Abstract
In a previous study, to identify genes of importance for hepatocellular carcinogenesis, and especially for processes involved in malignant transformation, the authors investigated differences in gene expression between adenomas and carcinomas by DNA microarray. In the present study, the authors investigated AW434047, one of the sequences that was upregulated in carcinomas. The investigation led to the identification of a novel gene, which the authors named hepatocyte malignant transforming factor (HMTF), of unknown function whose expression was increased in hepatocellular carcinomas. Northern blot and in situ hybridization also demonstrated high levels of HMTF in rat hepatocellular carcinoma (HCC) cell lines, lymphocytes in the spleen, colon mucosal epithelia, spermatocytes, and granule cells of the hippocampus. Reduction of HMTF by RNA interference (RNAi) in N1 cells, an HCC cell line, caused suppression of cell proliferation, invasion, and migration. Suppression of proliferation appeared to be due to cell cycle arrest without increased apoptosis. Decreased HMTF expression resulted in down-regulation of STAT3, PCNA, and cyclin D1 and upregulation of p27. These results suggest that HMTF is a new marker for rat HCC and is involved in HCC cell proliferation and may also be linked to cell proliferation in the spleen, colon, brain, and testis.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cancers worldwide and is a major cause of death, especially in Asia (Jemal et al. 2004). Most HCCs are considered to develop from preexisting adenomas (Eguchi et al. 1992; Hussain et al. 2002; Pitot et al. 1996). To prevent and control HCC, it is important to elucidate carcinogenic steps, especially those pertinent to malignant transformation, at both the histopathological and molecular levels.
Previously, using DNA microarray analysis, we compared gene expression in hepatocellular adenomas and carcinomas (Suzuki et al. 2009). We detected 196 genes whose expression was changed (Supplemental Table 1, available at http://tpx.sagepub.com/content/by/supplemental-data) and established that during hepatocellular carcinogenesis, acquisition of malignant potential requires specific signaling pathways related to high cell proliferation and associated with TP53. Several of the genes we identified were upregulated in carcinomas, but their function was unknown. One of these genes (GenBank ID: AW434047) is an expressed sequence tag (EST) whose sequence is similar (BLAST search, http://blast.ncbi.nlm.nih.gov/), to a hypothetical mRNA (GenBank ID: XM_347003) that encodes the hypothetical protein RGD1566118 (GenBank ID: XP_347004) in the Rat Genome Database project (de la Cruz et al. 2005; Twigger et al. 2002). In the present study we investigated AW434047.
Initially, we attempted to isolate the full-length sequence of XM_347003 from rat HCC cell lines. Our attempts, however, failed; therefore, we utilized a GeneRacer kit to determine the sequence of the mRNA transcript associated with AW434047. We registered the sequence obtained with GenBank under the name rat hepatocyte malignant transforming factor (HMTF; GenBank ID: GU969272). (XM_347003 and XP_347004 were removed from NCBI after the registration of HMTF.) Since microarray analysis in our previous study indicated that expression of HTMF (AW434047) was significantly higher in carcinomas than in normal liver tissue, glutathione S-transferase placental form (GST-P) foci, or adenomas (Supplemental Table 1 and Fig. 1A), we investigated the expression and function of HTMF in rat HCCs and its expression in normal tissues.
