Abstract
Attempts to characterize and formally qualify biomarkers for regulatory purposes have raised questions about how histological and histopathological methods impact the evaluation of biomarker performance. A group of pathologists was asked to analyze digitized images prepared from rodent kidney injury experiments in studies designed to investigate sources of variability in histopathology evaluations. Study A maximized variability by using samples from diverse studies and providing minimal guidance, contextual information, or opportunities for pathologist interaction. Study B was designed to limit interpathologist variability by using more uniform image sets from different locations within the same kidneys and allowing pathologist selected interactions to discuss and identify the location and injury to be evaluated but without providing a lexicon or peer review. Results from this study suggest that differences between pathologists and across models of disease are the largest sources of variability in evaluations and that blind evaluations do not generally make a significant difference. Results of this study generally align with recommendations from both industry and the U.S. Food and Drug Administration and should inform future studies examining the effects of common lexicons and scoring criteria, peer review, and blind evaluations in the context of biomarker performance assessment.
Introduction
Novel biomarkers of tissue injury have been identified and subsequently qualified by the U.S. Food and Drug Administration (FDA) for assessing preclinical safety within a well-defined context of use (Dieterle et al. 2010). Efforts to evaluate the clinical utility of some of these biomarkers are currently underway (Koyner and Parikh 2013). The search for new biomarkers continues, and some newer candidates, such as microRNAs (Rouse, Rosenzweig, and Thompson 2014), have been shown to reflect mild reversible injury at very early times. In some situations, researchers have proposed that biomarkers may even be providing predictive signals of disease or injury before changes actually occur (Ozer and Teitelbaum 2009). This conclusion assumes that the methods used to assess biomarker performance are robust and truly represent what is happening in the organ of interest (Shea, Stewart, and Rouse 2014).
Light microscopy–based histopathology functioned as the reference standard for identifying true positive and true negative cases and anchoring the assessment of the novel nephrotoxicity biomarkers submitted to the FDA’s Center for Drug Evaluation and Research as part of its formal drug development tool qualification process. During the qualification review, concerns were raised about the potential for histopathology practices to influence biomarker assessment and the interpretation of biomarker data. These concerns centered on the sensitivity of light microscopy methods, the adequacy of sampling methods, possible knowledge bias in histopathology evaluation, and potential variability among pathologists or groups of pathologists. These concerns would be compounded if the biomarkers were proposed for use in predicting injury prior to the observation of tissue pathology. If anchored to histopathology, a biomarker change detected prior to an observation of changes in tissue morphology would appear to be nonspecific, although in reality it might be very specific in addition to being highly sensitive. In cases where the proposed context of biomarker use is the early prediction of toxicity (Ozer 2010), the metrics used to assess biomarker performance need to be carefully evaluated.
The need for consistency and standards in the biomarker review process has resulted in a “best practices” document generated by members of the Society of Toxicologic Pathology (STP) for the conduct of histopathology analyses in biomarker studies (Burkhardt et al. 2011) as well as an FDA issued Draft Guidance for Industry entitled “Use of Histology in Biomarker Qualification Studies” (U. S. FDA 2011). However, few objective data exist that assess the potential impact of histopathology methods on biomarker evaluation. To address this gap, a series of studies were conducted to assess how histopathology methodology may impact biomarker evaluation. The following 2 critical issues were considered in designing these studies: (1) the traditional paradigm of recording “changes above control” may not work well for biomarker evaluation because spontaneous or “control” changes may be associated with biomarker change, and (2) the methodology used to assess biomarkers should, like the biomarkers themselves, be “fit for purpose” and not consist of inflexible standards applied across all applications. This study was not designed and does not presume to define a preference or even alternatives for pathologists’ handling of background lesions, nor does it begin to define various fit for purpose methodologies for biomarker assessment. The more elementary goal of this work was to demonstrate and quantify the premise that if left unaddressed, different pathologists’ interpretations (including those of background changes) can significantly alter biomarker performance assessment. Further, the study hoped to demonstrate that alternative methods, such as blind and open evaluations, might each have circumstances in which it is the most fit for purpose.
Materials and Methods
Histopathology Assessment
Three pathologists employed by the FDA provided independent evaluations for this project. Pathologists were of different tenures and varied career paths, but all pathologists were experienced in the evaluation of a wide range of nonclinical toxicology and drug safety studies and the microscopic identification of morphologic changes in animal tissues associated with toxicity in multiple organs. The work of the pathologists was divided into 2 large tasks that are illustrated in Figure 1.

Depiction of the experimental design including study A and study B. Study A evaluated 4 image sets each containing 23 to 27 images created from slides generated in four separate experiments that resulted in kidney injury. The experiments yielded kidney injury from gentamicin exposure, diabetes with or without contrast media, hypertension with or without contrast media, or salt-induced nephropathy with or without contrast media. Evaluation conditions for study A were controlled to maximize interpathologist variability. Study B evaluated 5 image sets containing 17 to 20 images created from slides generated from the same 20 kidneys representing 20 different animals injured in a cisplatin study. Each image set contained an image from a different location within the kidney and only 1 image from each kidney; some image sets did not have an image from all 20 kidneys. Evaluation conditions from study B were controlled to reduce interpathologist variability compared to study A.
1. Study A was designed to allow maximal interpathologist variability in blind (to treatment vs. control animals, dose group, and biomarker data) and nonblind (to treatment and dose group) evaluations of “diverse” histopathology image sets: a digital image set representing both cross sections and longitudinal sections was generated from kidneys sampled from male rats from each of the 4 different experiments in which kidney injury was produced by 4 diverse etiologies, namely, (1) gentamicin toxicity in Sprague-Dawley [(SD)] rats, (2) streptozotocin-induced diabetic nephropathy in SD rats with and without contrast media, (3) hypertension-related kidney injury in spontaneous hypertensive rats [(SHRs)] with and without contrast media, and (4) nephropathy in Dahl salt-sensitive [(DSS)] rats with and without contrast media. Image sets 1 and 4 incorporated a wider range and a more severe degree of injury. Image set 2 was intermediate, while image set 3 exhibited minimal changes. For each set of test images, the pathologists were provided a set of “training images” that represented the range of injuries that would be seen in the test set. However, the training images were not labeled as to severity and a prestudy standard was not created to gauge the accuracy of the pathologists in the study. The specific range of injuries typical of background changes and of various disease model strains and treatments was different from one image set to the next in study A and included tubular injury, glomerular injury, and vascular injury. The range of severity varied greatly from one image set to the next. For blind evaluations, only the sex of rats (all male) was provided for training sets. For open evaluations, rat strain, treatment, dosage, and disease model were also provided with training set images.
The pathologists were simply asked to “rank” the slides based on the severity of injury. Consequently, each pathologist had to identify the range of injury contained within the training set, according to a scale defined by their career training and life experiences. No lexicon, consensus scoring criteria, or opportunity for interaction among the pathologists was permitted for this task. Images available for study A were limited to those available from retrospective studies; therefore, training images were incorporated into test sets. Although differences in agreement on these images within the test set were not noted relative to the nontraining set images, the authors must point out that some bias could have been introduced by the inclusion of these test images especially for intrapathologist variability. Intuitively, the anticipated bias would have been toward increased agreement, which was very high across all images. Subsequently, high intrapathologist agreement in study B would suggest that this may not have been a significant bias. Image sets were made simultaneously available to pathologists for 2 to 4 weeks, one set at a time. Each set was evaluated 3 times, with no less than a 6-week washout period between the analyses of the same image set. The first and the final evaluation were both done blind to treatment, disease model, dosage, and rat strain. The second or middle evaluation was done with the knowledge of rat strain, treatment, dosage, and disease model. In each evaluation, the pathologists were asked to use their general knowledge and experience to score kidney injury on a 0 to 5 whole number interval scale across the digital images within a set, with 5 equaling the most profound changes conceivable (and not necessarily the most profound changes observed within the image set). Specific diagnosis or diagnoses were not requested, but pathologists were asked to use their knowledge and experience to score morphological changes beyond their individual interpretation of “normal” in the given image set. No instructions were given as to how to handle “background” or spontaneous changes since in each image set control animals were not identified for 2 of the 3 evaluations. Each pathologist determined individually how they would use their interpretation of background changes in their analysis. 2. Study B was designed to limit the variability through blind and nonblind evaluations of image sets created from a single cisplatin-induced renal injury experiment. The pathologists evaluated 5 image sets of kidney obtained from rats treated with varying doses of cisplatin or with saline. For this evaluation, pathologists were asked to meet as a group to identify and discuss the type and location of kidney injury induced by cisplatin treatment in rats. The goal of this interaction was to provide each pathologist with a loose framework of information to test whether this fact would minimize variability between pathologists. The intent was to create a common thought about treatment-related changes that would promote harmonization in scoring between pathologists. During the meeting as a group, pathologists looked at glass slides, starting with a training set of histological sections from the study to gauge their analysis to the specific morphological changes associated with cisplatin toxicity and to discuss the characteristics of the lesion location and severity. Changes ranged from minimal to marked tubular epithelial cell degeneration/necrosis at the corticomedullary junction (S3 or P3 segment) in treated rats. Once familiar with the nuanced differences between the images and slides, the pathologists evaluated digital images of the training slides as a group, to determine whether the digital images would yield a similar assessment as direct evaluation of the glass slide. Ultimately, the pathologists were satisfied that digital images would allow an adequate assessment for the purposes of this study. These training slides and images were not included in future test sets. Subsequently, pathologists who participated in the group meeting worked independently, assessing each digital image of cisplatin-treated rats, providing lesion scores ranging from 0 to 5 on a whole number interval scale. Pathologists were asked to provide this assessment based on their training and experience including that obtained in pathologist meetings conducted as part of study B. Although blind to other information, pathologists were aware that cisplatin was the agent used. Pathologists were, however, unaware of the fact that the 5 image sets were from the same kidneys sampled at different locations within the organ. Each image set was evaluated twice, once blind to biomarkers and dose groups and once with knowledge of dose groups but blind to biomarker data. Again, a minimal 6-week washout period was observed.
