Abstract
Decades of research, including the 1996 Nobel Prize in Medicine, confirm the evolutionary and immunological importance of CD8 T lymphocytes (TCD8+) that target peptides bound by the highly variable major histocompatibility complex class I (MHC-I) proteins. However, their perceived importance has varied dramatically over the past decade. Regardless, there remains myriad reasons to consider the diversity of MHC-I alleles and the TCD8+ that target them as enormously important in infectious disease research. Thus, understanding these molecules in the best animal models of human disease could be a necessity for optimizing the translational potential of these models. Knowledge of macaque MHC has substantially improved their utility for modeling HIV and could aid in modeling other viruses as well, both in the context of distribution of alleles across treatment groups in vaccine trials and in deciphering mechanisms of immune control of pathogens for which specific MHC alleles demonstrate differential impacts on disease.
Keywords
Macaques, due primarily to their relative evolutionary closeness to humans, provide an ideal animal model for a number of infectious diseases. Analysis of the fully sequenced rhesus macaque genome suggests that humans and macaques diverged approximately 25 million years ago (Rhesus Macaque Genome Sequencing and Analysis Consortium et al. 2007). On average, the nucleotide sequence of the macaque genome is between 90% and 95% homologous to that of the human genome. The macaque immune system is likewise similar to that of humans and, with notable exceptions, responds similarly to a number of infectious agents. It stands to reason, then, that the underlying genetic architecture of the human and macaque immune systems must be largely overlapping. One of the most important genome loci involved in immunity to pathogens is the major histocompatibility complex (MHC). This region encodes the classical MHC-I and MHC-II genes as well as nonclassical MHC genes such as MHC-E, which has received much recent attention (Davis et al. 2016; Hansen et al. 2016; Pietra et al. 2010; van Meijgaarden et al. 2015) and others. It also contains genes that encode proteins involved in a number of other immunological processes, such as the transporter associated with antigen processing and tumor necrosis factor α (Horton et al. 2004; Shiina et al. 2009).
