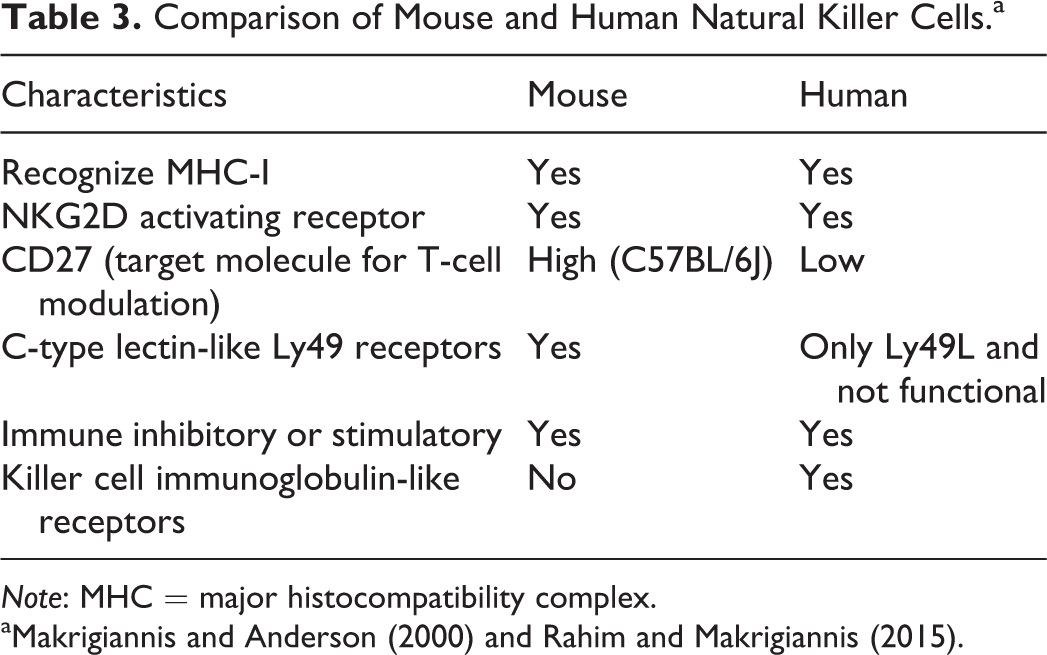

a Makrigiannis and Anderson (2000) and Rahim and Makrigiannis (2015).
Ly49 C-type Lectin Family Activating and Inhibitory Receptors on NK Cells.a
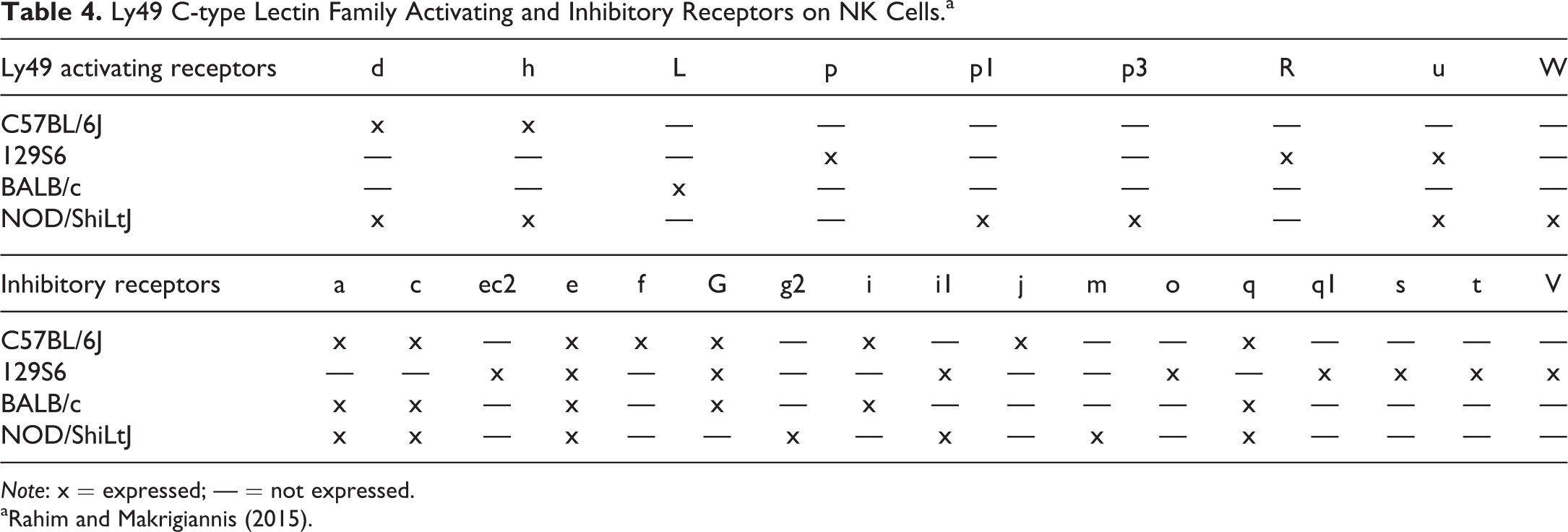
The γδ T cells are a heterogeneous population of cells which are infrequent in the circulation as compared to alpha–beta (αβ) T cells but are common in tissues, particularly in mucosal surfaces (Malinarich et al. 2010; Chennupati et al. 2010; Vantourout and Hayday 2013). These T cells have tissue-specific homing based on distinct site-specific effector functions and are localized primarily to epithelial and mucosal surfaces but are present variably within other tissues (Vantourout and Hayday 2013). Although γδ T cells are part of the adaptive immune system, they