Abstract
Methods of toxicity testing, barely changed for several decades, need to be improved. One way forward would be to use a small battery of inbred strains instead of the single outbred stock currently used in toxicity screening. Inbred strains are more stable, more uniform, more repeatable, and better defined than outbred stocks. Genetic variation would be observed as the difference between strains. Safety could be based on the most susceptible strain. Sometimes it may be possible to identify the genes involved. Mechanisms could be explored using gene expression profiling of susceptible and resistant strains. Two committees of toxicologists have concluded that the use of inbred strains, by controlling interindividual variability, would reduce the number of animals needed in toxicity screening, although both “preferred” outbred stocks. This preference appears to have been based on intuition rather than scientific principles. Data from a previously published study on the response to chloramphenicol in an outbred stock and four inbred strains is used to explain the advantages of the multistrain design. Toxicologists, safety pharmacologists, regulatory authorities, and pharmaceutical companies should take a critical look at the types of animals they use if they want to reduce the attrition rate of new drugs.
Keywords
Introduction
Inbred, “genetically defined” strains of mice and rats are more stable, more uniform, more repeatable, and better defined than the “genetically undefined” outbred stocks used in most toxicity testing (the term “stock” is usually used for outbred animals and “strain” for inbred strains). Experiments in which they are used should be more powerful with more accurate dose-response relationships and fewer false negative results than those done using outbred stocks. More than one strain can be used in a factorial experimental design without increasing the total number of animals. Strain differences give an indication of genetic variation in response, and safety can be assessed using the most sensitive strain. Further investigation could lead to the identification of susceptibility genes, a better understanding of mechanisms and possibly new drug targets. Inbred strains also provide a stable genetic background for mutants and genetic alterations, which are already widely used in toxicological research.
The failure to make use of this valuable resource in toxicological screening highlights the enormous inertia in the science of toxicity testing. At a time of extraordinary scientific progress, methods have hardly changed in several decades (Food and Drug Administration [FDA] 2004). The large attrition rate of investigative new drugs (INDs) following clinical trials, with 27% of 1,099 INDs being discontinued because of toxicity (Caldwell et al. 2001), may be one consequence of this failure. Toxicologists face a major challenge in the twenty-first century. They need to embrace the new “omics” techniques and ensure that they are using the most appropriate animals if their discipline is to become a more effective tool in drug development. They may need to work closely with the relatively new discipline of safety pharmacology (Pugsley, Authier, and Curtis 2008), which appears to be able to innovate with less regulatory bureaucracy.
Here, I review the properties of inbred strains and discuss how and why they, rather than outbred stocks, should be used in assessing the toxicity of new chemical entities (NCEs) and INDs.
A Brief History of Inbred Strains of Laboratory Animals
The first inbred strains of mice, rats, and guinea pigs were developed soon after the rediscovery of Mendel’s laws of inheritance in 1900. There are now more than 150 rat strains and several hundred mouse strains (Beck et al. 2000).
It was soon recognized that inbred strains were of value in many areas of research. Thus, in 1937, Dr. C. C. Little wrote, “Just as the purity of the chemical assures the pharmacist of the proper filling of the doctor’s prescription, so the purity of the mouse stock can assure a research scientist of a true and sure experiment… In experimental medicine today. .. the use of in-bred genetic material… is just as necessary as the use of aseptic and anti-septic precautions in surgery” (Rader 2004, 147). These sentiments have been echoed by several geneticists since then (Festing 2010).
More than twenty Nobel prizes have been awarded for work that probably would have been impossible without inbred strains (Festing and Fisher 2000). They have been used to identify the genes responsible for graft rejection, in the discovery of immunological tolerance, and in explaining the mechanisms of antibody diversity. They were used in the development of monoclonal antibodies, in the development of embryonic stem cells, in the discovery of viral and cellular oncogenes, and in the identification of the many genes associated with cancer and other spontaneous and induced disease. Their use is essential in the generation of “knockout” mice in which a single specified gene can be inactivated by homologous recombination and of “knock-in” mice in which a foreign gene can be inserted in a predefined location within the genome. Modern genomic research is heavily dependent on these strains (Nguyen and Xu 2008).
Definition and Properties of Inbred Strains
Inbred strains are produced by at least twenty generations of brother × sister mating with all offspring being derived from a single pair in the twentieth or a subsequent generation. Inbreeding is never absolutely complete, but after about forty generations only a handful of about twenty-five thousand loci will continue to segregate.
Strains are designated by uppercase letters, for example, CBA, sometimes with numbers such as F344 (not “Fischer 344”!). Generally, these designations have no particular meaning. Substrains are indicated by a slash and a substrain designation, which usually indicates the origin of the strain. For example, C57BL/6J is substrain number 6 of strain C57BL, maintained at the Jackson Laboratory (J). The nomenclature of both inbred strains and gene loci is strictly controlled by international nomenclature committees. Full details are given at www.informatics.jax.org.
F1 hybrids, the first-generation cross between two inbred strains, have all the useful properties of inbred strains except that they are not homozygous and do not breed true. They are more robust than pure lines and can be used in most situations where an inbred strain could be used. They are usually designated by an abbreviated version of the parental strain designations with “F1” appended, for example, B6D2F1 is an F1 hybrid from a cross between a female C57BL/6 and a male DBA/2 mouse. For brevity, the term “inbred strain” used here will be taken to include F1 hybrids unless otherwise stated.