Providing an adequate period between evaluations was considered an important part of the design. However, there is little hard data as to what period would be adequate. Although it seemed unlikely that the pathologists would not recognize the injury reflected in the images, the authors felt that an adequate washout would help ensure that the pathologists did not identify the exact metadata of previously viewed images and thereby potentially bias their scoring process. The 6-week washout was selected as adequate to limit the recall of exact images after consultation with nonassociated pathologists.
Animal Experiments
All animal procedures were performed in accordance with the U.S. Public Health Service Guide for the Care and Use of Laboratory Animals (IACUC protocol WOAP 2009-100) in an Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC)-accredited facility (PHS Assurance #A4300-01). Male SD rats (Harlan Laboratories, Frederick, MD) were housed in standard plastic caging and maintained with food and water ad libitum on a 12-hr light–dark cycle at 22°C to 24°C.
Rats were divided into 5 cohorts of 6 rats each and 1 group containing 3 rats. On day 0, the cohorts containing 6 rats received a single intraperitoneal injection of saline or 0.5, 1.0, 2.0, 3.0, or 5.0 mg/kg cisplatin (Sigma, St. Louis, MO). Based on the changes seen in pilot studies, 3 days posttreatment was selected for sacrifice because the intent was to detect early tissue changes. At this time point, the proximal tubular epithelial cells of some treated rats demonstrated mild to moderate changes, ranging from degenerative vacuolization to acute necrosis, primarily in the corticomedullary junction and the outer medulla, particularly at higher doses. However, based on light microscopy, it was observed that many treated rats had no changes in kidney morphology. The changes detected in rats were consistent with previous reports in type, location, and degree (Gautier et al. 2010; Vinken et al. 2012; Wadey et al. 2013). Urine for urinary biomarker measurement was collected from rats placed in individual metabolism cages on a refrigerated rack (Tecniplast, Buguggiate, Italy) for approximately 16 hr immediately prior to euthanasia. Rats were anesthetized via isoflurane inhalation 72 hr following intraperitoneal cisplatin injection and euthanized by exsanguination, with serum harvesting for chemistry analysis. In an attempt to minimize artifacts and promote evaluation consistency, perfusion fixation via the abdominal aorta with freshly mixed 4% paraformaldehyde/0.1% glutaraldehyde solution was completed immediately following euthanasia. Both kidneys were harvested with one half of each kidney being placed in formalin for routine histology processing and hematoxylin and eosin (H&E) staining. The other half of each kidney was separately retained for stereology processing and evaluation, which was described in more detail in an earlier publication (Shea, Stewart, and Rouse 2014).
Urinary Biomarkers
Prior to and independent of histopathology review, the urinary concentrations of 7 kidney injuries associated with biomarkers were determined using commercially available multiplex assays for the MesoScale Discovery Platform (Meso Scale Discovery, Gaithersburg, MD). Urine was thawed, centrifuged, and aliquoted for storage at −80°C until analyzed. Kidney injury molecule-1 (Kim-1), albumin (Alb), lipocalin-2 (Lcn-2), clusterin (Clu), osteopontin (Opn), mu and alpha glutathione S-transferases (μGst and αGst), and renal papillary antigen-1 (Rpa-1) were quantified in urine on the electrochemiluminescence platform using manufacturer’s instructions. Two urinary biomarkers, Kim-1 and αGst, demonstrated significantly increased urinary concentrations following cisplatin treatment. In this study, Kim-1 was more strongly correlated to preliminary histopathology than αGst. Further, in a previous kidney injury biomarker study, Kim-1 also demonstrated a more sensitive, consistent, and reliable response to proximal tubular injury (Rouse et al. 2011) and displayed superior immunohistochemical (IHC) staining significant to quantitative studies in these samples (Shea, Stewart, and Rouse 2014). Based on these findings, Kim-1 was chosen as the urinary biomarker to be assessed using the various qualitative methods described in this study and quantitative methods reported previously (Shea, Stewart, and Rouse 2014).
Histopathology Processing
Description of tissue processing for this project has been previously published (Shea, Stewart, and Rouse 2014). Briefly, tissues were shipped in formalin to Experimental Pathology Laboratories (EPL, Sterling, VA) where they were processed, cut into 3-µm sections with 2 to 4 serial sections being H&E stained every 600 µm (step sectioning) until the paraffin tissue block was exhausted. Therefore, not all resulting digital images (e.g., pole sections) contained all renal regions—cortex, medulla, and papilla. Stained slides containing longitudinal and transverse sections were returned. Subsequently, digital images were generated by scanning whole slides at 40× using the Aperio ScanScope CS slide scanner (Vista, CA). These images were then available for evaluation in Imagescope by Aperio (Vista, CA), allowing localization and high-resolution zooming throughout the image.
Digital Image Sets
In study A, a digital image set was created from the standard H&E-stained study slides (central longitudinal and transverse sections for each animal) from each of the 4 separate experiments designed to evaluate Kim-1 in acute kidney injury. All slides from each of the experiments were included if the corresponding Kim-1 data were available for the animal. Typically, controls represented approximately one-fourth of the total slides and subsequently of the images comprising an image set. Images were viewable on ImageScope from 1× to 40×.
In study B, 5 digital image sets were created from step (600 µm) sections from 20 kidneys. Each image set contained an image from a different location (step section) within each of the kidneys. Kidneys were selected for inclusion based on equal representation of cisplatin dose and on the availability of urinary Kim-1 concentration data. Sacrifice of rats 3 days posttreatment and prior to full development of cisplatin toxicity was purposeful, which resulted in rats exposed to all doses that did not exhibit morphological change or Kim-1 changes.
Pathologist Qualifications
Initially, a much larger and diverse group of pathologists were envisioned as the evaluators for these studies. However, the logistics of forming and maintaining this group were beyond the resources of this preliminary work. Therefore, pathologist selection was based on FDA employment in a pathology role, documented training in pathology (doctor of veterinary medicine [DVM] or doctor of medicine [MD]), 15 or more years of experience in toxicologic and/or investigative pathology, willingness to volunteer services, and supervisory approval to participate.
Statistics
Contributions to variability in image set scoring were determined using PROC GENMOD in SAS (SAS, Cary, NC) and/or LogXact for sparse data (Cytel, Cambridge, MA). All models used to determine significant impact on variability investigated interaction terms, and the majority of them were not statistically significant. To simplify interpretations, results presented in this study ignored these interaction terms. The contributors to variability examined in studies A and B were pathologist, blind/open evaluation, and treatment.
Agreement represents the percentage agreement based on every opportunity for any 2 evaluators to agree. Concordance is a measure of interrater agreement that also accounts for random agreement (Brennan and Prediger 1981) and is represented by a value designated as the κ statistic. Concordance in this study based on Randolph’s free-marginal κ values (Randolph 2005) was derived from Randolph’s 2008 Online Kappa Calculator (http://justusrandolph.net/kappa/). The κ values range from −1.0 to 1.0 and represent a chance-adjusted measure of agreement, where 0 is the same as chance, 1.0 is complete agreement, and −1.0 is complete disagreement. A percentage agreement value is the κ statistic without an adjustment for the probability of random agreement. Multiple versions of the κ statistic exist (Warrens 2010) and the utility of the κ statistic is influenced by the nature of the data (Sim and Wright 2005). Therefore, interpretation of κ values is also circumstantially variable. However, the calculator used for this study indicates that a κ score ≥0.70 would be considered adequate agreement. Percentage agreement and concordance (as κ statistic) were selected as one of the few quantitative methods available and commonly used to assess interrater agreement.
Receiver–operating characteristic (ROC) analysis (SigmaPlot 12.5, San Jose, CA) was used to comparatively assess evaluations in the context of the urinary biomarker, Kim-1. This analysis compares assay performance by generating an area under the curve (AUC) described by plotting the assay’s sensitivity (y axis) against 1 − specificity (x axis). Details of calculation and interpretation of the ROC analysis are beyond the scope of this article, although references are provided for those with more interest (Zweig and Campbell 1993; Fawcett 2006). True negatives and positives have to be defined by an accepted standard, in this case: histopathology evaluations. In this study, 0 scores were considered true negatives and anything other than 0 was considered true positives. Interpretation of this graph would be that the larger the AUC, the better predictive value of the biomarker. Significant deviation from pathologist to pathologist in histopathology evaluations might then be reflected by significant change in the performance of Kim-1 (AUC).