can also be directly activated by pathogen-associated molecular patterns and danger-associated molecular patterns allowing for a rapid innate immunity-like response to pathogens or cell damage. These cells respond rapidly with the release of cytokines (e.g., IFN-γ, TNF-α, and IL-17) and chemokines, induce cytolysis of target cells via perforin and granzymes, and interact with other immune and epithelial cells. Although functionally similar between mice and humans (Table 5), the TCRγ and TCRδ loci are highly divergent, and orthologs between the 2 species are not comparable (Vantourout and Hayday 2013). In addition, the
Brief Comparison of Mouse and Human Gamma–delta (γδ) T Cells.a
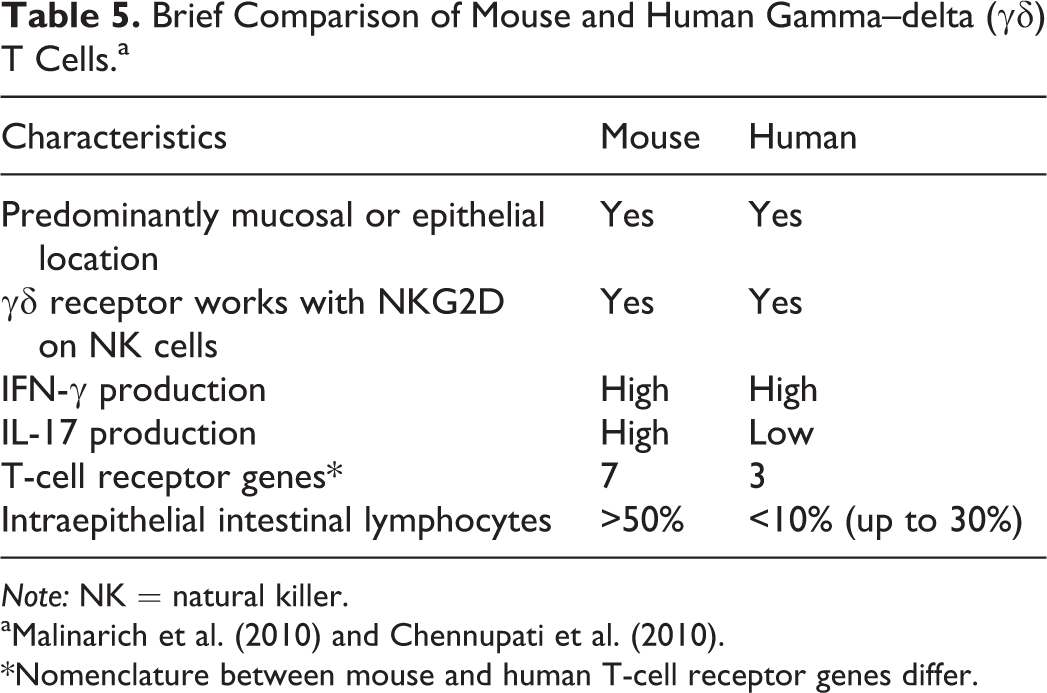
*Nomenclature between mouse and human T-cell receptor genes differ.