Inbred strains are like immortal clones of genetically identical individuals. Once a strain has been brother × sister mated for more than twenty generations, its properties become fixed, with all individuals being homozygous (not F1 hybrids) and genetically identical or “isogenic.” The only way an inbred strain can change genetically is as a result of new mutations. These are relatively rare for most characters of interest, with many of them being “quiet” (Stevens et al. 2007), possibly because they affect characters like disease resistance, which are not relevant under laboratory conditions. This immortality of the genotype means that information on the strain can be accumulated over long periods of time, with the reasonable assumption that the strain will not change. Selective breeding, say for increased body weight, litter size, or susceptibility to a chemical, is completely ineffective in changing the characteristics of an inbred strain, in strong contrast to outbred stocks.
To detect the effect of a test chemical or give a reliable dose-response curve, an experiment needs a high signal/noise ratio. The noise is expressed as the standard deviation. This can be minimized by using genetically uniform inbred animals that are free of disease maintained in well-controlled and optimized conditions. The magnitude of the signal depends on the sensitivity of the individual strain or stock. This can be maximized by choosing a sensitive strain. However, as there is no way of predicting which strain or stock is likely to be most sensitive, a sensible strategy is to use more than one strain using a factorial experimental design.
Inbred strains of mice (Beck et al. 2000) and rats (www.informatics.jax.org) (Festing 1979) have a wide range of spontaneous diseases and susceptibility to xenobiotics (Kacew and Festing 1996). Some strains are long-lived with little spontaneous early pathology, while others develop diseases such as cancer, diabetes, obesity, and atherosclerosis and can be used as models of these conditions. Some strains were selected before they were fully inbred for characters like hypertension (de Jong 1984), “emotionality” (Broadhurst 1975), learning ability (Amit and Smith 1992), and sensitivity to carcinogens (Benavides et al. 2000; Fischer et al. 1987; Stern et al. 1998). A small collection of inbred strains of rats or mice will usually differ for nearly all biological characteristics as a result of the chance fixation of the genes present in the stock from which they were derived. Genetic markers such as SNPs (single nucleotide polymorphisms) can be used to select strains likely to differ in a wide range of responses (Petkov et al. 2004; Mashimo et al. 2006). Similar techniques can be used for genetic quality control. It is relatively easy using such methods to confirm that each animal is of the correct inbred strain. It is much more difficult to do this with outbred stocks.
Geneticists have developed many protocols for biochemical, anatomical, physiological, pathological, and behavioral phenotyping to study the effects of mutations and genetic modifications in mice and rats (Crawley et al. 1997; Brown, Chambon, and de Angelis 2005). This allows for the rapid accumulation of reliable data. Several databases of strain characteristics are freely available to all investigators (see www.informatics.jax.org).
Mouse and rat informatics is well developed. The complete DNA sequence of eleven standard laboratory mouse strains and four wild-derived strains (Frazer et al. 2007) is now available, and these strains will be used in investigating genetic variability in response to xenobiotics, with the aim being to identify genes associated with interindividual variation in humans (http://ntp.niehs.nih.gov, see “Mouse Genome Resequencing Project”).
Derived Inbred Strains
In addition to “straight” inbred strains, there are several types of “derived” strains developed for special purposes. It is beyond the scope of this review to discuss these strains in any detail, but a brief outline of some of them is given because their availability can further enhance the value of using inbred strains.
A pair of inbred strains that differ only at a single genetic locus are said to be “coisogenic.” These arise either as a result of a mutation within an inbred strain or when a gene is knocked out or otherwise modified, but it is kept on the original genetic background. The effect of the mutation or modification can then be studied in comparison with the background strain without complications arising from the segregation of other genes. This situation is often approximated in the development of “congenic” strains where a mutation or GA is backcrossed to an inbred strain so that its effect can be studied in comparison with the chosen background strain.
“Recombinant inbred” (RI) strains are developed from a cross between two inbred strains, followed by at least twenty generations of brother × sister mating retaining twenty to thirty or more separate strains. These are used in genetic mapping and identification of polymorphic genes. “Consomic” or chromosome substitution (CS) strains have been developed by backcrossing a whole chromosome from one inbred strain into another strain. If the phenotypes of a matched pair of CS strains differ, then that indicates that the difference is due to genes on the chromosome by which they differ. Crosses can then be used to identify the gene or genes responsible.
Characteristics of Outbred Stocks
There has been relatively little work done to characterize outbred stocks, probably because they are not widely used by geneticists, and until recently there have been few genetic markers which could be used in such studies. Outbred stocks are “genetically undefined” in the sense that each animal is genetically unique and the genes it caries are unknown. Although the phenotypic variation within an outbred stock is usually larger than in an inbred strain, on average they are not as variable phenotypically as a collection of different inbred strains, particularly if these were derived from different parental stocks.
The phenotypic characteristics of an outbred stock can change relatively rapidly as a result of random genetic drift in gene frequency, selective breeding (say for large litter size, body weight, blood pressure, etc.) and/or genetic contamination, which may go undetected because genetic quality control is rarely done (Festing 1974; Papaioannou and Festing 1980). Sprague-Dawley rats from one commercial breeder became “fat, frail and dying young” over a short period, although it is not clear whether this was due to genetic drift or some environmental factor (Nohynek et al. 1993).