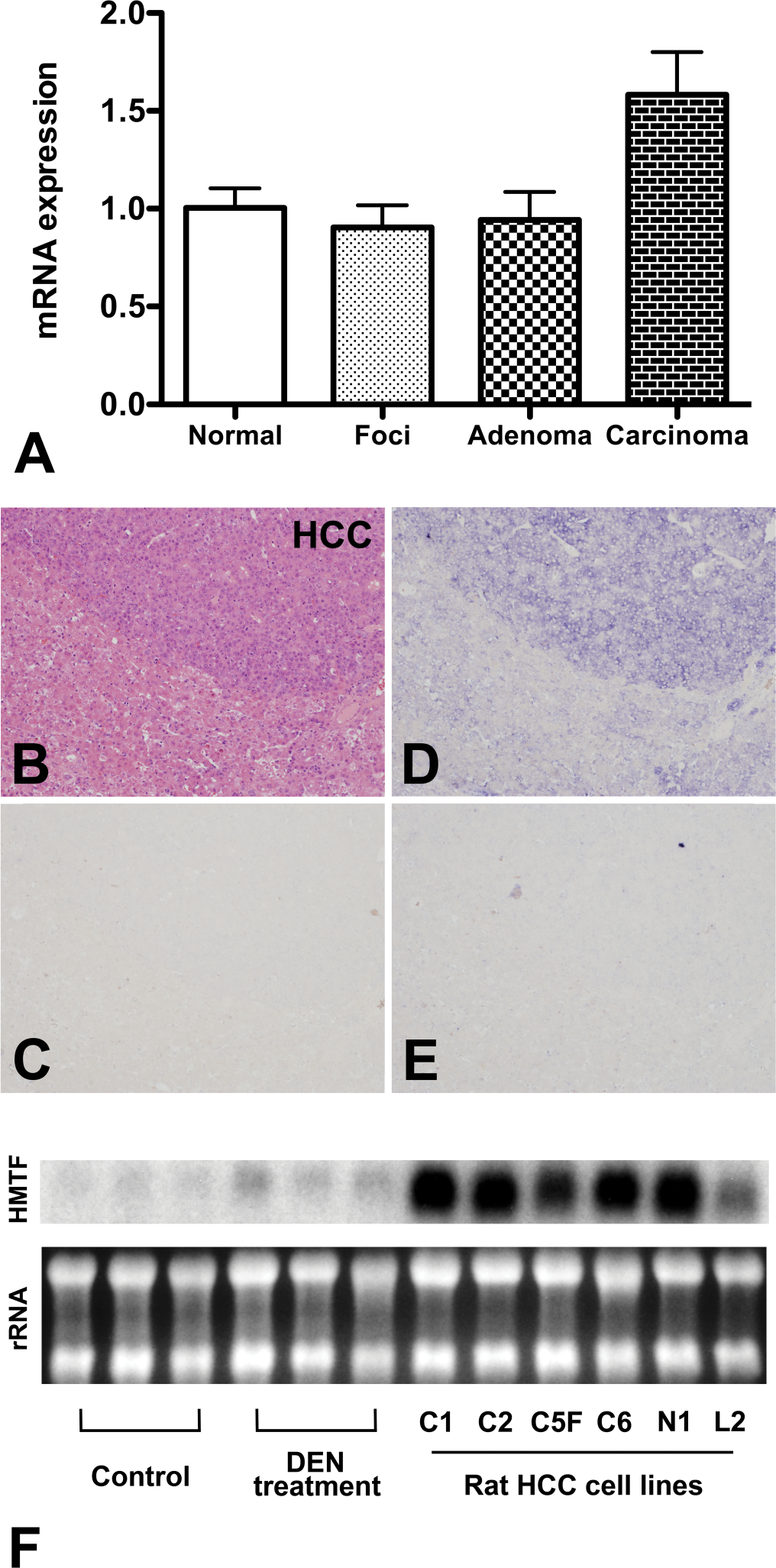
Expression and localization of hepatocyte malignant transforming factor (HMTF) in the rat liver. (A) RNA levels of AW434047 determined by previous microarray analysis. Normal: normal liver tissue (six-week-old nontreated rat); Foci: GST-P positive foci (DEN-treated for twelve weeks); Adenoma: adenomas (DEN-treated for twelve weeks); Carcinoma: carcinomas (DEN-treated for eighteen weeks). (B–E) Carcinoma and surrounding liver tissue in rats treated with DEN for eighteen weeks. B: H & E. HCC indicates the HCC area; C: sense rat HMTF fragment; D: antisense rat HMTF fragment; E: antisense rat HMTF fragment after RNAase treatment; (F) HMTF RNA expression detected by Northern blotting. Control: normal liver (twenty-four-week-old rats); DEN treatment: DEN-treatment for eighteen weeks (twenty-four-week-old rats); Rat HCC cell lines (C1, C2, C5F, C6 N1 and L2); rRNA: ribosomal RNA. For the expression data in panel A, the Normal was set to 1.0.
Materials and Methods
Animals
Details of the animal experiments were previously reported (Suzuki et al. 2009). Six-week-old male F344 rats (Charles River Japan Inc., Atsugi, Japan) continuously received either 0 or 50 ppm diethylnitrosamine (DEN; Tokyo Kasei Kogyo Co. Ltd. Tokyo, Japan) in their drinking water for up to eighteen weeks and were sacrificed six, twelve, or eighteen weeks after starting DEN treatment (six, six, and nine animals, respectively). The livers were immediately excised for analysis. Slices were fixed in 10% buffered formalin for immunohistochemical examination and hematoxylin and eosin staining, and the remaining liver tissue was immediately frozen in liquid nitrogen and stored at –80°C until processed for RNA and protein extraction.
Normal organs of twelve-week-old male rats were excised and frozen in liquid nitrogen and stored at –80°C until processing. The remaining tissues were fixed in 10% buffered formalin for in situ hybridization and hematoxylin and eosin staining. The definition of GST-P foci is GST-P immunohistochemically positive foci exclusive of adenoma and carcinoma.
All animal experiments were performed under protocols approved by the Institutional Animal Care and Use Committee of Nagoya City University School of Medical Sciences.
Diagnosis of Liver Lesions
Liver lesion diagnosis was based on “Proliferative and Nonproliferative Lesions of the Rat and Mouse Hepatobiliary System” (Thoolen et al. 2010).
Cell Lines
Rat HCC cell lines (C1, C2, C5F, C6, N1, and L2 cells) were established in our laboratory from a DEN plus N-nitrosomorpholine–induced hepatocellular carcinoma in a male F344 rat (Ogawa et al. 2001). In our present study, the N1 cell line was employed because it proved to be the most highly metastatic among the six cell lines (Ogawa et al. 2001).