Each haplotype on the human MHC-I locus contains a single A, B, and C allele. In contrast, the MHC-I locus in macaques is astoundingly complex, with each individual haplotype expressing multiple A alleles and potentially more than 10 B alleles (Otting et al. 2005). However, the majority of these alleles are expressed at very low levels if at all, and the major alleles, those expressed at the highest level, are the ones most involved in immune responses to pathogens (Budde et al. 2011). Thus, in terms of functionality, the MHC-I region in macaques parallels that of humans more than it appears upon sequencing. Figure 1 depicts the MHC-I region from the human and macaque genomes.
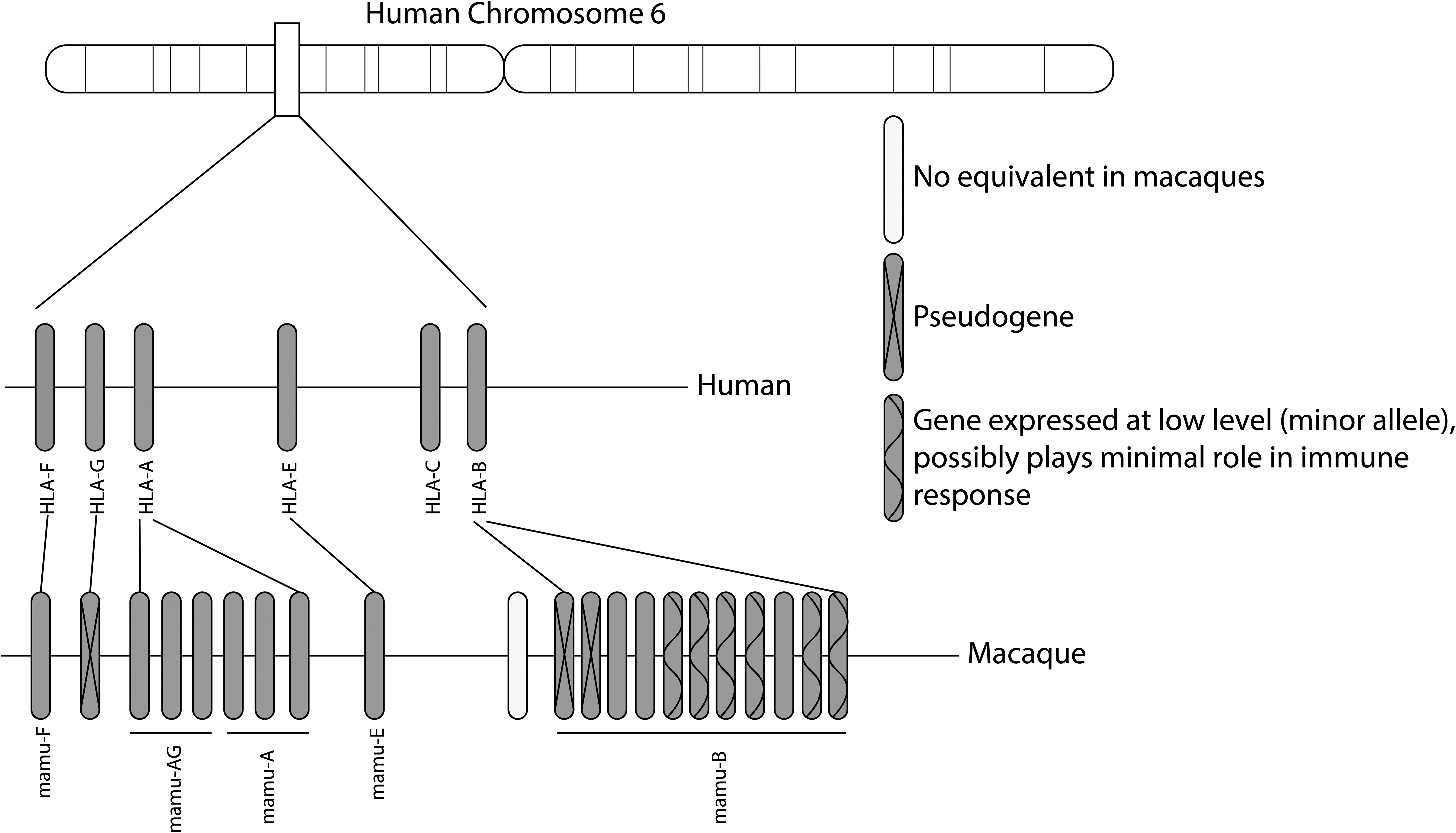
The organization of the major histocompatibility complex class I region in the human and macaque genome. Each human chromosome expresses a single A, B, C, E, and F allele. Macaques express equivalent E and F alleles, but the A and B loci have expanded resulting in expression of multiple alleles from each locus. However, many of these alleles are expressed at very low levels and are termed minor alleles. These minor alleles appear to play a minimal role in the antiviral immune response.
MHC-I molecules serve to present endogenously generated peptides to circulating CD8+ T lymphocytes (TCD8+). These peptides can be derived from self-proteins, including those expressed in an altered state during tumorigenesis, or from intracellular pathogens such as viruses. The MHC-I gene family is among the most polymorphic in the human genome. Given the predominant function of these molecules in peptide presentation, it is not surprising that the sequence differences that define different alleles are most commonly located in the peptide-binding region of the proteins, suggesting natural selection to maximize the ability to bind a diverse array of peptides from equally diverse pathogens (Hughes and Nei 1988).
Interestingly, a number of particular human MHC-I alleles are statistically correlated with enhanced capacity to control chronic viral infections, including HIV (Goulder and Watkins 2008) and hepatitis C virus (HCV; Neumann-Haefelin et al. 2010), although HCV shows stronger associations with MHC-II alleles (Scotto et al. 2003) and surprisingly with acute hantavirus infections (Torresilla et al. 2013). Likewise, particular MHC-I alleles in macaques infected with simian immunodeficiency virus (SIV) show strong correlations with significantly reduced viral load (Budde et al. 2012; Loffredo et al. 2007; O’Connor et al. 2010; Yant et al. 2006).