Stock names such as Sprague-Dawley (SD) or Wistar can be misleading because stocks with the same name from different breeders will have different characteristics (Ito et al. 2007) and there is no standard Wistar or SD stock with which to compare them. A morphometric analysis of skeletal shape, which is highly inherited, showed that although named inbred strains from different breeders were very similar, it was not possible to distinguish between Wistar and SD stocks because stocks with the same name but from different breeders were often quite different (Lovell and Festing 1982). Recently, outbred CD-1 mice from the same breeder but different animal rooms were found to be sufficiently different so that the colony from which any individual came could generally be determined using genetic markers. This was attributed to founder effects (Aldinger et al. 2009). However, the stock was as genetically heterogeneous as a wild population.
Lack of any standards and the unreliability of stock names may make it difficult for investigators to repeat each other’s work. For example, failure to repeat work on the effects of bisphenol A (Sekizawa 2008) may be because many different strains and stocks have been used with stock names being an unreliable indicator of genotype. It is notable that Good Laboratory Practice (GLP) requires quality control of many variables such as contamination of the diet but no quality control of the animals that are used.
Why Do Toxicologists Continue to Use Outbred Stocks?
Two expert committees have recognized that the use of inbred strains would have advantages in toxicity testing. In 1971 the Food and Drug Advisory Committee stated that cancer researchers are extremely fortunate compared to many other biologists in having available to them many inbred strains with known incidences of neoplastic disease. In many instances there is also a considerable background of information on the sensitivity of these strains to the effects of known carcinogens. It is thus possible to test unknown compounds that resemble known carcinogens chemically in relatively small groups of animals and to derive meaningful results. (p. 29)
Attempts are made to minimise within-species and within-strain variability in toxicological studies on animal models, e.g. by using defined or in-bred strains where available, to improve the ability of the study to detect and characterise effects and to reduce the statistical variance in the dose response.
there are nevertheless, many investigators who prefer a randomly bred animal on the basis that the genetic heterogeneity of such animals is more akin to the situation in man. There is of course much to be said for this viewpoint provided if it is recognized that often more animals are required under these conditions. (p. 29)
The VUT report went on to state that a potential disadvantage of such tight controls of experimental conditions is that this approach reduces the chance of detecting an adverse effect that occurs only in a sub-group of experimental animals. The use of larger groups of more outbred animals might increase the chance of detecting such groups, but this could not be guaranteed. (Committee on Toxicity 2007, 29)
The VUT committee was asked to consider the use of several inbred strains. They stated, It has been proposed that replacement of a single strain of outbred rodent with an equivalent number selected from several different inbred strains may provide a more robust model of genetic heterogeneity present in humans (Festing et al. 2001) although the appropriate uncertainty factors for such a design have yet to be defined. There would be practical problems in maintaining sufficient stable inbred strains. (Toxicity Committee 2007, 35)
In an unpublished letter to people who had given evidence to the VUT committee, the Committee on Toxicity (2007) also stated that “they consider it more representative of variation in the human population to use strains of animals with a range of genotypes (as in current testing regimes).”
Clearly, there is a strong intuitive feeling among many toxicologists that by using outbred stocks they are in some way “representing” human genetic variation and that this compensates for the need to use more animals or have a less sensitive test. But as explained above, all it does is to obscure genetic variation and reduce test sensitivity. Genetic variation can only be detected when related animals or those with a similar genotype respond in the same way. It is impossible to detect genetic variation in the experimental designs currently used in toxicity testing. Gene association studies may become possible in the future, but these usually require hundreds or even thousands of animals. Toxicologists rarely see the effects of genetic variation because they use the wrong animals, so they are unable to take it into account when interpreting their experimental results.
What is needed is an experimental design that controls interindividual variability so the test is powerful and able to detect all responses to the test chemical, bearing in mind that some responses may not represent toxicity and some may be specific to the species used and not relevant to humans. At the same time, the test needs to ensure that safety is not assessed on a group of animals that are genetically resistant to the test chemical. If it can also show whether there is genetic variation in response, with the possibility of identifying the susceptibility genes, this would be a bonus. These multiple objectives cannot be achieved using outbred stocks, but many of them can be met by using inbred strains, as explained below.