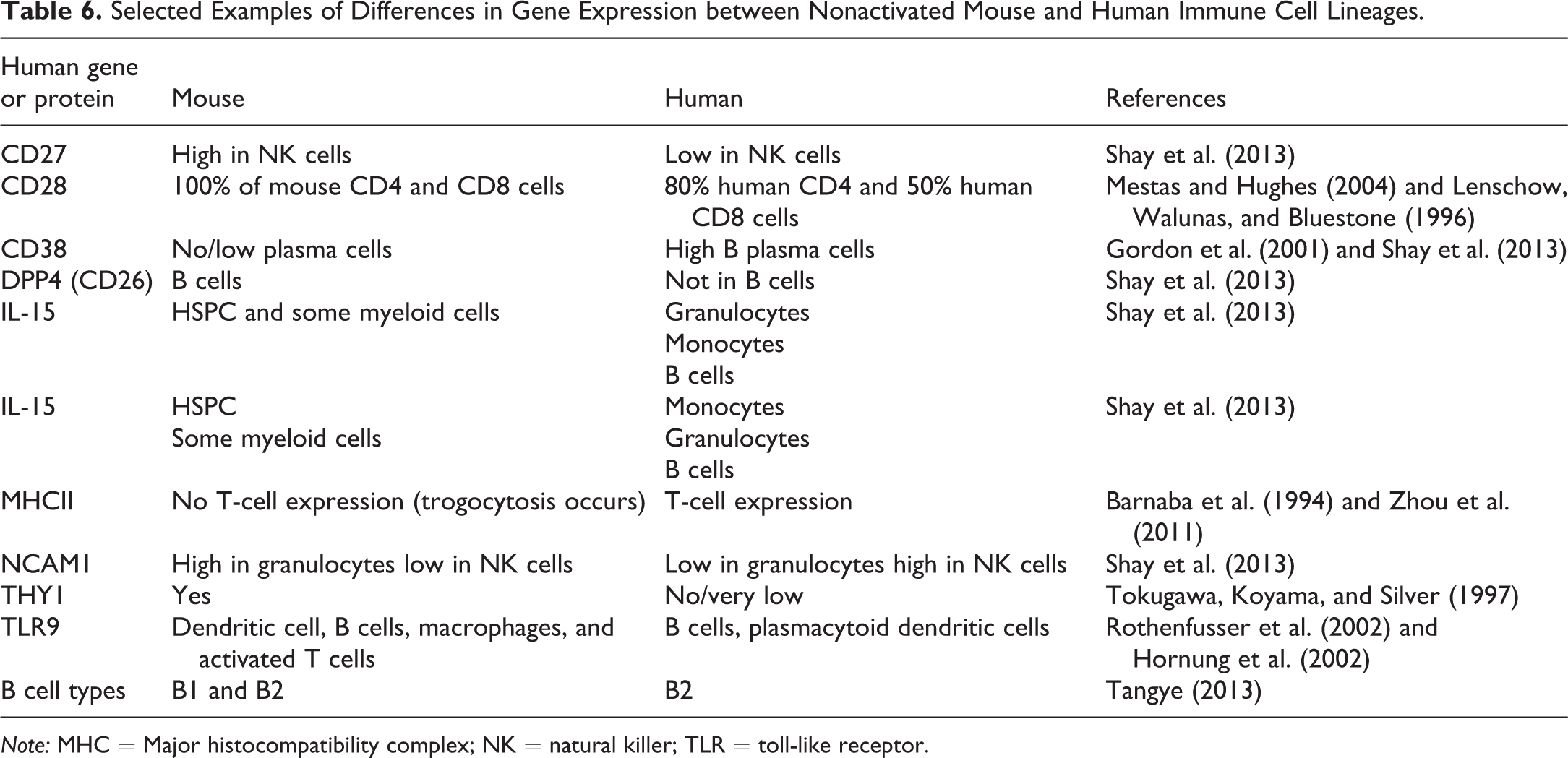
Recent work through the Immunological Genome Project (ImmGen; http://www.immgen.org) using genome-wide mRNA profiling for distinct immune cell lineages has allowed for detailed comparisons between mouse (C57BL/6J) and human immune cells and immune responses (Tables 6 and 7). Evaluation of these databases revealed that there is notable conservation in transcriptional profiles and lineage-specific gene expression between the species and conservation gene expression in activated T cells (Shay et al. 2013). Gene duplications of orthologous genes appear to have resulted in numerous paralogs in either species; these may function similarly to the orthologous gene but also may have an altered or new functions (Shay et al. 2013). In the immune regulators evaluated, seventy-six were expressed in mouse but not human common lineages. In addition, thirty-seven mouse immune regulators were not expressed in any human immune cell sample. Of particular interest were differences in early and late gene expression patterns in activated T cells. For example, activated human CD4 T cells have a more rapid and robust Th17 response than mice both in early (1–4 hr) and late (20–48 hr) time points. Activated human CD4 T cells also highly express
Selected Examples of Differences in Gene Expression between Nonactivated Mouse and Human Immune Cell Lineages.
Selected Examples of Divergence of Upregulated Gene Expression in Activated CD4+ T Cells between Mice and Humans.
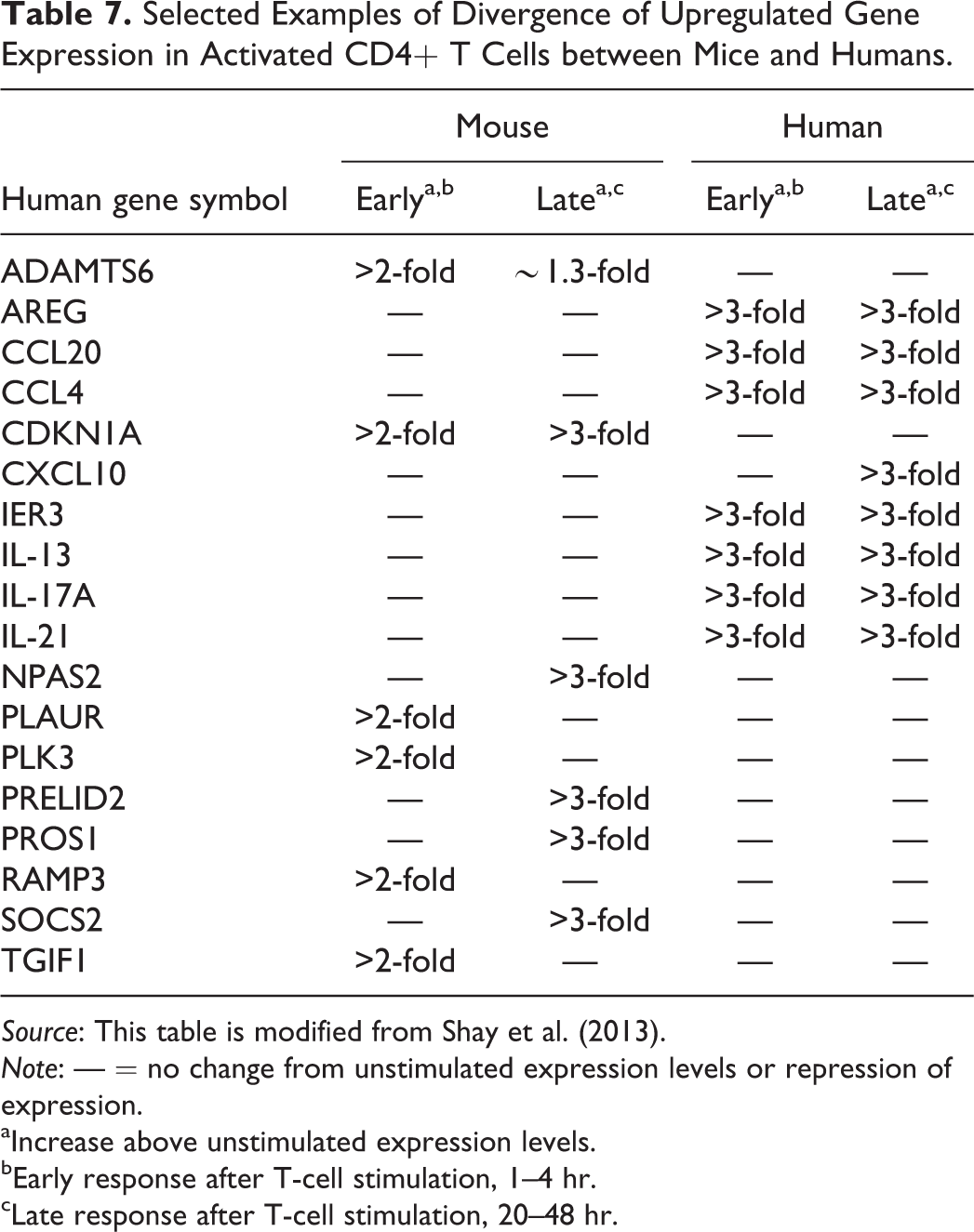
aIncrease above unstimulated expression levels.