Results
Study A: Maximal Variability Study
In the first study, no standardized lexicon or criteria were provided prior to analysis, and the pathologists were asked not to confer before, between, or during the evaluations. Table 1 presents the degree of interpathologist agreement and the κ value for the evaluations (blind, open, and blind) in each of the image sets for all scoring and for present/absent scoring. Overall, these scores ranged between 16% and 48% agreement and κ between 0.00 and 0.37. Present/absent scores were segregated and examined independently of other scoring levels because these are the evaluations that would more significantly impact a definition of true positives (≥1) and true negatives (<1) in a typical ROC analysis, particularly for low threshold signals. Two evaluations in present/absent scoring had concordance values equal to 1.0 because there were no absent (0) scores in the evaluations. The remaining 10 κ statistics ranged between 0.28 and 0.94.
Study A: interpathologist agreement and concordance.
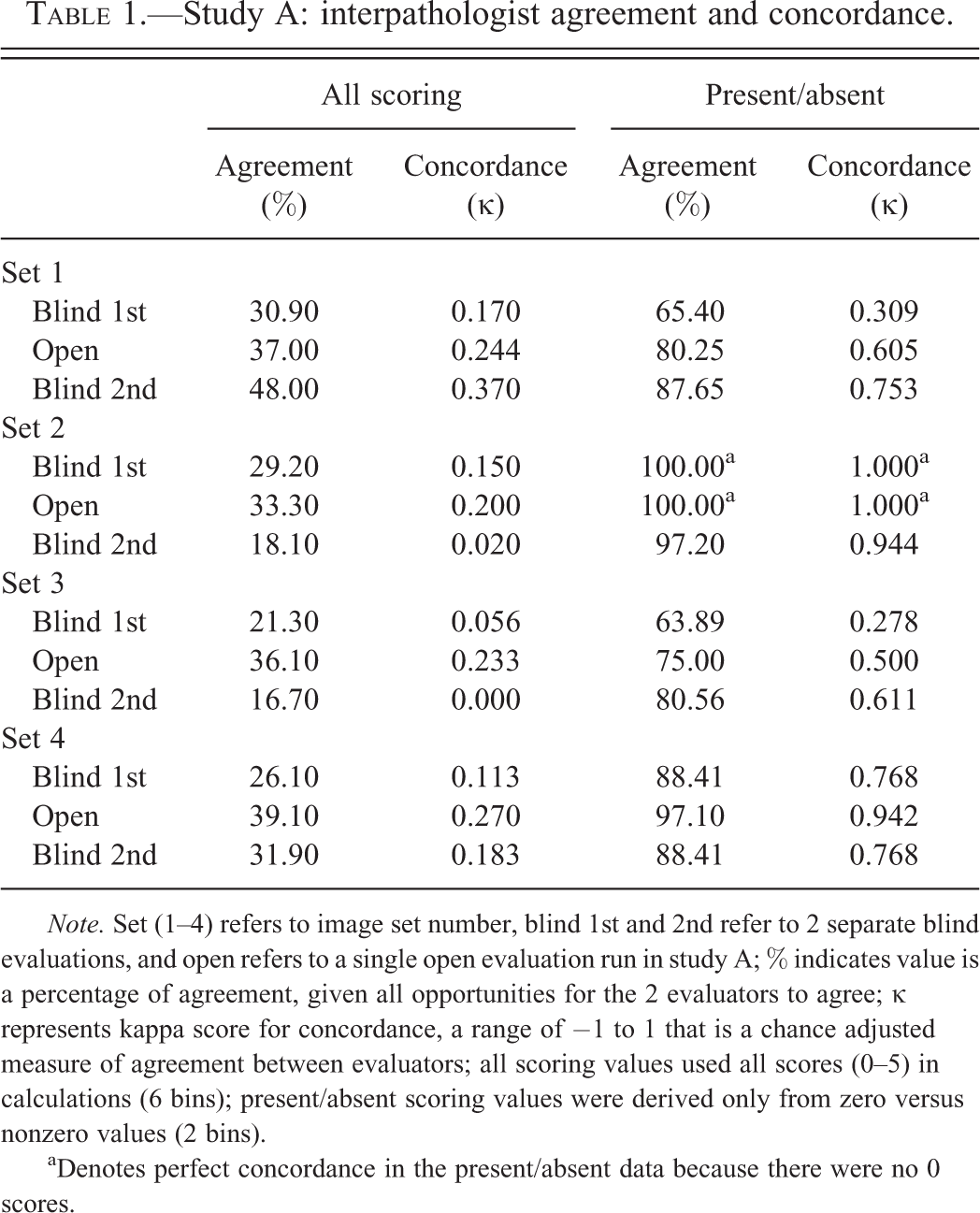
Note. Set (1–4) refers to image set number, blind 1st and 2nd refer to 2 separate blind evaluations, and open refers to a single open evaluation run in study A; % indicates value is a percentage of agreement, given all opportunities for the 2 evaluators to agree; κ represents kappa score for concordance, a range of −1 to 1 that is a chance adjusted measure of agreement between evaluators; all scoring values used all scores (0–5) in calculations (6 bins); present/absent scoring values were derived only from zero versus nonzero values (2 bins).
aDenotes perfect concordance in the present/absent data because there were no 0 scores.
Table 2 demonstrates intrapathologist (or self) agreement and concordance across all evaluations (blind, open, and blind) of the same image sets by the same evaluator. Values are shown for both all scores and present/absent scores. As might be anticipated, intrapathologist evaluations were in much higher agreement and concordance than interpathologist assessments. All scoring κ values ranged between 0.20 and 0.95. Again, concordance (κ ≥ 0.70) was not reached in intrapathologist evaluations for all scoring. In this case, present/absent κ scores of 1.0 were recorded in 4 evaluations because there were no 0 scores and the remaining 8 values ranged between 0.33 and 0.94, with a skew toward the upper values. In total, 10 of the 12 present/absent scoring concordance values were considered adequate. Table 3 depicts significant contributors to scoring variability for the evaluation of each image set by all pathologists. Treatment had a significant influence on scoring variability in 9 of the 12 evaluations, the pathologist was a significant contributor in 7 of the 12 evaluations, and blind evaluation was significant in 1 of the 8 evaluations.
Study A: intrapathologist agreement and concordance.
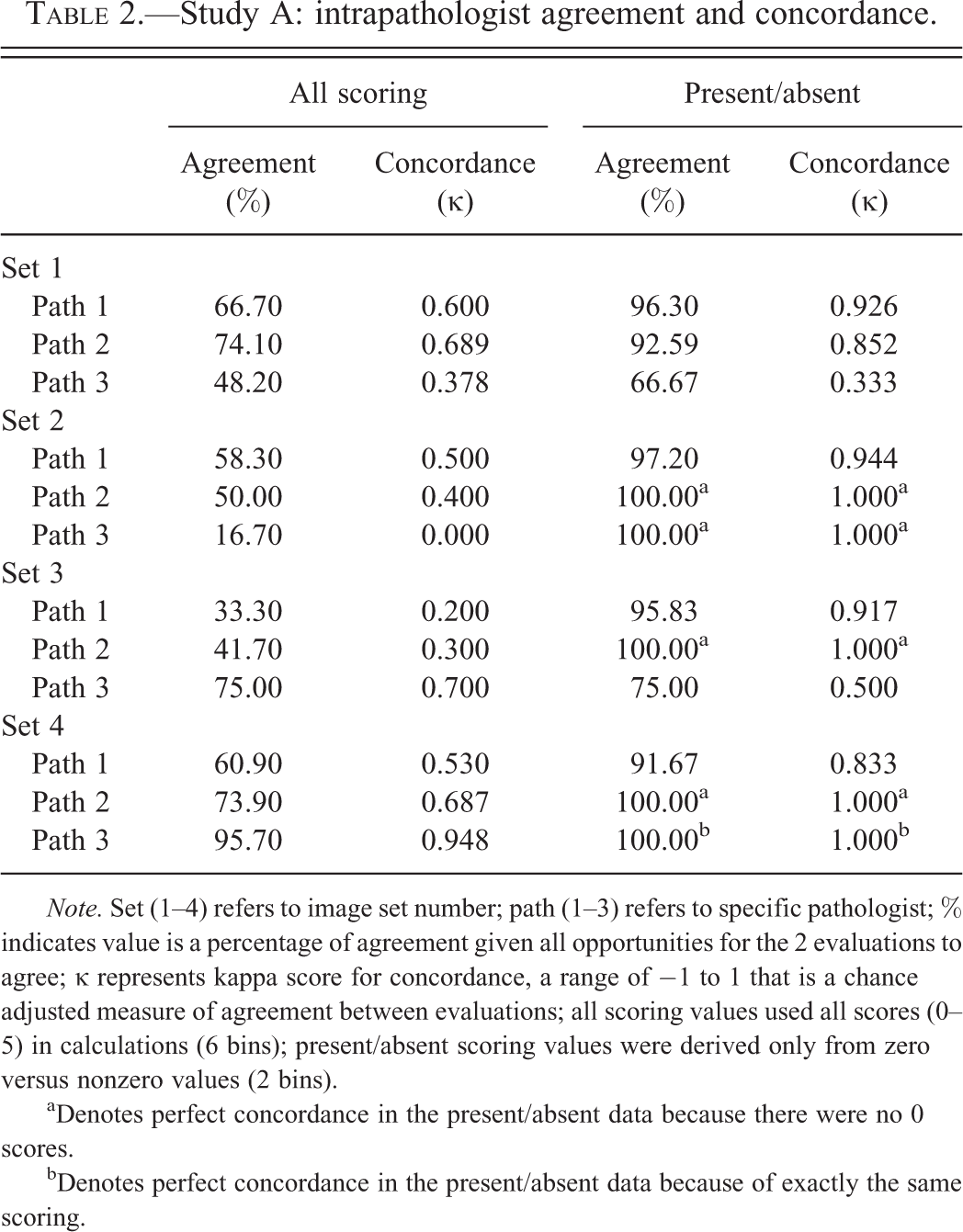
Note. Set (1–4) refers to image set number; path (1–3) refers to specific pathologist; % indicates value is a percentage of agreement given all opportunities for the 2 evaluations to agree; κ represents kappa score for concordance, a range of −1 to 1 that is a chance adjusted measure of agreement between evaluations; all scoring values used all scores (0–5) in calculations (6 bins); present/absent scoring values were derived only from zero versus nonzero values (2 bins).