An Improved Design for Toxicity Testing
Genetic variation should be present in the test population because it is present in human populations, but it needs to be controlled so that it does not reduce the sensitivity of the experiment. And it needs to be clearly visible. This can be achieved by using a battery of inbred strains in a “factorial” design in which both treatment and strain are varied simultaneously. Such designs are efficient because they provide more information with the same number of subjects. According to R. A. Fisher (1960), If the investigator… confines his attention to any single factor we may infer either that he is the unfortunate victim of a doctrinaire theory as to how experimentation should proceed, or that the time, material or equipment at his disposal is too limited to allow him to give attention to more than one aspect of his problem… Indeed in a wide class of cases (by using factorial designs) an experimental investigation, at the same time as it is made more comprehensive, may also be made more efficient if by more efficient we mean that more knowledge and a higher degree of precision are obtainable by the same number of observations. (p. 94)
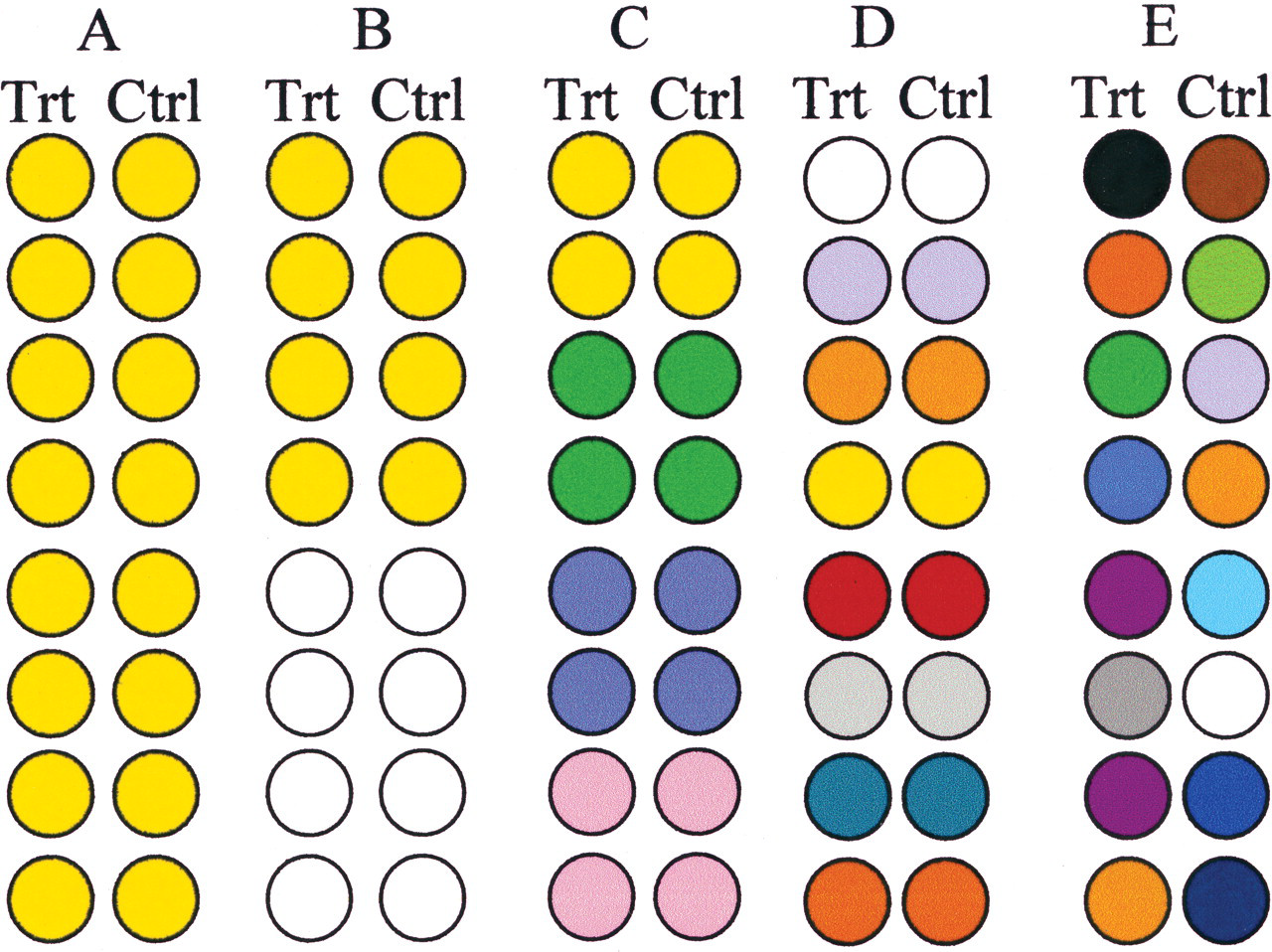
Diagram to show a single and three multistrain experiments and an experiment using an outbred stock with eight treated and eight control animals in each case. Each circle represents an animal and each color a different genotype. All these experiments are valid, can be statistically analyzed using a one- or two-way analysis of variance (for measurement data), and could be extended to multiple dose levels. All except 1E are balanced with treated and control groups being genetically identical. See text for a discussion of their properties.
Figure 1A represents an experiment using a single inbred strain with eight subjects (animals) per treatment group. This would be a moderately powerful experiment because the within-strain variation is minimized, but it is unable to show whether there is any genetic component to the treatment response. Figure 1E represents an experiment of the same size using a single outbred stock. In this case, treatment means are estimated by averaging across unknown individual genotypes. Interindividual variability is poorly controlled; treated and control groups are genetically different; and the more genetic variation there is in each group, the greater the “noise” and the less sensitive the experiment will be. The experiment provides no information on genetic variability, so it confers no benefits but reduces sensitivity. The main weakness in both these experiments is that the chosen strain or stock may be resistant to the test chemical, so neither of them can be recommended.
Figures 1B and 1C are factorial designs using the same total number of animals, but with two or four inbred strains, respectively. The genetic variation is controlled by using inbred strains, so there is low “noise” and the experiments should be powerful. If the strains differ in response to the test chemical, this shows that the response is under genetic control. The conclusions can then be based on the more sensitive strain with a high signal/noise ratio. The design in Figure 1B would be preferred if the aim is to get a good estimate of the response of the two strains used, whereas the design in Figure 1C would be used to reveal as wide a range of genetic variation as possible but with strain means being less well estimated. Both will give a formal test of whether strains differ in response to the xenobiotic, so these two designs provide more information compared with the designs involving a single inbred strain or outbred stock, without using any more animals. In general, the more strains that are used, the more powerful the experiment will be for both quantitative and binary (present/absent) characters (Haseman and Hoel 1979; Felton and Gaylor 1989). The number of strains that could be used depends on practical considerations. Four or five strains could probably be managed with current 28-day and 90-day studies involving eighty animals.