RNA Extraction, Reverse Transcription, and Quantitative PCR
Total RNAs from cultured cells were also isolated with ISOGEN (Wako, Tokyo, Japan) according to the manufacturer’s instructions.
cDNAs were prepared using superscript II (Life Technologies, Carlsbad, CA) according to the manufacturer’s instructions. The resulting cDNAs were used for polymerase chain reaction (PCR; PCR was done in triplicate), and data collection was performed on a LightCycler (Roche Diagnostics, Mannheim, Germany). All quantitative values were normalized to endogenous cyclophilin levels, and expression by nontreated cells was used as the reference. HMTF primers were 5'-GACCCAGGAGCACAATGAAT-3' and 5'-CTGGCAACAAGGCAGTAGTA-3'. Cyclophilin Reverse Transcription (RT)-PCR primers were 5'-TGCTGGACCAAACACAAATG-3' and 5'-GAAGGGGAATGAGGAAAATA-3'.
Northern Blot Analysis for HMTF
Ten µg aliquots of total RNAs from the liver tissues of twenty-four-week-old control rats, rats treated with DEN for eighteen weeks (twenty-four-week-old rats), rat HCC cell lines, and each normal organ were separated by electrophoresis in 1% agarose-formaldehyde gels and transferred to nylon membranes (Hybond-N+; GE Healthcare UK Ltd, Buckinghamshire, England) for autoradiography (Hyperfilm MP; GE Healthcare UK Ltd). The HMTF probe was a cDNA (269 to 542 of HMTF cDNA) constructed with the HMTF primers depicted previously and labeled with [alpha-32P]dCTP (GE Healthcare UK Ltd).
In Situ Hybridization
A fragment of the HMTF cDNA, corresponding to nucleotides 269 to 542, was cloned into the vector pGEM-T (Promega, Madison, WI, USA), and antisense and sense HMTF riboprobes were prepared using pGEM-T/HMTF as the template with a digoxigenin RNA labeling kit (Roche Diagnostics). After deparaffinization, sections were treated with 3 µg/ml proteinase K for twenty min at 37°C, postfixed in 4% paraformaldehyde for ten min, and then placed in hybridization buffer containing digoxigenin-labeled antisense riboprobes for sixteen to eighteen hr at 42°C. After hybridization, the sections were washed in 2 × SSC/50% formamide for thirty min at 42°C, then TNE buffer (10 mM Tris-HCl [pH 8.0], 500 mM NaCl,1 mM EDTA) for ten min at 37°C. Thereafter, the sections were treated with 10 µg/ml of RNase (Roche Diagnostics) for thirty min at 37°C, rinsed in buffer A (100 mM maleic acid, 150 mM NaCl, pH 7.5) for ten min, and then incubated with buffer C (1% blocking reagent [Roche Diagnostics] dissolved in buffer A). Horse-radish peroxidase conjugated sheep anti-digoxigenin Fab fragments (1:100, Roche Diagnostics) were applied for thirty min. The slides were then washed in buffer A followed by incubation with biotinylated tyramide in amplification diluent (NEN Life Science Products, Inc., Boston, MA, USA) for ten min, washed three times in buffer A for five min, and then incubated with alkaline phosphate conjugated streptoavidin (1:5000, Roche Diagnostics) for thirty min. After sequential washing in buffer A containing 0.2% Tween 20 and buffer B (100 mM Tris-HCl [pH 9.5], 100 mM NaCl, 50 mM MgCl2), hybridization signals were detected with 5-bromo-4-chloro-3-indolyl-phosphate (BCIP)/4-nitro blue tetrazolium chloride (NBT) solution (Roche Diagnostics). The specificity of the in situ hybridization (ISH) was assessed by parallel hybridization of the sections with sense riboprobes.
Determination of the 5’ End Sequence of HMTF mRNA
To determine the 5’ end sequence of HMTF mRNA, a GeneRacer kit (Life Technologies) was used according to the manufacturer’s instructions using the AW434047 specific primer 5'-TTCAGACATCGCCCTTCCTT-3'. The resultant HMTF cDNA was cloned into a pGEM-T vector (Promega), and its sequence was determined with a BigDye® Terminator v1.1 Cycle Sequencing Kit (Life Technologies) and an ABI PRISM® 310 Genetic Analyzer (Life Technologies) following the manufacturer’s instructions.
HMTF RNA-Interference
N1 cells were cultured with Dulbecco’s Modified Eagle’s Medium (DMEM; Life Technologies) containing 10% fetal bovine serum and 1% Penicillin-Streptomycin (Life Technologies). An HMTF siRNA was designed and obtained from RNAi Co. Ltd. (Tokyo, Japan) with the sequence 5'-ACCACCGUCUGCCUUCUUCCUGUUU-3'. Stealth RNAiTM siRNA Negative Control Med GC Duplex (GE Healthcare UK Ltd) was used as a control. N1 cells were transfected with the siRNAs using Lipofectamine RNAiMAX (Life Technologies).