aDenotes perfect concordance in the present/absent data because there were no 0 scores.
bDenotes perfect concordance in the present/absent data because of exactly the same scoring.
Study A: sources of variability.
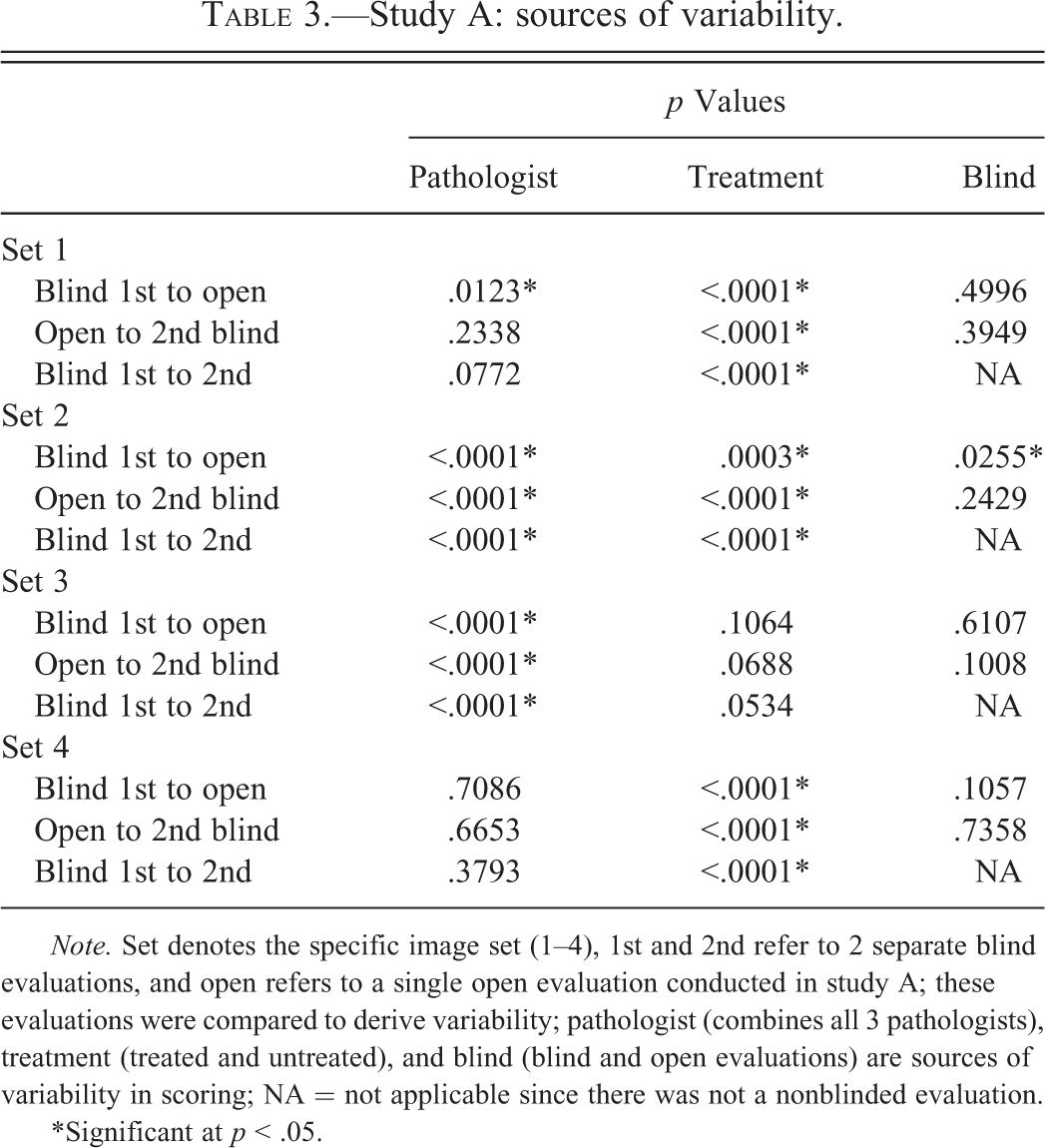
Note. Set denotes the specific image set (1–4), 1st and 2nd refer to 2 separate blind evaluations, and open refers to a single open evaluation conducted in study A; these evaluations were compared to derive variability; pathologist (combines all 3 pathologists), treatment (treated and untreated), and blind (blind and open evaluations) are sources of variability in scoring; NA = not applicable since there was not a nonblinded evaluation.
*Significant at p < .05.
Figure 2A shows the curves produced when each evaluator’s data were used in an ROC analysis of urinary Kim-1 for their first 2 evaluations (blind then open) of image set 1. ROC analysis was not performed with the open evaluation of pathologist 2 because true negatives were not recorded (there were no 0 scores). Likewise, either one or more of the pathologists reported no 0 values in evaluations of the remaining 3 image sets of this phase of the project. Thus, ROC analysis comparisons are not available for these image sets. Figure 2 presents results and illustrates the potential for very different biomarker interpretations based on the different evaluations by different pathologists, and AUC ranges from 0.68 to 0.90.
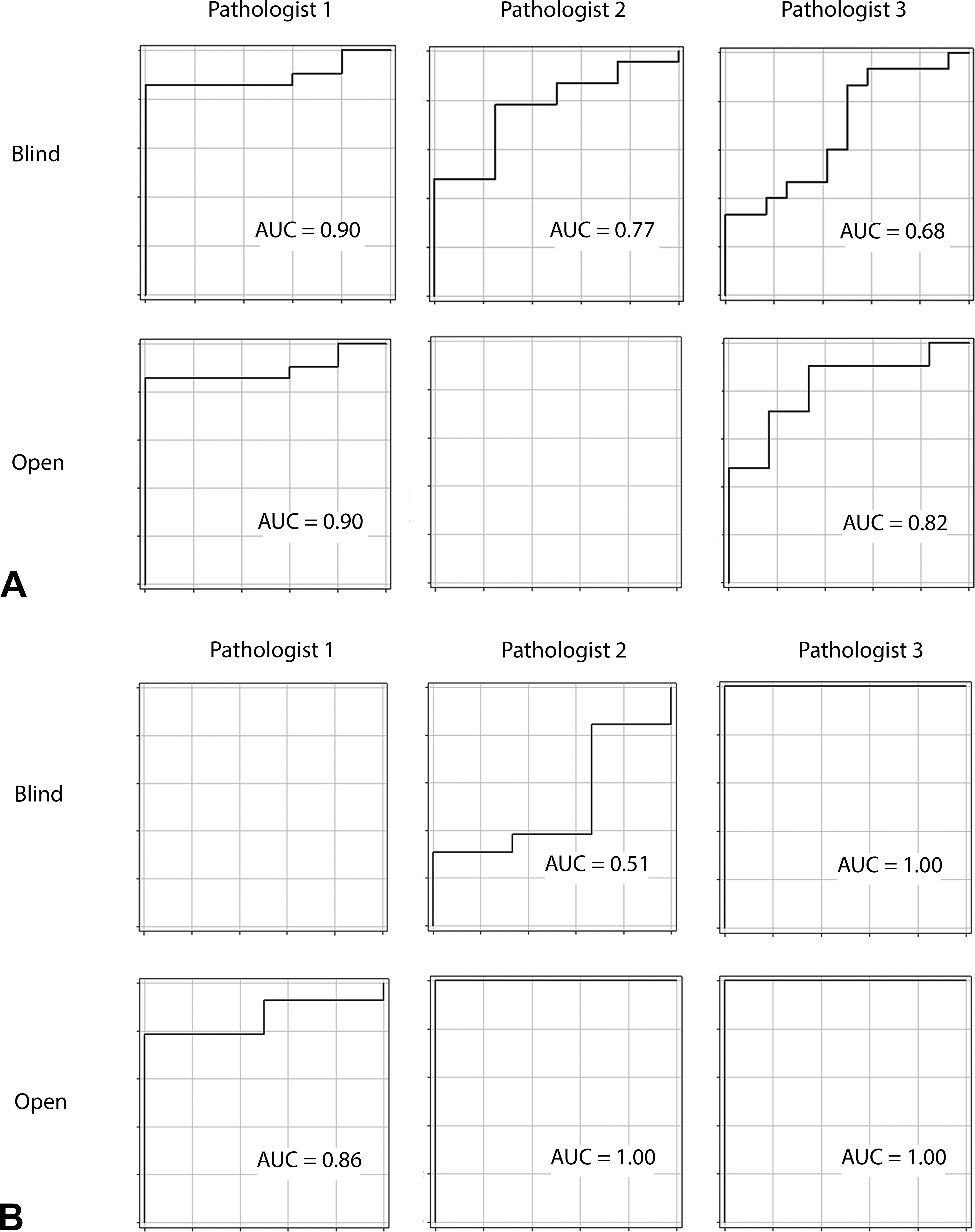
(A) Representation of receiver–operating characteristic (ROC) curves generated from study A, image set 1. In all graphs, y axis = sensitivity and x axis = 1 − specificity. Graphs with missing curves indicate an inability to complete the ROC analysis because no true negatives (0 scores) were identified. Although there were 2 blind evaluations, the first blind evaluation was the one used to compare to the open evaluation. (B) Representation of ROC curves generated from the study B, image set 1. Graphs missing curves indicate an inability to complete the ROC analysis because no true negatives (0 scores) were identified. Pathologist designation (1, 2, 3) was consistent throughout the study; Blind = evaluations done without knowledge of treatments or groups; Open = evaluations done with knowledge of treatments and groups (all evaluations were blind to biomarker data); AUC = area under the curve, providing a relative measure of assay performance based on histopathology-defined true positives and true negatives.