Figure 1D shows a design with eight inbred strains. This is equivalent to a monozygous twin study in humans. This would be a powerful experiment because treated and control groups are genetically identical, although it gives no formal test of whether strains differ in response. It is not as useful as experiments 1B or 1C.
These hypothetical examples show that it is possible to design multistrain experiments using no more animals than when using a single strain/stock. Such designs will usually be more powerful than ones involving a single strain/stock. Safety levels can be based on the most sensitive strain, and they give some indication of whether there is genetic variation in response. Gene expression profiling of the susceptible and resistant strains may help in identifying susceptibility loci.
Example: A Comparison of an Experiment Involving Four Inbred Strains with One Involving a Single Outbred Stock Using Real Data
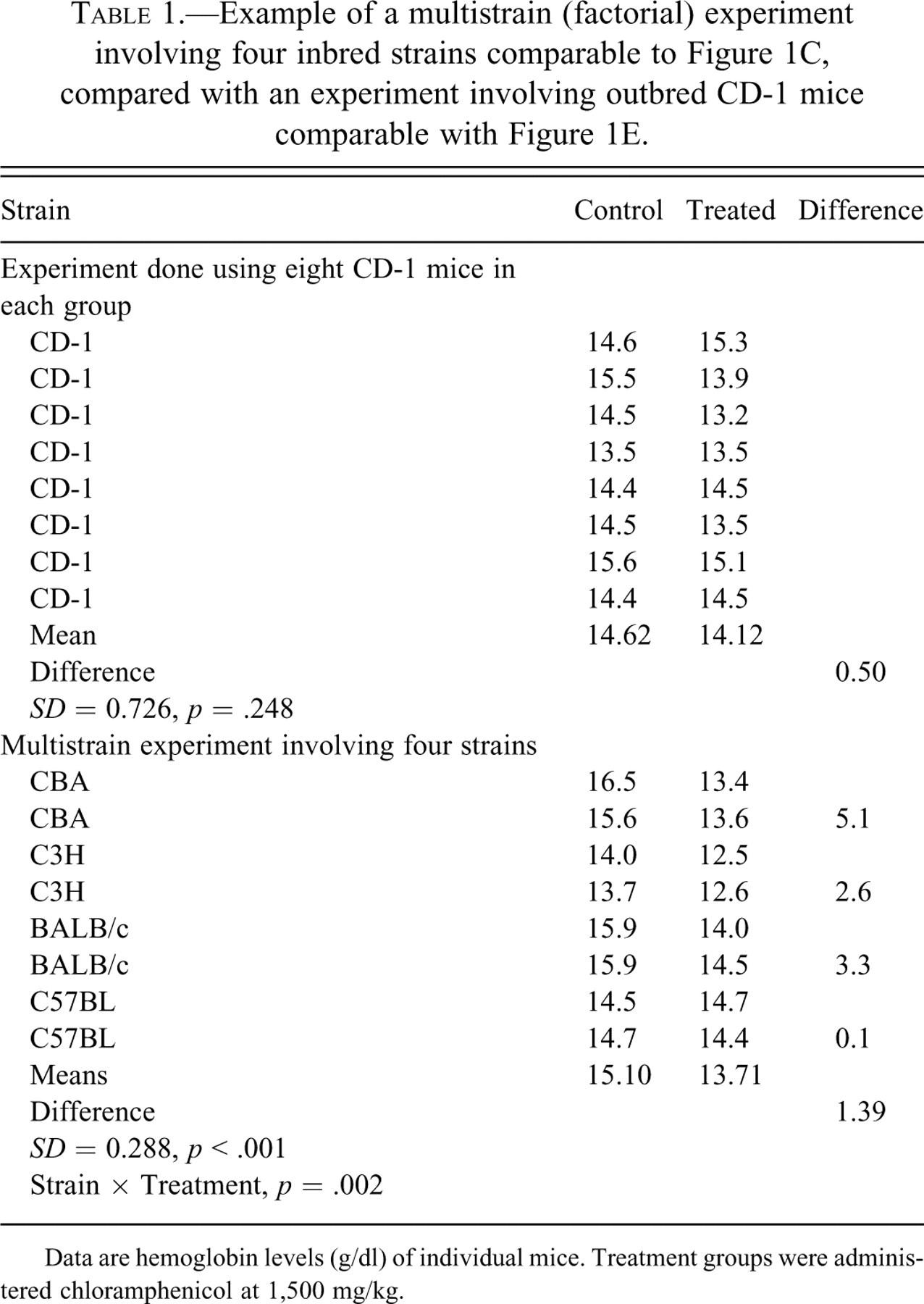
There has been no report of an experiment set up specifically to compare the multistrain study with a similar one using a single outbred stock. The data used here comes from the only experiment known to the author that used several inbred strains and an outbred stock in a multidose experiment with a toxicological end-point (Festing, Diamanti, and Turton 2001). Table 1 shows hemoglobin levels in two experiments analogous to those in Figures 1C and 1E, that is, a 2 (treatments) × 4 (strains) factorial design with two animals per subgroup and a single factor design using the same numbers of CD-1 outbred mice. For simplicity, only data for the control mice and those given 1,500 mg/kg of chloramphenicol is shown, but the results reflect what was found when considering all dose levels and all hematological parameters. In the original study there were eight mice of each inbred strain at each dose. To make the two experiments comparable, two mice of each of the four inbred strains were selected at random, with the results shown in the table.