Understanding MHC-associated control of viral replication provides a framework for rational design of vaccines that induce TCD8+ cells. The ultimate goal of vaccines in general is to prevent infection at the port of entry, suggesting that neutralizing antibodies should be the most critical immunological component of a vaccine. However, one of the most effective vaccines ever created, the yellow fever vaccine 17D, induces a broad immunological response that includes antibodies and TCD8+ cells (Muyanja et al. 2014; Querec et al. 2009; Wieten et al. 2016), all of which likely contribute to viral prevention and clearance. In addition, a number of vaccines could be enhanced by induction of effective TCD8+ cells, which can target all viral proteins, rather than just the envelope glycoprotein, the predominant target of neutralizing antibodies. In the case of HIV, the envelope protein is highly variable, rendering the induction of broadly neutralizing antibodies, those that neutralize isolates of the virus that might be encountered in the real world, incredibly difficult. Thus, a vaccine that can target the conserved regions of other viral proteins would be of enormous benefit. Similarly, there is considerable interest in creation of vaccines that can target phylogenetically divergent but related viruses. Many related virus harbor substantial conservation in their capsid and polymerase proteins but lack such sequence conservation in their surface glycoproteins. Hence, a broadly effective vaccine in such a context must almost certainly induce TCD8+ cells.
Here, I argue for an integrative approach to understanding the mechanisms of immunological control of viral infections based on similarities between macaque and human genetic associations with such control. The most striking association between an MHC-I allele and control of HIV-1 replication is that of HLA-B27 (Goulder and Watkins 2008). HLA-B27 presents HIV-1-derived peptides that conform to a very specific sequence motif. Specifically, position 2 of the peptide is nearly always an arginine (R), and position 9 (or the C terminal residue) is most often a leucine (L). Interestingly, position 1 is very frequently an R or a K, similar acidic residues, which forms a diacidic motif that impacts peptide stability (Herberts et al. 2006). HLA-B27 is also associated with control of HCV (Neumann-Haefelin et al. 2010) and Puumala hantavirus (Mustonen et al. 1998), suggesting a generalized and highly effective mechanism of control of these very divergent viruses. Of note, the MHC-I allele Mamu-B*08, expressed by a subset of rhesus macaques of Indian origin, is associated with very strong control of SIV replication (Loffredo et al. 2007; Mudd et al. 2012a, 2012b). Mamu-B*08 and HLA-B27 have nearly identical peptide binding preferences (Loffredo et al. 2009). Together, these data suggest that TCD8+ responses that target virus-derived peptides bound by these MHC-I molecules are uniquely potent at restricting viral replication. Intriguingly, primary isolates of hantaviruses induce strikingly similar patterns of disease in macaques that they do in humans, suggesting an ideal animal model for identifying correlates of TCD8+ mediated control of viral replication across multiple viruses and multiple species (human and macaque). One could envision a study wherein macaques that express or do not express Mamu-B*08 are infected with these hantaviruses and their capacity to control the virus compared. Immunological parameters of the antiviral TCD8+ could be compared in a systems approach, including transcriptomics and cytokine production upon antigen encounter. These data could be compared between macaques infected with SIV and Puumala virus and humans with HIV-1 and Puumala virus. Such a data set could suggest immunological mechanisms of control.
TCD8+ cells are clearly important in control of viral infections, but they are often overlooked in the context of vaccines. I would argue that we simply have yet to identify the most critical aspects of an effective TCD8+ response. At the heart of such an examination is an in-depth understanding of the MHC-I allelic repertoire expressed by both humans and the animal models used to model particular infectious diseases and the peptides they produce.
Footnotes
Acknowledgment
This work is dedicated to the memory of Dr. Austin Hughes whose work in the evolution of MHC inspired the author to pursue a career studying how MHC diversity impacts immune responses.
Author Contributions
All authors (NM) contributed to conception or design; data acquisition, analysis, or interpretation; drafting the manuscript; and critically revising the manuscript. All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