Cell Proliferation, Cell Cycle, and Apoptosis Assays
N1 cells were seeded into twenty-four-well plates at a density of 2 × 104 cells per well in 500 µl medium and incubated for twenty-four hr at 37°C. Then the cells were transfected with siRNAs. After twenty-four-hr transfection, the culture supernatant was removed and replaced with fresh complete culture medium (DMEM) and culture was continued. For cell proliferation, live cells were counted using Trypan blue staining (Life Technologies) one and three days after addition of fresh culture medium. For cell cycle analysis, one day after addition of fresh culture medium, cells were harvested and stained with propidium iodide (Guava® cell cycle reagent, Guava Technologies, Hayward, CA, USA) according to the Guava® Cell Cycle Assay protocol. Cell cycle phase distributions were determined on a Guava® PCA Instrument using CytoSoft Software. For analysis of apoptosis, one day after addition of fresh culture medium, cell suspensions were prepared and stained with Guava® Nexin reagent for twenty min and immediately processed with the Guava® PCA system (Guava Technologies) according to the manufacturer’s protocol. Cells were gated based on forward scatter (size), and results reported as the percentage of gated cells positive for both annexin V and 7-AAD. Each assay was carried out in triplicate.
Chemoinvasion and Migration Assays
Cell culture inserts coated with Matrigel Matrix (BD Biocoat Matrigel invasion chambers; BD Biosciences, Bedford, MA) for invasion assays and the BD Falcon twenty-four-multiwell insert system (BD Biosciences) for migration assays, were placed in twenty-four-well plates filled with DMEM containing 0.1% bovine serum albumin (BSA) and 10 µg/ml rat HGF as a chemoattractant. Twenty-four hours after transfection with siRNA, cells (1 × 105) were suspended in DMEM containing 0.1% BSA and seeded into the cell culture inserts and incubated at 37 °C for forty-eight hours. Cells moving through the Matrigel Matrix (invasion) and cells migrating through uncoated inserts (migration) were then counted using Trypan blue staining. The number of nontreated, invading/migrating cells was used as the reference. Each assay was carried out in triplicate.
Immunoblot Analysis
After transfection and culture for twenty-four hr, N1 cells were washed twice with ice-cold phosphate buffered saline (PBS) and harvested by scraping into ice-cold PBS. After centrifugation at 1,500 rpm for ten min, the supernatant was removed and cell pellets were lysed with RIPA buffer (Pierce Biotechnology, Rockford, IL, USA) containing protease inhibitors (Pierce Biotechnology) for fifteen min on ice. The insoluble matter was removed by centrifugation at 12,000 rpm, for twenty min at 4°C. Supernatants were transferred to new tubes and the debris pellet was discarded. Protein concentrations were determined with a Coomassie PlusTM-The Better Bradford Assay Kit (Pierce Biotechnology).
Samples were mixed with 2× sample buffer (Bio-Rad Laboratories, Hercules, CA, USA), heated for five minutes at 95°C, and then subjected to 10% SDS-PAGE. The separated proteins were transferred onto nitrocellulose membranes followed by blocking with 5% nonfat milk powder (w/v) in 1× TBS [10 mmol/L Tris-HCl (pH7.5), 100 mmol/L NaCl, 0.1% Tween-20] for one hr at room temperature. Membranes were probed with antibodies against cyclin D1, cyclin E, p21, p27 (Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA), p53 (Epitomics, Inc, Burlingame, CA, USA), the signal transducers and activators of transcription 3 (STAT3), phospoh-STAT3 (pSTAT3; Y705), nuclear factor-kappa B (NF-kB) (Cell Signaling, Technology Inc., Danvers, MA, USA), and c-MYC (Novocastra Laboratories Ltd., Newcastle upon Tyne, UK) in 5% nonfat dry milk, 1× TBS, 0.1% Tween-20 at 4°C overnight, followed by incubation with peroxidase-conjugated secondary antibodies. Binding was visualized with an enhanced chemiluminescence detection system (GE Healthcare UK Ltd). To confirm equal protein loading, each membrane was stripped and reprobed with anti-beta-actin (b-actin) antibody (Sigma-Aldrich, Co., St. Louis, MO, USA).
Statistical Analysis
All data are presented as mean ± standard deviation values. Statistical comparisons were performed with one-way ANOVA followed by Dunnett’s test. Correlations were assessed by Spearman correlation coefficient analysis. Significance was concluded when p < .05. All statistical analyses were performed using GraphPad Prism 5 (GraphPad Software, Inc., La Jolla, CA).
Results
Expression and Sequence of HMTF mRNA in Normal Hepatocytes and Malignant Tumors
We utilized a GeneRacer kit to determine the sequence of the mRNA transcript associated with the EST AW434047. The sequence is registered with GenBank under the name rat HMTF (GenBank ID: GU969272).
In the rat liver treated with DEN for eighteen weeks, HMTF expression, determined by ISH, was readily detected in carcinoma cells (Fig. 1B & D). No hybridization was detected with a sense RNA probe (Fig. 1C) or after RNAse treatment before hybridization with the anti-sense RNA probe (Fig. 1E). In the same liver, there was less staining of HMTF in GST-P positive foci and adenomas (supplemental Fig. 1).