Data are hemoglobin levels (g/dl) of individual mice. Treatment groups were administered chloramphenicol at 1,500 mg/kg.
In the CD-1 experiment, the response (difference between treated and control means) was 0.50 units and the pooled standard deviation was 0.725, giving a signal/noise ratio of 0.5/0.725 = 0.69 with a nonsignificant p-value of .25. In contrast, in the multistrain experiment, the response was 1.39 units averaged across the strains and the pooled standard deviation was 0.29, giving a signal/noise ratio of 1.39/0.29 = 4.79 and the difference was highly significant (p < .001). Moreover, there was also a highly significant Strain × Treatment interaction. At this dose level, three of the strains responded, with strain CBA being the most sensitive but C57BL being resistant, clearly showing genetic variation in susceptibility. So in this example the outbred stock was both more variable (higher noise) and less sensitive (lower signal) than the study involving the inbred strains, and it gave no indication of genetic variation in response. Neither outbred CD-1 nor inbred C57BL mice alone were able to detect the effect of the compound at this dose level, although C57BL was sensitive at higher dose levels (see below).
Many characters of interest to toxicologists and pathologists such as cancer, necrosis, inflammation, and apoptosis are detected histologically and are scored as present/absent or normal/abnormal. Most of these will have a polygenic threshold mode of inheritance, that is, some animals will be genetically more susceptible than others, probably due to several quantitative trait loci (QTLs), but the condition will only appear beyond a certain threshold dose. In some cases this threshold is exceeded because of environmental influences, and background pathology is observed. Inbred strains will differ in susceptibility, but within a strain there is also nongenetic variation so that not all animals will necessarily show the same pathology, or some will show it sooner than others.
Intuitively, it would seem reasonable to use an outbred stock with a wide range of genotypes when screening for such characters. But toxicity testing requires a controlled experiment (i.e., there needs to be an untreated or vehicle treated control group) because absence of background pathology can never be ruled out. In a controlled experiment treated and control animals should be as similar as possible, and this can best be achieved by making them genetically identical. There are four reasons intuition in favoring outbred stocks is misleading in this case. First, using an outbred stock would be at the expense of extra noise so that quantitative characters would be less well estimated with more false negative results. Second, if there is spontaneous background pathology, it may be unequally distributed, leading to more false positive or negative results. For example, if there were three genetically determined spontaneous tumors, they might all be in the treated groups (a 1/8 chance). This is a chance effect, not due to the test compound, but it might take some explaining to a regulator. Third, outbred stocks are not as variable as a collection of inbred strains, so a wider range of possible sensitivities would be obtained by using several inbred strains. Fourth, the experiment will give no indication of genetic variation in response.
The situation where there is a discrete character with no background pathology is illustrated in Figure 2 using more of the data shown in Table 1. This shows hemoglobin levels in response to chloramphenicol at six dose levels. A horizontal dotted line has been drawn at 13.00 g/dl, which is well below the normal range. Suppose all animals below that level were classified as “anemic,” and all that could be observed was whether the hemoglobin levels were more or less than 13 g/dl, that is, that full data shown in Figure 2 was not visible.
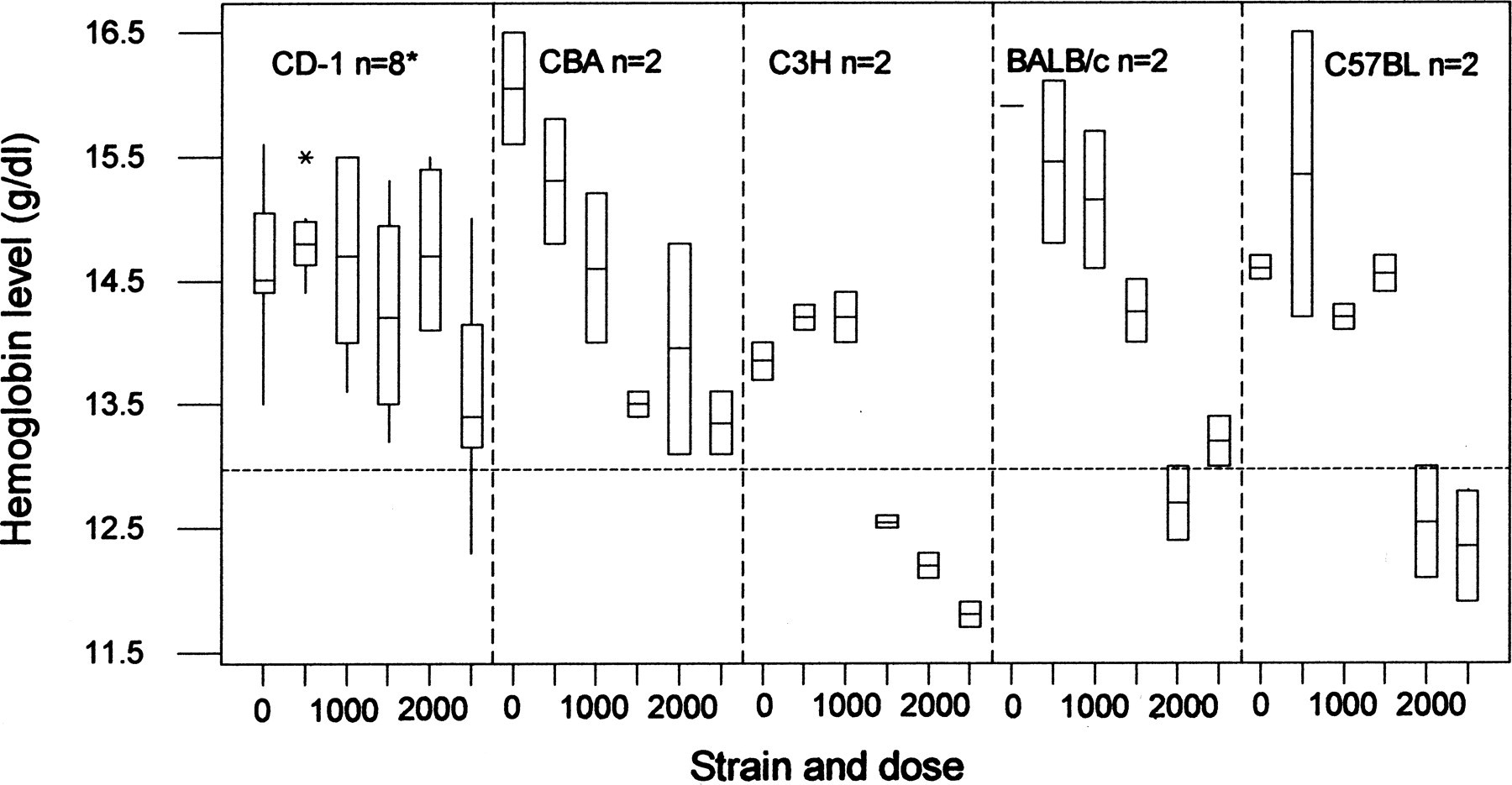