Study B: Limited Variability Study
In the second study, similar data were collected from image sets with more knowledge, as described in Materials and Methods section. Table 4 contains interpathologist agreement and concordance data from their blind and open reads of 5 separate image sets derived from the sample kidney step sections (one set from each of the 5 different locations within the same kidney). The κ values across all scoring, ranging between 0.02 and 0.66, were slightly higher in study B than those from study A. Present/absent κ values ranged between 0.00 and 0.60.
Study B: interpathologist agreement and concordance.
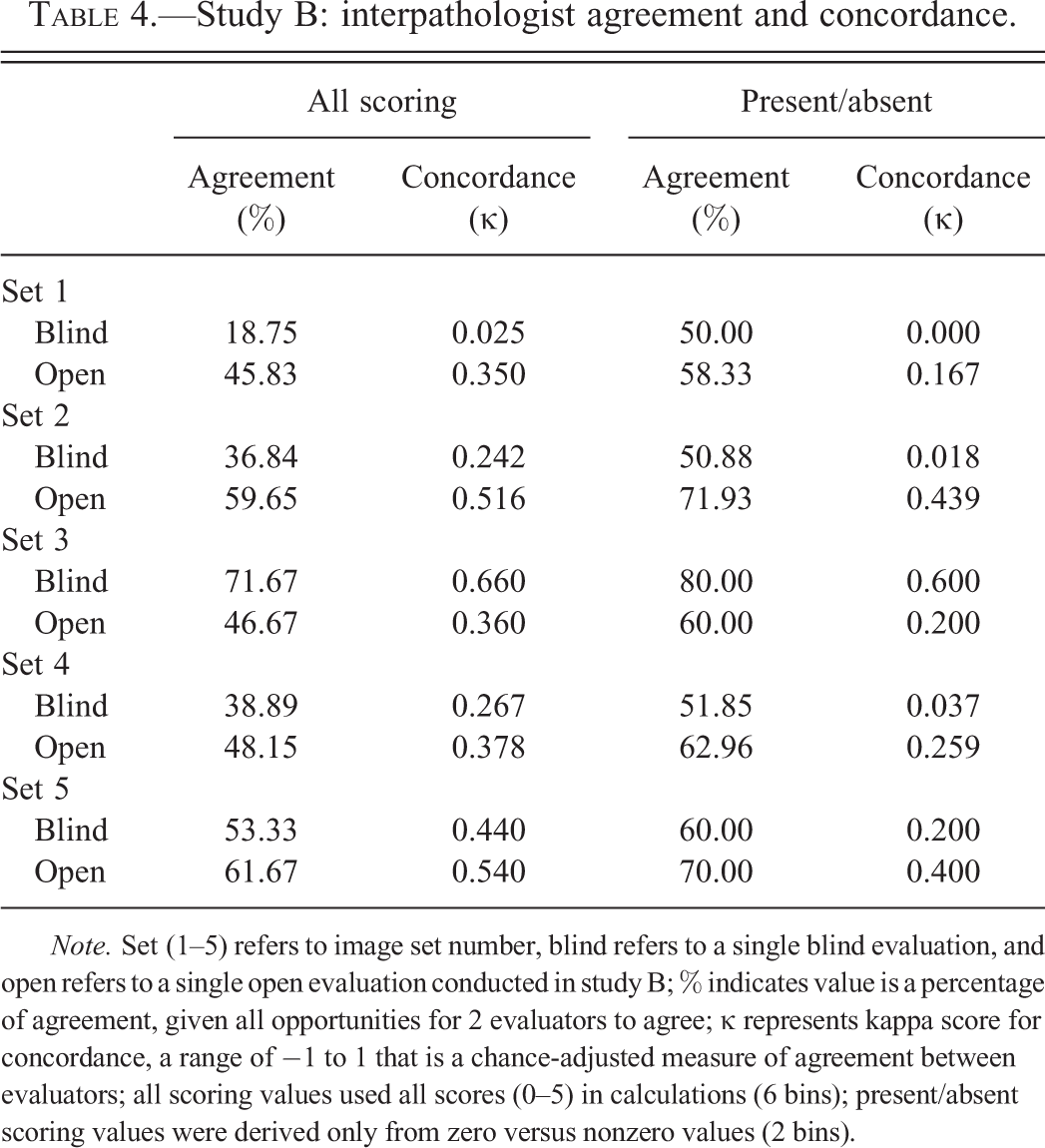
Note. Set (1–5) refers to image set number, blind refers to a single blind evaluation, and open refers to a single open evaluation conducted in study B; % indicates value is a percentage of agreement, given all opportunities for 2 evaluators to agree; κ represents kappa score for concordance, a range of −1 to 1 that is a chance-adjusted measure of agreement between evaluators; all scoring values used all scores (0–5) in calculations (6 bins); present/absent scoring values were derived only from zero versus nonzero values (2 bins).
Table 5 contains intrapathologist agreement and concordance data generated in study B. The table reveals that intrapathologist κ values between 0.05 and 1.00 were slightly higher than those described in study A. Table 6 shows the significance of pathologist, treatment, and blind evaluation as contributing factors to variability in study B of the project. In a single circumstance, blind evaluation was a significant source of variability in scoring. As might be expected, treatment was a consistent source of variability in all evaluations. The pathologist was also a significant source of variability in each evaluation.
Study B: intrapathologist agreement and concordance.