Northern blotting demonstrated HMTF RNA expression in the livers of DEN treated rats and in the rat HCC cell lines (Fig. 1F) with bands localized around 1 kb. HMTF expression was higher in the livers of twenty-four-week-old rats treated with DEN for eighteen weeks (these livers contained HCC) than liver tissue from untreated twenty-four-week-old rats. HMTF expression was particularly high in all rat HCC cell lines tested (C1, C2, C5F, C6, N1, and L2). These data confirm the high expression of HMTF in HCC originally indicated by microarray analysis.
Knock-Down of HMTF Expression in N1 Cells
To investigate HMTF function in HCC, HMTF siRNA was used in N1 cells. HMTF expression at the mRNA level was significantly suppressed after transfection of HMTF siRNA (1.00 ± 0.06, 0.92 ± 0.11, and 0.47 ± 0.05, nontreated, control siRNA-treated, and HMTF siRNA-treated cells, respectively, at day 1) (Fig. 2A). At day 4 no statistically significant effects were evident (1.00 ± 0.14, 1.14 ± 0.11, and 0.80 ± 0.11, nontreated, control siRNA-treated, and HMTF siRNA-treated cells, respectively, data not shown).
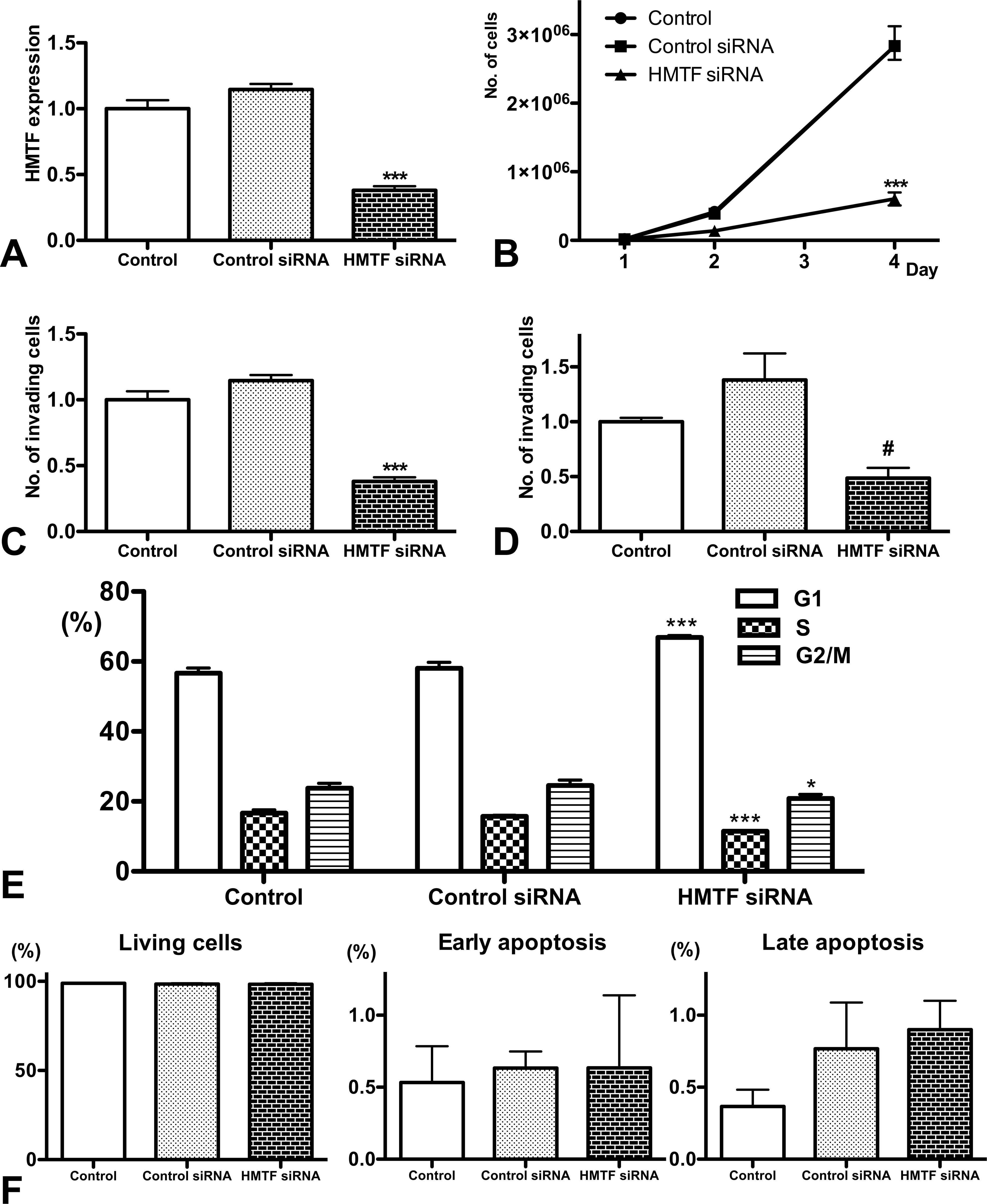
Functions of hepatocyte malignant transforming factor (HMTF) as assessed by RNA interference. (A) Quantitative RT-PCR. (B) Cell growth assay. (C) Invasion assay. (D) Migration assay. (E) Percentages of cells in the different cell cycle phases. (F) Percentages of living cells and cells undergoing early and late apoptosis. Control: nontreated. Control siRNA: treated with negative control siRNA. HMTF siRNA: treated with HMTF siRNA. For the data in panels A, C, and D, the control was set to 1.0. *, ***, significantly different from control and control siRNA at p < .05 and p < .001, respectively. #, significantly different from control at p < .05 and from control siRNA at p < .001. All values are means ± standard deviations.