Box and whisker plot of hemoglobin levels in mice treated with 0-2,500 mg/kg chloramphenicol. The CD-1 group involves a total of 47 mice (approximately 8 in each dose group), with the four inbred strains CBA, C3H, BALB/c, and C57BL totaling 48 mice (2 per dose by strain subgroup, hence no whiskers to the boxes). The horizontal dotted line is set at the level of 13 g/dl, notionally the level at which the mice might be diagnosed as “anemic.” Note that there is more variation among the inbred strains than in the outbred stock, giving a broader genetic base. Data extracted from Festing, Diamanti, and Turton (2001).
To answer the question of whether there is a dose-related increased incidence of anemia, the three low dose levels can be compared with the three high dose levels. In the CD-1, 0/24 mice were anemic in the lowest three dose levels and 1/23 in the highest levels. So there is no evidence of a dose-related increase, with only a single mouse being anemic. Pooling across the four inbred strains, in the three low dose levels there were 0/24 anemic, and in the three high levels 10/24 were anemic, a clear dose-related effect. Looking at the individual strains, in the CBA there were 0/12 anemic mice, whereas in the C3H there were 6/12 anemic, giving evidence of genetic variation in susceptibility. Although full quantitative data such as shown in Figure 2 would not be available in most cases, here it seems that the extra sensitivity of C3H is because it has low basal levels of hemoglobin. The figure also shows that the total variation across a set of inbred strains is greater than that found in a single outbred stock, but as most of it is between-strain variation it does not reduce the power of the experiment.
This is only a single example, but this is the sort of result that is expected with discrete characters when using multiple inbred strains rather than a single outbred stock. The multistrain design is the most sensitive, and any genetic variation can easily be detected.
Objections to the Use of Multistrain Designs
Some investigators have suggested that a multistrain study is impractical. Strains may not always be readily available in the numbers wanted. However, several inbred strains of mice are already bred in large numbers, and commercial breeders could supply large numbers of inbred rats if there is a demand for them. Inbred strains are also more expensive than outbred stocks. But the cost of animals is trivial compared with all the other costs of a toxicity test.
Pathologists rely on historical data, and this may not be available for some strains, so it will have to be accumulated. However, with many pathologists working on genetically identical animals, this data should accumulate rapidly. And data on genetically defined inbred strains should be more valuable once it has been collected.
Possibly the most serious objection is that the regulators might not accept data based on multistrain experiments. So research is badly needed to evaluate the idea and provide strong evidence to the regulators that it is a better method than the current one. The costs of such research would be relatively trivial and would be totally insignificant in comparison with the costs of developing the high-throughput in vitro methods proposed by the National Academies committee (National Research Council 2007). Maybe the FDA would support it as part of its critical path initiative.
Incorporation of Inbred Strains into Existing Designs
A typical 28-day toxicity test involves four dose levels, two sexes, and ten animals per group. Dose levels are usually chosen in relation to the MTD, which is the dose that shows no adverse effect in the most sensitive animals. This could be determined much as is currently done, using small numbers of animals of each strain. It would probably not be sensible to have different doses for each strain, at least for short-term tests.
If inbred strains were only used to replace the ten outbred animals (which may not be the optimum strategy), this could be done using, say, five inbred strains with two animals of each strain by sex group at each dose level. This could be organized on a “per strain” basis. So eight male animals of the first strain would be assigned at random to the four dose levels, with the same being done with the females. These sixteen animals would form the first “miniexperiment.” The next four strains would then be tested in sequence with the data being combined for the statistical analysis.