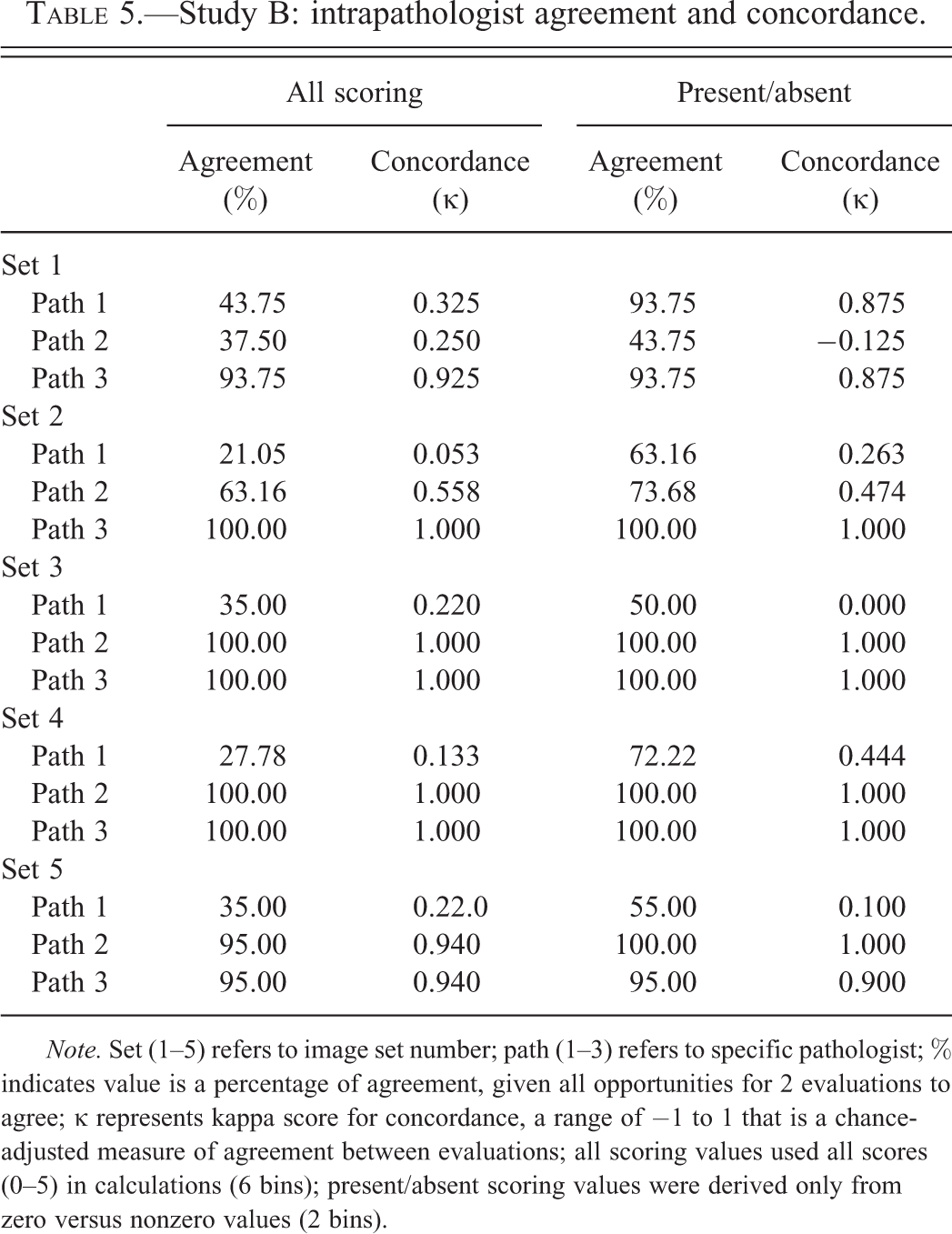
Note. Set (1–5) refers to image set number; path (1–3) refers to specific pathologist; % indicates value is a percentage of agreement, given all opportunities for 2 evaluations to agree; κ represents kappa score for concordance, a range of −1 to 1 that is a chance-adjusted measure of agreement between evaluations; all scoring values used all scores (0–5) in calculations (6 bins); present/absent scoring values were derived only from zero versus nonzero values (2 bins).
Study B: sources of variability.
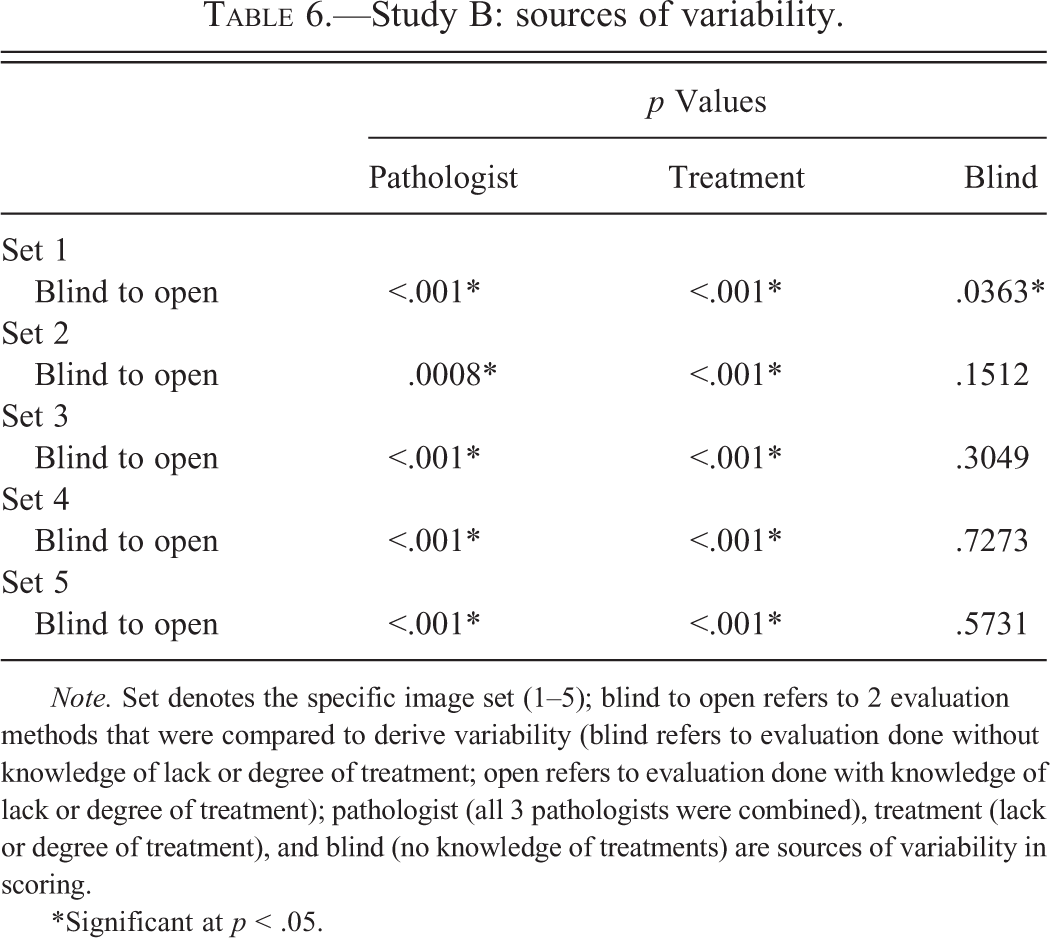
Note. Set denotes the specific image set (1–5); blind to open refers to 2 evaluation methods that were compared to derive variability (blind refers to evaluation done without knowledge of lack or degree of treatment; open refers to evaluation done with knowledge of lack or degree of treatment); pathologist (all 3 pathologists were combined), treatment (lack or degree of treatment), and blind (no knowledge of treatments) are sources of variability in scoring.
*Significant at p < .05.
Figures 2B, 3, and 4 depict ROC curves generated from study B that show improved consistency in biomarker performance compared to that seen in study A. But these curves also demonstrate sufficient inconsistency to alter the perception of biomarker performance (Note: difference in AUC for pathologist 1 vs. pathologists 2 and 3). Since the treatments in study B were the same for each image set evaluated, treatment as a predictor of histopathology was incorporated into the ROC analysis. Notably, for pathologist 1, treatment as a predictor of histopathology change improved greatly from the blind to the open evaluations for each image set and became the superior predictor in the open evaluations. For the other pathologists, treatment as a predictor of morphological change was generally consistent between the blind and the open evaluations.
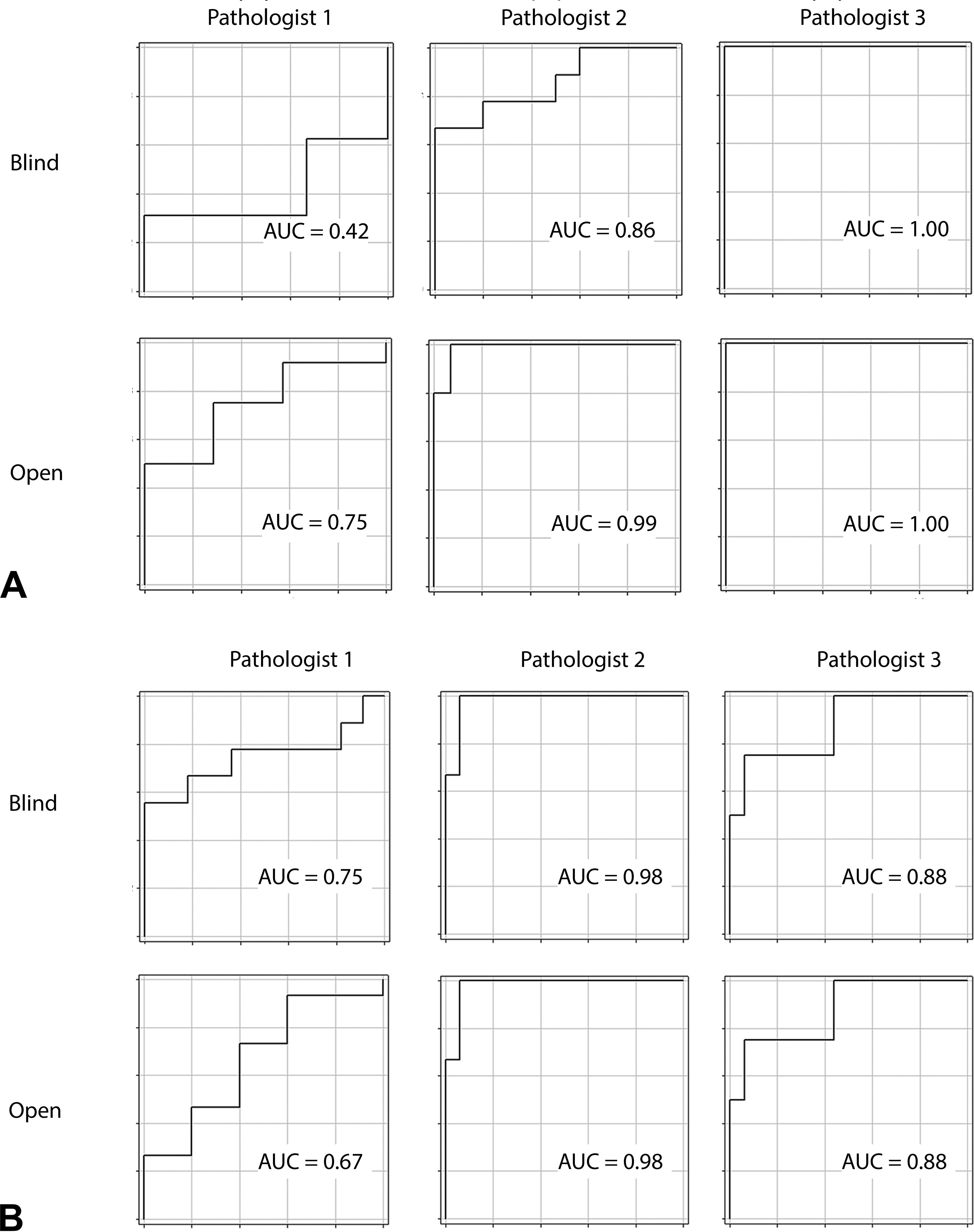
(A) Representation of receiver–operating characteristic (ROC) curves generated from the study B, image set 2. (B) Representation of ROC curves generated from the study B, image set 3. Pathologist designation (1, 2, 3) was consistent throughout the study; Blind = evaluations done without knowledge of treatments or groups; Open = evaluations done with knowledge of treatments and groups (all evaluations were blind to biomarker data); AUC = area under the curve, providing a relative measure of assay performance based on histopathology defined true positives and true negatives.