RNAi-mediated HMTF knock-down led to significant suppression of cell growth (79%; Fig. 2B), invasion (62%; Fig. 2C), and migration potential (52%; Fig. 2D) in N1 cells. The growth suppression was associated with G1 arrest of cell cycle (Fig. 2E) and not with the induction of early (annexin V-positive cells) or late apoptosis (annexin V- and 7-AAD-positive cells) (Fig. 2F).
HMTF-Associated Signaling Pathways
For determination of HMTF-regulated genes, analysis by immunoblot revealed that cyclin D1 and PCNA were down-regulated and p27 up-regulated in HMTF siRNA-treated cells (Fig. 3 ). There were no differences in expression of p53, p21, or cyclin E among the groups (Fig. 3). Total and activated STAT3 levels were also reduced in the HTMF siRNA-treated cells. No differences in the expression of c-myc or NF-kB were found among the groups. The protein blots were quantified using ImageJ (National Institute of Mental Health, Maryland, USA), and the data are presented in supplemental Fig. 2.
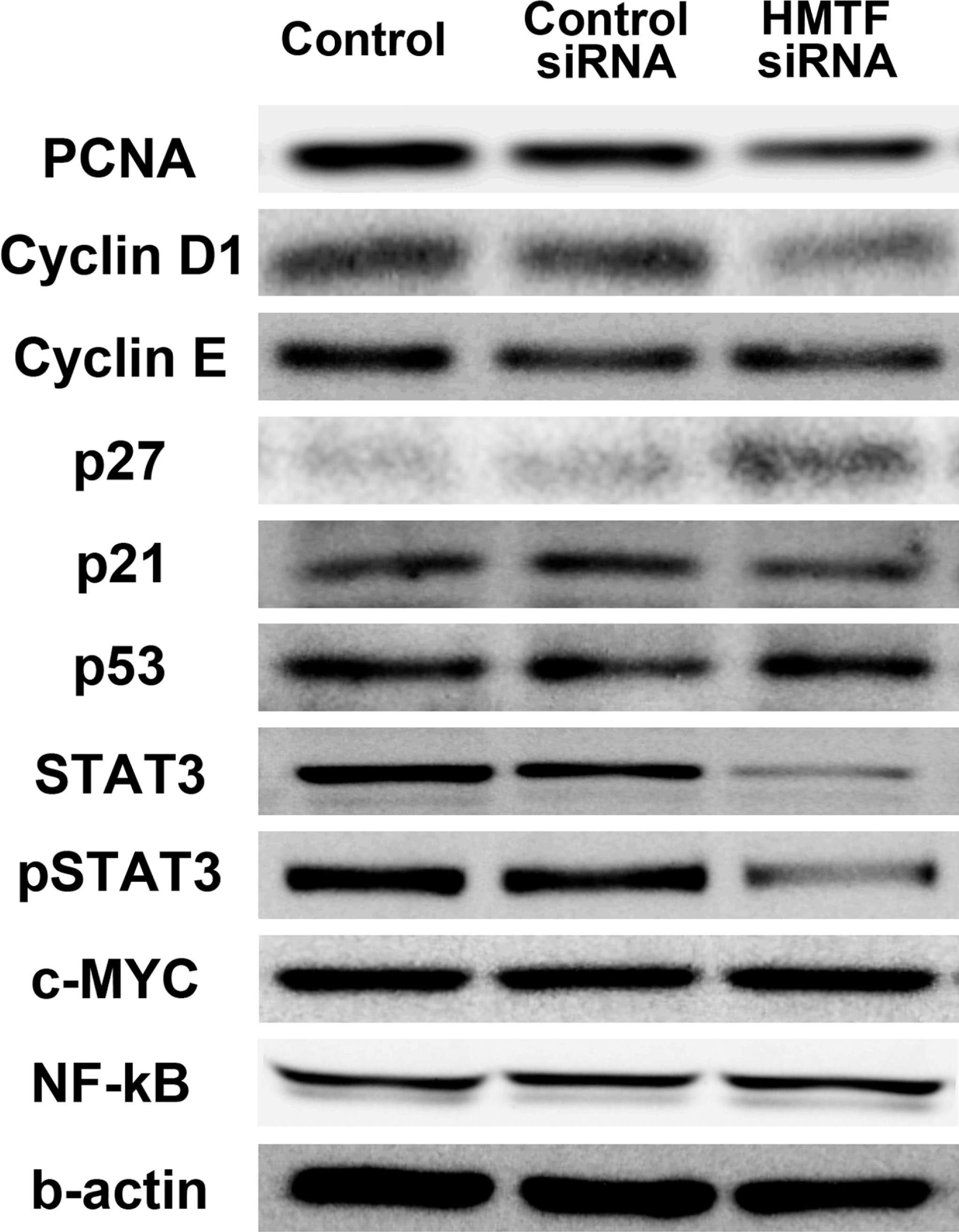
Changes in protein expression in N1 cells treated with siRNAs. Control: nontreated. Control siRNA: treated with negative control siRNA. Hepatocyte malignant transforming factor (HMTF) siRNA: treated with HMTF siRNA.