Strains could be chosen from among well-established laboratory strains, omitting any with unwanted background pathology. A battery of strains that are genetically as different as possible could be selected on the basis of known genetic markers. For example, mouse strains fall into seven major families (Petkov et al. 2004) based on single nucleotide genetic markers. Similar data is available for rat inbred strains (Smits et al. 2004). Statistical analysis presents no problem. Factorial designs are widely used, and methods of statistical analysis are well developed.
More flexible strategies could be used if the experiments were used as part of in-house safety pharmacology prior to the formal regulatory testing.
Discussion
Toxicity is at a crossroad. It is widely recognized that current methods of toxicity testing are inadequate (FDA 2004), and there is a substantial backlog of industrial/environmental chemicals of unknown hazard that need to be assessed. A committee of the U.S. National Academy of Sciences has suggested that toxicity testing in the twenty-first century should mostly be done using in vitro methods with high-throughput screening to identify toxicity pathways rather than apical end points (National Research Council [NRC] 2007). This will involve a considerable financial investment, it will take ten to twenty years to perform the necessary research, and there are a formidable number of barriers to overcome (Hartung 2009), so it may not be successful, particularly for the testing of pharmaceuticals. The suggestions discussed here provide an alternative way forward through modifications of existing methods. They are neither new nor particularly radical, especially when compared with the NRC proposals. More than fifty years ago, Russell and Burch suggested that toxicity testing, as usual…, is the scene of some confused thought, which may be delaying the exploitation of statistical methods. We have not infrequently heard the opinion expressed that… in toxicity tests you need a thoroughly heterogeneous mass of animals, and plenty of them. The physician, it is argued, is going to deal with patients with a very wide range of sensitivities to a given toxic action. There is a vague feeling that since this variation is quite uncontrolled, that of the test animals ought to be uncontrolled too… The fallacy consists in supposing that in order to obtain a wide inductive base a heterogeneous stock should be used… The proper procedure is, of course to use several different homogeneous samples, by using a plurality of pure lines (or preferably F1 crossbreds)… for otherwise the experimenter deprives himself of the possibility of making a relatively precise estimate of the error. (Russell and Burch 1959, 112)
Maybe the potential benefits of a multistrain study need to be reemphasized using a simple (and therefore not entirely realistic) example. The chloramphenicol study used here to illustrate a multistrain experiment showed no evidence of a decline in the white cell lineage at any dose level up to the highest dose used (2,500 mg/kg) in the CD-1 or BALB/c mice. So if this were an IND being tested using CD-1 mice, white blood cell (WBC) counts might not be a parameter of obvious interest in the clinical development of the compound. However, in the multistrain study there was a large, statistically significant decline in WBC counts in two of the inbred strains at this dose level. So the multistrain experiment would flag up that here is a character that needs to be carefully monitored during clinical development. Gene expression profiling of bone marrow cells in treated and untreated resistant (BALB/c) and susceptible (C3H) strains of mice might give some insight into mechanisms of this WBC decline. If a WBC response was judged to be of potential clinical importance, then the genetic basis of the difference between BALB/c and C3H could be explored. One approach would be to cross the two strains and intercross the F1 hybrids to produce about 100-200 F2 hybrids. These could be treated with chloramphenicol and the response recorded. The mice could then be typed using a large number of SNP markers. Any association between a marker and susceptibility would imply that the SNP or a gene linked to it is associated with response. This would not be a prohibitively difficult or expensive study, and it would almost certainly show whether susceptibility was due to a single or to multiple genes. Thus, the toxicologist would be able to supply important information to the clinicians that would not have been available when using an outbred stock, particularly if the stock were resistant.
Unfortunately, the regulators are not yet encouraging innovation. Although the FDA has brought attention to the failure of current preclinical toxicity tests, improving them does not seem to have high priority. Among the sixty projects listed in the Critical Path Initiative report for 2008, none is directly aimed at improving current animal toxicity testing methods. Better safety pharmacology testing (Pugsley, Authier, and Curtis 2008) using inbred strains within each pharmaceutical company prior to the formal regulatory testing may be an alternative.
It is time for toxicologists to think more critically about the design of their experiments and the animals they use. Maybe a relatively small change such as using inbred strains would result in considerable savings in the cost of drug development.
Conclusions
Too many potential new drugs pass the preclinical testing in animals but are subsequently rejected following clinical trials because of lack of efficacy and toxicity. This increases the cost of drug development. There have been substantial advances in mammalian genetics in the past couple of decades that depend on, or have been facilitated by, the use of inbred strains of mice and rats. These provide a resource that is underused by toxicologists, who appear never to have given much critical thought to the type of animals they use. Compared with outbred stocks, inbred strains are more stable, better defined, more uniform, have more extensive background data, and have a wider international distribution than outbred stocks. A toxicity screen using small numbers of inbred animals of several strains could be used without increasing the total numbers. This would broaden the genetic base of tests and reveal genetic variation that is not seen when using a single outbred stock. Safety margins could then be based on the most sensitive strains. Where large strain differences are identified, the mode of inheritance could be determined using the large number of genetic markers that are now available. In association with “omics” techniques such as gene expression profiling, the use of inbred strains could transform preclinical testing of new drugs in the twenty-first century. Now is the time for the pharmaceutical industry and the regulatory organizations to investigate these suggestions in more detail.