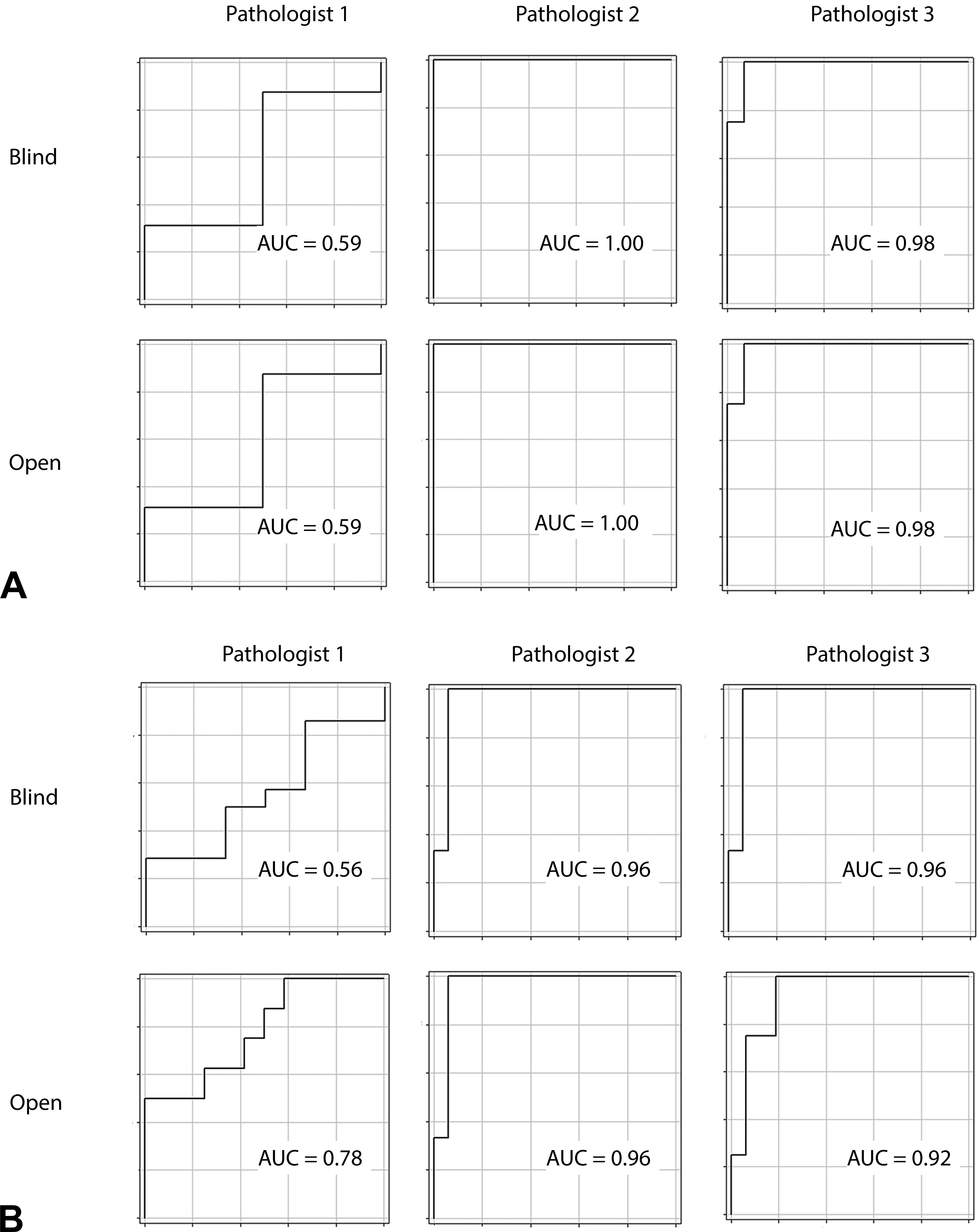
(A) Representation of receiver–operating characteristic (ROC) curves generated from the study B, image set 4. (B) Representation of ROC curves generated from the study B, image set 5. Pathologist designation (1, 2, 3) was consistent throughout the study; Blind = evaluations done without knowledge of treatments or groups; Open = evaluations done with knowledge of treatments and groups (all evaluations were blind to biomarker data); AUC = area under the curve, providing a relative measure of assay performance based on histopathology defined true positives and true negatives.
Discussion
These studies were intended to provide an estimate of the degree of interpathologist variability and the potential influence of knowledge bias (blind vs. nonblind) that might exist when assessing biomarker performance. Evaluations by different pathologists yielded different results in ROC analyses reflected by extremely different AUC values. In a biomarker assessment, this would translate to different interpretations of biomarker performance. The true concern in an ROC analysis, especially within the context of the experiments described in this article, is differentiation of 0, or true negative from true positive, represented by values greater than 0. Therefore, agreement and concordance analyses in this study were repeated using only changes to or from 0 values. In many circumstances, this binary evaluation might be the appropriate approach to biomarker evaluation. However, it must be recognized that not all definitions of true positive or true negative may be based on a strict binary (presence vs. absence) and that the subjective values attributed to spontaneous changes by different evaluators hold the potential to impact categorization as positive or negative. Further, in more complex and comprehensive evaluations, a true negative score might be defined statistically (Shea, Stewart, and Rouse 2014) and then binned into a binary determinant for ROC analysis. However, complete biomarker evaluation also captures the magnitude of biomarker response to the magnitude of morphological change and the temporal relationship between the biomarker and change (Vonderscher 2008). Thus, for biomarker assessment, concordance across all possible scores cannot be summarily dismissed to focus on present/absent scores. Although the interpretation of κ values is subject to debate and variability (as described in Materials and Methods section), the calculator used for this study uses κ ≥ 0.70 as an indicator of adequate concordance for medical diagnosis, a value that is consistent with the other references mentioned in Materials and Methods section.
In study A, none of the κ values denoted adequate concordance (κ ≥ 0.70) for consensus evaluation. Interpathologist concordance findings were improved when present/absent scores were the determinants, suggesting that pathologists agreed more consistently on injury versus noninjury than they did on the relative severity of injury. In total, 7 of the 12 present/absent κ values exceeded the 0.70 threshold for acceptable concordance. Intuitively, this seems a reasonable finding. Nevertheless, this was not a consistent finding even within this study. In study B, an acceptable level of interpathologist concordance was not achieved for either all scoring or present/absent scoring in any of the evaluations. Present/absent scoring yielded decreased interpathologist concordance relative to analyses that included gradients of injury. This was an unexpected finding, given the attempts to better harmonize scoring during study B and highlights the lack of agreement in what constituted injury even after the group discussion. This emphasizes the contribution of interpathologist differences to outcome in spite of efforts to minimize this source of variability. Adequate present/absent scoring intrapathologist concordance was recognized in 9 of the 15 evaluations. Intrapathologist present/absent scoring usually yielded increased concordance in both studies, indicating that pathologists were generally more consistent within themselves on their opinions of injury versus noninjury than they were with severity of injury. These findings demonstrate that interpathologist differences can play a large role in evaluation variability and that intrapathologist variability is of much less concern, although exceptions exist in support of knowledge bias.