mRNA Expression and Localization of HMTF in Rat Normal Organs
HMTF RNA expression was detected in the spleen, colon, brain, and especially the testis of nontreated control rats (Fig. 4A). ISH performed for these four organs demonstrated HMTF expression in the lymphocytes in the spleen, mucosal epithelia of the colon, granule cells of hippocampus, and spermatocytes of the testis (Fig. 4B).
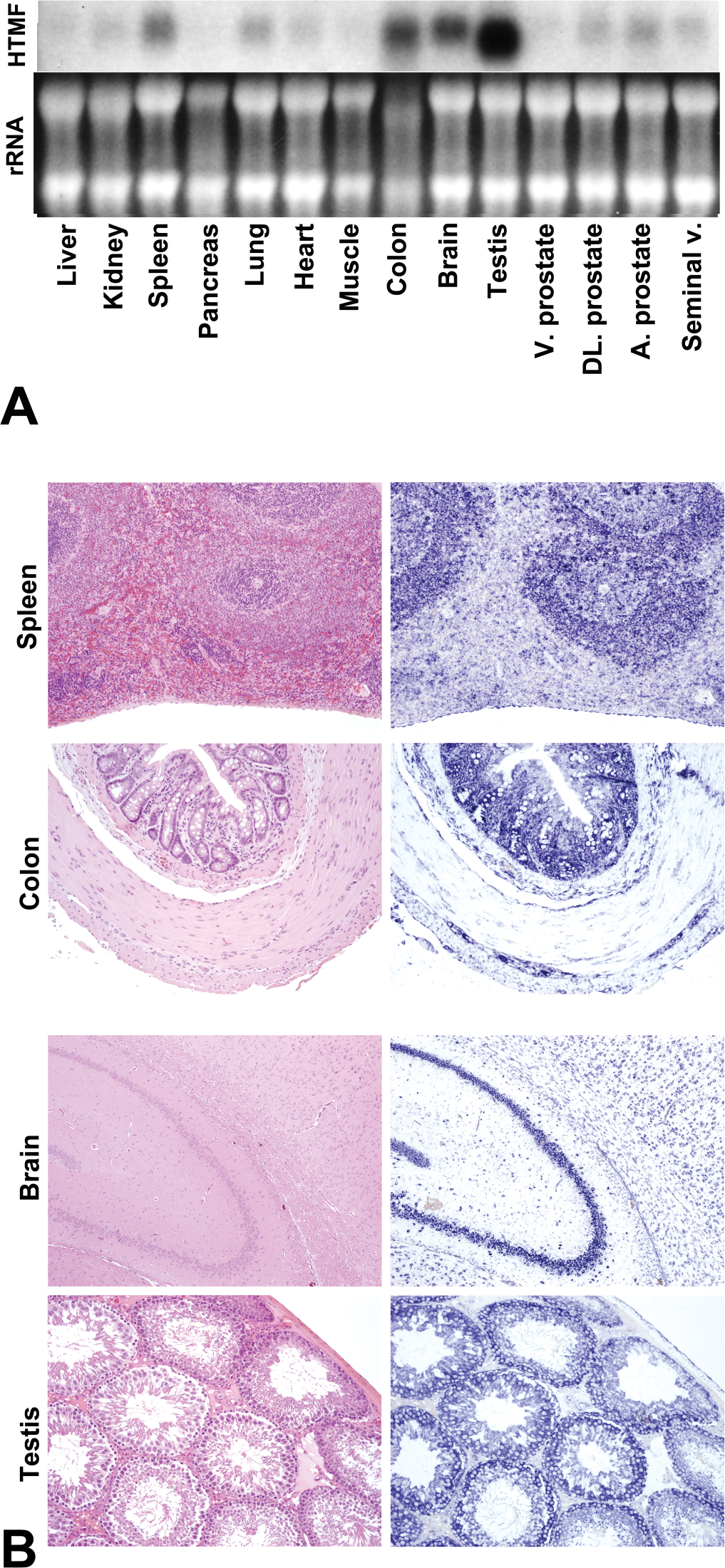
Expression and localization of hepatocyte malignant transforming factor (HMTF) in normal organs by Northern blotting (A), histology with H&E (left) and in situ hybridization with antisense HMTF (right; B). rRNA: ribosomal RNA.