bEarly response after T-cell stimulation, 1–4 hr.
cLate response after T-cell stimulation, 20–48 hr.
Hidden Confounders of Translational Research
An underaddressed factor in mouse model data reproducibility and comparability is related to husbandry. As mentioned earlier, it is common practice to use different mouse suppliers for the same strain even within a single lab as well as to use supplier sourced wild-type controls in GEM studies. Laboratory diets vary in content and sources of fat, carbohydrate, micronutrient, as well as amounts of phytoestrogens. These variations can have fairly rapid or direct effects (Barnard et al. 2009) such as inducing obesity in susceptible mice or, with increased phytoestrogens, alterations in reproductive function (Brown and Setchell 2001; Kudwa et al. 2007) and study results in evaluations of endocrine disruptors (Thigpen et al. 2013; Camacho et al. 2016). These dietary variations may also contribute to longer-term multigenerational effects. For example, high-fat diets have been demonstrated to drive epigenetic changes that are manifested for multiple generations (Seki et al. 2012; Williams et al. 2014). Frequently, investigators are not aware of diets being fed or potential impacts of diet options on their research (Jorgensen, Grunnet, and Quistorff 2015; Boots et al. 2016). Finally, the microbiome has been demonstrated to have a potentially tremendous impact on immunological studies (Stappenbeck and Virgin 2016; Goodrich et al. 2016). An example that has been well described is the requirement for
Conclusions
Mouse models are valuable for understanding the disease mechanisms and testing therapeutics, however, care must be taken when choosing or developing these models to ensure optimal translational value derived from studies using those models. A number of questions must be considered: (1) what aspect of the human disease is being addressed with the model? (2) does the model recapitulate the human immune response in the disease condition or in a particular stage of a disease? (3) is the mouse strain appropriate to model the immune question of interest? (4) how can mouse strain(s) help to answer the research question of interest? (5) would adding another mouse model give a fuller picture? and (6) how could the genetic background of the mouse model impact the study outcome? In addition, rigorous study parameters must be maintained and fully reported in order to meaningfully contribute to the scientific knowledge, comparability, and translatability derived from mouse studies (Kilkenny et al. 2010). Finally, it is essential for comparative pathologists and investigators to work together to design and interpret animal models of disease.
Footnotes
Acknowledgments
Many thanks to Cory Brayton, Paul Snyder, Piper Treuting, and Magali Guffroy for their contributions and support in the development and review of this article.
Author’s Contribution
The author (RS) contributed to conception or design; data acquisition, analysis, or interpretation; drafting the manuscript; and critically revising the manuscript. The author gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