Although inconsistent, ROC AUC values were generally higher based on open evaluations compared to blind evaluations. Comparing ROC analyses based on evaluations from different pathologists demonstrated that the differences in these evaluations are sufficient to impact the assessment of Kim-1 performance as an injury biomarker in the kidney. These findings are consistent with the STP best practices document (Burkhardt et al. 2011) that identifies interobserver variability as one of the largest sources of study variance. In study B, modifying methods to allow some common study and discussion of the specific injury to be evaluated provided some increase in concordance but did not align scoring between pathologists. Post-study interview of the pathologists suggests that within study B, this was a function of 1 pathologist identifying changes as “injury” that the other 2 pathologists considered background change. This negatively impacted agreement and concordance but is an occurrence that would typically be addressed by a common lexicon with detailed scoring criteria followed by peer review, a recommendation common to both the STP document and the FDA’s draft guidance.
In addition, this study showed that, as described in the STP best practices document, blind (to treatment and dose group) evaluation only infrequently contributed to significant image scoring variability. In study B, it was noted that scoring for pathologist 1 correlated strongly with treatment in open evaluations but did not in blinded evaluations. This did not imply improved accuracy as this scoring was independent of urinary biomarker concentrations to which all pathologists remained blinded throughout the entire experiment. This finding infers but does not prove a knowledge bias influence on scoring and stands out only because there was not a similar occurrence with the other 2 pathologists. However, this intrapathologist variability could arise from knowledge expansion, acquired experience, and confidence as well as other unrecognized factors. It was noted that open evaluations provided greater interpathologist concordance and agreement across all pathologists possibly by allowing a more uniform interpathologist identification and segregation of spontaneous changes. Open evaluations to treatment and dose provided assessments of and correlations (not shown) to the Kim-1, a biomarker previously associated with cisplatin-induced injury (Gautier et al. 2010; Vinken et al. 2012; Wadey et al. 2013), that were generally of slightly greater value albeit not statistically significant. These data do not distinguish one approach (blind or open) as inherently superior to the other and thereby can support both the theory that open evaluations increase pathologist precision and the practical use of open evaluations as is standard practice (Crissman et al. 2004) in preclinical safety studies and the hypothesis of the FDA draft guidance on histopathology for biomarker qualification that blind evaluations would not adversely affect biomarker assessment while minimizing bias.
The range and severity of the tissue injury presented in the slides affected scoring. The more severe the treatment-induced changes, the more significant the treatment was found to be. Image set 3 in study A represented a treatment with extremely minimal changes in the kidney (also reflected in minimal biomarker release). Consequently, this was the only evaluation in which treatment was not significant. In study A, image sets 1 and 4 were taken from studies that had a more marked range of injury. This improved the pathologists’ ability to agree on scoring almost certainly through consensus on the more severe injury images. For image sets 1 and 4, pathologist variability was less with the significance of interpathologist variability increasing with diminishing severity of injury as evidenced in image sets 2 and 3. Since study B dealt with image sets created from the same set of samples, the influence of injury severity could not be confirmed, or even evaluated, in that portion of the project. This finding further supports the need for a common lexicon and criteria such as those proposed by the efforts for International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice (Frazier et al. 2012) and the initial use of “training slides” for the specific morphological changes to be evaluated. Both FDA and STP capture this need in their recommendations although their approaches are different. STP recommends an initial open evaluation of controls and treatment groups, with subsequent masked (blind) evaluations as deemed necessary. Alternatively, the FDA suggests that lesion identification and definition should be the function of exploratory studies with open evaluations and that subsequent confirmatory studies should feature blind evaluations with, at most, review of training slides or images derived from the study but separate from those ultimately used to assess biomarker performance.
This study has several weaknesses that minimized the power of the experimental design and our ability to make specific recommendations about possible histopathology best practices for biomarker qualification. First, the study would have benefited from a larger number of pathologists evaluating the slide sets. The 3 pathologists involved in the study came with their specific background education, experience, and expertise. This affected the power of the study and could have impacted the variability determinations. Second, additional blind and open evaluations of the same image sets would have added strength to the findings and increased the power to detect differences or more correctly to be assured that undetected differences do not truly exist. Third, this study was too small to definitively address sampling location, sample adequacy, and sample number issues. In study B, sections from different areas of the same kidneys were used to create different image sets with the intention of examining the impact of sample location and sample numbers on scoring ultimately applied to the whole organ. While this investigation is needed and it initially appeared that multiple sections from different locations might be of some value, the pathologists pointed out weaknesses in the conceived design that make any statements regarding impact of sample location largely conjecture. Specifically, the step sectioning used resulted in some images that did not contain the corticomedullary junction required to determine cisplatin injury These images were by default given a 0 score, which could have resulted in a false negative in the ROC analysis. But issues of sampling remain critical and merit investigation under well-designed and controlled circumstances. Fourth, a consensus lexicon and scoring system and peer review could have furthered pathologist concordance and minimized variability. Fifth, alternative staining (periodic acid–Schiff [PAS]) or techniques (electron microscopy) would have likely impacted the pathologists’ sensitivity and may or may not have affected interpathologist variability. Nevertheless, this study found significant results and has provided initial data for parameters not often quantitatively addressed in assessing biomarker performance as well as yielding findings that are consistent with the STP best practices and/or the FDA draft guidance.
The ability of qualitative data to adequately identify true positives and true negatives can be limited as demonstrated in this study. Histopathology in several instances in this study was unable to provide both true positives and true negatives. The authors do not presume via this study to identify the source of this inability to define true positive and true negative. In these cases, it may be that there were no true negatives (zero scores), or it could be that qualitative light microscopy was not sensitive enough to make the distinction, or it could be an artifact of blind evaluation in which a more “sensitive” interpretation was applied to assure that significant lesions were not overlooked, or in specific cases it could have been inadequate sampling (the sample did not contain the required structures for evaluation). Individual pathologist’s predilection for handling “spontaneous” changes could also lead to this outcome. This project demonstrates that pathologists’ interpretation and definition of background changes can profoundly impact scoring and therefore biomarker assessment. This influence was more prominent in study B where other contributors to interevaluator variability were minimized. To control this source of variability, pathologists working in peer groups need to strictly define how background changes are identified and handled in scoring true positive versus true negatives and/or all changes need to be recorded and other methods (possibly statistical) applied to that data to define true positives and true negatives.
In many cases, the sensitivity of a biomarker is inherently limited by the biomarker itself and pathologist interpretation has no significant influence. However, if interpathologist variability can influence biomarker performance assessment, the sensitivity of the biomarker becomes limited by the sensitivity of the pathologist. Further, should biomarkers be identified that signal prior to observable histopathology change, then histopathology will not be useful as the reference standard for determination of true positives and true negatives. Alternatives to ROC analysis and/or statistical approaches to larger qualitative data sets may be required to quantify biomarker performance in these situations where no true negatives are detected, and, collectively, these facts suggest that each biomarker assessed should be examined carefully and methods adopted that are fit for purpose much as the novel biomarkers they hope to describe.
The results of this study demonstrate that in uncoordinated efforts variability in histopathology scoring between pathologists in both blind and open evaluation can be large enough to bias performance assessments for a biomarker, but that even modest efforts at providing a common framework for these evaluations can have a positive effect on concordance. In spite of this improvement with a more coordinated approach, sufficient variability remained between pathologists to influence biomarker assessment. This implies that in collaborative biomarker assessment efforts, more stringent and detailed lexicon development and peer review will help further minimize interpathologist variability. But even these practices do not address issues that may arise due to inadequate sampling or when there is an attempt to assemble data from multiple independent sources where one peer group evaluation may not align with that of another peer group. Investigations of optimum sample numbers and locations with verified adequate organ/tissue sections should be undertaken to determine their impact or lack of impact on biomarker assessment, especially given that this study failed in this regard.
In conclusion, the data are consistent with previous STP and FDA recommendations that evaluations of biomarker performance should utilize common lexicons and scoring criteria with peer review to ensure that the reporting or prediction of tissue injury is robust and reliable. This study did highlight the need for more information on the impact of lexicons and scoring system development (defining a well-controlled study) on pathologist variability. The data revealed that blind evaluation was not generally a significant source of variability between evaluations and that open evaluations did not significantly increase concordance or improve biomarker assessment. These findings support a “fit for use” approach as opposed to a standard method for all circumstances. The increasing sensitivity and early appearance of novel molecular biomarkers also suggest that assessment should be specifically designed for and with knowledge of the specific biomarker and that new paradigms might be beneficial in assessing biomarkers relative to background or spontaneous changes.
Footnotes
Acknowledgments
Alan Knapton, Scott Pine, and Dr. David Peters were instrumental in completing the animal phase of this study. The authors would particularly like to acknowledge Dr. Patricia Harlow, Dr. Elizabeth Hausner, and Dr. Melanie Blank who provided guidance in the conception and execution of this work.
Author Contribution
Authors contributed to conception or design (RR, SS, TC); data acquisition, analysis, or interpretation (RR, MM, SF, SM, KS, JZ, SS, TC); drafting the manuscript (RR, MM, SF, SM, KS, JZ, SS, TC); and critically revising the manuscript (RR, SF, SM, KS). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Authors’ Notes
This article reflects the views of the authors and should not be construed to represent FDA’s views or polices. Research materials supporting this article are available from the corresponding author.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