Discussion
In the present study, we focused on a new marker of malignant transformation of rat hepatocytes, HMTF. HMTF is expressed in rat HCCs but not normal liver. Importantly, in all six rat HCC cell lines, HMTF was more highly expressed than in primary liver carcinoma levels, confirming its association with malignancy. HMTF was associated not only with cell proliferation but also with invasion and migration in the N1 rat HCC cell line. These data all indicate HMTF is a marker of malignant potential in HCCs.
Our data also demonstrate that HMTF is linked with proliferation in the N1 rat HCC cell line. It is reasonable that acquisition of malignant potential requires specific signaling pathways related to cell proliferation and p53 (Martin and Dufour 2008; Qin and Tang 2002), as demonstrated in our previous study (Suzuki et al. 2009). In the present study, regulation of cell proliferation by HMTF was clearly associated with changes in cell cycle elements but not apoptosis. Suppression of HMTF expression did not affect the expression of p53, and cell cycle arrest by HMTF siRNA appears independent of p53-mediated pathways.
STAT3 is a transcription factor and binds to specific DNA-response elements in the promoter of target genes and regulates the expression of genes that mediate transformation, apoptosis, proliferation, cellular invasion, and angiogenesis (Aggarwal et al. 2009). It has been reported to be important for both human and rat hepatocarcinogenesis and associated with PCNA, cyclin D1, Bcl-xL, and c-MYC (Cui et al. 2008; Sun et al. 2008; Yu et al. 2007). In this study, HMTF siRNA reduced both STAT3 and phospho-STAT3 expression, and this might suppress cyclin D1 and PCNA and cell proliferation. The STAT3 pathway may also enhance invasion and migration by inducing the expression of matrix metalloproteinases (Aggarwal et al. 2009). On the other hand, while c-MYC and NF-kB are known to be important factors for HCCs (Arsura and Cavin 2005; Qin and Tang 2002), they were not affected by HMTF siRNA. These data indicate that the main function of HMTF is regulation of the STAT3 pathway in HCCs.
Detectable levels of expression of HMTF were not found in normal liver, but lymphocytes in the spleen, mucosal epithelia of the colon, and spermatocytes, which are known to have a high cell proliferative potential, all proved positive. Granule cells of the hippocampus are reported to be associated with neurogenesis in the brain (Aimone, Wiles, and Gage 2006). Therefore, HMTF may be associated with cell proliferation in normal organs.
Homologous sequences of HTMF mRNA in different species including human and mouse were not detected by Basic Local Alignment Search Tool (BLAST search). Rat HMTF is located within the 20p12 region of RGD1566118 (UniGene ID: Rn.128917), where the genes for heat shock 70 kDa protein 1B and Neu1 are located. A homologous location is within 6p21.3 of human and 17 B1 of mouse, respectively (detected by Map Viewer; NCBI); however, no homologous genes for HMTF appear to be present in the human or mouse genomes, indicating that HMTF is not conserved across species.
The hypothetical protein of HMTF is RGD1566118 (predicted), isoform CRA_a [Rattus norvegicus] (GenBank ID: EDL83471), which consists of a thirty-four amino acid sequence registered in the Rat Genome Database project (de la Cruz et al. 2005; Twigger et al. 2002). No homologous proteins were detected by BLAST search with “Non-redundant protein sequence.” However, “Chain A, Solution Structure Of The Pdz Domain Of G-protein signaling 3 (RGS3)” (PDB: 1WHDA) was detected by BLAST search with “Protein Data Bank proteins.” RGS proteins are regulators of G protein-coupled receptors (GPCRs), which play major roles in signal transduction, and some RGS proteins have PDZ domains that interact with GPCRs (Ishii and Kurachi 2003; Zhong and Neubig 2001). For determination of the structural function of HMTF, further studies are needed focusing on interactions and binding affinity with GPCRs.
In conclusion, the present results indicate that HMTF is a novel marker of malignant potential in rat hepatocarcinogenesis and is associated with up-regulated cell proliferation through STAT3, p27, and cyclin D1 pathways. HMTF may also be linked to cell proliferation in normal organs.
Footnotes
Acknowledgment
The authors would like to thank Dr. David B. Alexander for helpful review of this manuscript.
The author(s) declared no potential conflicts of interests with respect to the authorship and/or publication of this article.
This work was supported in part by a Grant-in-aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology, Japan; a Grant-in-aid from the Ministry of Health, Labour and Welfare for the Second Term Comprehensive 10-Year Strategy for Cancer Control, Japan; funds for Development of a High Precision Summary Toxicity (Hazard) Assessment System for New Energy and Industrial Technology Development (NEDO), Japan; funding from Aichi Cancer Research Foundation; and a grant from the Society for Promotion of Toxicological Pathology of Nagoya, Japan.
